Abstract
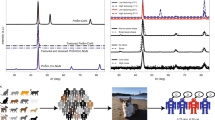
Accurate crystal structure determination underpins materials discovery, yet powder X-ray diffraction (XRD) analysis still depends on expert-driven, iterative fitting that limits scalability for high-throughput and autonomous experiments. We introduce XRD-Crystal Contrastive Pretraining (XCCP), a physics-guided contrastive learning framework that aligns PXRD patterns with candidate crystal structures in a shared embedding space to enable efficient structure retrieval and symmetry inference. XCCP employs a dual-expert XRD encoder with a Kolmogorov-Arnold Network (KAN) projection head. A low-angle branch captures long-length-scale signatures, while a wide-angle branch encodes dense, symmetry-governed fingerprints. Attribution and perturbation analyses show that the KAN head concentrates evidence on physically meaningful Bragg reflections rather than background-dominated regions, improving robustness to peak-shape variations. We further introduce similarity-based confidence scores to flag potentially unreliable predictions in open-set settings. Without elemental priors, XCCP achieves 46.42% top-1 accuracy for structure retrieval and 60.85% accuracy for space-group identification. When chemical composition is available for elemental pre-screening, performance increases to 88.98% and 93.39%, respectively. XCCP also generalizes to compositionally similar multi-principal element alloys and enables zero-shot transfer to experimental patterns. These results establish XCCP as an interpretable, confidence-aware, and scalable paradigm for XRD analysis, enabling high-throughput screening, rapid candidate shortlisting, and integration with autonomous laboratory workflows.
Similar content being viewed by others
Data availability
The opXRD-related experimental dataset used in the present work can be found in https://huggingface.co/datasets/caobin/opxrd_hkust_expdata, and the CIFs for 22 MPEAs can be found in https://github.com/George-JieXIONG/Materials-Dataset/tree/main/XRD-Files.
References
Altomare, A. et al. EXPO2009: structure solution by powder data in direct and reciprocal space. J. Appl. Crystallogr. 42, 1197–1202 (2009).
Huang, T. C. Powder diffraction: theory and practice—a new and comprehensive book on the methods and applications of powder diffraction. Powder Diffr. 24, 2–3 (2012).
Long, C. J., Bunker, D., Li, X., Karen, V. L. & Takeuchi, I. Rapid identification of structural phases in combinatorial thin-film libraries using X-ray diffraction and non-negative matrix factorization. Rev. Sci. Instrum. 80, 103902 (2009).
Leonardi, A. & Bish, D. L. Interactions of lattice distortion fields in nano polycrystalline materials revealed by molecular dynamics and X-ray powder diffraction. Acta Mater. 133, 380–392 (2017).
Vishina, A., Eriksson, O. & Herper, H. C. Stable and metastable rare-earth-free permanent magnets from a database of predicted crystal structures. Acta Mater. 261, 119348 (2023).
Albesa-Jové, D. et al. Challenges in direct-space structure determination from powder diffraction data: a molecular material with four independent molecules in the asymmetric unit. ChemPhysChem 5, 414–418 (2004).
Stanev, V. et al. Unsupervised phase mapping of X-ray diffraction data by nonnegative matrix factorization integrated with custom clustering. npj Comput. Mater. 4, 43 (2018).
Hua, Y. et al. Outstanding comprehensive piezoelectric properties in KNN-based ceramics via co-optimization of crystal structure and grain orientation. Acta Mater. 297, 121378 (2025).
McCusker, L. B., Von Dreele, R. B., Cox, D. E., Louër, D. & Scardi, P. Rietveld refinement guidelines. J. Appl. Crystallogr. 32, 36–50 (1999).
Rietveld, H. M. The Rietveld method: a retrospection. Z. f.ür. Kristallogr. 225, 545–547 (2010).
Marks, L. D. A standard format for reporting atomic positions in measured or calculated surface structures: the CIF file. Surf. Sci. 604, 878–881 (2010).
Brown, I. D. CIF (Crystallographic Information File). A standard for crystallographic data interchange. J. Res. Natl. Inst. Stand. Technol. 101, 341–346 (1996).
Liu, Y. et al. Machine learning in materials genome initiative: a review. J. Mater. Sci. Technol. 57, 113–122 (2020).
Long, T. et al. Inverse design of crystal structures for multicomponent systems. Acta Mater. 231, 117898 (2022).
Guo, G. B. et al. Ab initio structure solutions from nanocrystalline powder diffraction data via diffusion models. Nat. Mater. 24, 1726–1734 (2025).
Agrawal, A. & Choudhary, A. Perspective: materials informatics and big data: Realization of the “fourth paradigm” of science in materials science. APL Mater. 4, 053208 (2016).
Pei, C. H. et al. A novel model to predict oxidation behavior of superalloys based on machine learning. J. Mater. Sci. Technol. 235, 232–243 (2025).
Su, T. H., Cao, B., Hu, S. B., Li, M. S. & Zhang, T. Y. CGWGAN: crystal generative framework based on Wyckoff generative adversarial network. J. Mater. Inform. 4, 20 (2024).
