Abstract
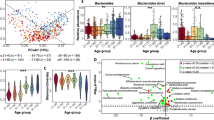
Biological aging has been associated with altered risk of aging-related diseases, but the contribution of the gut microbiota to this process remains poorly understood. Here, we constructed an interpretable gut microbiota age clock using metagenomic data from 8115 fecal samples across five continents. We discovered a key microbial perturbation occurring at 56–60 years of chronological age, which was validated in an independent cohort of 2263 metagenomes. This perturbation was associated with a decline in ecological stability and substantial changes in the abundance of core species. Notably, the association between gut microbiota age and diseases was identified to be significantly altered before and after this inflection time. Moreover, within-species analyses uncovered phylogenetic divergence for seven age-related species, such as Escherichia coli, alongside functional alterations in older individuals, including enhanced cell motility, carbohydrate metabolism and horizontal gene transfer. Overall, our global gut microbiome atlas uncovers a critical age transition phase, highlighting opportunities for microbiota-based therapies and offering novel insights into evolutionary dynamics during aging.
Similar content being viewed by others
Data availability
The curatedMetagenomicData resource is publicly accessible via https://doi.org/10.18129/B9.bioc.curatedMetagenomicData. The raw metagenomic sequencing data of the Chinese Gut Microbial Reference (CGMR) cohort are available in the China National Gene Bank (CNGB) under accession number CNP0004122.
Code availability
The bioinformatic analysis process used in this study is thoroughly described in Methods. Customized visualization and statistical analysis scripts are available on our team’s GitHub repository https://github.com/ZJYY-HY-team/Microbiome-aging-paper.
References
Guo, J. et al. Aging and aging-related diseases: from molecular mechanisms to interventions and treatments. Signal Transduct. Target Ther. 7, 391 (2022).
Niccoli, T. & Partridge, L. Ageing as a risk factor for disease. Curr. Biol. 22, R741–R752 (2012).
Kennedy, B. K. et al. Geroscience: linking aging to chronic disease. Cell. 159, 709–713 (2014).
Ahadi, S. et al. Personal aging markers and ageotypes revealed by deep longitudinal profiling. Nat. Med. 26, 83–90 (2020).
Rutledge, J., Oh, H. & Wyss-Coray, T. Measuring biological age using omics data. Nat. Rev. Genet. 23, 715–727 (2022).
Horvath, S. DNA methylation age of human tissues and cell types. Genome Biol. 14, 3156 (2013).
Almanzar, N. et al. A single-cell transcriptomic atlas characterizes ageing tissues in the mouse. Nature 583, 590–595 (2020).
Li, Y. et al. Large language model-based biological age prediction in large-scale populations. Nat. Med. 31, 2977–2990 (2025).
Martino, C. et al. Microbiota succession throughout life from the cradle to the grave. Nat. Rev. Microbiol. 20, 707–720 (2022).
Almeida, A. et al. A new genomic blueprint of the human gut microbiota. Nature 568, 499–504 (2019).
Kadyan, S. et al. Microbiome-based therapeutics towards healthier aging and longevity. Genome Med. 17, 75 (2025).
Lopez-Otin, C., Blasco, M. A., Partridge, L., Serrano, M. & Kroemer, G. Hallmarks of aging: an expanding universe. Cell. 186, 243–278 (2023).
Elliott, M. L. et al. Disparities in the pace of biological aging among midlife adults of the same chronological age have implications for future frailty risk and policy. Nat. Aging 1, 295–308 (2021).
Tian, Y. E. et al. Heterogeneous aging across multiple organ systems and prediction of chronic disease and mortality. Nat. Med. 29, 1221–1231 (2023).
Ghosh, T. S., Shanahan, F. & O’Toole, P. W. The gut microbiome as a modulator of healthy ageing. Nat. Rev. Gastroenterol. Hepatol. 19, 565–584 (2022).
Ram, U. et al. Age-specific and sex-specific adult mortality risk in India in 2014: analysis of 0.27 million nationally surveyed deaths and demographic estimates from 597 districts. Lancet Glob. Health 3, e767–e775 (2015).
Lehallier, B. et al. Undulating changes in human plasma proteome profiles across the lifespan. Nat. Med. 25, 1843–1850 (2019).
Shen, X. et al. Nonlinear dynamics of multi-omics profiles during human aging. Nat. Aging. 4, 1619–1634 (2024).
Liu, W.-S. et al. Plasma proteomics identify biomarkers and undulating changes of brain aging. Nat. Aging 5, 99–112 (2024).
He, Y. et al. Regional variation limits applications of healthy gut microbiome reference ranges and disease models. Nat. Med. 24, 1532–1535 (2018).
Claesson, M. J. et al. Gut microbiota composition correlates with diet and health in the elderly. Nature 488, 178–184 (2012).
