Abstract
In advanced HER2-positive breast cancer, the standard taxane-trastuzumab-pertuzumab (THP) regimen faces competition from new therapies, emphasizing the need for biomarkers to guide treatment. This study evaluates the HER2DX ERBB2 mRNA score as a prognostic predictor, aiming to tailor treatment strategies. We retrospectively analyzed 94 patients treated with the THP regimen between 2010 and 2024. The HER2DX ERBB2 mRNA score was categorized as low (n = 14), medium (n = 20), or high (n = 60), and its correlation with progression-free survival (PFS) and overall survival (OS) was assessed using Cox regression models. The median follow-up was 31.5 months. Patients with ERBB2-high scores had significantly better median PFS (33.9 vs. 10.6 months, hazard ratio [HR] = 0.40, 95% CI: 0.24–0.69, p < 0.001) and OS (not reached vs. 30.8 months, HR = 0.26, 95% CI: 0.13–0.49, p < 0.001) compared to ERBB2-low patients. Based on these findings, further validation of this biomarker in tumor samples from the CLEOPATRA phase III trial is ongoing, which could help optimize treatment strategies in this population.
Similar content being viewed by others
Introduction
In the first-line treatment of advanced HER2-positive (HER2+) breast cancer, the standard regimen of taxane-trastuzumab-pertuzumab (THP) provides a median progression-free survival (PFS) of approximately 18–21 months and a median overall survival (OS) of around 57–65 months1,2. While THP remains a cornerstone therapy, antibody–drug conjugates (ADCs) are increasingly challenging this standard. ADCs, such as trastuzumab deruxtecan (T-DXd), have demonstrated superior outcomes compared to standard treatments in patients previously treated with chemotherapy and anti-HER2 therapy3,4,5. The ongoing DestinyBreast-09 trial (NCT04784715)6, exploring THP versus T-DXd with or without pertuzumab in the first-line setting, reflects this shift. These developments underscore the critical need for robust biomarkers to guide treatment decisions in advanced HER2+ breast cancer.
Efforts to identify prognostic factors in patients treated with THP have yielded only a few clinical and analytical markers—such as non-visceral disease, good performance status, oligometastatic disease, and low baseline neutrophil/lymphocyte ratio—linked with longer PFS7,8. However, the correlation between these factors and OS remains unclear, necessitating novel approaches in the context of precision oncology.
Transcriptomics in HER2+ breast cancer has significantly advanced the understanding of breast cancer’s complexity and heterogeneity9,10,11, enabling more precise diagnostic tools. To date, no biomarker has been validated for predicting prognosis or treatment efficacy in advanced HER2+ breast cancer. However, in early-stage HER2+ breast cancer, the 27-gene HER2DX genomic assay, which provides a pathological complete response (pCR) likelihood score and a risk score, has demonstrated both prognostic and predictive value12,13,14,15,16,17,18,19. In addition, HER2DX provides the ERBB2 mRNA score, offering a continuous assessment of ERBB2 mRNA levels associated with HER2 protein expression, reported on a scale from 1 to 9915. It categorizes patients into three groups: ERBB2-low (1–32), ERBB2-medium (33–50), and ERBB2-high (51–99). These cutoffs were trained to predict clinical HER2 status based on the American Society of Clinical Oncology/College of American Pathologists (ASCO/CAP) guidelines20. The 33 cutoff in the score distinguishes between ERBB2-low and ERBB2-medium/high, correlating with clinical HER2-negative versus HER2+ status, while the 51 cutoff separates ERBB2-medium from ERBB2-high, corresponding to the lower 33% of HER2DX ERBB2 expression levels in HER2+ breast cancer15.
Given the potential of the HER2DX ERBB2 mRNA score as a predictive biomarker, particularly in the context of THP treatment, our study aims to validate the ability of the HER2DX ERBB2 mRNA score to predict survival outcomes in patients with advanced HER2+ breast cancer treated with first-line THP. By identifying patients who may benefit more from THP versus emerging therapies like T-DXd, we seek to refine therapeutic approaches in this evolving landscape.
