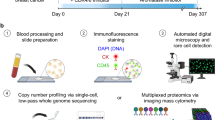
Abstract
The presence of bone marrow (BM) disseminated tumor cells (DTCbm) identifies early-stage breast cancer patients at increased risk of recurrence and poorer overall survival. However, limitations in detecting DTCbm by standard immunohistochemical approaches have hampered clinical application. To address this gap, we developed a flow cytometry-based method, DTC-Flow, that enables the sensitive and efficient detection and molecular characterization of breast cancer DTCbm. Our analysis identified HER2 as a sensitive marker for detecting breast cancer cells, including those lacking HER2 amplification are claudin-low. DTC-Flow using a HER2/EpCAM/CD45 marker panel enabled >90% cancer cell recovery and sensitivity of one cancer cell per million nucleated BM cells across a range of breast cancer subtypes. Molecular analyses of DTC-Flow-sorted DTCbm from metastatic patients suggested a quiescent state and demonstrated their close genomic relationship to primary/metastatic tumors, as well as continued genetic evolution. In early-stage breast cancer patients, DTC-Flow detected DTCbm with greater sensitivity than cytokeratin-based immunohistochemical approaches. Our data support the development of DTC-Flow as a sensitive and specific platform to identify breast cancer patients harboring DTCbm and better understand the biology of minimal residual disease. Ultimately, this platform could enable the selection of personalized therapeutic approaches based on molecular features of DTCbm, monitoring of DTCbm to assess the efficacy of such therapies, and the development of novel therapeutic approaches targeting unique biological vulnerabilities of DTCs in order to eradicate these cells before they can give rise to lethal recurrent cancers.
Similar content being viewed by others
Introduction
Breast cancer is the most frequently diagnosed cancer and the leading cause of cancer mortality among women worldwide1. As a consequence of its high incidence and favorable 5-year survival rates, breast cancer is by far the most prevalent cancer in the world, with ~4 million breast cancer survivors alive today in the United States alone2. Most survivors reside within the surveillance period that follows definitive treatment of their primary tumors. Unfortunately, up to 30% of early-stage breast cancer patients eventually develop recurrent cancers, sometimes 20 years or more after treatment of their primary tumor3,4. As recurrent metastatic breast cancer is typically incurable, this event is responsible for the majority of mortality from this disease5,6.
Current guidelines recommend only breast imaging to detect local recurrence or new cancers, along with symptom-based testing for distant disease in breast cancer survivors7. Additionally, few validated approaches to prevent recurrence are available during the surveillance period beyond adjuvant endocrine therapy to inhibit the estrogen receptor (ER) pathway and adjuvant CDK4/6 inhibition. Consequently, most patients live with the uncertainty of recurrence and the anxiety and reduced quality of life that accompany it8,9.
Recurrent metastatic tumors arise from disseminated residual tumor cells (DTCs) that survive therapy and persist in their host as minimal residual disease (MRD). These cells escape from their primary tumor, seed secondary organs, and can persist in their hosts for years before giving rise to recurrent metastatic disease. Consistent with this, the presence of bone marrow DTCs (DTCbm), which occurs in at least 30% of early-stage breast cancer patients, is strongly associated with increased risk of recurrence and reduced breast cancer-specific (BCSS) and overall survival (OS), whether detected before or after adjuvant therapy10,11,12,13,14,15,16,17,18,19,20.
Since eradicating MRD in early-stage breast cancer patients would, by definition, eliminate recurrent breast cancer and its associated mortality, identifying and targeting unique vulnerabilities of these cells represents a critical therapeutic goal. Indeed, some evidence suggests improved recurrence-free survival (RFS) in patients who convert from a DTCbm-positive to DTCbm-negative state after additional chemotherapy20. This finding supports the important possibility that DTCs may not only identify breast cancer patients at increased risk of recurrence, but may also constitute a therapeutic target as well as a pharmacodynamic marker to monitor the efficacy of treatments that target MRD.
It is notable that while routine assessment of MRD has long been incorporated into standard clinical practice for a number of hematological malignancies21, MRD testing is not currently recommended for breast cancer patients in the surveillance period22. Despite this, the identification in recent years of tractable approaches to prevent breast cancer recurrence by targeting unique biological properties of MRD in preclinical models23,24,25,26,27,28—and, more recently, in early-stage patients29—highlights the importance of developing improved methods for detecting and characterizing breast cancer DTCs.
At present, remarkably little is known about the biology of MRD in breast cancer patients, which represents a barrier to the rational design of DTC-targeted therapies. For example, the unusually long latency for tumor recurrence in some patients, which may span decades, has led to the hypothesis that DTCs may exist in a dormant state. This hypothesis is bolstered by preclinical evidence23,27,30,31,32,33 as well as inferences from mathematical models that constant proliferation in residual tumor cells is difficult to reconcile with the long latencies and stochastic kinetics with which breast cancers recur34,35. Nevertheless, direct evidence in patients for dormancy or the molecular characteristics of DTCs is extremely limited36,37,38,39, principally due to the challenges of identifying and isolating these ultra-rare cells40.
The most commonly used research method for detecting DTCbm is based on the immunohistochemical (IHC) detection of epithelial cytokeratins (CKs) in cytospin preparations of BM aspirates (BMAs) subjected to density gradient centrifugation41. While DTC-IHC has been used successfully in the research setting to assess risks of recurrence and mortality across populations of breast cancer survivors, this method has several important limitations that have impeded its widespread clinical implementation. Among these are the limited volume of BMA that can feasibly be screened, given the requirement for manual slide evaluation by specially trained pathologists, its poor sensitivity, and its reliance on the expression of epithelial markers that may preclude the detection of DTCs that have undergone an epithelial-to-mesenchymal-like transition (EMT)42,43,44. In addition, the multistep nature of DTC-IHC protocols contributes to low and variable rates of recovery of both BM mononuclear cells45 and rare spiked tumor cells46,47. A further drawback of DTC-IHC beyond its low sensitivity is its limited compatibility with downstream molecular analysis, which will inevitably be required for identifying the mechanisms that underlie DTC survival and recurrence so that these may be leveraged to develop ‘personalized’ therapeutic approaches.
To address these limitations, we now describe the development and application of a flow cytometry-based method, DTC-Flow, for the sensitive and efficient detection, isolation, and characterization of DTCbm in breast cancer patients. This method utilizes a “stain-lyse-no wash” workflow to minimize sample handling and cell loss, enabled by the prototype BD FACSFocus™ Pre-Enrichment System (hereafter “FACSFocus”)48. Compared to DTC-IHC, analysis of BMAs spiked with breast cancer cells by DTC-Flow demonstrated lower rates of cell loss and increased sensitivity, detection of a broader range of breast cancer subtypes, including those with mesenchymal phenotypes, and compatibility with downstream molecular analyses. As proof-of-concept, we first utilized DTC-Flow to isolate and characterize DTCbm from patients with metastatic breast cancer. Downstream molecular analyses of sorted DTCs for protein expression, gene expression, exomic mutations, and copy number demonstrated both the specificity of the DTC-Flow assay and its utility for investigating DTC biology. Second, application of DTC-Flow to early-stage breast cancer patients identified a higher percentage of patients harboring BM MRD and detected a larger number of DTCbm in DTC-positive patients compared to DTC-IHC.
Results
Combined HER2 and EpCAM staining identifies human breast cancer cells across subtypes
We sought to develop a multicolor flow cytometry/fluorescence-activated cell sorting (FACS) method for detecting DTCbm in breast cancer patients that would employ cell surface markers to identify human breast cancer cells representing each of the major breast cancer subtypes, while avoiding fixation and permeabilization steps that could impede the subsequent genomic analysis of identified tumor cells. The cell surface protein EpCAM (epithelial cell adhesion molecule, CD326) has frequently been used as a biomarker for the detection of circulating tumor cells (CTCs) in peripheral blood samples, and serves as the molecular target for capture in the FDA-approved CellSearch® assay for identifying CTCs in cancer patients49. As assessed by flow cytometry, antibodies against EpCAM efficiently labeled both luminal and basal human breast cancer cell lines (Fig. 1a, b). In contrast, EpCAM expression was extremely low or absent in breast cancer cell lines of the claudin-low subtype, which is a molecular subtype of triple-negative breast cancer characterized by decreased expression of epithelial markers and increased expression of mesenchymal markers characteristic of EMT42. This suggests that DTC detection approaches based solely on EpCAM are unlikely to be adequate for detecting claudin-low and/or post-EMT DTCs, consistent with previous reports50,51.
Flow cytometric analysis of EpCAM (a, b) and HER2 (c, d) expression in human breast cancer cell lines representative of the major breast cancer subtypes. Histogram plots display cells stained with APC anti-EpCAM (a) or APC anti-HER2 (c) antibodies in red and unstained control cells in blue, along with the percentage of EpCAM-positive or HER2-positive cells for each cell line. The numbers at the top right of each histogram plot are the fold-change in median fluorescence intensity for stained cells compared with unstained cells for each cell line. Bar graphs display cell surface expression levels (median fluorescence intensity) for EpCAM (b) and HER2 (d) in an exemplar experiment for a panel of cell lines representing luminal, basal, and claudin-low breast cancer subtypes. Hatched bars in (d) indicate cell lines with known HER2 amplification and/or overexpression (HER2+). (BMC = CD45– bone marrow cells). (e) Summary data depicting the percentage of cells within each tested human breast cancer cell line that express detectable cell surface HER2 (white), EpCAM (gray), or both markers (black). Data shown are mean percentages across an average of four independent experiments per cell line.
We next evaluated the extent to which breast cancer cell lines could be identified by expression of HER2 (ERBB2), which is amplified and overexpressed in approximately 15% of human primary breast cancers and is associated with poor prognosis52. As expected, staining for HER2 efficiently labeled multiple cell lines with HER2 amplification and overexpression (e.g., MDA-MB-453, BT-474, HCC1954), but not CD45– bone marrow cells (BMCs) from healthy donors (Fig. 1c, d). Notably, endogenous levels of HER2 expression in breast cancer cell lines lacking HER2 amplification were readily detectable by flow cytometry in ~94–100% of cells, including in claudin-low cell lines. This finding is consistent with the expression of HER2 in normal mammary epithelium53, as well as the substantial proportion of “HER2-negative” breast cancers that express low, but detectable, levels of HER2 (HER2 IHC score of 1+ or 2+)54.
Imaging flow cytometry (ImageStream®) confirmed the detectable expression of HER2 in HER2-unamplified breast cancer cell lines, including each of the claudin-low lines tested, as well as high levels of HER2 expression in HER2-amplified/overexpressing cell lines (HCC1954, MDA-MB-453) and absent HER2 expression in the T lymphoblast cell line, Jurkat (Supplementary Fig. 1A). ImageStream analysis also confirmed the absence of expression of both CKs and EpCAM in most claudin-low cell lines.
In combination, staining for EpCAM and HER2 was sufficient to identify >90% of cells within each of the 14 human breast cancer cell lines evaluated (Fig. 1e), including the claudin-low cell lines Hs 578T and BT-549 that do not express CKs at levels detectable by IHC (Supplementary Fig. 1B).
