Abstract
The GRIN family is implicated in neurological disorders, such as global developmental delay (GDD) and epilepsy. We reviewed 31 patients with GRIN-related neurodevelopmental disorders at Seoul National University Hospital; all exhibited profound GDD, with 58.1% unable to walk independently and 74.2% unable to speak meaningful words. In a pooled analysis with the GRIN portal data (https://grin-portal.broadinstitute.org/), patients with missense or in-frame variants had significantly higher rates of profound GDD (74.3% vs. 30.4%, p < 0.001) and movement disorders (69.0% vs. 41.4%, p < 0.01) than those with protein-truncating variants. Furthermore, missense or in-frame variants in the M3 and M4 helices of the transmembrane domain were significantly associated with profound GDD (M3 helix: adjusted odds ratio [aOR] 8.48; 95% confidence interval [CI] 2.79–25.76; M4 helix: aOR 3.14; 95% CI 1.39–7.09) compared to those in other domains. Our findings highlight the importance of detailed variant characterization to inform personalized treatment strategies.
Similar content being viewed by others
Introduction
The N-methyl-D-aspartate (NMDA) receptor, a ligand-gated ion channel, regulates excitatory neurotransmission and contributes to higher cortical functions, such as learning and memory, by coordinating neuronal network activity within the synapse1,2. The GRIN gene family encodes two glycine-binding GluN1 subunits (GRIN1) and two glutamate-binding GluN2/3 subunits (GRIN2A, GRIN2B, GRIN2C, GRIN2D, and GRIN3) of the NMDA receptor. Among the GRIN genes, four are currently listed in the Online Mendelian Inheritance in Man (OMIM) database as causative genes for GRIN-related neurodevelopmental disorders (NDDs): GRIN1 (OMIM #614254), GRIN2A (OMIM #616139), GRIN2B (OMIM #613970), and GRIN2D (OMIM #617162)1,3,4. All disorders associated with these four genes follow an autosomal dominant inheritance pattern. The distribution of variant types varies by database, with more than 65% being missense and protein-truncating variants (PTVs) accounting for approximately 20%5.
The NMDA receptor subunits exhibit a modular domain architecture comprising amino-terminal domains (ATDs), ligand-binding domains (LBDs), transmembrane domains (TMDs), and intracellular carboxy-terminal domains (CTDs)6. The TMD functions as the pore-forming region, consisting of three transmembrane helices (M1, M3, and M4) and a re-entrant loop (M2). Although the molecular structures of the NMDA receptor subunits share common features2, each GRIN variant presents distinct neurological phenotypes, primarily attributed to differences in expression patterns. GRIN2B variants are associated with profound global developmental delay (GDD) and autism spectrum disorder (ASD), as GluN2B subunits are expressed during early postnatal development. In contrast, GRIN2A variants are highly associated with epilepsy, as GluN2A expression increases significantly upon birth7.
With advancements in next-generation sequencing (NGS), loci corresponding to specific domains of the NMDA receptor have been identified6. Recent findings by Xu et al.8 showed that the gain-of-function (GoF) variants in the M3 helix of the TMD are associated with neurological disorders, such as epilepsy and GDD. Additionally, the M4 helix plays a critical role in receptor function, including channel opening and transmembrane modulation1. Given that each domain encodes a protein with a distinct function, it is possible to infer the relationship between specific domains and phenotypes among the diverse neurological phenotypes of GRIN. Accordingly, we aimed to analyze the association between certain GRIN domains, particularly the M3 and M4 helices of the TMD, and neurological phenotypes. The recently developed GRIN Portal (https://grin-portal.broadinstitute.org/) has facilitated the exploration of genotype–phenotype correlations and the phenotypic features of GRIN variants. However, a comprehensive characterization of neurological phenotypes in relation to specific variant subtypes and locations remains limited.
This study was conducted in 31 Korean pediatric patients with pathogenic GRIN variants who visited a tertiary center with chief complaints of GDD or seizures. In this study, we conducted an in-depth description of the neurological phenotypes observed in these patients. Through a pooled analysis of previously reported GRIN variants from the GRIN Portal, we investigated the genotype–phenotype correlations in GRIN-related NDDs, with a particular focus on the relationship between variant location and the presence or severity of neurological phenotypes.