Results
Clinicopathological features of the patients
A total of 94 patients diagnosed with HER2+ metastatic breast cancer and treated with THP were included in this study. The baseline characteristics of the cohort are summarized in Table 1. The median follow-up was 31.5 months (1.5–162.5 months). Our cohort consisted of hormone receptor-positive cases (61.7%), patients with visceral disease (73.4%), brain metastasis (13.8%), and de novo metastatic disease (53.2%). Most patients were treated with paclitaxel (n = 79, 84%), and the remaining with docetaxel (n = 15, 16%). Based on pre-specified cutoffs for the HER2DX ERBB2 mRNA score, the cohort distribution was 14.9% in the low group, 21.3% in the medium group, and 63.8% in the high group (Fig. 1A). No significant differences in HER2DX ERBB2 mRNA score were observed between primary (n = 60) and metastatic tumor (n = 34) tissues as a categorical or continuous variable (Fig. 1B, C). However, the HER2DX ERBB2 mRNA score was statistically significantly higher in patients with de novo metastatic disease than those with recurrent disease (Fig. 1D, E). The HER2DX ERBB2 score, analyzed as a continuous variable, was significantly associated with HER2 IHC (Supplementary Fig. 1a). Among HER2 IHC 3+ tumors (n = 63), the distribution of HER2DX ERBB2 scores as a categorical variable showed that 76.2% were classified as high, 17.5% as medium, and 6.3% as low. In HER2 IHC 2+ tumors (n = 24), the distribution was 50.0% high, 20.8% medium, and 29.2% low. Notably, in HER2 IHC 0 (n = 3) and 1+ (n = 2) tumors, no cases were classified as high (Supplementary Fig. 1b).
A Distribution of HER2DX ERBB2 mRNA score groups across the 94 patients. B Distribution of HER2DX ERBB2 mRNA score groups in primary and metastatic tumor. P-value was determined using Fisher’s exact test. C HER2DX ERBB2 mRNA score in primary and metastatic tumors. P-value was determined using an unpaired t-test. D Distribution of HER2DX ERBB2 mRNA score groups in de novo and relapsed disease. The P-value was determined using Fisher’s exact test. E HER2DX ERBB2 mRNA score in de novo and relapsed disease. The P-value was determined using an unpaired t-test.
HER2DX ERBB2 mRNA score and clinical endpoints
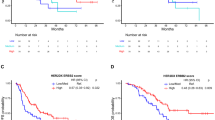
The median PFS and OS were 27.1 months (95% CI: 16.1–43.7 months) and 62.7 months (95% CI: 47.9–NR months), respectively. The HER2DX ERBB2 mRNA score, when analyzed as a continuous variable, showed a significant association with both PFS (HR = 0.69 [95% CI: 0.53–0.88], p = 0.028) and OS (HR = 0.58 [95% CI: 0.43–0.77], p < 0.001) (Fig. 2A–D). When compared to the HER2DX ERBB2-low group, the HER2DX ERBB2-high group demonstrated significantly better outcomes, with a median PFS of 33.9 months versus 10.6 months (HR = 0.40 [95% CI: 0.24–0.69], p < 0.001) and a median OS that was not reached versus 30.8 months (HR = 0.26 [95% CI: 0.13–0.49], p < 0.001). Eight of nine (88.8%) patients without progression for over 5 years, and all patients without death over 8 years (n = 4) had HER2DX ERBB2-high scores.
A Association of HER2DX ERBB2 mRNA score high, medium, and low groups and PFS. B Association of HER2DX ERBB2 mRNA score high and low/medium groups and PFS. C Association of HER2DX ERBB2 mRNA score high, medium, and low groups and OS. D Association of HER2DX ERBB2 mRNA score high and low/medium groups and OS.
In univariate analyses, clinical variables associated with PFS included de novo metastasis (HR = 0.56 [95% CI: 0.33–0.95], p = 0.031), visceral metastasis (HR = 2.1 [95% CI: 1.06–4.20], p = 0.033), bone-only disease (HR = 0.29 [95% CI: 0.11–0.81], p = 0.018), brain metastasis (HR = 2.4 [95% CI: 1.24–4.51], p = 0.009), and the number of metastatic sites (≥3 vs <3) (HR = 3.19 [95%CI: 1.86–5.46], p < 0.001) (Table 2), while the clinical variables associated with OS included visceral metastasis (HR = 2.5 [95% CI: 1.05–5.95], p = 0.031), brain metastasis (HR = 2.4 [95% CI: 1.2–5.05], p = 0.016), and the number of metastatic sites (≥3 vs <3) (HR = 2.8 [95% CI: 1.5–5.24], p = 0.001) (Table 3). Notably, HER2 IHC (3+ vs others) was significantly associated with better PFS (HR = 2.3 [95% CI: 1.32–3.96], p = 0.003) and OS (HR = 4.3 [95% CI: 2.25–8.06], p < 0.001).