Formulation of the DTC-Flow assay to detect spiked tumor cells in BMAs
A major obstacle to detecting DTCs in BMAs has been the extremely low frequency of these cells. This has led to the use of pre-enrichment methods that, in turn, are invariably associated with some degree of cell loss. To address this limitation, we employed a novel in-line pre-enrichment device, the BD FACSFocus™, in combination with the BD Influx™ cell sorter to develop a streamlined workflow coupling magnetic depletion of CD45+ hematopoietic cells from BMAs with acoustic washing and focusing for removal of unbound antibodies and debris resulting from red blood cell lysis. This platform enables a “stain-lyse-no wash” protocol, thereby permitting enrichment of DTCs while minimizing hands-on sample processing steps and their accompanying cell loss. Performance characteristics of the FACSFocus pre-enrichment device have previously been described48,55.
As an initial proof-of-principle for the FACS-based identification of breast cancer DTCbm (DTC-Flow), we employed a staining panel consisting of HER2 and EpCAM for DTC identification, CD45 for magnetic depletion and exclusion of normal BM hematopoietic cells, and DAPI for assessment of cell viability (Fig. 2a). Viable DTCs are identified as CD45–, DAPI– cells expressing HER2 and/or EpCAM (Fig. 2b). Gating cutoffs for HER2 and EpCAM levels in putative DTCs were established based on the evaluation of control BMAs from healthy donors (Fig. 2c, shaded area represents “background” staining found in control BMAs). Gating cutoffs were set above the highest fluorescence levels observed in eight control BMAs on a HER2 vs. EpCAM biaxial flow plot in order to minimize false-positive events in control samples. Notably, assessment of background levels of HER2 and EpCAM expression in healthy donor control BMAs reproducibly revealed the presence of an “EpCAM-low” population of CD45– BMCs (Fig. 2c). This finding further highlights the limitations of relying solely on EpCAM staining for DTCbm detection, and the importance of excluding such potential false-positive cells from analysis.
a Schematic depicting the DTC-Flow assay “stain/lyse/no wash” sample processing protocol. b Overall DTC-Flow gating strategy. The third dot plot shows the establishment of DTC gating cutoffs for HER2 and EpCAM. The polygonal gate shown in blue reflects the DTC gating cutoffs, defined by background staining levels in healthy donor control bone marrow aspirates (BMAs) (blue shaded area, “non-DTCs”), that were used to classify each bone marrow cell (BMC) as a DTC or not a DTC. The quadrant gates shown in black, defined using control BMAs in which HER2 and EpCAM antibodies were omitted from the staining protocol, were used to define cells as positive or negative for EpCAM and HER2 staining. c Dot plots showing background levels of HER2 and EpCAM expression (blue shaded area) in CD45– BMCs from representative control donors. Arrows indicate the “EpCAM-low” normal BMC population present in most control BMAs. d Dot plots showing HER2 and EpCAM expression levels in representative human breast cancer cell lines, relative to the gating cutoff established for CD45– cells from control BMAs. Percentages of cells from each cell line with HER2 and/or EpCAM levels above the control bone marrow gating cutoff are shown.
Evaluating HER2 and EpCAM levels in human breast cancer cell lines relative to background staining in normal BMCs enabled the identification of cell lines representing multiple breast cancer subtypes (Fig. 2d). Claudin-low breast cancer cells were detected by DTC-Flow as HER2-low EpCAM-low, although not all cells expressed HER2 at levels sufficient to permit detection above background staining of CD45– control BMCs (Fig. 2d).
DTC-Flow sensitively detects ultra-rare spiked tumor cells in BMAs
To test the performance of the DTC-Flow assay, we evaluated control BMAs (0.2 mL whole BMA per sample) spiked with human breast cancer cell lines at tumor cell frequencies ranging from 1:4000 to 1:4,000,000. At a 1:10,000 spiking ratio, DTC-Flow detected >90% of spiked tumor cells (mean recovery = 93.7%, n = 4; Fig. 3a), with <0.5% loss of spiked tumor cells and BM granulocytes and lymphocytes to the FACSFocus waste chamber during the acoustic washing process (Supplementary Fig. 2A, B).
a Assessment of the percentage of spiked tumor cells successfully recovered using DTC-Flow assay, as compared to samples without pre-enrichment. Average recovery of spiked tumor cells was 93.7 ± 6.1% (mean ± standard deviation, n = 4). b Assessment of spiked tumor cell recovery at 3-log range of spiking levels, with linear regression analysis. c Flow cytometry dot plots demonstrating detection of spiked human breast cancer cells (MDA-MB-453, orange boxed area) in control bone marrow at spiking ratios of 1 tumor cell per 100,000 nucleated bone marrow cells (BMCs) (left panel) or per 1 million BMCs (right panel). d Assessment of DTC-Flow assay performance at low spiking ratios. Spiking ratios shown are tumor cells per given number of nucleated BMCs. e Comparison of spiked tumor cell recovery (1:10,000 spiking ratio) for the DTC-Flow assay as compared to the current standard DTC detection method, consisting of Lymphoprep™-based mononuclear cell isolation followed by IHC detection of cytokeratin expression (CK-IHC). (*p = 0.0221; paired two-tailed t-test, n = 3).
Assessment of spiked tumor cell recovery across a 3-log range of spiking frequencies demonstrated a linear relationship between the number of tumor cells spiked and the number detected by DTC-Flow, with a slope of 0.89 (Fig. 3b). DTC-Flow reproducibly and efficiently detected tumor cells spiked at frequencies as low as one tumor cell per million nucleated BMCs (Fig. 3c, d). We observed a 100% success rate in detecting spiked “DTCs” in five different samples, each containing four tumor cells spiked into four million nucleated BMCs (number of detected spiked tumor cells ranged from one to four, as indicated in Fig. 3d). While DTC-Flow was able to detect spiked tumor cells at frequencies as low as 1:4,000,000, the success rate for spiked cell detection dropped to an average of 60% for spiking ratios below 1:1,000,000 (Fig. 3d), though we were able to detect a single-spiked tumor cell in three independent experiments. Notably, the percentage of recovered tumor cells spiked at a ratio of 1:10,000 was ~2-fold greater for DTC-Flow compared to DTC-IHC performed in parallel on paired samples (Fig. 3e). This suggests greater sensitivity for DTC-Flow compared to DTC-IHC.
To confirm the accuracy of spiked tumor cell identification by DTC-Flow and demonstrate its compatibility with downstream molecular analysis, we performed single-cell gene expression analyses following index sorting of three spiked human breast cancer cell lines, in addition to the EpCAM-low population of normal CD45– BMCs (Supplementary Fig. 3A). Control BMAs were spiked with T47D, HCC1954, and MDA-MB-231 cells and subjected to DTC-Flow followed by single-cell RNA-seq (scRNA-seq). Cells from each of the three spiked breast cancer cell lines mapped to distinct clusters on a t-SNE plot generated from gene expression data; moreover, all spiked tumor cells clustered distinctly from the EpCAM-low CD45– BMC population (Supplementary Fig. 3B). Importantly, comparison of single-cell gene expression data from spiked tumor cells isolated by DTC-Flow to unspiked tumor cells not subjected to DTC-Flow revealed that spiked cells clustered alongside unspiked cells for each cell line (Supplementary Fig. 3C). This indicates that single-cell gene expression is not substantially altered by DTC-Flow sample processing.
As anticipated, spiked tumor cells isolated by DTC-Flow expressed CKs and other known breast cancer cell markers (Supplementary Fig. 3D). In contrast, EpCAM-low BMCs lacked expression of epithelial cell markers, including HER2, and instead expressed high levels of erythroid markers (HBB, HBA1, and HBA2). Further, each of the three spiked breast cancer cell lines could be distinguished based on their distinct gene expression patterns. For example, the basal/HER2+ cell line HCC1954 was identified based on its expression of the basal CK KRT5 along with high levels of HER2 expression, whereas claudin-low MDA-MB-231 cells expressed lower levels of CK along with elevated levels of the mesenchymal marker VIM (Supplementary Fig. 3D).
Additionally, the tumor cell identity of sorted, spiked MDA-MB-453 tumor cells detected by DTC-Flow was confirmed by single-cell whole genome amplification, followed by mutation-specific qPCR. This demonstrated the presence of the PIK3CA c.3140A>G mutation in single-spiked MDA-MB-453 tumor cells recovered by DTC-Flow, but not in CD45+ BMCs (Supplementary Fig. 3E). Together, these analyses demonstrate the potential for downstream molecular analysis of cells isolated by DTC-Flow, as well as the accuracy with which tumor cells and non-tumorigenic BMCs can be identified using this methodology.
Application of DTC-Flow for detecting, isolating, and characterizing DTCbm in patients with metastatic breast cancer
Having demonstrated the ability of the DTC-Flow assay to sensitively and specifically identify spiked breast cancer cells in BMAs from healthy donor controls, we next applied this method to clinical BMAs from two patients with ER + /PR + /HER2– metastatic breast cancer enrolled in the METAMORPH (“Metastatic Markers of Recurrent Tumor Phenotype”) study56. Patient/tumor characteristics are shown in Supplementary Table 1.
DTC-Flow detected abundant DTCs within BMAs from each patient, ranging from ~200 to 600 DTCs per milliliter of BMA, which corresponded to a frequency of ~1 DTC per 30,000 nucleated BMCs (Fig. 4a). Magnetic depletion of CD45+ BMCs coupled with removal of debris from red blood cell lysis using the FACSFocus instrument enabled marked enrichment of DTCs within assayed BMAs (>1000-fold enrichment relative to total cells; ~4-fold enrichment relative to nucleated cells) (Supplementary Table 2).
a Flow cytometry dot plots showing the level of expression of HER2 and EpCAM in bone marrow (BM) DTCs (DTCbm) (unshaded area, arrows) detected by DTC-Flow in two patients with ER+/PR+ metastatic breast cancer. The table below indicates DTC counts for each patient along with DTC frequencies relative to total nucleated BM cells (BMCs) prior to FACSFocus pre-enrichment. b ImageStream analysis of HER2, EpCAM, and CK expression in sorted DTCbm from patient B compared to CD45 + BM cells and human breast cancer cell lines. Acquisition channels and fluorochromes used are indicated below the images. (BF = brightfield; scale bars = 10 µm). c Heat map depicting targeted qPCR analysis of mRNA expression in sorted DTCbm pools (~10 cells each) from two metastatic breast cancer patients, along with sorted ~10-cell aliquots of representative human breast cancer cell lines. Sorted Jurkat T cells and normal BMC cDNA served as positive controls for BMC markers. Values shown are threshold cycles (Ct) for the indicated genes. d Plot displaying mRNA expression levels of proliferation/cell cycle marker genes in DTCbm versus average expression in four cultured human breast cancer cell lines (HCC1954, T47D, MDA-MB-231, and Hs 578 T) (ND = not detected). Data are presented as mean ± standard deviation of three technical replicates. (*p < 0.05, **p < 0.01, ***p < 0.001 vs. breast cancer cell line average; one-way ANOVA with Bonferroni’s multiple comparisons test).
DTCs identified in both patients were HER2+ EpCAM-low and displayed similar and relatively homogeneous levels of expression of HER2 and EpCAM as assessed by DTC-Flow (Fig. 4a). Sorted DTCs were further subjected to downstream imaging and genomic analyses to confirm tumor cell identity and characterize biological properties. As anticipated, ImageStream analysis confirmed the absence of CD45 expression and presence of a nucleus in sorted DTCbm (Fig. 4b). ImageStream further demonstrated expression of EpCAM, HER2, and CK within each DTC at levels comparable to those within the T47D luminal cell line, and higher than those observed in the claudin-low breast cancer cell lines MDA-MB-231 and Hs 578 T. By DTC-Flow and ImageStream analysis, HER2 expression in DTCs isolated from each patient was lower than in the HER2-amplified cell line, HCC1954, consistent with the non-amplified HER2 status (IHC/FISH) of these patients’ cancers. The presence of detectable HER2 expression in these DTCbm is consistent with our ability to detect endogenous, non-amplified HER2 levels in breast cancer cell lines (Fig. 1c, d and Supplementary Fig. 1A).
