Abstract
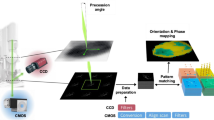
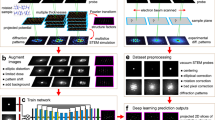
Four-dimensional scanning transmission electron microscopy (4D-STEM) is a high-throughput automated data acquisition technique with great potential for real-time data collection and analysis in automated STEM. However, its practical implementation is limited by challenges in data preprocessing, which hinder the timely and accurate interpretation of the large amounts of data it generates. Issues like pervasive noise, beam center drift, and elliptical distortions during high-throughput acquisition inevitably degrade diffraction patterns, leading to systematic errors in quantitative measurements. Conventional calibration algorithms are often material-specific and fail to provide a robust, generalizable solution. In this work, we introduce 4D-PreNet, an end-to-end deep-learning pipeline that integrates attention-enhanced U-Net and ResNet architectures to simultaneously perform denoising, center calibration, and ellipse calibration. The network is trained on extensive simulated datasets that cover a broad range of noise levels, drift magnitudes, and distortion types, thereby enabling generalization to experimental data obtained under different acquisition conditions. Quantitative evaluations demonstrate that 4D-PreNet reduces mean squared error by up to 50% in denoising and achieves sub-pixel center localization with average errors below 0.04 pixels. Compared to conventional algorithms, 4D-PreNet shows improved noise suppression and accurate restoration of diffraction features, enabling reliable real-time analysis of 4D-STEM data and supporting automated STEM workflows.
Similar content being viewed by others
Data availability
The datasets generated and analyzed during the current study are not publicly available due to the internal data sharing policies of the participating institutions, but are available from the corresponding author on reasonable request. The code for implementing the 4D-STEM preprocessing framework—including denoising, beam center calibration, and ellipse calibration networks—is currently under preparation. It will be released via a public GitHub repository upon publication to support reproducibility and community use.
Code availability
The code for implementing the 4D-STEM preprocessing framework—including denoising, beam center calibration, and ellipse calibration networks—is currently under preparation. It will be released via a public GitHub repository upon publication to support reproducibility and community use.
References
Roccapriore, K. M. et al. Automated experiment in 4d-stem: Exploring emergent physics and structural behaviors. ACS Nano 16, 7605–7614 (2022).
Kalinin, S. V. et al. Machine learning in scanning transmission electron microscopy. Nat. Rev. Methods Prim. 2, 11 (2022).
Spurgeon, S. R. et al. Towards data-driven next-generation transmission electron microscopy. Nat. Mater. 20, 274–279 (2021).
Ophus, C. A fast image simulation algorithm for scanning transmission electron microscopy. Adv. Struct. Chem. Imaging 3, 13 (2017).
Zeltmann, S. E. et al. Patterned probes for high precision 4D-stem Bragg measurements. Ultramicroscopy 209, 112890 (2020).
Wang, Z. et al. 4d-misr: A unified model for low-dose super-resolution imaging via feature fusion. AI Sci. 1, 015003 (2025).
Savitzky, B. H. et al. Image registration of low signal-to-noise cryo-stem data. Ultramicroscopy 191, 56–65 (2018).
Zhang, C., Han, R., Zhang, A. R. & Voyles, P. M. Denoising atomic resolution 4D scanning transmission electron microscopy data with tensor singular value decomposition. Ultramicroscopy 219, 113123 (2020).
Wu, C. H., Reynolds, W. T. & Murayama, M. A software tool for automatic analysis of selected area diffraction patterns within digital micrograph™. Ultramicroscopy 112, 10–14 (2012).
Lábár, J. L. & Das, P. P. Pattern center and distortion determined from faint, diffuse electron diffraction rings from amorphous materials. Microsc. Microanal. 23, 647–660 (2017).
Pekin, T. C. et al. Optimizing disk registration algorithms for nanobeam electron diffraction strain mapping. Ultramicroscopy 176, 170–176 (2017).
Jones, L. et al. Managing dose-, damage- and data-rates in multi-frame spectrum-imaging. Microscopy67, i98–i113 (2018).
Yuan, R., Zhang, J. & Zuo, J.-M. Lattice strain mapping using circular hough transform for electron diffraction disk detection. Ultramicroscopy 207, 112837 (2019).
Savitzky, B. H. et al. Py4dstem: A software package for four-dimensional scanning transmission electron microscopy data analysis. Microsc. Microanal. 27, 712–743 (2021).
Kim, J. et al. Self-supervised machine learning framework for high-throughput electron microscopy. Sci. Adv. 11, eads5552 (2025).
Lobato, I. et al. Deep convolutional neural networks to restore single-shot electron microscopy images. npj Comput. Mater. 10, 1188 (2024).
Ziletti, A., Kumar, D., Scheffler, M. & Ghiringhelli, L. M. Insightful classification of crystal structures using deep learning. Nat. Commun. 9, 2775 (2018).