In multivariable analyses, the HER2DX ERRB2-high group remained significantly associated with better PFS (HR = 0.41 [95% CI: 0.20–0.85], p = 0.016) and OS (HR = 0.38 [95% CI: 0.17–0.85], p = 0.018), while HER2 IHC (3+ vs others) lost its significance (PFS HR = 0.82 [95% CI: 0.38–1.76], p = 0.61) and (OS HR = 0.47 [95% CI: 0.21–1.07], p = 0.072)(Tables 2 and 3).
Finally, in this cohort, the ORR was 79.14%. The HER2DX ERBB2 mRNA score was not associated with ORR (OR = 1.21 [95% CI: 0.89–1.65], p = 0.235). ORR by HER2DX ERBB2 mRNA categories was 78.5% for the low group, 63.2% for the medium group, and 96.1% for the high group (OR HER2DX ERBB2 high vs low/med = 2.37 [95% CI: 0.85–6.75], p = 0.100).
Discussion
This study is, to our knowledge, the first to evaluate the association between the HER2DX ERBB2 mRNA score and clinical outcomes in HER2+ advanced breast cancer patients treated with first-line THP chemotherapy. Our findings confirm that a higher HER2DX ERBB2 mRNA score is significantly associated with improved PFS and OS, highlighting the prognostic value of HER2DX ERBB2 mRNA expression in this setting.
Previous research by Brasó-Maristany et al.21 evaluated the role of HER2DX ERBB2 in metastatic breast cancer, demonstrating that higher ERBB2 mRNA levels were significantly associated with better outcomes in patients treated with ado-trastuzumab emtansine (T-DM1). The study showed that higher ERBB2 mRNA levels correlated with longer PFS and OS, regardless of HER2 IHC levels, hormone receptor status, brain metastasis, and line of therapy21. Our findings in the first-line THP setting further support the prognostic value of HER2DX ERBB2 mRNA score, indicating that ERBB2 mRNA is a robust biomarker across multiple lines of therapy in metastatic HER2+ breast cancer.
In our cohort, all patients who remained progression-free beyond 5 years and alive beyond 8 years had high HER2DX ERBB2 mRNA scores, underscoring the clinical significance of this biomarker. Identifying long-term responders is crucial for optimizing treatment strategies. A high HER2DX ERBB2 mRNA score could help clinicians personalize treatment by continuing THP in responders, while those with lower scores might benefit from more aggressive upfront treatments or alternative therapies. This approach could minimize unnecessary toxicity and improve the overall quality of life for patients.
The ongoing DESTINY-Breast09 trial could further reshape clinical practice in HER2+ metastatic breast cancer6. This phase III trial is recruiting 1134 patients and is comparing trastuzumab deruxtecan (T-DXd) to THP in the first-line setting, with PFS as the primary endpoint. Given T-DXd’s promising results in later lines of therapy3,4,5, it may become the new standard of care for first-line treatment if the trial proves its superiority. HER2DX ERBB2 mRNA score could help clinicians stratify patients who are more likely to benefit from T-DXd, particularly those with lower ERBB2 scores who may not respond well or do well to THP. The results of DESTINY-Breast09, combined with HER2DX ERBB2 mRNA score stratification, could lead to more personalized treatment approaches and improved outcomes for HER2+ metastatic breast cancer patients. This is particularly relevant when considering differences in adverse events and treatment-related quality of life between T-DXd and THP. The HER2DX assay could play a crucial role in identifying long-term responders to THP, offering a potentially better-tolerated treatment option, particularly in the context of maintenance therapy. Furthermore, the recent results from the PATINA trial, which demonstrated the benefit of maintenance palbociclib following THP induction in HER2+/hormone receptor-positive disease, underscore the potential of THP as a viable treatment strategy, even as new evidence for T-DXd emerges22.