Targeted qRT-PCR analysis confirmed the expression of multiple luminal CKs within DTCs isolated from each patient (Fig. 4c), as well as expression of ESR1 (estrogen receptor-α), PGR (progesterone receptor), and ERBB2 (HER2) consistent with the ER/PR/HER2 status of the corresponding primary and metastatic tumors (Supplementary Table 1). Importantly, DTCs isolated by DTC-Flow did not express detectable levels of the BM hematopoietic cell markers CD45 (also known as PTPRC), CD43 (also known as SPN), or HBD, further supporting their tumor cell identity and the purity of sorted DTCs.
ImageStream analysis of DTCbm isolated from metastatic patients also allowed us to evaluate their size (Supplementary Fig. 4A). Mean DTCbm diameters were similar for the two patients analyzed (16.9 µm, 17.2 µm) and were significantly larger than the mean diameters of BM granulocytes (11.7 µm), monocytes (11.3 µm), or lymphocytes (10.1 µm; p < 0.001), although there was some overlap in the size distributions for DTCbm and BMCs. Notably, DTCs were significantly smaller in diameter than the cultured breast cancer cell line T47D (mean = 20.6 µm; p < 0.001), possibly reflecting differences in cell size that might be anticipated for actively proliferating cells in culture versus cells in a potentially non-proliferative state in BM (Supplementary Fig. 4A).
ImageStream analysis also demonstrated the presence of DTC clusters, most frequently consisting of 2–3 DTCs, which were identified at frequencies of 11% and 23% in the two patients analyzed and could be detected prior to Influx sorting (Supplementary Fig. 4B–D). This finding implies that the processing methods employed here likely permit the preservation and detection of small clusters of DTCs, which is of interest given the reported association of CTC clusters with increased metastatic potential57.
Gene expression analysis of DTCbm suggests a quiescent state
The compatibility of DTC-Flow with downstream molecular analysis enables its use as a tool to probe the biology of DTCbm in breast cancer patients. As breast cancers, particularly those that are ER+, can recur 20 years or more after treatment of the primary tumor4, it has been hypothesized that DTCs may persist in a dormant state prior to “re-awakening” in some patients to give rise to recurrent, metastatic tumors. Multiple models for the cellular basis of this clinical latency period have been proposed, including cellular dormancy (characterized by reversible DTC quiescence33), as well as tumor mass dormancy (i.e., angiogenic or immune-mediated dormancy), in which ongoing DTC proliferation is balanced by cell death due to immune surveillance or inadequate vascular supply58. Nevertheless, relatively little direct evidence exists regarding the cell cycle status of DTCbm in breast cancer patients.
Despite the presence of actively growing metastatic tumors in each of the two patients evaluated, we found that DTCbm isolated from each patient expressed proliferation-associated genes at low levels characteristic of non-proliferating cells (Fig. 4d). Ki-67 antigen (MKI67), which is expressed in all phases of the cell cycle but not in quiescent cells that have exited the cell cycle59, was not expressed at detectable levels in DTCbm from patient A and in patient B was expressed in DTCbm at levels 10- to 20-fold lower than those observed in actively growing human breast cancer cell lines. Consistent with this, DTCbm from each patient expressed extremely low levels of the cell cycle-regulated genes TK1 (thymidine kinase 1, levels of which peak in S phase of the cell cycle60) and CCNB1 (cyclin B1, expressed predominantly in G2/M61). In contrast to MKI67, TK1, and CCNB1, levels of the G1 marker CCND1 (cyclin D1)62 were only ~2-fold lower in DTCs from each patient compared to actively proliferating breast cancer cell lines. Taken together, these findings suggest that the majority of DTCbm in both patients were quiescent, consistent with limited prior studies of DTCbm in patients with primary breast tumors36,39,63.
Mutation and copy number profiles of DTCbm are closely related to their respective primary and metastatic tumors
To further evaluate the genomic landscape of DTCs, we performed whole-exome sequencing (WES) on DTCbm from the above metastatic patients and their corresponding primary or metastatic tumors, compared to germline DNA. For patient A, who presented with Stage IV synchronous metastatic disease, several mutations were found in both sorted DTCbm and the corresponding metastasis that were not present in germline control DNA from sorted CD45+ BMCs (Fig. 5a). These included a nonsense mutation in CDH1 (p.Q255*; Catalogue of Somatic Mutations in Cancer; genomic mutation ID COSV55727283), encoding E-cadherin, which was estimated to be clonally mutated within DTCs (allele frequency = 1.00). In addition, a well-characterized activating mutation in exon 20 of PIK3CA (p.H1047R) was detected both in DTCbm (allele frequency = 0.73) and the metastatic tumor in this patient, which was confirmed by targeted Sanger sequencing of PIK3CA exon 20 (Fig. 5b). Notably, this PIK3CA mutation was detected at different frequencies relative to the WT allele in separate DTC pools, suggesting potential heterogeneity in the overall DTCbm population. Overall, three of five non-synonymous COSMIC mutations detected in the synchronous metastatic tumor at allele frequencies ≥0.10 were also found in DTCbm from the same patient. Strikingly, a mutation in ESR1 (p.D538G) known to confer resistance to endocrine therapy64 was detected in the metastatic tumor (allele frequency = 0.28; estimated to be ~clonal after adjusting for tumor cellularity), but was not found in DTCbm from this patient. This suggests that the pool of detected DTCbm may have been seeded by the primary tumor rather than the metastatic tumor.
Non-synonymous COSMIC mutations detected by whole-exome sequencing of bone marrow (BM) DTCs (DTCbm) from patient A (~200 cell pool, a) or patient B (~10-cell pool, c), along with the corresponding primary or metastatic tumor. Predicted functional impact and pathogenic status of each mutation are per the COSMIC database. Numbers in parentheses indicate the variant allele frequency of each mutation. Mutations were not detected in germline DNA from sorted CD45 + BM cells or buffy coat cells. Mutations annotated as Pathogenic in COSMIC are in bold. Discordances between DTC and tumor mutation status are in blue. b PIK3CA sequencing demonstrating that DTCbm from patient A exhibit the mutation c.3140A>G, which was also present in the corresponding metastatic tumor. Copy number (CN) plots generated from sWGS data from DTCbm, primary or metastatic tumor FFPE gDNA, or CD45+ normal BM cells from patients A (d) and B (e). Insets show regions of CN gain (red arrows) and CN loss (blue arrows) shared between DTCbm and the corresponding primary or metastatic tumors, but not CD45 + BM cell germline DNA. For patient B, repeat sequencing libraries were prepared from the same WGA products for CD45+ normal BM cells and DTCbm.
Whole-exome sequencing was also performed on the primary tumor and DTCbm isolated from patient B (Fig. 5c), which similarly revealed a high degree of concordance for mutations. Clonal mutations in MYH9 (p.E495K) and TRIM24 (p.E34D) were detected in both DTCbm and the primary tumor, as were two separate missense mutations in PIK3CA, one in exon 9 (p.E545K; DTC allele frequency = 0.33) and one in exon 20 (p.T1025S; DTC allele frequency = 0.68). In addition, two subclonal non-synonymous COSMIC mutations were detected within DTCbm (with DTC allele frequencies of 0.16 and 0.13) that were not detected in the primary tumor, including HSP90AB1 p.R719H and RAC1 p.D63G. Potential explanations for this finding include the possibilities that additional mutations were acquired in some DTCs after their dissemination from the primary tumor, that the sampled population of DTCbm arose from a primary tumor subclone that was relatively rare or was otherwise not sampled within the primary tumor sections analyzed, or that a portion of these DTCbm were seeded from the metastatic tumor that had undergone subsequent genomic evolution.
We additionally performed shallow whole genome sequencing (sWGS) and copy number analysis on pools of DTCbm isolated from these two metastatic breast cancer patients. Analogous to the substantial overlap observed for mutation profiles, we identified a high degree of overlap in copy number alterations between DTCs isolated from each patient and their corresponding primary or metastatic tumors (Fig. 5d, e). For example, comparison of the copy number profile for DTCbm isolated from patient A to that of their metastatic tumor yielded a Pearson correlation coefficient of 0.74, while the correlation coefficient for the comparison of DTCbm isolated from patient B to their primary tumor was 0.48. Further, DTCbm from each patient exhibited specific chromosomal gains and losses (e.g., amplification of 1q, loss of 16q) frequently observed in breast cancer65,66, as did the corresponding primary and metastatic tumors. As anticipated, copy number profiles for DTCs and CD45+ normal BMCs were poorly correlated (Pearson correlation coefficient r = −0.17 for patient A; r = −0.02 for patient B). When taken in combination with the above WES data, these findings support the tumor cell identity of DTCbm detected in these patients by DTC-Flow and demonstrate a high degree of genomic similarity between DTCbm and their corresponding primary or metastatic tumors, thereby confirming their close evolutionary relationship.
Identification of disseminated minimal residual disease in early-stage breast cancer patients by DTC-Flow
Having demonstrated the utility of DTC-Flow for isolating and characterizing DTCs from metastatic breast cancer patients, as well as proof-of-concept for its specificity and increased sensitivity in bone marrow spiking experiments, we next explored the feasibility of DTC-Flow for evaluating BMAs from early-stage breast cancer patients. To accomplish this, we analyzed 42 BMAs obtained through the PENN-SURMOUNT surveillance study (“PENN-Surveillance Markers of Utility for Recurrence After (Neo)Adjuvant Therapy for Breast Cancer”; NCT02732171)67. PENN-SURMOUNT is a single center, prospective study of women with early-stage breast cancer within 5 years of diagnosis who have completed surgery, (neo)adjuvant chemotherapy and adjuvant radiation therapy for their primary breast cancer and are at increased risk for recurrence due to lymph node positivity, triple-negative subtype, ER+ cancer with Oncotype DX recurrence score ≥25, or residual disease after neoadjuvant chemotherapy. Patient characteristics of the 42-patient PENN-SURMOUNT cohort are shown in Supplementary Table 3.
BMAs from this patient cohort were divided and analyzed in parallel using both standard DTC-IHC and DTC-Flow assays. As for metastatic patients, flow cytometry gating cutoffs were established using control BMAs from healthy donors. Of note, the HER2 fluorescence cutoff for DTCs in PENN-SURMOUNT BMAs was increased due to the reproducible presence of a large population of CD45-negative normal BM cells expressing low levels of HER2 that was generally absent in healthy donor control marrows (Supplementary Fig. 5A, B). Further, in anticipation of finding substantially lower DTC frequencies in early-stage compared to metastatic patients, we established DTC count thresholds for each DTC HER2;EpCAM phenotype that would be sufficient to eliminate any false-positive calls in healthy donor control BMAs (thresholds defined as mean false-positive count in control BMAs + 3 standard deviations). These thresholds were: EpCAM + > 0, HER2 + > 1.1, or HER2+ EpCAM + > 2.7 per 0.5 mL bone marrow (Supplementary Fig. 5B, C).