Results
Identification of GRIN mutations
We included 31 pediatric patients with 28 distinct GRIN variants, including 22 missense variants, one nonsense variant, two in-frame variants, and three large deletions (Table 1). The most prevalent GRIN variant was GRIN2B, followed by GRIN1, GRIN2A, and GRIN2D. Among these, 22 patients were confirmed as de novo mutations through trio analysis and were confirmed as pathogenic or likely pathogenic according to the American College of Medical Genetics (ACMG) criteria. One child (Seoul National University Hospital [SNUH] 31) with a GRIN2D variant was confirmed to have two discrete de novo missense variants in cis: p.Asn830Ser and p.Asn844His. Three recurrent GRIN2B variants were identified within the cohort: c.2459G>C (found in SNUH 13 and SNUH 19), c.2539C>T (found in SNUH 15 and SNUH 27), and c.2287G>A (found in SNUH 29 and SNUH 30, who are monozygotic twins).
Among the 25 GRIN variants (missense, in-frame, and PTVs) identified, 13 were not previously reported in ClinVar. All three deletions involved GRIN2B, with deletion sizes of 6.6 Mb (SNUH 14), 257 kb (SNUH 16), and 4.2 Mb (SNUH 18), respectively. When stratifying the 25 GRIN variants based on their localization within the NMDA receptor, 15 variants (six in S1 and nine in S2) were located in the LBD, whereas seven variants (three in M3 and four in M4) were found in the TMD. One mutation was identified in each of the following regions: CTD, M2-M3 linker, and S1-M1 linker.
Clinical characteristics of the SNUH cohort patients
The neurological phenotypes of patients are summarized in Table 2. Except for three patients (SNUH 1, SNUH 10, and SNUH 12) who presented with a chief complaint of seizures, the majority of patients visited the pediatric neurology clinic for GDD. Eighteen patients (58.1%) visited the pediatric neurology clinic before 12 months of age, and eight patients (25.8%) exhibited clinical seizures. The median observation period was 3 years (interquartile range [IQR]: 1.5–6.5 years). GDD was severe or profound, with 18 patients (58.1%) unable to walk independently and 23 (74.2%) unable to speak meaningful words. Additionally, nine patients (29.0%) had microcephaly (head circumference at age <3%), whereas one had macrocephaly (head circumference at age >97%). Eight patients had abnormal brain magnetic resonance imaging (MRI) findings, including one patient with polymicrogyria and two patients with delayed myelination. Lastly, one patient showed hypothyroidism (SNUH 10).
Eight patients in the SNUH cohort experienced clinical seizures, and their phenotypes are summarized in Table 3. The median age of seizure onset was 14 months (IQR: 9–31 months). Seizure types and electroencephalogram (EEG) findings varied among patients. Three patients achieved seizure freedom, with only one patient successfully tapering off antiseizure medication (ASM). Notably, four patients had abnormal EEG findings without clinical seizures. The most frequently prescribed initial ASM was valproic acid. However, drug-resistant epilepsy was common, with persistent epileptiform discharges observed in most patients during the last follow-up EEG. Responsive therapies included vigabatrin, valproic acid, oxcarbazepine, and the ketogenic diet.
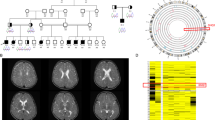
Representative cases of GRIN-related NDDs are presented in Fig. 1. Figure 1a shows a patient who visited our clinic with a chief complaint of GDD, along with seizures and microcephaly. Initial brain MRI revealed delayed myelination, whereas follow-up imaging revealed diffuse brain atrophy. Trio whole-exome sequencing (WES) identified that the patient had a de novo missense mutation in the GRIN1 gene (c.2443G>C) located within the M4 helix of the TMD. The patient expired with an unknown etiology at the age of 2 years and 11 months of age. Figure 1b shows monozygotic twins with the chief complaint of GDD. Notably, both patients presented very similar MRI findings, including nodular T2 hyperintensities in the brainstem (SNUH 29 and SNUH 30) and cerebellum (SNUH 30). They both presented with profound GDD with an inability to speak meaningful words and walk independently.