Interestingly, HER2 IHC was significantly associated with better OS in univariate analyses, but it lost significance when the HER2DX ERBB2 mRNA score was included in multivariable analyses. This suggests that HER2DX ERBB2 mRNA score may provide a more accurate assessment of HER2 status compared to IHC. While IHC has been the standard for HER2 assessment, it relies on subjective interpretation and can be affected by tissue fixation and antibody variability20,23. In contrast, HER2DX offers a standardized and quantitative measure of ERBB2 mRNA24, potentially improving the precision of HER2 assessment.
The limitations of this study include its retrospective nature and relatively small sample size. Retrospective data collection introduces inherent biases, and the study’s conduct across two Spanish institutions may limit generalizability to other populations. Additionally, the lack of centralized HER2 testing could have introduced variability in IHC interpretation. Nonetheless, although not centralized, it was performed by expert breast pathologists. Although HER2 ISH testing was not consistently reported for all patients, we acknowledge its importance as the gold standard for HER2 status assessment. However, the lack of consistent FISH results reflects real-world clinical practice, where HER2 IHC 3+ is often sufficient for determining HER2 positivity. Future prospective studies with larger sample sizes and diverse populations are warranted to validate these findings. Another potential limitation is that tumor response was assessed by the treating physicians per clinical practice. With other tools, such as RECIST criteria, an association between HER2DX ERBB2 mRNA score and ORR could be found. We acknowledge the lack of information regarding other biomarkers (e.g. PIK3CA mutatations) that have been associated with poorer outcomes in first-line HER2+ MBC25. Such biomarkers are not currently performed by clinical practice, precluding us from having access to this information. Finally, subsequent treatments after THP were not included in the analyses, potentially confounding the association with OS.
Despite these limitations, this study provides valuable insights into the utility of the HER2DX ERBB2 mRNA score in HER2+ advanced breast cancer treated with THP. By offering a standardized, quantitative measure of HER2 status, HER2DX has the potential to improve upon conventional HER2 assessment methods and help tailor treatment strategies for patients.
In conclusion, the HER2DX ERBB2 mRNA score is significantly associated with improved survival in patients receiving first-line THP for HER2+ advanced breast cancer. The ongoing DESTINY-Breast09 trial may further influence clinical practice, and incorporating HER2DX into treatment decision-making could optimize treatment strategies and enhance personalized care. Based on the results of this study, the HER2DX ERBB2 mRNA score is also being tested in the CLEOPATRA phase III trial1, which led to the approval of pertuzumab.
Methods
Study population
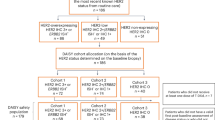
We conducted a retrospective, multi-center observational study in patients treated with the standard taxane, trastuzumab, and pertuzumab (THP) regimen in the first-line setting for advanced HER2+ breast cancer between 2010 and 2024 from two Spanish institutions (Hospital Clinic of Barcelona, and Hospital Universitario 12 de Octubre in Madrid). Demographic and clinical–pathological characteristics were collected from electronic medical records. Eligible patients met the following criteria: histological confirmation of breast cancer with positive HER2 status, defined as either 3+ immunohistochemistry (IHC) or HER2 amplification by in situ hybridization (ISH); locally advanced disease not amenable to curative treatment or metastatic disease; and treatment with the THP regimen in the first-line setting. Both newly diagnosed and recurrent cases were included.
Clinical HER2 status
HER2 status was locally assessed using IHC and/or ISH, following the ASCO/CAP guidelines at the time of diagnosis20,23. IHC was performed using an anti-HER-2/neu (4B5) Rabbit Monoclonal Primary Antibody kit (Ventana Medical Systems Inc., Oro Valley, AZ, USA). HER2 ISH was conducted using the FDA-approved XL ERBB2 (HER2/NEU) AMP (MetaSystems Probes, Altlußheim, Germany).