Cao, B. et al. XQueryer: an intelligent crystal structure identifier for powder X-ray diffraction. Natl. Sci. Rev. 12, nwaf421 (2025).
Xie, T. & Grossman, J. C. Crystal graph convolutional neural networks for an accurate and interpretable prediction of material properties. Phys. Rev. Lett. 120, 145301 (2018).
Liu, Z. et al. BraggNN: fast X-ray Bragg peak analysis using deep learning. IUCrJ 9, 104–113 (2021).
Zheng, X., Zhang, X., Chen, T. T. & Watanabe, I. Deep learning in mechanical metamaterials: from prediction and generation to inverse design. Adv. Mater. 35, e2302530 (2023).
Schleder, G. R., Padilha, A. C. M., Acosta, C. M., Costa, M. & Fazzio, A. From DFT to machine learning: recent approaches to materials science–a review. J. Phys. Mater. 2, 032001 (2019).
Choudhary, K. et al. Recent advances and applications of deep learning methods in materials science. npj Comput. Mater. 8, 59 (2022).
Zhang, S. Y. et al. Crystallographic phase identifier of a convolutional self-attention neural network (CPICANN) on powder diffraction patterns. IUCrJ 11, 634–642 (2024).
Dong, H. et al. A deep convolutional neural network for real-time full profile analysis of big powder diffraction data. npj Comput. Mater. 7, 74 (2021).
Lee, B. D. et al. A deep learning approach to powder X-Ray diffraction pattern analysis: addressing generalizability and perturbation issues simultaneously. Adv. Intell. Syst. 5, 2300140 (2023).
Maffettone, P. M. et al. Crystallography companion agent for high-throughput materials discovery. Nat. Comput. Sci. 1, 290–297 (2021).
Lee, J.-W., Park, W. B., Lee, J. H., Singh, S. P. & Sohn, K.-S. A deep-learning technique for phase identification in multiphase inorganic compounds using synthetic XRD powder patterns. Nat. Commun. 11, 86 (2020).
Pan, T., Jin, S., Miller, M. D., Kyrillidis, A. & Phillips, G. N. A deep learning solution for crystallographic structure determination. IUCrJ 10, 487–496 (2023).
Szymanski, N. J. et al. Adaptively driven X-ray diffraction guided by machine learning for autonomous phase identification. npj Comput. Mater. 9, 31 (2023).
Wang, H. et al. Rapid identification of X-ray diffraction patterns based on very limited data by interpretable convolutional neural networks. J. Chem. Inf. Model. 60, 2004–2011 (2020).
Chen, L. et al. Crystal structure assignment for unknown compounds from X-ray diffraction patterns with deep learning. J. Am. Chem. Soc. 146, 8098–8109 (2024).
Cao, B. et al. SimXRD-4M: big simulated X-ray diffraction data and crystal symmetry classification benchmark. In Proc. International Conference on Learning Representations (ICLR) (2025).
Salgado, J. E., Lerman, S., Du, Z., Xu, C. & Abdolrahim, N. Automated classification of big X-ray diffraction data using deep learning models. npj Comput. Mater. 9, 214 (2023).
Park, W. B. et al. Classification of crystal structure using a convolutional neural network. IUCrJ 4, 486–494 (2017).
Vecsei, P. M., Choo, K., Chang, J. & Neupert, T. Neural network based classification of crystal symmetries from x-ray diffraction patterns. Physical Review B 99, https://doi.org/10.1103/PhysRevB.99.245120 (2019).
Lee, B. D. et al. Powder X-Ray diffraction pattern is all you need for machine-learning-based symmetry identification and property prediction. Adv. Intell. Syst. 4, 2200042 (2022).
Xie, T. et al. CSU-MS2: a contrastive learning framework for cross-modal compound identification from MS/MS spectra to molecular structures. Anal. Chem. 97, 13350–13360 (2025).
Chen, C. & Ong, S. P. A universal graph deep learning interatomic potential for the periodic table. Nat. Comput. Sci. 2, 718 (2022).
Choudhary, K. & DeCost, B. Atomistic line graph neural network for improved materials property predictions. npj Comput. Mater. 7, 185 (2021).
Deng, B. W. et al. CHGNet as a pretrained universal neural network potential for charge-informed atomistic modelling. Nat. Mach. Intell. 5, 1031–1041 (2023).
Zeni, C. et al. A generative model for inorganic materials design. Nature 639 624–632 (2025).
Lai, Q. S. et al. End-to-end crystal structure prediction from powder X-Ray diffraction. Adv. Sci. 12, 2410722 (2025).
Zhang, X., Wang, X. T. & Hu, S. B. Structural fingerprinting of crystalline materials from XRD patterns using atomic cluster expansion neural network and atomic cluster expansion. Appl. Sci. 15, 5851 (2025).
Li, Q. et al. Powder diffraction crystal structure determination using generative models. Nature Communications 16, https://doi.org/10.1038/s41467-025-62708-8 (2025).