a Clinical information of SNUH 2, who presented with a chief complaint of profound GDD at four months of age. The patient was found to have a de novo GRIN1 missense variant located in the M4 helix of TMD (c.2443G>C, p.Gly815Arg). An initial brain MRI acquired at four months of age showed delayed myelination, while follow-up imaging at two years old showed diffuse brain atrophy. b Clinical information of SNUH 29 and SNUH 30, monozygotic twins presenting with a chief complaint of profound GDD. Quartet whole-exome sequencing analysis identified a de novo GRIN2B missense variant (c.2287G>A, p.Gly763Ser) in both twins. Brain MRI acquired at 12 months old revealed nodular T2 hyperintensities in the brainstem (SNUH 29, SNUH 30) and cerebellum (SNUH 30). TMD transmembrane domain, GDD global developmental delay, MRI magnetic resonance imaging.
Association between neurologic features and GRIN variant subtypes
The most prominent neurological phenotype was profound GDD (239 out of 349, 68.5%), with the highest prevalence in GRIN1 (98 out of 123, 79.7%), followed by GRIN2D (14 out of 20, 70.0%) and GRIN2B (96 out of 138, 69.6%). Seizure prevalence varied by gene type and was the highest in patients with GRIN2D (20 out of 20, 100%) and GRIN2A (56 out of 68, 82.4%) variants, followed by those with GRIN1 (71 out of 123, 57.7%) and GRIN2B (49 out of 138, 35.5%) variants (Fig. 2a and Supplementary Table 1). The neurological phenotypes of GRIN missense and in-frame variants are shown in Supplementary Fig. 1. The prevalence of neurological phenotypes was comparable across all GRIN gene types when limited to missense and in-frame variants. For instance, the prevalence of profound GDD in GRIN2A variants was the lowest at 45.6%, but increased to 60.4% when limited to missense and in-frame variants, making neurological phenotype prevalence comparable across GRIN gene types. Additionally, the prevalence of neurological phenotypes such as profound GDD (74.3% vs. 30.4%, p < 0.001), movement disorder (MD) (69.0% vs. 41.4%, p < 0.01), and malformation of cortical development (MCD) (21.4% vs. 0%, p < 0.01) was significantly higher in patients with missense or in-frame variants than in those with PTVs (Fig. 2b). The above analyses were conducted using only variants classified as pathogenic or likely pathogenic.
Prevalence of neurological phenotypes stratified by a gene type and b variant type. *p < 0.05. ASD autism spectrum disorder, GDD global developmental delay, MD movement disorder, CVI cortical visual impairment, MCD malformation of cortical development, PTV protein-truncating variant.
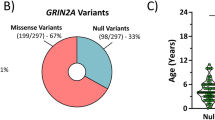
The distribution and characteristics of GRIN variants are summarized in Fig. 3. Among the 505 GRIN variants pooled from the ClinVar and SNUH cohorts, GRIN2A variants were the most frequent (211/505, 41.8%), followed by GRIN2B (199/505, 39.4%) (Fig. 3a). PTVs were most prevalent in patients with GRIN2A (99/214, 46.3%), followed by those with GRIN2B (56/196, 28.6%). In contrast, patients with GRIN1 variants exhibited a lower prevalence of PTVs (7.5%), whereas no PTVs were reported among those with GRIN2D variants (Fig. 3b). Mapping of the missense and in-frame variant loci revealed clustering within regions encoding the M3 and M4 helices of the TMD (Fig. 3c and Supplementary Fig. 2).
a A pie chart illustrating the proportion of GRIN variants categorized by gene type. b The number and proportion of pathogenic variants (PTVs vs. missense or in-frame variants in other domains vs. missense or in-frame variants in the M3–M4 helices). c Genomic locations of pathogenic variants identified in GRIN1, GRIN2A, GRIN2B, and GRIN2D. The blue density plots represent the domain-specific distribution of missense or in-frame variants from the GRIN Portal and the SNUH cohort. Lollipops in the upper row indicate variant information from the SNUH cohort. Lollipops in the bottom row indicate pathogenic/likely pathogenic missense or in-frame variants reported in ClinVar. PTV protein-truncating variant.