RNA Extraction and HER2DX assay
Patients without sufficient archival tissue samples for HER2DX ERBB2 mRNA score assay were excluded. One formalin-fixed paraffin-embedded (FFPE) tumor sample per patient was selected: if available, a biopsy of the metastatic site nearest in time to the start of THP was preferred. The standardized HER2DX assay was performed in a central lab (Barcelona, Spain). The HER2DX assay quantifies mRNA expression levels of 27 target genes and 5 normalization genes with constitutive expression (GAPD, PUM1, ACTB, RPLP0, and PSMC4). These 27 target genes are grouped into four distinct gene signatures: immune infiltration (CD27, CD79A, HLA-C, IGJ, IGKC, IGL, IGLV3-25, IL2RG, CXCL8, LAX1, NTN3, PIM2, POU2AF1, and TNFRSF17), tumor cell proliferation (EXO1, ASPM, NEK2, and KIF23), luminal differentiation (BCL2, DNAJC12, AGR3, AFF3, and ESR1), and HER2 amplicon expression (ERBB2, GRB7, STARD3, and TCAP). HER2DX integrates these molecular profiles with clinical characteristics, including tumor and nodal stage, to generate three key scores: long-term prognosis (risk score), likelihood of achieving a pathological complete response (pCR score), and ERBB2 mRNA expression (ERBB2 score)24. The HER2DX ERBB2 score was calculated based on the ERBB2 mRNA levels. Pre-established cutoffs, derived from prior validation studies, were used to categorize scores as low, medium, or high15.
Endpoints and statistical analyses
The primary objective was to assess the association between HER2DX ERBB2 mRNA score, as both a continuous variable and categorized (low, medium, high), with PFS and OS. Secondary objectives included evaluating the association of HER2DX ERBB2 mRNA score with the overall response rate (ORR) according to the physician´s local report. PFS was defined as the time from the initiation of THP treatment to disease progression or death, whichever occurred first. OS was defined as the time from the start of THP treatment to death from any cause. Missing data were not imputed, and patients with incomplete clinical data were excluded from the respective analyses. Univariate and multivariable Cox proportional hazard regression analyses were performed to assess associations with PFS and OS, while logistic regression was used to evaluate ORR. Sensitivity analyses were performed to ensure the robustness of the findings, and proportional hazard assumptions were tested for the Cox models. Statistical analyses were conducted using R v4.2.2, with a two-sided p-value < 0.05 considered statistically significant.
Data availability
Data are available upon reasonable request.
References
Swain, S. M. et al. Pertuzumab, trastuzumab, and docetaxel for HER2-positive metastatic breast cancer (CLEOPATRA): end-of-study results from a double-blind, randomised, placebo-controlled, phase 3 study. Lancet Oncol. 21, 519–530 (2020).
Miles, D. et al. Final results from the PERUSE study of first-line pertuzumab plus trastuzumab plus a taxane for HER2-positive locally recurrent or metastatic breast cancer, with a multivariable approach to guide prognostication. Ann. Oncol. 32, 1245–1255 (2021).
André, F. et al. Trastuzumab deruxtecan versus treatment of physician’s choice in patients with HER2-positive metastatic breast cancer (DESTINY-Breast02): a randomised, open-label, multicentre, phase 3 trial. Lancet 401, 1773–1785 (2023).
Cortés, J. et al. Trastuzumab Deruxtecan versus Trastuzumab Emtansine for Breast Cancer. N. Engl. J. Med. 386, 1143–1154 (2022).
Hurvitz, S. A. et al. Trastuzumab deruxtecan versus trastuzumab emtansine in patients with HER2-positive metastatic breast cancer: updated results from DESTINY-Breast03, a randomised, open-label, phase 3 trial. Lancet 401, 105–117 (2023).
Tolaney, S. M. et al. 328TiP Phase III study of trastuzumab deruxtecan (T-DXd) with or without pertuzumab vs a taxane, trastuzumab and pertuzumab in first-line (1L), human epidermal growth factor receptor 2–positive (HER2+) metastatic breast cancer (mBC): DESTINY-Breast09. Ann. Oncol. 32, S507–S508 (2021).
Arciero, C. A. et al. ER+/HER2+ breast cancer has different metastatic patterns and better survival than ER-/HER2+ breast cancer. Clin. Breast Cancer 19, 236–245 (2019).