Using these gating cutoffs and thresholds, DTC-Flow identified 27 of 42 patients (64%) as DTC+ (Fig. 6a, b), with total counts for DTC+ patients ranging from 1 to 21 DTCs per 0.5 mL BMA, with a median of 6 DTCs per 0.5 mL BMA in those patients who were DTC+ (Fig. 6c). In contrast, only 12 patients (29%) were identified as DTC+ by DTC-IHC (Fig. 6b). The majority of DTCs identified by DTC-Flow were EpCAM+ or HER2+ EpCAM+ (Fig. 6d).
a Flow cytometry dot plots showing the level of expression of HER2 and EpCAM in bone marrow aspirates (BMAs) analyzed by DTC-Flow in four early-stage PENN-SURMOUNT patients. Putative DTCs (unshaded area) are indicated by arrows. b Comparison of DTC-IHC vs. DTC-Flow results for PENN-SURMOUNT patient BMAs (n = 42). Concordant IHC and Flow results are shown in black, while discordant DTC results are shown in red. DTC-Flow identified 83% (10 of 12) of those BMAs screened as DTC-positive by IHC, while DTC-IHC only identified 37% (10 of 27) of the samples screened as DTC-positive by DTC-Flow. c Box plot showing distribution of DTC counts for DTC-positive patients as identified by either DTC-IHC (n = 10) or DTC-Flow (n = 27). DTC counts for the IHC assay are per 5 million bone marrow mononuclear cells, while Flow DTC counts are per 0.5 mL BMA. (Boxes extend from the 25th to 75th percentiles; center lines = median; whiskers = maximum and minimum values). d DTC counts in BMAs as determined by DTC-Flow (per 0.5 mL BMA). Total DTC counts are shown for PENN-SURMOUNT patient BMAs (blue dots) and healthy donor control BMAs (pink dots) across HER2; EpCAM phenotypes. Overall median counts (including DTC– samples) are indicated by lines.
Notably, 10 of the 12 patients classified as DTC+ by DTC-IHC were also classified as positive by DTC-Flow (Fig. 6b). In addition, DTC-Flow identified 17 patients as DTC+ who were classified as negative by DTC-IHC. Overall, 24% of patients were identified as DTC+ by both assays, 31% were identified as DTC– by both assays, 40% were identified as DTC-Flow+/DTC-IHC–, and only 5% were classified as DTC-IHC+/DTC-Flow–. Discordant cases were significantly enriched with DTC-IHC– DTC-Flow+ patients (McNemar’s test, p = 0.0013), which is consistent with DTC-Flow being a more sensitive assay than DTC-IHC. Beyond identifying a higher percentage of patients as DTC+, and thus potentially at increased risk for breast cancer recurrence, DTC-Flow also identified a greater number of DTCs in those patients classified as DTC+ by both assays (Fig. 6c). These findings are consistent with what would be expected if the sensitivity of DTC-Flow was higher than that for DTC-IHC, as would be anticipated based upon assay performance characteristics determined in tumor cell spiking experiments (Fig. 3).
Discussion
Incurable, recurrent metastatic breast cancers arise from the reservoir of residual tumor cells that survive therapy and persist in their host. Consistent with this, patients harboring BM MRD are at increased risk of recurrence. This, in turn, suggests that eradicating MRD could provide an effective means to prevent breast cancer recurrence and its associated mortality. Achieving this goal, however, requires the development of markedly improved methods for detecting and molecularly characterizing DTCs in breast cancer patients. Such methods would enable the more accurate identification of those at increased risk of recurrence, a better understanding of DTC biology, and the design of rational therapeutic interventions capable of targeting these cells, as well as permit DTC burden to be used as a pharmacodynamic marker for monitoring the efficacy of DTC-targeted therapies.
We have developed a flow cytometry/FACS-based assay for DTCbm, DTC-Flow, which begins to address these needs. Use of a novel pre-enrichment device coupled with in-line cell sorting enabled >90% recovery of human breast cancer cells spiked into BMAs, the sensitive detection of spiked tumor cells at frequencies as low as one tumor cell per million nucleated BMCs, and increased tumor cell recovery compared to IHC-based methods requiring manual density gradient centrifugation. Our studies further demonstrated that DTC-Flow is compatible with downstream molecular analysis at the protein, mRNA, and DNA levels, and each of these methods was successfully employed to confirm the specificity of the DTC-Flow assay and characterize DTCs from patients with metastatic breast cancer. These analyses permitted the unambiguous identification of DTCbm as tumor cells based on their expression of multiple epithelial markers, the absence of expression of BM hematopoietic cell markers, concordant ER/PR/HER2 receptor status, and the presence of multiple mutations and copy number alterations that were shared with their corresponding primary or metastatic tumors. In addition, the presence of clonal or near-clonal mutations in the small pools of DTCs analyzed supported the relative purity of their isolation.
Importantly, DTC-Flow was able to detect DTCbm in non-metastatic, early-stage breast cancer patients, in whom DTC frequencies were markedly lower than in metastatic patients. DTC-Flow also detected a higher number of DTCbm within each positive patient compared to IHC-based detection, suggesting that it may offer enhanced dynamic range for evaluating MRD-targeted therapies in early-stage patients.
As anticipated for an assay with increased sensitivity, patients who were DTC-IHC+ were also typically DTC-Flow+, whereas DTC-Flow frequently identified patients as DTC+ who were negative by the IHC assay but who could potentially be at greater risk of recurrence. While the DTC-Flow positivity rate in early-stage breast cancer patients (64%) is higher than that observed for DTC-IHC (both in the same cohort of PENN-SURMOUNT patients, as well as in the literature11), it is notable that a similar positivity rate (67%) was observed in a cohort of 45 breast cancer patients when DTCs were assessed by CK staining after several weeks of in vitro tissue culture expansion (compared to a 37% DTC positivity rate by standard DTC-IHC analysis of the same patients)68. Taken in combination with our findings, these findings suggest that the proportion of early-stage breast cancer patients who are DTC+ may substantially exceed the ~30% level commonly referenced in the literature11,15. Further single-cell molecular characterization will be needed to confirm the tumor cell identity of DTCbm detected by DTC-Flow in the non-metastatic setting.
Several prior studies have evaluated flow cytometry-based methods for detecting DTCbm in breast cancer patients69,70,71 using assays that relied upon expression of epithelial markers (EpCAM, CKs) in DTCbm, which our findings suggest may constitute a potential limitation of these methods. Consistent with previous reports50,51, we found that human breast cancer cell lines of the claudin-low subtype typically cannot be detected using these markers. The inability to detect breast cancer cells with mesenchymal phenotypes, including those that have undergone an EMT, has important clinical implications since EMT has been suggested to play a role in metastasis72,73, the acquisition of therapy resistance, and stem cell-like properties44,74,75. Indeed, we have previously reported EMT to be a feature of mammary tumor dormancy and recurrence in some genetically engineered mouse models33,76. Consistent with the possibility that EMT-like transitions may also occur in patients, we found that DTCbm isolated from metastatic breast cancer patients displayed lower levels of classical epithelial markers such as EpCAM and CKs than human breast cancer cell lines, a finding consistent with a prior report of EMT in cell lines derived from DTCbm in breast cancer patients43.
Notably, the DTCs that we detected in metastatic breast cancer patients expressed EpCAM levels comparable to those found within an “EpCAM-low” population of normal CD45– BMCs that we reproducibly detected in healthy donor control BMAs. As additional evidence that the EpCAM-low BMC population identified in healthy donors does not represent DTCs, single-cell RNA-sequencing of sorted EpCAM-low BMCs revealed expression of erythroid-specific genes. Indeed, early erythroid progenitor cells in the BM have been reported to express low levels of epithelial markers such as EpCAM77,78. This finding likely precludes the accurate identification of DTCbm by single epithelial markers such as EpCAM70,79, and further emphasizes the value of HER2 as an epithelial marker for DTCbm.
Consistent with the expression of HER2 in normal mammary epithelium and its critical role in mammary gland morphogenesis in mice53,80,81, we found that the addition of HER2 as a cell surface marker allowed us to differentiate between DTCbm and the above non-tumorigenic EpCAM-low BMC population. While HER2 is perhaps best known for its amplification in ~15% of human breast cancers in which it drives aggressive tumor behavior52, we found that virtually all human breast cancer cell lines assayed, as well as DTCbm from metastatic breast cancer patients, expressed detectable levels of HER2, albeit at levels far lower than those observed in HER2-amplified breast cancer cells. This finding is noteworthy in light of increasing recognition of the clinical importance of low levels of HER2 expression, including the potential for targeting HER2-low breast cancers using anti-HER2 antibody-drug conjugates82. Accordingly, we found that the combined use of HER2 and EpCAM as markers was sufficient to identify all human breast cancer cell lines analyzed, including several claudin-low cell lines lacking expression of epithelial CKs. As the ability to distinguish claudin-low breast cancer cells from normal BMCs can likely be further improved, an additional advantage of DTC-Flow is that this method can readily accommodate additional markers to either improve the specificity of DTC identification by excluding rare CD45– non-tumorigenic BMC populations, or improve its sensitivity by enabling the detection of DTCs with mesenchymal phenotypes.
An additional important possibility is that the non-hematopoietic/stromal composition of BM may differ in healthy control donors compared to cancer patients, as has been suggested83,84. These differences may necessitate the use of additional negative selection markers to exclude potentially confounding BMC populations and improve the specificity of DTCbm detection – for instance, by excluding the population of CD45– BMCs expressing low levels of HER2 that we detected in most early-stage breast cancer patients. It will also be necessary to ensure that the incorporation of additional markers for BMC exclusion or depletion does not decrease sensitivity for DTC detection, i.e., by excluding or removing DTCs that exist in clusters with BMCs.
Another important feature of DTC-Flow is that it is scalable, thereby enabling the analysis of greater numbers of BMCs as a means to further increase the sensitivity of detecting patients who are DTC+, as well as estimating the absolute number of DTCbm present. Indeed, for the breast cancer patients evaluated in this study, we analyzed an average of 25 million nucleated BMCs, which represents a ~10-fold increase over the standard DTC-IHC method that typically involves screening of a restricted number of nucleated BMCs (often only two million)41, thereby limiting the detection of ultra-rare DTCs. Consistent with this, prior studies have shown that increasing the number of BM mononuclear cells screened by IHC results in an increased fraction of patients who are identified as DTC+85,86. Of note, while screening an increased number of BMCs by pan-CK DTC-IHC increases the percentage of early-stage breast cancer patients with detectable DTCs, it may also reduce the correlation between DTC positivity and recurrence risk86. This suggests either a dose-response relationship between DTCbm abundance and risk of recurrence, or the potential existence of a threshold DTC frequency/burden associated with increased recurrence risk. Whether this limitation also applies to the DTC-Flow assay remains to be determined in future studies, including long-term follow-up of DTC-Flow+ vs. DTC-Flow– early-stage breast cancer patients, as does whether predictive value could be optimized based on criteria including number of BMCs analyzed, thresholds for DTC counts, and/or biological properties of detected DTCs.
The ability to collect pure populations of DTCs, including single cells, for a variety of downstream molecular analyses, represents yet another important advantage of DTC-Flow over DTC-IHC. Indeed, beyond simply enumerating DTCs, molecular analysis of these cells is essential for answering critical questions regarding MRD biology, including elucidating the properties characteristic of the dormant state or EMT, as well as the pathways that sustain DTC survival and persistence. In this regard, it has been reported that expression of the orphan nuclear receptor NR2F1 by breast cancer DTCbm may be an important determinant of the dormancy phenotype and that the proportion of DTCs expressing this marker may correlate with risk of relapse36,87. In addition, genomic analysis of DTCs in patients has the potential to identify mutations or copy number alterations in key oncogenes and tumor suppressor genes that may, in turn, hold value for predicting the natural history of disease and its likely response to therapy. Thus, defining the molecular phenotype of DTCs in individual patients may enable the informed selection of personalized therapies based on the unique characteristics of their disease.