Figure 4 shows the phenotypic profiles of the GRIN Portal and SNUH cohorts (Supplementary Table 2). We examined 349 patients with GRIN-related NDDs, including 123 with GRIN1 variants, 68 with GRIN2A variants, 138 with GRIN2B variants, and 20 with GRIN2D variants. Patients with missense or in-frame variants in the M3–M4 helices of the TMD had the highest prevalence of various neurological phenotypes, except for those with ASD (Fig. 4a). Patients with M3 helix variants had the highest risk of profound GDD (adjusted odds ratio [aOR]: 8.48, 95% confidence interval [CI]: 2.79–25.76), ataxia (aOR: 5.33, 95% CI: 1.34–21.12), and cortical visual impairment (CVI) (aOR: 3.89; 95% CI: 1.60–9.48). Patients with M4 helix variants had the highest risk of MD (aOR: 11.40, 95% CI: 2.58–50.37) (Fig. 4b).
a Comparisons among M3–M4 helices, other domains, and PTVs. b Prevalence and risk of neurological phenotypes across missense or in-frame variant domains (other domains, M3 helix, and M4 helix of each GRIN gene) presented as odds ratios with 95% confidence intervals, adjusted for GRIN gene subtypes. ap < 0.05 for comparisons between missense or in-frame variants in M3–M4 helices and those in other domains. bp < 0.05 for comparisons between in-frame variants in the M3–M4 helices and PTVs. cp < 0.05 for comparisons between missense or in-frame variants in other domains and PTVs. *p < 0.05. GDD global developmental delay, ASD autism spectrum disorder, MD movement disorder, CVI cortical visual impairment, MCD malformation of cortical development.
Discussion
This study investigated an in-depth analysis of the clinical phenotypes associated with GRIN-related NDDs in the SNUH and GRIN Portal cohorts. The most consistent finding across the cohort was profound GDD. The neurological manifestations were diverse, including ASD and MD, with genotype-specific patterns and significant associations with variant localization within the NMDA receptor.
Profound GDD was the most common phenotype observed, which is consistent with the findings of previous studies9,10. However, earlier studies often focused solely on investigating patients exhibiting intellectual disability and speech problems, without considering the severity of GDD. Our findings highlight that most patients present severe developmental challenges from infancy, with the majority failing to achieve basic milestones, such as independent walking and speaking simple words by 12 months of age. Notably, some patients experienced developmental regression, further underscoring the profound impact of GRIN variants on neurodevelopment. Furthermore, most children exhibited severe cognitive impairment that could not carry out full-scale intelligence quotient testing. Additionally, brain MRI imaging, performed as part of the diagnostic process, did not reveal consistent structural abnormalities.
In addition to profound GDD, phenotypic manifestations varied by gene. Seizures were more prominent in patients with GRIN2A and GRIN2D variants, whereas MD was more prominent in patients with GRIN1 variants. Sapuppo et al.11 demonstrated that GRIN2A variants were linked to Landau-Kleffner syndrome, whereas GRIN2B variants were linked to West syndrome. However, we were unable to identify any specific epilepsy patterns in the SNUH cohort. This study highlights the neurological phenotypic overlap among pediatric patients with GRIN variants and suggests genotype-specific differences in detailed phenotypes. Notably, the differences in variant type distribution, such as the PTVs primarily observed in patients with GRIN2A and GRIN2B variants, may have contributed to the clinical differences between the genes (Fig. 2 and Supplementary Fig. 1).
Previous studies have shown that patients with GRIN PTVs have a significantly lower risk of severe intellectual disability compared to those with missense variants3,12. Notably, Santos-Gomez et al. investigated the clinical characteristics of patients with GRIN2A and GRIN2B PTVs, reporting that GRIN2A variants were associated with seizures, whereas GRIN2B variants were linked to ASD12. In line with our findings, patients with missense or in-frame variants exhibited a broader range of neurological phenotypes—including not only GDD and seizures, but also MD and MCD—compared to those with PTVs3,12. By integrating genotypic and phenotypic information from 349 patients with GRIN variants (including 46 with PTVs), our study expands the current understanding of GRIN-related NDD phenotypes according to variant type.