Ding, N. et al. Prognostic value of baseline neutrophil/lymphocyte ratio in HER2-positive metastatic breast cancer: exploratory analysis of data from the CLEOPATRA trial. Breast Cancer Res. 26, 9 (2024).
Prat, A. et al. A multivariable prognostic score to guide systemic therapy in early-stage HER2-positive breast cancer: a retrospective study with an external evaluation. Lancet Oncol. 21, 1455–1464 (2020).
Prat, A. et al. Molecular features and survival outcomes of the intrinsic subtypes within HER2-positive breast cancer. J. Natl Cancer Inst. 106 (2014).
Schettini, F. & Prat, A. Dissecting the biological heterogeneity of HER2-positive breast cancer. Breast 59, 339–350 (2021).
Bueno-Muiño, C. et al. Assessment of a genomic assay in patients with ERBB2-positive breast cancer following neoadjuvant trastuzumab-based chemotherapy with or without pertuzumab. JAMA Oncol. 9, 841–846 (2023).
Guarneri, V. et al. HER2DX genomic test in HER2-positive/hormone receptor-positive breast cancer treated with neoadjuvant trastuzumab and pertuzumab: a correlative analysis from the PerELISA trial. eBioMedicine 85, 104320 (2022).
Llombart-Cussac, A. et al. HER2DX genomic assay in HER2-positive early breast cancer treated with Trastuzumab and Pertuzumab: a correlative analysis from PHERGain phase II trial. Clin. Cancer Res. 30, 4123–4130 (2024).
Prat, A. et al. Development and validation of the new HER2DX assay for predicting pathological response and survival outcome in early-stage HER2-positive breast cancer. eBioMedicine 75, 103801 (2022).
Tarantino, P. et al. Adjuvant Trastuzumab Emtansine Versus Paclitaxel Plus Trastuzumab for Stage I Human Epidermal Growth Factor Receptor 2—positive breast cancer: 5-year results and correlative analyses from ATEMPT. J. Clin. Oncol. 42, 3652–3665 (2024).
Villacampa, G. et al. Prognostic value of HER2DX in early-stage HER2-positive breast cancer: comprehensive analysis of 757 patients in the Sweden Cancerome Analysis Network Breast dataset (SCAN-B). ESMO Open 9, 102388 (2024).
Villacampa, G. et al. Association of HER2DX with pathological complete response and survival outcomes in HER2-positive breast cancer. Ann. Oncol. 34, 783–795 (2023).
Waks, A. G. et al. Assessment of the HER2DX assay in patients with ERBB2-positive breast cancer treated with neoadjuvant Paclitaxel, Trastuzumab, and Pertuzumab. JAMA Oncol. 9, 835–840 (2023).
Wolff, A. C. et al. Recommendations for human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists Clinical Practice Guideline Update. J. Clin. Oncol. 31, 3997–4013 (2013).
Brasó-Maristany, F. et al. HER2DX ERBB2 mRNA expression in advanced HER2-positive breast cancer treated with T-DM1. J. Natl Cancer Inst. 115, 332–336 (2022).
Metzger, O. et al. AFT-38 PATINA: A randomized, open label, phase III trial to evaluate the efficacy and safety of palbociclib + anti-HER2 therapy + endocrine therapy vs anti-HER2 therapy + endocrine therapy after induction treatment for hormone receptor-positive/HER2-positive metastatic breast cancer. 2024 San Antonio Breast Cancer Symposium. Abstract P2-03-20 (SESS-18111). Presented December 10, 2024.
Wolff, A. C. et al. Human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists Clinical Practice Guideline Focused Update. J. Clin. Oncol. 36, 2105–2122 (2018).
Tolaney, S. M. et al. HER2DX genomic test in early-stage HER2-positive breast cancer. ESMO Open 9, 103987 (2024).
Baselga, J. et al. Biomarker analyses in CLEOPATRA: a phase III, placebo-controlled study of pertuzumab in human epidermal growth factor receptor 2-positive, first-line metastatic breast cancer. J. Clin. Oncol. 32, 3753–3761 (2014).