While there are currently few therapies proven to specifically target DTCs in breast cancer patients harboring MRD, it has been suggested that these cells may be resistant to traditional forms of chemotherapy88,89 as a consequence of their hypothesized quiescent state and/or stem cell-like phenotypes90,91. Indeed, we found that DTCbm from patients with metastatic breast cancer expressed low levels of proliferation markers, including MKI67, TK1, CCND1, and CCNB1, consistent with prior reports that DTCs in early-stage breast cancer patients are largely Ki-67-negative39,63. Beyond demonstrating that tumor cells detected by DTC-Flow were unlikely to represent cells from actively proliferating metastatic lesions in bone, our data are consistent with a model in which DTCbm may largely be quiescent, even in patients with frank metastatic growth. This, in turn, supports a model of cellular dormancy as opposed to tumor mass dormancy (i.e., angiogenic or immune-mediated) in the BM58. Quiescence in DTCbm may also affect the utility of CTCs and/or ctDNA for detecting dormant MRD in breast cancer patients, since the short half-lives for each of these markers in the circulation (1–2.4 h for CTCs and <1 h for ptDNA92,93) necessitate their constant replenishment, which may not occur in the absence of proliferating tumor cells.
Since DTCs give rise to metastatic recurrences, genomic characterization of these cells has the potential not only to identify targetable mutations and pathways but also to elucidate the biological processes by which cells disseminate from the primary tumor and eventually give rise to metastases. In this regard, we observed substantial overlap in mutation and copy number profiles between DTCs and their corresponding primary or metastatic tumors for each of the breast cancer patients evaluated in this study. Strikingly, all mutations detected in the primary tumor in patient B were also detected in the corresponding DTCbm isolated from a BMA performed nearly three years after initial diagnosis. Additional subclonal mutations were detected in DTCs from this patient that were not identified in the primary tumor, suggesting the continued genomic evolution of some DTCs post-dissemination.
For patient A, all mutations detected in isolated DTCbm were also found in the metastasis; conversely, three of five COSMIC mutations in the metastatic tumor were detected in DTCbm, whereas a clonal ESR1 mutation (p.D538G) known to confer endocrine therapy resistance64 was detected in the metastasis, but not DTCbm. Notably, patient A initially presented with synchronous metastasis and was treated with the aromatase inhibitor letrozole for approximately three years prior to metastatic biopsy and DTCbm assessment. Thus, the presence of a clonal ESR1 mutation in the metastasis, but not the isolated DTCbm, following three years of anti-estrogen therapy, suggests that the DTCbm isolated from this patient may have been seeded from the primary tumor. Moreover, the apparent acquisition of this ESR1 mutation in metastatic cells but not DTCbm over the three-year course of treatment, despite selective pressure in both tumor cell populations, is intriguing insofar as it may provide some additional evidence that DTCbm may exist in a long-term quiescent state, since such a state would not formally be compatible with the acquisition and expansion of new mutations.
Several models of breast cancer metastatic progression have been proposed based on preclinical cancer models or genomic analyses of human primary and metastatic tumors. While the linear progression model postulates that acquisition of metastatic potential occurs late in the process of tumor development, the parallel progression model proposes that tumor cell dissemination occurs early in tumorigenesis94. The relative importance of linear versus parallel progression models of metastasis, or their combination95, has important clinical implications both for the use of primary tumor characteristics as the basis for selection of therapies targeting disseminated disease, and for the use of primary tumor mutation profiles for detection of ctDNA in liquid biopsies for MRD assessment. In this regard, our detection of additional subclonal mutations in DTCbm that were not present in the antecedent primary tumor is consistent with recent findings suggesting that substantial and biologically impactful genomic evolution occurs in human breast cancer cells following their dissemination from the primary tumor96. Nevertheless, the high degree of concordance in mutations that we found, along with the substantial degree of overlap in copy number alterations, is most consistent with a linear progression model of breast cancer metastasis, although longitudinal assessment and evaluation of substantially more patients will be needed to resolve this question. Clonality of these DTCs may also have been influenced by the prior therapies received by each patient (patient A had received treatments including endocrine therapy and a CDK4/6 inhibitor, while patient B also received endocrine therapy, along with earlier chemotherapy treatment), as treatment has previously been associated with a higher degree of clonality in metastases97. Longitudinal assessment would also be needed to elucidate therapy-induced clonal evolution of the BM DTC population.
In summary, we have demonstrated the ability of DTC-Flow to sensitively detect breast cancer DTCbm and isolate these cells to high purity for molecular characterization of their biological properties. Application of this method to a large number of breast cancer patients in the MRD setting, in combination with long-term follow-up for cancer recurrence, will be required to assess its prognostic significance. Nevertheless, our initial studies support the utility of DTC-Flow for detecting breast cancer MRD and demonstrate its potential for enabling improved understanding of DTC biology and the development of DTC-targeting therapeutic options for breast cancer survivors at risk of recurrence.
Methods
Cell lines
Human breast cancer cell lines were obtained from the ATCC Breast Cancer Cell Panel, with the exception of SUM149 human breast cancer cells, which were obtained from Stephen Ethier (Medical University of South Carolina). Jurkat human T cell line was kindly provided by Richard Schretzenmair (Penn Cytomics and Cell Sorting Resource Laboratory).
Antibodies/Reagents
APC, PerCP-Cy5.5, and FITC anti-human EpCAM (clone EBA-1), APC and PE anti-human HER2/neu (clone Neu 24.7), APC anti-human CD45 (clone 2D1), and PE and PE-Cy7 anti-human CD45 (clone HI30) antibodies, as well as APC IMag™ magnetic particles, were purchased from BD Biosciences (San Jose, CA, USA). DAPI was from MilliporeSigma (St. Louis, MO, USA). eFluor® 660 anti-human pan-CK (AE1/AE3) antibody was from Thermo Fisher Scientific (eBioscience™; Waltham, MA, USA). Unconjugated anti-pan-CK (AE1/AE3) for IHC and the corresponding isotype control antibody (mouse monoclonal IgG1; cat. # NCL-L-AE1/AE3) were from Leica Biosystems (Newcastle, UK). Red blood cell (RBC) lysis buffer was purchased from G-Biosciences (St. Louis, MO, USA).
Bone marrow aspirates and breast cancer patient cohort
Control BMAs from healthy donors were purchased from AllCells (Alameda, CA, USA) and from the University of Pennsylvania Stem Cell and Xenograft Core (Philadelphia, PA, USA; RRID:SCR_010035). BMAs were acquired from patients with metastatic breast cancer who provided informed consent as part of the METAMORPH observational study (Metastatic Markers of Recurrent Tumor Phenotype for Breast Cancer; NCT02530008; study registration 8/20/2015) conducted at the Rena Rowan Breast Center at the University of Pennsylvania and from patients with early-stage, non-metastatic breast cancer enrolled in the PENN-SURMOUNT surveillance study (“PENN-Surveillance Markers of Utility for Recurrence After (Neo)Adjuvant Therapy for Breast Cancer”; NCT02732171; study registration 4/8/2016). Study protocols and procedures were reviewed and approved by the appropriate Institutional Review Board (METAMORPH = University of Pennsylvania IRB 1 #818874, University of Pennsylvania Cancer Center #30113; PENN-SURMOUNT = IRB 5 #824098, University of Pennsylvania Cancer Center #28115), and the studies were performed in accordance with the Helsinki Declaration of 1975. Approximately 10 mL of BMA fluid was obtained from the posterior iliac crest using a Jamshidi needle, stored at room temperature (RT) in EDTA-containing tubes, and analyzed by DTC-Flow within 24 h of isolation.
Processing for analytical flow cytometry
For analysis of cell surface expression of EpCAM or HER2, the indicated human breast cancer cell lines were harvested by trypsinization, washed, and resuspended in FACS Buffer (PBS with 1% BSA, 1 mM EDTA, and 25 mM HEPES), prior to staining with the indicated antibodies according to the manufacturer’s protocol. For analysis of normal BMCs, RBC lysis buffer was incubated with BMAs at RT, and BMCs were washed with FACS Buffer prior to staining with APC anti-HER2 or APC anti-EpCAM antibodies in combination with PE anti-CD45 and DAPI (5 µg/mL). Data acquisition was performed using an Attune NxT flow cytometer (Thermo Fisher Scientific), with data analysis performed using FlowJo software (BD). Median APC fluorescence intensities were determined after gating for viable (DAPI–), CD45–, non-debris singlet events, while the percentage of either EpCAM+ or HER2+ cells was determined using gates based on unstained control cells.
DTC-Flow sample processing, acquisition, and sorting
BMAs filtered through a 70 µm cell strainer were incubated with APC anti-CD45 antibody at RT, followed by PE anti-HER2 and FITC anti-EpCAM antibodies and APC IMag™ particles. Following staining and RBC lysis (15 min at RT), DAPI was added to each sample immediately prior to flow analysis and cell sorting. For spiking experiments, a known number of unstained cells from the indicated human breast cancer cell line were sorted directly into 0.2 mL whole BMA from a healthy control donor prior to sample processing. Tumor cells were spiked using the BD Influx cell sorter (sorted viable singlets, using 1.0 drop single-cell sort mode). For experiments in which a small number of tumor cells (10 cells or fewer) were spiked, the spiked cells were sorted into the bone marrow sample one at a time. A target sort count of 1 was set in the Influx Sortware, and successful sorting was confirmed visually by observing a sort “flash” at the appropriate deflection site in the Influx sort stream monitor window. For experiments in which larger numbers of tumor cells were spiked, we corrected for differences in sort efficiency by comparison to an equivalently spiked baseline sample that was not subjected to DTC-Flow; the number of tumor cells detected in this baseline sample was used as the true number of spiked cells successfully sorted into bone marrow.
Stained, lysed BMA samples were analyzed and sorted using a BD Influx cell sorter equipped with an in-line pre-enrichment device (BD FACSFocus™), specific details of which have been described48,55. Briefly, the BMA sample first flows through a magnetic column to enable removal of APC anti-CD45-labeled hematopoietic cells, followed by passage through a microfluidic chip in which acoustic focusing/washing enables separation and removal of cellular debris and unbound antibodies, prior to entering the BD Influx for analysis and cell sorting. Putative DTCs were gated in BD FACS™ Sortware as live (DAPI–), CD45– events with levels of HER2 and/or EpCAM greater than the staining background observed in healthy control BMAs. To ensure consistency of gating across PENN-SURMOUNT samples, target median fluorescence intensity (MFI) values were established for each scatter and fluorescence channel using SPHERO™ Supra Rainbow Midrange Fluorescent Particles (Spherotech; Lake Forest, IL, USA). Each day, PMT voltage settings were adjusted in order to match these target MFI values, enabling the use of the same gates in Sortware across samples.