We analyzed the detailed responses to ASM in the SNUH cohort. Our results suggest that medications, such as vigabatrin and valproic acid, may be effective options for managing seizures in GRIN-related NDDs. However, due to the lack of ASM response data in the GRIN Portal, a more detailed analysis based on variant subtypes was not available. To enable personalized medicine and tailor therapeutic options for GRIN-related NDDs, a comprehensive database containing detailed clinical information must be developed.
The differences in the prevalence of neurological phenotypes among GRIN gene subtypes may be attributed to the distinct spatiotemporal expression patterns of the GRIN gene2. GluN2A, encoded by GRIN2A, is initially expressed postnatally and becomes abundant during brain maturation. On the other hand, GluN2B, encoded by GRIN2B, is widely expressed during embryogenesis and remains highly prominent in the forebrain during early development. Nevertheless, as shown in Fig. 2a and Supplementary Fig. 1, gene-specific characteristics were not clearly defined, and the prevalence of each phenotype appeared more comparable when restricted to missense and in-frame variants.
As depicted in Fig. 3c, pathogenic and likely pathogenic variants tended to be located adjacent to the TMD domain helices M3 and M4, particularly in GRIN1 and GRIN2D3. Myers et al.7 showed that this critical region is under greater selection pressure and is highly intolerant to genetic variations. Korinek et al. demonstrated that missense variants located adjacent to the ABD and TMD tend to affect glutamate or glycine affinity13. These changes in affinity may contribute to either GoF or loss-of-function (LoF) of NMDA receptors, potentially leading to various neurological symptoms. We observed that the prevalence of profound GDD did not differ significantly by GRIN gene subtype but varied considerably by missense or in-frame variant domains. Notably, the severity and diversity of neurological phenotypes were more pronounced in missense or in-frame variants within the M3 and M4 helices than in the PTVs. This finding is in line with that of a recent study by Xu et al.8, which predicted that most variants within the M3 helix would be dysfunctional and pathological. Among the 48 variants located in the M3 domain, functional analysis revealed that 28 exhibited GoF, whereas only nine showed LoF, collectively suggesting that these variants enhanced channel gating activities. The M3 helix, which is directly linked to the ABD domain, plays a pivotal role in controlling channel opening and closing8,14. Although studies on the role of the M4 domain in channel function are currently limited to receptor modulation, our findings emphasize the clinical significance of the M4 domain in the neurological phenotypes of GRIN-related NDDs.
Our study, which included a large cohort of 349 patients, supports previous findings on the clinical relevance of missense variants in the M3 and M4 helices1,8. Patients with missense variants in affected domains exhibited a higher incidence of neurological phenotypes, such as profound GDD, MD, and ataxia, than those with variants in other domains. Additionally, we observed distinct phenotypic differences between the two helices, with the risk of seizures and ataxia being more prominent in the M3 helix. Although the underlying mechanisms remain unclear, our findings highlight the importance of domain-level interpretation, which may support phenotype prediction and the implementation of precision medicine in GRIN-related NDDs.
Based on the characteristics of the GRIN gene, previous studies have explored the use of NMDA receptor blockers (e.g., ketamine and memantine) in patients with GRIN-related NDD15,16. Nevertheless, as both GoF and LoF variants of GRIN2A result in prolonged NMDAR-mediated synaptic current decay, their clinical applications warrant further investigation. Additionally, with regard to the functional differences among the domains, previous studies have suggested that the M3 helix increases the potency of glutamate and glycine, whereas the M4 helix interacts with the M1 and M3 helices to modulate receptor activity. However, further research is required to better understand these mechanisms1,8,14.
As demonstrated in representative cases, patients with GRIN-related NDDs typically present with profound GDD during infancy, which is characterized by limited developmental progress. Previous studies have reported that the MRI findings of patients with GRIN-related NDDs include cortical atrophy and increased white matter signal intensity in the periventricular white matter17,18. To our knowledge, the MRI findings shown in Fig. 1b have not been previously reported. We hypothesize that these features may be attributed to a genetic etiology and could be distinctive characteristics of GRIN variants. Notably, identical findings were observed in the monozygotic twins, further supporting the genetic basis of these manifestations.