Shen, L. et al. Mapping the gut microbial structural variations in healthy aging within the Chinese population. Cell Rep. 43, 114968 (2024).
Chen, L. et al. The long-term genetic stability and individual specificity of the human gut microbiome. Cell. 184, 2302–2315 e2312 (2021).
Sato, Y. et al. Novel bile acid biosynthetic pathways are enriched in the microbiome of centenarians. Nature. 599, 458–464 (2021).
Crost, E. H. et al. The mucin-degradation strategy of Ruminococcus gnavus: The importance of intramolecular trans-sialidases. Gut Microbes 7, 302–312 (2016).
Van Rossum, T., Ferretti, P., Maistrenko, O. M. & Bork, P. Diversity within species: interpreting strains in microbiomes. Nat. Rev. Microbiol. 18, 491–506 (2020).
Xiao, Y. et al. Human gut-derived B. longum subsp. longum strains protect against aging in a D-galactose-induced aging mouse model. Microbiome 9, 180 (2021).
Pasolli, E. et al. Accessible, curated metagenomic data through ExperimentHub. Nat. Methods 14, 1023–1024 (2017).
Ma, S. et al. Population structure discovery in meta-analyzed microbial communities and inflammatory bowel disease using MMUPHin. Genome Biol. 23, 208 (2022).
Fu, J. et al. Ageing trajectory of the gut microbiota is associated with metabolic diseases in a chronological age-dependent manner. Gut 72, 1431–1433 (2022).
Olecka, M. et al. Nonlinear DNA methylation trajectories in aging male mice. Nat. Commun. 15, 3074 (2024).
Marquez, E. J. et al. Sexual-dimorphism in human immune system aging. Nat. Commun. 11, 751 (2020).
Huang, P. et al. Gut microbial genomes with paired isolates from China illustrate probiotic and cardiometabolic effects. Cell Genom. 4, 100559 (2024).
Chen, L. et al. Integrative multiomics analysis reveals host-microbe-metabolite interplays associated with the aging process in Singaporeans. Gut. Microbes. 14, 2070392 (2022).
Ghosh, T. S. et al. Mediterranean diet intervention alters the gut microbiome in older people reducing frailty and improving health status: the NU-AGE 1-year dietary intervention across five European countries. Gut 69, 1218–1228 (2020).
Coelho, L. P. et al. Multivariable association discovery in population-scale meta-omics studies. PLoS Comput. Biol. 17, e1009442 (2021).
Sloan, W. T., Woodcock, S., Lunn, M., Head, I. M. & Curtis, T. P. Modeling taxa-abundance distributions in microbial communities using environmental sequence data. Micro. Ecol. 53, 443–455 (2007).
Zorea, A. et al. Plasmids in the human gut reveal neutral dispersal and recombination that is overpowered by inflammatory diseases. Nat. Commun. 15, 3147 (2024).
Cheng, M. et al. A genome and gene catalog of the aquatic microbiomes of the Tibetan Plateau. Nat. Commun. 15, 1438 (2024).
Pandit, S. N., Kolasa, J. & Cottenie, K. Contrasts between habitat generalists and specialists: an empirical extension to the basic metacommunity framework. Ecology 90, 2253–2262 (2009).
Mei, Z. et al. Strain-specific gut microbial signatures in type 2 diabetes identified in a cross-cohort analysis of 8,117 metagenomes. Nat. Med. 30, 2265–2276 (2024).
Chen, S. et al. Bifidobacterium adolescentis regulates catalase activity and host metabolism and improves healthspan and lifespan in multiple species. Nat. Aging. 1, 991–1001 (2021).
Rees, D. C., Johnson, E. & Lewinson, O. ABC transporters: the power to change. Nat. Rev. Mol. Cell Biol. 10, 218–227 (2009).
Hall, A. B. et al. A novel Ruminococcus gnavus clade enriched in inflammatory bowel disease patients. Genome Med. 9, 103 (2017).
Rocha, E. P., Cornet, E. & Michel, B. Comparative and evolutionary analysis of the bacterial homologous recombination systems. PLoS Genet. 1, e15 (2005).
Lerner, A., Matthias, T. & Aminov, R. Potential effects of horizontal gene exchange in the human gut. Front. Immunol. 8, 1630 (2017).
Sun, H. et al. Regulation of flagellar motility and biosynthesis in enterohemorrhagic Escherichia coli O157:H7. Gut Microbes 14, 2110822 (2022).
Bafei, S. E. C. & Shen, C. Biomarkers selection and mathematical modeling in biological age estimation. npj Aging 9, 13 (2023).
Robinson, O. et al. Determinants of accelerated metabolomic and epigenetic aging in a UK cohort. Aging Cell 19, e13149 (2020).
Belsky, D. W. et al. Quantification of the pace of biological aging in humans through a blood test, the DunedinPoAm DNA methylation algorithm. Elife 9, e54870 (2020).