Cytospin preparation and pan-cytokeratin IHC
Isolation of mononuclear cells (MNCs) from BMAs, including spiked tumor cells, was performed by density gradient centrifugation using Lymphoprep™ (STEMCELL Technologies; Vancouver, BC, Canada), following the manufacturer’s protocol. BM MNCs resuspended in PBS were used to prepare cytospins using a Shandon cytospin centrifuge (Thermo Fisher Scientific), followed by fixation in 95% ethanol. IHC staining for pan-CK (AE1/AE3) or the corresponding isotype control antibody was performed at the Pathology Clinical Service Center-Anatomic Division at the University of Pennsylvania, using a Leica Bond III autostainer (Leica Biosystems). The MCF7 cell line was used as a positive control, and a negative mouse isotype control was included in each staining run. For assessment of tumor cell recovery in spiking experiments, identification of CK+ tumor cells was performed by a trained investigator (NNCS) without knowledge of the DTC-Flow results, following standard morphological criteria for DTC identification41. For analysis of PENN-SURMOUNT BMAs, five stained cytospin slides (106 BM MNCs each) were evaluated for each patient by a trained pathologist (MF) blinded to patient identifiers.
Transcriptional analysis of DTCs
Targeted qPCR Panel: For targeted gene expression analysis, small pools (~10 cells) of DTCs or human cell line controls (HCC1954, T47D, MDA-MB-231, Hs 578T, and Jurkat) were sorted directly into chilled PCR tubes (MicroAmp, Thermo Fisher Scientific) containing 10 µL of Single Cell Lysis mix (including DNase I) from the Single Cell-to-CT™ kit (Thermo Fisher Scientific). Cell pool sizes are estimated; sort efficiency varied between ~60% and over 90%, as assessed by a colorimetric plate-sorting assay98. Cell lysis was performed as directed, and samples were stored at −80 °C. Reverse transcription was performed per the standard kit protocol.
Pre-amplification was performed using a mixture of up to 87 pooled TaqMan® Gene Expression assays (listed in Supplementary Table 4). RNA was isolated from human cell lines and from mononuclear cells from healthy control BMAs using a Qiagen RNeasy Mini kit (Germantown, MD, USA), and 1 µg RNA was used as input for cDNA synthesis using the High-Capacity cDNA Reverse Transcription kit (Applied Biosystems/Thermo Fisher Scientific). A positive control mixture of equal quantities of cDNAs from human breast cancer cell lines and control BMA was used to generate a standard curve for real-time qPCR analysis. Pre-amplification products and cDNAs were diluted 20-fold prior to qPCR analysis using the same TaqMan® gene expression assays included in the pre-amplification reaction. Real-time qPCR was performed using a ViiA 7 real-time instrument (Thermo Fisher Scientific; 384-well plate; 5 μL total reaction volume). Technical triplicates were included for each sample, along with nuclease-free water (no-template control). Data were analyzed using Microsoft Excel and GraphPad Prism.
Whole transcriptome amplification (WTA) and RNA-sequencing: A BD Influx cell sorter was used to sort single cells (spiked tumor cells or BMCs) directly into chilled 96-well plates from the BD Precise™ WTA kit, with each well containing cell lysis buffer, dNTPs, and an RT primer containing an 8-bp cell-specific index and 8-bp unique molecular indices for labeling of individual transcripts. Precise™ plates were stored at -80 °C until processing. Single-cell cDNAs were pooled following reverse transcription, and WTA was performed using 20 cycles of PCR amplification. PCR plate-specific indices (equivalent to Illumina indexed adapters D701 = ATTACTCG and D702 = TCCGGAGA) were added during library amplification, and final libraries were quantified using the Qubit™ dsDNA HS assay kit (Thermo Fisher Scientific) and KAPA Universal Library Quantification Kit (Roche; Indianapolis, IN, USA). Quality was further assessed using a 2100 Bioanalyzer with High Sensitivity DNA kit (Agilent; Santa Clara, CA, USA). Libraries from two Precise WTA plates were pooled and loaded at 1.1 pM with 20% PhiX spike-in into a NextSeq 500 Mid-Output cartridge (Illumina; San Diego, CA, USA).
Sequencing was performed using paired-end, 75 bp reads, and demultiplexing and mapping of reads (STAR aligner) was performed as part of the BD Precise WTA analysis pipeline (Seven Bridges Genomics; Charlestown, MA, USA). Additional data analysis was performed using BD Data View™ software. t-SNE plots were calculated after filtering data to include only the most highly variable genes using the Data View “Select variable genes” function (1013 total genes included in analysis; initial_dims = 50, perplexity = 30). Wells with fewer than 200 total unique molecular indices were removed from analysis as poor-quality cells or sorting failures. Median read count for the cells passing filtering criteria was ~800,000 reads per cell, with median molecular index counts of ~20,000 per cell.
Imaging flow cytometry (ImageStream®)
For ImageStream analysis of human breast cancer cell lines, cells were trypsinized, washed, and resuspended in FACS Buffer prior to staining the cell suspension. DTCs or CD45+ BMCs were sorted directly into chilled 1.5 mL Protein LoBind tubes (Eppendorf North America; Framingham, MA, USA) containing 25 µL FACS Buffer; tubes were quickly spun down immediately after cell sorting and kept on ice prior to staining. Samples were stained at RT with a cocktail of antibodies recognizing cell surface markers (FITC anti-EpCAM, PE anti-HER2, and PE-Cy7 anti-CD45). Cells were then fixed and permeabilized using the BD IntraSure™ kit and stained with eFluor® 660 anti-pan-CK (AE1/AE3). Samples were washed once with FACS Buffer containing 0.5 µg/mL DAPI, centrifuged for 3 min at 200 × g, and the supernatant removed. Cells were resuspended immediately prior to analysis on the ImageStream (Amnis, now part of Cytek Biosciences; Bethesda, MD, USA). Single-color compensation controls were prepared using UltraComp eBeads™ compensation beads (Thermo Fisher Scientific) for FITC, PE, PE-Cy7, and eFluor® 660 and using the human breast cancer cell line MCF7 for DAPI.
Image data were acquired at 40× magnification using a 2-camera, 12-channel ImageStream imaging flow cytometer and INSPIRE™ software (Amnis®/Luminex; Austin, TX, USA). Brightfield images were acquired in channels 1 and 9 (intensity = 800), while fluorescence images were acquired under the following conditions: FITC anti-EpCAM – channel #2 (488 nm laser at 60 mW; 480-560 nm emission), PE anti-HER2 – channel #3 (488 nm laser at 60 mW; 560–595 nm emission); PE-Cy7 anti-CD45 – channel #6 (488 nm laser at 60 mW; 745–800 nm emission); DAPI – channel #7 (405 nm laser at 7.0 mW; 430–505 nm emission); and eFluor® 660 anti-pan-CK – channel #11 (658 nm laser at 3.0 mW; 640–745 nm emission). A cell classifier was set for brightfield channel #1 (area lower limit of 50) to reduce the detection of debris. To enable data comparison and quality control between patients, four human breast cancer cell lines (HCC1954, MCF7, MDA-MB-231, and Hs 578T) were processed and analyzed in each experiment under identical conditions to DTCs. Post-acquisition compensation and data analysis were performed using IDEAS® software (Amnis®/Luminex). All image display mapping adjustments were applied equally to all images and data files. For determining cell diameters, an image mask was defined using the Erode function (M01, 2), and the mask diameter was measured for each image using the Feature Manager.
Whole genome amplification, mutation analysis, and whole-exome sequencing
For genomic analysis, small pools of DTCs (~10–200 cells) were sorted into chilled PCR tubes containing RNase-free PBS, as were identically sized pools of CD45+ BMCs as a germline DNA control. Samples were centrifuged at 300 × g for 10 min and stored at −80 °C. Whole genome amplification (WGA) was performed using a REPLI-g® Single Cell kit (Qiagen). Amplified DNA was quantified using the Qubit™ dsDNA HS assay kit (Thermo Fisher Scientific), and DNA quality was assessed using both the Ampli1™ QC kit (Menarini Silicon Biosystems; Huntingdon Valley, PA, USA) and QIAseq™ DNA QuantiMIZE assay kit (Qiagen). For METAMORPH patients, genomic DNA was isolated from banked peripheral blood buffy coat cells (when available) using the Gentra® Puregene® Blood Kit (Qiagen). Genomic DNA was also isolated from FFPE blocks of primary tumors and/or metastatic biopsies using the AllPrep® DNA/RNA FFPE Kit (Qiagen). Macrodissection guided by hematoxylin & eosin-stained sections was performed for metastatic tumor biopsies prior to DNA isolation to maximize tumor content. FFPE DNA samples were treated with Uracil-DNA Glycosylase (UDG; New England Biolabs; Ipswich, MA, USA) for 2 h at 37 °C, then 10 min at 95 °C. All DNA samples were purified using the DNA Clean & Concentrator-5 kit (Zymo Research; Irvine, CA, USA).
Mutation-specific qPCR was performed for the detection of the PIK3CA c.3140A>G mutation, using the TaqMan® mutation detection assay Hs00000831_mu and corresponding PIK3CA wild-type reference assay Hs00001025_rf. qPCR was performed using a ViiA 7 real-time instrument (Thermo Fisher Scientific) following the manufacturer’s recommended protocol. Detection of PIK3CA exon 9 and 20 mutations in DTCs was performed using the Ampli1 PIK3CA Seq kit (Menarini Silicon Biosystems), followed by Sanger sequencing performed by the Nucleic Acid/Protein Research Core Facility at the Children’s Hospital of Philadelphia.
WES was performed by Beijing Genomics Institute (BGI; Shenzhen, China). Library preparation was performed using SureSelectXT (Agilent) using a low-input workflow (200 ng input DNA) and exon capture with SureSelect All Exon V5 (50 Mb). Sequencing was performed on a HiSeq 4000 system (Illumina) at 100PE to reach a target data size of 10 GB per sample after the removal of low-quality reads. WES reads were trimmed for low-quality base-pairs, de-duplicated, and aligned to hg38. Variants were annotated using snpEff99. For DTC and CD45+ BMC WGA samples, WES variants were called using GATK HaplotypeCaller100, while WES variants in matched tumor samples were obtained using a consensus of 4 mutation callers (Mutect1101, Mutect2102, Freebayes103, and Varscan104) for patient B. For patient A, a matched buffy coat gDNA sample was not available, so variants in the metastatic tumor sample were obtained using Mutect2 in single-sample mode102. Variants were further filtered, limiting analysis to those non-synonymous variants listed in the COSMIC (Catalogue of Somatic Mutations in Cancer) database, which were absent in germline DNA (CD45+ BMCs and/or buffy coat gDNA), and present in DTCs and/or matched tumor samples at variant allele frequencies of 0.1 or greater, with read depths of 20 or greater for DTC samples and 30 or greater for tumor samples.
Shallow whole genome sequencing (sWGS)
WGA was performed using the Ampli1™ WGA kit (Menarini Silicon Biosystems). Amplified DNA was quantified using the Qubit™ dsDNA HS assay kit (Thermo Fisher Scientific), and DNA quality was assessed using both the Ampli1™ QC kit (Menarini Silicon Biosystems) and QIAseq™ DNA QuantiMIZE assay kit (Qiagen). WGA products were sheared to a 350 bp target size using a Covaris S220 sonicator. Libraries for sWGS were prepared using the NeoPrep TruSeq® Nano DNA Library Prep kit (Illumina), quality control was performed using a 2100 Bioanalyzer with High Sensitivity DNA kit (Agilent), and libraries were quantified using the KAPA Universal Library Quantification Kit (Roche). Sequencing was performed on a NextSeq 500 system (Illumina) to an average of 5 million mapped reads (75 bp single-end) per sample. Reads from sWGS were trimmed of low-quality base-pairs, aligned, and duplicates were removed. Copy number estimates were generated using QDNAseq105, in which read counts were corrected based on sequence mappability and GC content, and problematic genomic regions were removed based on blacklist status from 1000 Genomes106. Copy number estimates were segmented into bins of 15 kbp each. For each sample, there were around 167,000 bins in total across all chromosomes. The QDNAseq function’s output log2 copy ratio estimates (to an assumed diploid reference) and the Pearson correlation between any pair of samples were computed using these values.