The strength of our study lies in the detailed phenotyping of patients with GRIN variants from the SNUH cohort, including 13 with previously unreported variants. Our findings expand our understanding of the phenotypic spectrum and phenotype-genotype correlations in GRIN-related NDDs. To our knowledge, this study is the first to present a series of cases involving pediatric patients with GRIN variants and neurological phenotypes in an Asian population. Our study has some limitations, including the lack of functional validation and the small sample size, which limit the generalizability of the findings to the full spectrum of GRIN-related NDDs. However, our detailed phenotyping, including ASM responses and developmental milestones, provides clinically relevant insights for the personalized management of patients with GRIN-related NDDs.
This study included a total of 349 patients by combining the SNUH cohort and the GRIN Portal dataset. Although our study represents one of the largest datasets of GRIN-related NDD, there remains considerable potential to incorporate a much larger number of patients and variants. As observed in the SNUH cohort, where more than 50% of the variants were novel, a substantial number of pathogenic variants likely remain unreported. Furthermore, a considerable proportion of pathogenic or likely pathogenic variants previously submitted to ClinVar have not yet been incorporated into the GRIN Portal. Even among those included in the GRIN Portal, over 200 variants were excluded due to insufficient clinical information. In particular, the rate of missing data for ASD, MD, ataxia, and CVI was notably higher than that of seizures and GDD. These findings highlight the need to develop a more comprehensive database that encompasses a broader spectrum of genotypes and phenotypes. Further research utilizing large-scale datasets is warranted to better elucidate genotype–phenotype correlations in GRIN-related NDDs.
In conclusion, this study presents the genetic and clinical presentations of GRIN-related NDDs. GRIN variants are associated with profound GDD accompanied by various neurological manifestations. Patients with pathogenic missense or in-frame variants in the M3 and M4 helices of the TMD exhibited significantly distinctive phenotypes. The evaluation of the specific variant domains of GRIN mutations provides valuable insights into the clinical characteristics.
Methods
Study participants
We enrolled 31 patients with genetically confirmed GRIN mutations (8 with GRIN1, 4 with GRIN2A, 18 with GRIN2B, and 1 with GRIN2D). All patients were evaluated at the Pediatric Neurology Clinic of SNUH. The study was performed in accordance with the ethical standards of the Declaration of Helsinki and approved by the Institutional Review Board of SNUH (H-2408-063-1559). The requirement for informed consent was waived, as the research involved minimal risk and used de-identified retrospective data. All participants were unrelated, except for a pair of monozygotic twins who shared the same GRIN2B pathogenic variant (SNUH 29 and SNUH 30).
GRIN variants were distributed across the GRIN1, GRIN2A, GRIN2B, and GRIN2D genes. The clinical information of the SNUH cohort was retrospectively reviewed using electronic medical records, including neurological phenotypes such as GDD, MD, and ASD. Neurodevelopmental status was evaluated at the initial and most recent visits. Additional clinical data included EEG and brain MRI results. The detailed seizure characteristics of eight patients with epilepsy, including semiology and ASM usage, were also collected.
Genetic diagnosis
The GRIN variants in the SNUH cohort were identified using NGS. Of them, 23 patients were analyzed by trio WES, five by singleton WES, and three by chromosomal microarray (CMA). WES was performed at our institution using the Illumina technology. The generated reads were aligned to the human reference genome (hg38) and processed according to the Genome Analysis Toolkit best practice guidelines19. Then, the variants were annotated using the ANNOVAR program, with a focus on rare protein-altering variants with a population allele frequency of <0.001% based on the Genome Aggregation Database (gnomAD)20,21. The Human Gene Mutation Database and ClinVar database were searched for previously reported variants22,23. Variants were interpreted using the criteria established by the American College of Medical Genetics and Genomics/Association for Molecular Pathology (ACMG/AMP) criteria24. All subsequent analyses, including genotype–phenotype correlations, were conducted using only variants classified as pathogenic or likely pathogenic. The detailed WES analysis process was described in our previous study25.
For subsequent analyses, the missense and in-frame variants were classified based on their respective domains. Domain information, such as ATDs, LBDs (S1 and S2), TMD (M1, M2, M3, and M4 helices), and CTD, was obtained from the UniProt database26.