Acknowledgements
A.P. received funding from the Breast Cancer Research Foundation (BCRF-22-198, BCRF-23-198 and BCRF-24-198), Beca Marta Santamaría, Fundación CRIS contra el Cancer PR_EX_2021-14, Agència de Gestó d’Ajuts Universitaris I de Recerca 2021 SGR 01156, Fundación Fero BECA ONCOXXI21, Asociación Cáncer de Mama Metastásico IV Premios M. Chiara Giorgetti, PI22/01017: funded by Instituto de Salud Carlos III (ISCIII) and co-funded by the European Union, and RESCUER: funded by European Union’s Horizon 2020 Research and Innovation Program under Grant Agreement No. 847912. F.B.-M. received funding from Fundación científica AECC Ayudas Investigador AECC 2021 (INVES21943BRAS). This study was funded by Reveal Genomics. The study was designed and performed by investigators from Hospital Universitario 12 de Octubre, Hospital Clinic Barcelona, and Reveal Genomics. All authors had full access to all data in the study and had final responsibility for the decision to submit for publication.
Author information
Authors and Affiliations
Contributions
A.P., R.S.B., and E.C. designed the study. All authors contributed to data collection and assembly, wrote and reviewed the article, and approved the final version for submission.
Corresponding author
Ethics declarations
Ethics
The study was conducted in accordance with Good Clinical Practice guidelines and the World Medical Association Declaration of Helsinki. Patient anonymity was maintained by de-identifying clinical and genomic data before analysis. All patients provided written informed consent. The study protocol was approved by independent ethics committees at each participating institution (Institutional Review Board from Hospital Clinic Barcelona and Hospital Universitario 12 de Octubre).
Competing interests
R.S.-B. reports advisory/consulting/speaker fees from Roche, AstraZeneca, Novartis, Lilly, Daiichi Sankyo, Pfizer, Eisai, GlaxoSmithKline, Reveal Genomics, and Gilead; travel expenses from Pfizer, AstraZeneca, Gilead, Novartis, and Roche. O.M.S. reports advisory/consulting fees from Reveal Genomics, Roche, and AstraZeneca, lecture fees from Daiichi Sankyo, Novartis, Pfizer, and Eisai, and travel expenses from Gilead and Novartis. E.S. declares personal fees for educational events and/or material from Novartis, Pfizer, Eisai, and Daiichi Sankyo; advisory fees from Pfizer and Seagen; and travel/accommodation expenses from Gilead, Daiichi Sankyo, Novartis, and Lilly. P.T. reports advisory and consulting fees from AstraZeneca, Daiichi-Sankyo, Adamed, Novartis, Pfizer, Lilly, Esteve, Gilead, Roche, and Reveal Genomics. T.P. reports advisory and consulting fees or speaker honoraria from Novartis, Astra Zeneca, Lilly, Pfizer, Veracyte, Gilead, and Roche, and support for attending meetings and/or travel from Gilead. A.M. reports advisory and consulting fees from AstraZeneca, Clovis (ended), GSK, MSD, PharmaMar, Pharma&, Immunogen. F.B-M. reports part-time employment from Reveal Genomics and has patents filed: PCT/EP2022/086493, PCT/EP2023/060810, EP23382703, and EP23383369. E.C. reports advisory/consulting/speaker fees from Roche, AstraZeneca, Novartis, Lilly, Daiichi Sankyo, Pfizer, and Gilead; travel expenses from Pfizer, AstraZeneca, MSD, Novartis, and Roche. A.P. reports advisory and consulting fees from AstraZeneca, Roche, Pfizer, Novartis, Daiichi Sankyo, Ona Therapeutics, and Peptomyc, lecture fees from AstraZeneca, Roche, Novartis, and Daiichi Sankyo, institutional financial interests from AstraZeneca, Novartis, Roche, and Daiichi Sankyo; stockholder and employee of Reveal Genomics; patents filed PCT/EP2016/080056, PCT/EP2022/086493, PCT/EP2023/060810, EP23382703, and EP23383369; editor of NPJ Breast Cancer journal. The remaining authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sánchez-Bayona, R., Martínez-Sáez, O., Romero-Romero, D. et al. HER2DX ERBB2 mRNA score in first-line advanced HER2-positive breast cancer treated with chemotherapy, trastuzumab, and pertuzumab. npj Breast Cancer 11, 37 (2025). https://doi.org/10.1038/s41523-025-00753-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41523-025-00753-8