Statistical analysis
All statistical analyses were performed using GraphPad Prism software, v10. For comparison of spiked tumor cell recovery by DTC-Flow vs. by DTC-IHC, we used a paired two-tailed t-test. For analyses involving multiple statistical comparisons using a single data set (i.e., gene expression comparisons, ImageStream cell size analysis), we performed one-way ANOVA followed by Bonferroni’s multiple comparisons test. p-values were considered significant at 0.05.
Data availability
The datasets generated and/or analyzed during the current study are not publicly available due to the need to ensure confidentiality of identifying and/or sensitive information regarding clinical trial participants, but are available from the corresponding author upon reasonable request.
References
Bray, F. et al. Global Cancer Statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 68, 394–424 (2018).
Giaquinto, A. N. et al. Breast Cancer Statistics, 2022. CA Cancer J. Clin. 72, 524–541 (2022).
Cossetti, R. J., Tyldesley, S. K., Speers, C. H., Zheng, Y. & Gelmon, K. A. Comparison of breast cancer recurrence and outcome patterns between patients treated from 1986 to 1992 and from 2004 to 2008. J. Clin. Oncol. 33, 65–73 (2015).
Pan, H. et al. 20-year risks of breast-cancer recurrence after stopping endocrine therapy at 5 years. N. Engl. J. Med. 377, 1836–1846 (2017).
Dawood, S., Broglio, K., Ensor, J., Hortobagyi, G. N. & Giordano, S. H. Survival differences among women with de novo stage IV and relapsed breast cancer. Ann. Oncol. 21, 2169–2174 (2010).
DeSantis, C. E. et al. Breast cancer statistics, 2019. CA Cancer J. Clin. 69, 438–451 (2019).
Khatcheressian, J. L. et al. Breast cancer follow-up and management after primary treatment: American Society of Clinical Oncology clinical practice guideline update. J. Clin. Oncol. 31, 961–965 (2013).
Carreira, H. et al. Associations between breast cancer survivorship and adverse mental health outcomes: a systematic review. J. Natl. Cancer Inst. 110, 1311–1327 (2018).
Luigjes-Huizer, Y. L. et al. What is the prevalence of fear of cancer recurrence in cancer survivors and patients? A systematic review and individual participant data meta-analysis. Psychooncology 31, 879–892 (2022).
Bidard, F. C. et al. Disseminated tumor cells of breast cancer patients: a strong prognostic factor for distant and local relapse. Clin. Cancer Res. 14, 3306–3311 (2008).
Braun, S. et al. A pooled analysis of bone marrow micrometastasis in breast cancer. N. Engl. J. Med. 353, 793–802 (2005).
Giuliano, A. E. et al. Association of occult metastases in sentinel lymph nodes and bone marrow with survival among women with early-stage invasive breast cancer. JAMA 306, 385–393 (2011).
Gruber, I. et al. Disseminated tumor cells as a monitoring tool for adjuvant therapy in patients with primary breast cancer. Breast Cancer Res. Treat. 144, 353–360 (2014).
Hall, C. et al. Disseminated tumor cells predict survival after neoadjuvant therapy in primary breast cancer. Cancer 118, 342–348 (2012).
Hartkopf, A. D. et al. Prognostic relevance of disseminated tumour cells from the bone marrow of early stage breast cancer patients - results from a large single-centre analysis. Eur. J. Cancer 50, 2550–2559 (2014).
Janni, W. et al. Persistence of disseminated tumor cells in the bone marrow of breast cancer patients predicts increased risk for relapse-a European pooled analysis. Clin. Cancer Res. 17, 2967–2976 (2011).
Schindlbeck, C. et al. Prognostic relevance of disseminated tumor cells in the bone marrow of patients with primary breast cancer-results of a standardized follow-up. Anticancer Res. 31, 2749–2755 (2011).
Wiedswang, G. et al. Isolated tumor cells in bone marrow three years after diagnosis in disease-free breast cancer patients predict unfavorable clinical outcome. Clin. Cancer Res. 10, 5342–5348 (2004).
Mathiesen, R. R. et al. Persistence of disseminated tumor cells after neoadjuvant treatment for locally advanced breast cancer predicts poor survival. Breast Cancer Res. 14, R117 (2012).
Naume, B. et al. Clinical outcome with correlation to disseminated tumor cell (DTC) status after DTC-guided secondary adjuvant treatment with docetaxel in early breast cancer. J. Clin. Oncol. 32, 3848–3857 (2014).
Ben Lassoued, A., Nivaggioni, V. & Gabert, J. Minimal residual disease testing in hematologic malignancies and solid cancer. Expert Rev. Mol. Diagn. 14, 699–712 (2014).
Harris, L. et al. American Society of Clinical Oncology 2007 update of recommendations for the use of tumor markers in breast cancer. J. Clin. Oncol. 25, 5287–5312 (2007).
Abravanel, D. L. et al. Notch promotes recurrence of dormant tumor cells following HER2/neu-targeted therapy. J. Clin. Invest. 125, 2484–2496 (2015).
Alvarez, J. V. et al. Par-4 downregulation promotes breast cancer recurrence by preventing multinucleation following targeted therapy. Cancer Cell 24, 30–44 (2013).
Feng, Y. et al. SPSB1 promotes breast cancer recurrence by potentiating c-MET signaling. Cancer Discov. 4, 790–803 (2014).
Payne, A. W., Pant, D. K., Pan, T. C. & Chodosh, L. A. Ceramide kinase promotes tumor cell survival and mammary tumor recurrence. Cancer Res. 74, 6352–6363 (2014).
Sreekumar, A. et al. B3GALT6 promotes dormant breast cancer cell survival and recurrence by enabling heparan sulfate-mediated FGF signaling. Cancer Cell 42, 52–69.e57 (2024).
Dwyer, S., Ruth, J., Seidel, H. E., Raz, A. A. & Chodosh, L. A. Autophagy is required for mammary tumor recurrence by promoting dormant tumor cell survival following therapy. Breast Cancer Res. 26, 143 (2024).
DeMichele, A. et al. Targeting dormant tumor cells to prevent recurrent breast cancer: a randomized phase 2 trial. Nat. Med. (in the press).
Bragado, P. et al. TGF-beta2 dictates disseminated tumour cell fate in target organs through TGF-beta-RIII and p38alpha/beta signalling. Nat. Cell Biol. 15, 1351–1361 (2013).
Gao, H. et al. The BMP inhibitor Coco reactivates breast cancer cells at lung metastatic sites. Cell 150, 764–779 (2012).
Ghajar, C. M. et al. The perivascular niche regulates breast tumour dormancy. Nat. Cell Biol. 15, 807–817 (2013).
Ruth, J. R. et al. Cellular dormancy in minimal residual disease following targeted therapy. Breast Cancer Res. 23, 63 (2021).
Demicheli, R., Retsky, M. W., Swartzendruber, D. E. & Bonadonna, G. Proposal for a new model of breast cancer metastatic development. Ann. Oncol. 8, 1075–1080 (1997).
Dalla, E., Sreekumar, A., Aguirre-Ghiso, J. A. & Chodosh, L. A. Dormancy in Breast Cancer. Cold Spring Harb. Perspect. Med. 13, a041331 (2023).
Borgen, E. et al. NR2F1 stratifies dormant disseminated tumor cells in breast cancer patients. Breast Cancer Res. 20, 120 (2018).
Gangnus, R., Langer, S., Breit, E., Pantel, K. & Speicher, M. R. Genomic profiling of viable and proliferative micrometastatic cells from early-stage breast cancer patients. Clin. Cancer Res. 10, 3457–3464 (2004).
Klein, C. A. et al. Genetic heterogeneity of single disseminated tumour cells in minimal residual cancer. Lancet 360, 683–689 (2002).
Pantel, K. et al. Differential expression of proliferation-associated molecules in individual micrometastatic carcinoma cells. J. Natl. Cancer Inst. 85, 1419–1424 (1993).
Agudo, J. et al. Targeting cancer cell dormancy. Nat. Rev. Cancer 24, 97–104 (2024).
Fehm, T. et al. A concept for the standardized detection of disseminated tumor cells in bone marrow from patients with primary breast cancer and its clinical implementation. Cancer 107, 885–892 (2006).
Prat, A. et al. Phenotypic and molecular characterization of the claudin-low intrinsic subtype of breast cancer. Breast Cancer Res. 12, R68 (2010).
Willipinski-Stapelfeldt, B. et al. Changes in cytoskeletal protein composition indicative of an epithelial-mesenchymal transition in human micrometastatic and primary breast carcinoma cells. Clin. Cancer Res. 11, 8006–8014 (2005).
Yu, M. et al. Circulating breast tumor cells exhibit dynamic changes in epithelial and mesenchymal composition. Science 339, 580–584 (2013).
Sorg, N. et al. Red blood cell depletion from bone marrow and peripheral blood buffy coat: a comparison of two new and three established technologies. Transfusion 55, 1275–1282 (2015).
Osborne, M. P. et al. Sensitivity of immunocytochemical detection of breast cancer cells in human bone marrow. Cancer Res. 51, 2706–2709 (1991).
Wulf, G. G. et al. Reverse transcriptase/polymerase chain reaction analysis of parathyroid hormone-related protein for the detection of tumor cell dissemination in the peripheral blood and bone marrow of patients with breast cancer. J. Cancer Res. Clin. Oncol. 123, 514–521 (1997).
Yu, L. et al. An integrated enrichment system to facilitate isolation and molecular characterization of single cancer cells from whole blood. Cytom. A 93, 1226–1233 (2018).
Cristofanilli, M. et al. Circulating tumor cells, disease progression, and survival in metastatic breast cancer. N. Engl. J. Med. 351, 781–791 (2004).
Ring, A. et al. EpCAM based capture detects and recovers circulating tumor cells from all subtypes of breast cancer except claudin-low. Oncotarget 6, 44623–44634 (2015).
Sieuwerts, A. M. et al. Anti-epithelial cell adhesion molecule antibodies and the detection of circulating normal-like breast tumor cells. J. Natl. Cancer Inst. 101, 61–66 (2009).
Andrulis, I. L. et al. neu/erbB-2 amplification identifies a poor-prognosis group of women with node-negative breast cancer. Toronto Breast Cancer Study Group. J. Clin. Oncol. 16, 1340–1349 (1998).
Ingthorsson, S. et al. HER2 induced EMT and tumorigenicity in breast epithelial progenitor cells is inhibited by coexpression of EGFR. Oncogene 35, 4244–4255 (2016).
Schettini, F. et al. Clinical, pathological, and PAM50 gene expression features of HER2-low breast cancer. NPJ Breast Cancer 7, 1 (2021).
Bhagwat, N. et al. An integrated flow cytometry-based platform for isolation and molecular characterization of circulating tumor single cells and clusters. Sci. Rep. 8, 5035 (2018).
Maxwell, K. N. et al. Comparative clinical utility of tumor genomic testing and cell-free DNA in metastatic breast cancer. Breast Cancer Res. Treat. 164, 627–638 (2017).
Aceto, N. et al. Circulating tumor cell clusters are oligoclonal precursors of breast cancer metastasis. Cell 158, 1110–1122 (2014).
Aguirre-Ghiso, J. A. Models, mechanisms and clinical evidence for cancer dormancy. Nat. Rev. Cancer 7, 834–846 (2007).