Pooled analysis of GRIN variants
We collected variants associated with known clinical phenotypes of the GRIN family (GRIN1, GRIN2A, GRIN2B, and GRIN2D) from the GRIN Portal (https://grin-portal.broadinstitute.org/; accessed July 1, 2024). The GRIN Portal dataset included 632 patients: 215 with GRIN1, 156 with GRIN2A, 229 with GRIN2B, and 32 with GRIN2D variants. Given that seizures and GDD are the most prominent neurological phenotypes of GRIN-related NDDs, patients without identifiable information on these features were excluded from the analysis. As a result, 321 patients (226 unique variants) remained in the final dataset. Among these 226 variants, 55 (24.3%) were not listed in the ClinVar database (as of December 2024). Notably, only 44.1% (216 out of 490) of the GRIN-related variants in ClinVar were represented in the GRIN Portal. Data on the presence of neurological features (seizure, GDD, MD, ASD, ataxia, CVI, MCD), and genotype were extracted and combined with the SNUH cohort.
Patients were categorized into two groups based on variant type: (1) PTVs and (2) missense or in-frame variants. In addition, we hypothesized that pathogenic and likely pathogenic variants are clustered in specific domains, particularly in the M3 and M4 helices of TMD, are highly associated with neurological phenotypes, and display genotype-specific characteristics. Missense or in-frame variants were subsequently stratified into three categories based on domain location: (1) variants in other domains, (2) variants in the M3 helix of the TMD, and (3) variants in the M4 helix of the TMD. Using data from the GRIN Portal and our institution, we analyzed the prevalence of each neurological phenotype in relation to genetic variants and domain-specific localizations.
Statistical analysis
The prevalence of each neurological phenotype by genotype was compared using Fisher’s exact test or the chi-squared test, as appropriate. Comparison among the three groups was performed using the chi-squared test, with Fisher’s exact test applied for post-hoc analysis. Logistic regression analysis was conducted to assess the association between neurological phenotypes and variant domains, with a focus on variants located in the M3 and M4 helices of the TMD.
The associations were expressed as aOR and 95% CIs, adjusted for gene subtype. All analyses were performed using R (version 4.0.4; R Foundation for Statistical Computing) and GraphPad Prism version 8.0 (GraphPad Software), with a significance level of p < 0.05 considered significant. GRIN variants from the SNUH cohort and ClinVar databases were visualized using the PeCan protein viewer27. The identified variants were annotated in accordance with the Human Genome Variation Society guidelines28.
Data availability
The datasets analyzed during the current study are available from the corresponding author upon reasonable request.
References
Langer, K., Müller-Längle, A., Wempe, J. & Laube, B. Analysis of M4 transmembrane segments in NMDA receptor function: a negative allosteric modulatory site at the GluN1 M4 is determining the efficiency of neurosteroid modulation. Front. Pharmacol. 12, 769046 (2021).
Paoletti, P., Bellone, C. & Zhou, Q. NMDA receptor subunit diversity: impact on receptor properties, synaptic plasticity and disease. Nat. Rev. Neurosci. 14, 383–400 (2013).
Platzer, K. et al. GRIN2B encephalopathy: novel findings on phenotype, variant clustering, functional consequences and treatment aspects. J. Med. Genet. 54, 460–470 (2017).
Strehlow, V. et al. GRIN2A-related disorders: genotype and functional consequence predict phenotype. Brain 142, 80–92 (2019).
XiangWei, W., Jiang, Y. & Yuan, H. De novo mutations and rare variants occurring in NMDA receptors. Curr. Opin. Physiol. 2, 27–35 (2018).
Lee, C.-H. et al. NMDA receptor structures reveal subunit arrangement and pore architecture. Nature 511, 191–197 (2014).
Myers, S. J. et al. Distinct roles of GRIN2A and GRIN2B variants in neurological conditions. F1000Research 8 (2019).
Xu, Y. et al. De novo GRIN variants in M3 helix associated with neurological disorders control channel gating of NMDA receptor. Cell. Mol. Life Sci. 81, 153 (2024).