Tanaka, T. et al. Plasma proteomic signature of age in healthy humans. Aging Cell 17, e12799 (2018).
Li, J. et al. Determining a multimodal aging clock in a cohort of Chinese women. Med 4, 825-848.e13 (2023).
Ding, Y. et al. Comprehensive human proteome profiles across a 50-year lifespan reveal aging trajectories and signatures. Cell 188, 5763–5784.e5726 (2025).
Wu, P. et al. Identifying tipping points during healthy brain aging through single-nucleus transcriptomic analysis. Adv. Sci. 12, e05779 (2025).
Hahn, O. et al. Atlas of the aging mouse brain reveals white matter as vulnerable foci. Cell 186, 4117–4133.e4122 (2023).
Wilmanski, T., Gibbons, S. M. & Price, N. D. Healthy aging and the human gut microbiome: why we cannot just turn back the clock. Nat. Aging 2, 869–871 (2022).
Wang, T. et al. Divergent age-associated and metabolism-associated gut microbiome signatures modulate cardiovascular disease risk. Nat. Med. 30, 1722–1731 (2024).
Dong, C. et al. Disentangling the age-related manner in the associations between gut microbiome and women’s health: a multi-cohort microbiome study. Gut Microbes 15, 2290320 (2023).
Groussin, M., Mazel, F. & Alm, E. J. Co-evolution and co-speciation of host-gut bacteria systems. Cell Host Microbe 28, 12–22 (2020).
Hayashi, F. et al. The innate immune response to bacterial flagellin is mediated by Toll-like receptor 5. Nature 410, 1099–1103 (2001).
Stecher, B. et al. Gut inflammation can boost horizontal gene transfer between pathogenic and commensal Enterobacteriaceae. Proc. Natl. Acad. Sci. USA 109, 1269–1274 (2012).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2006).
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Lundberg, S. M. et al. From local explanations to global understanding with explainable AI for trees. Nat. Mach. Intell. 2, 56–67 (2020).
Musaev, A., Grigoriev, D. & Kolosov, M. Adaptive algorithms for change point detection in financial time series. AIMS Math. 9, 35238–35263 (2024).
Valderrama Balaguera, J. C. Precipitation forecast estimation applying the change point method and ARIMA. Cogent Eng. 11, 2340191(2024).
Ross, G. J. Parametric and nonparametric sequential change detection in R: the cpm package. J. Stat. Softw. 66, 1–20 (2015).
Ross, G. J. & Adams, N. M. Two nonparametric control charts for detecting arbitrary distribution changes. J. Qual. Technol. 44, 102–116 (2017).
Libiseller, C. & Grimvall, A. Performance of partial Mann–Kendall tests for trend detection in the presence of covariates. Environmetrics 13, 71–84 (2002).
Shan, Q. et al. Do extreme climate events cause the degradation of Malus sieversii forests in China? Front. Plant Sci. 12, 608211 (2021).
Dini-Andreote, F., Stegen, J. C., van Elsas, J. D. & Salles, J. F. Disentangling mechanisms that mediate the balance between stochastic and deterministic processes in microbial succession. Proc. Natl. Acad. Sci. USA 112, E1326–E1332 (2015).
Carpenter, B. et al. Stan: a probabilistic programming language. J Stat Softw 76, 1 (2017).
Paradis, E., Claude, J. & Strimmer, K. APE: analyses of phylogenetics and evolution in R language. Bioinformatics 20, 289–290 (2004).
Gu, Z. Complex heatmap visualization. Imeta 1, e43 (2022).
Wu, T. et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Acknowledgements
This study was supported by funding from the National Natural Science Foundation of China (NSFC82300623, S.H, NSFC82302610, C.Q, NSFC82472340, Y.H, and NSFC82272391, Y.H.), the National Key R&D Program of China (2019YFA0802300, Y.H. and SQ2022YFA090032, Y.H.), Guangdong Provincial Clinical Research Center for Laboratory Medicine (2023B110008) and Guangdong Basic and Applied Basic Research Foundation (2023A1515012538, W.S.).
Author information
Authors and Affiliations
Contributions
W.S., C. Qi, Y.H., and S.H. contributed to conceptualization and funding. Z. Qi, L. Chen, X.C., W.W., W.M., H.Z., and Y.H. provided the validation data. J.F., J.Z., R.H., Q. Dong, and H. Mao curated and analyzed the data. J.F., J.Z., and R.H. prepared the manuscript. C. Qi, Y.H., S.H., and L. Chen reviewed and provided critical suggestions on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fu, J., Zhang, J., He, R. et al. A global metagenomic atlas of aging identifies a microbiota phase transition associated with disease risk. npj Biofilms Microbiomes (2026). https://doi.org/10.1038/s41522-026-00970-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41522-026-00970-4