Gerdes, J. et al. Cell cycle analysis of a cell proliferation-associated human nuclear antigen defined by the monoclonal antibody Ki-67. J. Immunol. 133, 1710–1715 (1984).
Sherley, J. L. & Kelly, T. J. Regulation of human thymidine kinase during the cell cycle. J. Biol. Chem. 263, 8350–8358 (1988).
Hwang, A., Maity, A., McKenna, W. G. & Muschel, R. J. Cell cycle-dependent regulation of the cyclin B1 promoter. J. Biol. Chem. 270, 28419–28424 (1995).
Baldin, V., Lukas, J., Marcote, M. J., Pagano, M. & Draetta, G. Cyclin D1 is a nuclear protein required for cell cycle progression in G1. Genes Dev. 7, 812–821 (1993).
Pantel, K. et al. In Metastasis: Basic Research and Its Clinical Applications Contributions to Oncology (eds Rabes, H. et al.) Vol. 44, 283–293 (S. Karger AG, 1992).
Merenbakh-Lamin, K. et al. D538G mutation in estrogen receptor-alpha: A novel mechanism for acquired endocrine resistance in breast cancer. Cancer Res. 73, 6856–6864 (2013).
Mesquita, B. et al. Frequent copy number gains at 1q21 and 1q32 are associated with overexpression of the ETS transcription factors ETV3 and ELF3 in breast cancer irrespective of molecular subtypes. Breast Cancer Res. Treat. 138, 37–45 (2013).
Rakha, E. A., Green, A. R., Powe, D. G., Roylance, R. & Ellis, I. O. Chromosome 16 tumor-suppressor genes in breast cancer. Genes Chromosomes Cancer 45, 527–535 (2006).
Bayne, L. et al. Abstract PD9-11: Identifying breast cancer survivors with dormant disseminated tumor cells: The PENN-SURMOUNT screening study. Cancer Res. 81 (4 Suppl), PD9-11 (2021).
Solakoglu, O. et al. Heterogeneous proliferative potential of occult metastatic cells in bone marrow of patients with solid epithelial tumors. Proc. Natl. Acad. Sci. USA 99, 2246–2251 (2002).
Gross, H. J., Verwer, B., Houck, D., Hoffman, R. A. & Recktenwald, D. Model study detecting breast cancer cells in peripheral blood mononuclear cells at frequencies as low as 10(-7). Proc. Natl. Acad. Sci. USA 92, 537–541 (1995).
Magbanua, M. J. M. et al. Genomic and expression profiling reveal molecular heterogeneity of disseminated tumor cells in bone marrow of early breast cancer. NPJ Breast Cancer 4, 31 (2018).
Simpson, S. J. et al. Detection of tumor cells in the bone marrow, peripheral blood, and apheresis products of breast cancer patients using flow cytometry. Exp. Hematol. 23, 1062–1068 (1995).
Kalluri, R. & Weinberg, R. A. The basics of epithelial-mesenchymal transition. J. Clin. Invest.119, 1420–1428 (2009).
Thiery, J. P. Epithelial-mesenchymal transitions in tumour progression. Nat. Rev. Cancer 2, 442–454 (2002).
Creighton, C. J. et al. Residual breast cancers after conventional therapy display mesenchymal as well as tumor-initiating features. Proc. Natl. Acad. Sci. USA 106, 13820–13825 (2009).
Mani, S. A. et al. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 133, 704–715 (2008).
Moody, S. E. et al. The transcriptional repressor Snail promotes mammary tumor recurrence. Cancer Cell 8, 197–209 (2005).
Brugger, W. et al. Expression of MUC-1 epitopes on normal bone marrow: implications for the detection of micrometastatic tumor cells. J. Clin. Oncol. 17, 1535–1544 (1999).
Lammers, R. et al. Monoclonal antibody 9C4 recognizes epithelial cellular adhesion molecule, a cell surface antigen expressed in early steps of erythropoiesis. Exp. Hematol. 30, 537–545 (2002).
Chery, L. et al. Characterization of single disseminated prostate cancer cells reveals tumor cell heterogeneity and identifies dormancy associated pathways. Oncotarget 5, 9939–9951 (2014).
Andrechek, E. R., White, D. & Muller, W. J. Targeted disruption of ErbB2/Neu in the mammary epithelium results in impaired ductal outgrowth. Oncogene 24, 932–937 (2005).
Schroeder, J. A. & Lee, D. C. Dynamic expression and activation of ERBB receptors in the developing mouse mammary gland. Cell Growth Differ. 9, 451–464 (1998).
Tarantino, P. et al. ESMO expert consensus statements (ECS) on the definition, diagnosis, and management of HER2-low breast cancer. Ann. Oncol. 34, 645–659 (2023).
Bandyopadhyay, S. et al. Mapping the cellular biogeography of human bone marrow niches using single-cell transcriptomics and proteomic imaging. Cell 187, 3120–3140.e3129 (2024).
Solomayer, E. F. et al. Influence of adjuvant hormone therapy and chemotherapy on the immune system analysed in the bone marrow of patients with breast cancer. Clin. Cancer Res. 9, 174–180 (2003).
Pantel, K. et al. Methodological analysis of immunocytochemical screening for disseminated epithelial tumor cells in bone marrow. J. Hematother 3, 165–173 (1994).
Wiedswang, G., Borgen, E., Karesen, R. & Naume, B. Detection of isolated tumor cells in BM from breast-cancer patients: significance of anterior and posterior iliac crest aspirations and the number of mononuclear cells analyzed. Cytotherapy 5, 40–45 (2003).
Sosa, M. S. et al. NR2F1 controls tumour cell dormancy via SOX9- and RARbeta-driven quiescence programmes. Nat. Commun. 6, 6170 (2015).
Braun, S. et al. Lack of effect of adjuvant chemotherapy on the elimination of single dormant tumor cells in bone marrow of high-risk breast cancer patients. J. Clin. Oncol. 18, 80–86 (2000).
Naumov, G. N. et al. Ineffectiveness of doxorubicin treatment on solitary dormant mammary carcinoma cells or late-developing metastases. Breast Cancer Res. Treat. 82, 199–206 (2003).
Balic, M. et al. Most early disseminated cancer cells detected in bone marrow of breast cancer patients have a putative breast cancer stem cell phenotype. Clin. Cancer Res. 12, 5615–5621 (2006).
Reuben, J. M. et al. Primary breast cancer patients with high risk clinicopathologic features have high percentages of bone marrow epithelial cells with ALDH activity and CD44(+)CD24lo cancer stem cell phenotype. Eur. J. Cancer 47, 1527–1536 (2011).
Chen, K. et al. Perioperative dynamic changes in circulating tumor DNA in patients with lung cancer (DYNAMIC). Clin. Cancer Res. 25, 7058–7067 (2019).
Meng, S. et al. Circulating tumor cells in patients with breast cancer dormancy. Clin. Cancer Res. 10, 8152–8162 (2004).
Klein, C. A. Parallel progression of primary tumours and metastases. Nat. Rev. Cancer 9, 302–312 (2009).
Kroigard, A. B. et al. Genomic analyses of breast cancer progression reveal distinct routes of metastasis emergence. Sci. Rep. 7, 43813 (2017).
Lawrence-Paul, M. R. et al. Rare subclonal sequencing of breast cancers indicates putative metastatic driver mutations are predominately acquired after dissemination. Genome Med. 16, 26 (2024).
Hu, Z., Li, Z., Ma, Z. & Curtis, C. Multi-cancer analysis of clonality and the timing of systemic spread in paired primary tumors and metastases. Nat. Genet. 52, 701–708 (2020).
Rodrigues, O. R. & Monard, S. A rapid method to verify single-cell deposition setup for cell sorters. Cytom. A 89, 594–600 (2016).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 6, 80–92 (2012).
Van der Auwera, G. A. & O’Connor, B. D. Genomics in the Cloud: Using Docker, GATK, and WDL in Terra (O’Reilly Media, 2020).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Li, H. Toward better understanding of artifacts in variant calling from high-coverage samples. Bioinformatics 30, 2843–2851 (2014).
Koboldt, D. C. et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576 (2012).
Scheinin, I. et al. DNA copy number analysis of fresh and formalin-fixed specimens by shallow whole-genome sequencing with identification and exclusion of problematic regions in the genome assembly. Genome Res. 24, 2022–2032 (2014).
Genomes Project, C. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Acknowledgements
This study was supported in part by grants from the National Cancer Institute (CA208273) and the Penn Medicine Abramson Cancer Center 2-PREVENT Breast Cancer Translational Center of Excellence and by a Postdoctoral Fellowship Award from Susan G Komen® (E. Chislock; PDF17483333). The funders played no role in study design, data collection, analysis and interpretation of data, or the writing of this manuscript. Some of the data for this manuscript were generated in the Penn Cytomics and Cell Sorting Shared Resource Laboratory at the University of Pennsylvania, which is partially supported by the Abramson Cancer Center NCI Grant (P30 016520); the research identifier number is RRid:SCR_022376. The authors thank Richard Schretzenmair of the Penn Cytomics and Cell Sorting Shared Resource Laboratory for guidance on imaging flow cytometry.
Author information
Authors and Affiliations
Contributions
Study concept and design: E.M.C., A.D., and L.A.C. Development of DTC-Flow methodology and conduct of flow cytometry/FACS/imaging cytometry experiments: E.M.C., G.W., J.G., T.B., M.S., X.J., W.D., A.M., W.M., C.H.P., J.S.M., and L.A.C. BD FACSFocus development: L.Y., K.J., S.B., C.H.P., and J.S.M. IHC assay/pathology: G.K.B., J.W., M.S., N.N.C.S., A.N., and M.F. Clinical team (patient enrollment, consents, bone marrow aspirates, biopsy acquisition, tissue samples, etc.): L.B., D.S.E., C.C., N.G., I.N., B.G., S.D., M.L., J.N., S.W.S., A.C., and A.D. Sample processing for genomic/molecular analysis: E.M.C., Y.C., G.W., J.G., G.K.B., and T.B. Bioinformatic and statistical analysis: T.C.P., D.P., and M.R.L.-P. Data analysis and interpretation (i.e., molecular biology, flow cytometry): E.M.C. and L.A.C. A.D. designed, implemented and oversaw the METAMORPH Study (“Metastatic Markers of the Recurrent Phenotype”) and PENN-SURMOUNT Study (“PENN-Surveillance Markers of Utility for Recurrence After (Neo)Adjuvant Therapy for Breast Cancer”). Manuscript drafting: E.M.C. and L.A.C. All authors reviewed and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
A.D. has institutional research funding from Novartis, Genentech, Pfizer, and NeoGenomics. L.A.C. has served as an expert consultant to Teva Pharmaceuticals, Eisai, Sanofi, Eli Lilly, Whittaker, Clark and Daniels, Wyeth, Imerys, Colgate, Becton Dickinson, Sterigenics, and the U.S. Department of Justice in litigation. L.Y., K.J., and S.B. are current or former employees of BD. L.Y. and BD are inventors on patents on the technology used in the FACSFocus device. E.M.C. and L.A.C. are inventors on a patent related to the DTC-Flow assay. Other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chislock, E.M., Pan, Tc., Pant, D. et al. DTC-Flow: a flow cytometry-based detection platform for characterizing bone marrow disseminated tumor cells in breast cancer. npj Breast Cancer 11, 139 (2025). https://doi.org/10.1038/s41523-025-00853-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41523-025-00853-5