Benke, T. A. et al. Clinical and therapeutic significance of genetic variation in the GRIN gene family encoding NMDARs. Neuropharmacology 199, 108805 (2021).
García-Recio, A. et al. GRIN database: a unified and manually curated repertoire of GRIN variants. Hum. Mutat. 42, 8–18 (2021).
Sapuppo, A. et al. GRIN2A and GRIN2B and their related phenotypes. J. Pediatr. Neurol. 21, 212–223 (2023).
Santos-Gómez, A. et al. Disease-associated GRIN protein truncating variants trigger NMDA receptor loss-of-function. Hum. Mol. Genet. 29, 3859–3871 (2020).
Korinek, M. et al. Disease-associated variants in GRIN1, GRIN2A and GRIN2B genes: insights into NMDA receptor structure, function, and pathophysiology. Physiol. Res. 73, S413 (2024).
Li, J. et al. De novo GRIN variants in NMDA receptor M2 channel pore-forming loop are associated with neurological diseases. Hum. Mutat. 40, 2393–2413 (2019).
Eiler, I. et al. GRIN1-related epilepsy in a neonate with response to memantine and vigabatrin. Ann. Child Neurol. Soc. 2, 299–302 (2024).
Xu, Y. et al. Recurrent seizure-related GRIN1 variant: molecular mechanism and targeted therapy. Ann. Clin. Transl. Neurol. 8, 1480–1494 (2021).
Pironti, E. et al. Electroclinical history of a five-year-old girl with GRIN1-related early-onset epileptic encephalopathy: a video-case study. Epileptic Disord. 20, 423–427 (2018).
XiangWei, W. et al. Heterogeneous clinical and functional features of GRIN2D-related developmental and epileptic encephalopathy. Brain 142, 3009–3027 (2019).
McKenna, A. et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Landrum, M. J. et al. ClinVar: improving access to variant interpretations and supporting evidence. Nucleic Acids Res. 46, D1062–D1067 (2018).
Stenson, P. D. et al. The Human Gene Mutation Database: towards a comprehensive repository of inherited mutation data for medical research, genetic diagnosis and next-generation sequencing studies. Hum. Genet. 136, 665–677 (2017).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405–423 (2015).
Lee, S. et al. TNNT1 myopathy with novel compound heterozygous mutations. Neuromuscul. Disord. 32, 176–184 (2022).
Consortium, U. UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res. 47, D506–D515 (2019).
McLeod, C. et al. St. Jude Cloud: a pediatric cancer genomic data-sharing ecosystem. Cancer Discov. 11, 1082–1099 (2021).
Den Dunnen, J. T. et al. HGVS recommendations for the description of sequence variants: 2016 update. Hum. Mutat. 37, 564–569 (2016).
Acknowledgements
We sincerely acknowledge the GRIN Portal database (https://grin-portal.broadinstitute.org/) for systematically compiling and making available comprehensive genetic and clinical information on patients with GRIN-related neurodevelopmental disorders. This research was supported by a grant from the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea (grant number: RS-2023-00265923), and by the SNUH Lee Kun-hee Child Cancer & Rare Disease Project, Republic of Korea (grant number: 22B-001-0100).
Author information
Authors and Affiliations
Contributions
Conceptualization: J.H.C., S.L., and J.-H.C.; methodology: J.M.K., H.-J.Y., and H.C.; investigation: J.H.C.; validation: S.L.; formal analysis: J.H.C., J.M.K., H.-J.Y., and H.C.; access and verification of the data: H.J.K., S.Y.K., B.C.L., K.J.K., and S.L.; data curation: J.H.C, S.L.; writing – original draft preparation: J.H.C., S.L.; writing – review and editing: W.K., S.Y.K., B.C.L., K.J.K., and J.-H.C.; visualization: J.H.C., S.L.; supervision: W.K, S.Y.K., B.C.L., K.J.K.; funding acquisition: S.L., B.C.L., J.-H.C. The authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cha, J.H., Kim, J.M., Yun, HJ. et al. Exploring gene-phenotype relationships in GRIN-related neurodevelopmental disorders. npj Genom. Med. 10, 40 (2025). https://doi.org/10.1038/s41525-025-00499-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41525-025-00499-z