Jia, Z., Li, Z., Wang, Z., Liu, Z. & Vong, C.-M. Joint prediction of SOH and RUL for lithium batteries considering capacity self recovery and model drift. IEEE Internet Things J. 12, 22187–22196 (2025).
Bandyopadhyay, Y., Avlani, H. & Zhuang, H. L. Kolmogorov–Arnold neural networks for high-entropy alloys design. Modell. Simul. Mater. Sci. Eng. 33, 035005 (2025).
Wu, X. Y., Song, X. Y., Yue, Y. F., Zheng, R. & Jiang, J. W. MOF-KAN: Kolmogorov-Arnold networks for digital discovery of metal-organic frameworks. J. Phys. Chem. Lett. 16, 2452–2459 (2025).
Liu, Z. et al. KAN: Kolmogorov–Arnold Networks. In Proc. International Conference on Learning Representations (ICLR) (2025).
Thakolkaran, P. et al. Can KAN CANs? Input-convex Kolmogorov-Arnold Networks (KANs) as hyperelastic constitutive artificial neural networks (CANs). Comput. Methods Appl. Mech. Eng. 443, 118089 (2025).
Wang, Y. et al. Kolmogorov–Arnold-Informed neural network: a physics-informed deep learning framework for solving forward and inverse problems based on Kolmogorov–Arnold networks. Comput. Methods Appl. Mech. Eng. 433, 117518 (2025).
Wu, Y. et al. Kolmogorov-Arnold network made learning physics laws simple. J. Phys. Chem. Lett. 15, 12393–12400 (2024).
Graewert, M. A. & Svergun, D. I. Impact and progress in small and wide angle X-ray scattering (SAXS and WAXS). Curr. Opin. Struct. Biol. 23, 748–754 (2013).
Ge, J. et al. Correlations of multiscale structural evolution and homogeneous flows in metallic glass ribbons. Mater. Res. Lett. 11, 547–555 (2023).
Lundberg, S. M. et al. From local explanations to global understanding with explainable AI for trees. Nat. Mach. Intell. 2, 56–67 (2020).
Sundararajan, M., Taly, A. & Yan, Q. Q. Axiomatic attribution for deep networks. In Proc. 34th International Conference on Machine Learning Vol. 70 (PMLR, 2017).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In Proc. IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 770–778 (IEEE, 2016).
Dosovitskiy, A. et al. An image is worth 16×16 words: Transformers for image recognition at scale. In Proc. International Conference on Learning Representations (ICLR) (2021).
Rincón, S., González, G., Macías, M. A. & Arbeláez, P. A new benchmark for machine learning applied to powder X-ray diffraction. Sci. Data 12, 1186 (2025).
Ong, S. P. et al. Python materials genomics (pymatgen): a robust, open-source Python library for materials analysis. Comput. Mater. Sci. 68, 314–319 (2013).
Zunger, A., Wei, S. H., Ferreira, L. G. & Bernard, J. E. Special quasirandom structures. Phys. Rev. Lett. 65, 353–356 (1990).
van de Walle, A., Asta, M. & Ceder, G. The alloy theoretic automated toolkit: a user guide. Calphad 26, 539–553 (2002).
Hollarek, D. et al. opXRD: Open Experimental Powder X-Ray Diffraction Database. Advanced Intelligent Discovery. e2500044 https://doi.org/10.1002/aidi.202500044 (2025).
Jain, A. et al. Commentary: The Materials Project: A materials genome approach to accelerating materials innovation. APL Mater. 1, 011002 (2013).
Fredericks, S., Parrish, K., Sayre, D. & Zhu, Q. PyXtal: a Python library for crystal structure generation and symmetry analysis. Comput. Phys. Commun. 261, 107810 (2021).
Acknowledgements
This work was financially supported by the Advanced Materials-National Science and Technology Major Project (Grant No. 2025ZD0620100), National Natural Science Foundation of China (Grant No. 52401015), Shanghai Pujiang Program (Grant No. 23PJ1403500) and Shanghai Artificial Intelligence Open Source Award Project. We also sincerely thank Mr. Bin Cao from The Hong Kong University of Science and Technology (Guangzhou) for his assistance in organizing and openly releasing the experimental database.
Author information
Authors and Affiliations
Contributions
C. Xu performed model training and drafted the manuscript. T. Su contributed to data preprocessing and statistical analysis, and revised the manuscript. J. Xiong conceived the research idea, performed data analysis, reviewed and revised the manuscript, and supervised the project. Y. Wu participated in the model design and training. S. Chen, S. Dong, and T. Jiang participated in the interpretation and discussion of the results. M. He analyzed the results, co-conceived the research idea, and revised the manuscript. T.Y. Zhang reviewed the manuscript and provided overall supervision.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xu, C., Su, T., Xiong, J. et al. KAN-enhanced contrastive learning: the accelerator of crystal structure identification from XRD patterns. npj Comput Mater (2026). https://doi.org/10.1038/s41524-026-02015-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41524-026-02015-y