Aguiar, J. A., Gong, M. L. & Tasdizen, T. Crystallographic prediction from diffraction and chemistry data for higher throughput classification using machine learning. Comput. Mater. Sci. 173, 109409 (2020).
Han, Y. et al. Deep learning stem-edx tomography of nanocrystals. Nat. Mach. Intell. 3, 267–274 (2021).
Zheng, H., Lu, X. & He, K. In situ transmission electron microscopy and artificial intelligence enabled data analytics for energy materials. J. Energy Chem. 68, 454–493 (2022).
Kaufmann, K. et al. Crystal symmetry determination in electron diffraction using machine learning. Science 367, 564–568 (2020).
Xu, W. & LeBeau, J. M. A deep convolutional neural network to analyze position averaged convergent beam electron diffraction patterns. Ultramicroscopy 188, 59–69 (2018).
Sadri, A. et al. Unsupervised deep denoising for four-dimensional scanning transmission electron microscopy. npj Comput. Mater. 10, 243 (2024).
Nordahl, G., Dagenborg, S., Sørhaug, J. & Nord, M. Exploring deep learning models for 4D-stem-dpc data processing. Ultramicroscopy 267, 114058 (2024).
Ge, M. et al. Automatic center identification of electron diffraction with multi-scale transformer networks. Ultramicroscopy 259, 113926 (2024).
Xie, Y. et al. Spatially resolved structural order in low-temperature liquid electrolyte. Sci. Adv. 9, eadc9721 (2023).
Martin, B. R. et al. abTEM: Transmission electron microscopy simulations with the atomic potential. Open Res. Eur. 2, 13015 (2022).
Ronneberger, O., Fischer, P. & Brox, T. U-Net: Convolutional networks for biomedical image segmentation. Med. Image Comput. Comput. -Assist. Inter. 9351, 234–241 (2015).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. Proc. IEEE Conf. Comput. Vis. Pattern Recognit. 770, 778 (2016).
Woo, S., Park, J., Lee, J.-Y. & Kweon, I. S. CBAM: Convolutional block attention module. Proc. Eur. Conf. Comput. Vis. 11211, 3–19 (2018).
Han, S., Mao, H. & Dally, W. J. Deep compression: Compressing deep neural networks with pruning, trained quantization and Huffman coding. Proc. Int. Conf. Learn. Represent. (2016).
Lehtinen, J. et al. Noise2Noise: Learning Image Restoration without Clean Data. Proc. Int. Conf. Mach. Learn. 80, 2965–2974 (2018).
Ede, J. M. Deep learning in electron microscopy. Mach. Learn.: Sci. Technol. 2, 011004 (2021).
Bengio, Y. Practical recommendations for gradient-based training of deep architectures. In Neural Networks: Tricks of the Trade (eds Montavon, G., Orr, G. B. & Müller, K.-R.) 437–478 (Springer, 2012).
Sun, K., Xiao, B., Liu, D. & Wang, J. Deep high-resolution representation learning for human pose estimation. Proc. IEEE Conf. Comput. Vis. Pattern Recognit. 5686, 5696 (2019).
Zhang, C. et al. Understanding deep learning (still) requires rethinking generalization. Commun. ACM 64, 107–115 (2021).
Ruiz, N., Chong, E. & Rehg, J. Fine-grained head pose estimation without keypoints. Proc. IEEE Conf. Comput. Vis. Pattern Recognit. Workshops 2074, 2083 (2018).
Jain, A. et al. Commentary: The materials project: A materials genome approach to accelerating materials innovation. APL Mater. 1, 011002 (2013).
Kingma, D. P. & Ba, J. Adam: A method for stochastic optimization. Proc. Int. Conf. Learn. Represent. (2014).
Shi, C. et al. Uncovering material deformations via machine learning combined with four-dimensional scanning transmission electron microscopy. npj Comput. Mater. 8, 114 (2022).
Acknowledgements
National Natural Science Foundation of China (12474186, 52401159, and 52425101) is acknowledged for funding this research.
Author information
Authors and Affiliations
Contributions
M.L. and Y.X. conceived and designed the study. M.L. and Z.M. developed the model, performed simulations, and drafted the manuscript. Z.M., S.C., and Y.X. co-supervised the research and contributed to model refinement and theoretical analysis. Z.L. and J.G. implemented the algorithm and conducted training. H.Z. and X.H. supported dataset preprocessing and validation. S.C. and C.C. assisted with simulation and data augmentation. J.D. provided materials science expertise. J.H. provided assistance with crystal orientation determination and strain calculations during the revision process. X.Z. provided additional datasets required for the revision and offered guidance on data analysis. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, M., Mao, Z., Liu, Z. et al. A Unified preprocessing framework for high-throughput diffraction pattern analysis. npj Comput Mater (2026). https://doi.org/10.1038/s41524-026-01993-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41524-026-01993-3