Abstract
Nineteen genomic regions have been associated with high-grade serous ovarian cancer (HGSOC). We meta-analyzed >22 million variants for 398,238 women from the Ovarian Cancer Association Consortium (OCAC), UK Biobank (UKBB) and Consortium of Investigators of Modifiers of BRCA1/BRCA2 (CIMBA) to identify novel HGSOC susceptibility loci. Eight novel variants were associated with HGSOC risk. An interesting discovery biologically was TP53 3’-UTR SNP rs78378222-T’s association with HGSOC (per-T-allele relative risk (RR) = 1.44, 95% CI:1.28–1.62, P = 1.76 × 10−9). Polygenic scores (PGS) were developed using OCAC and CIMBA data and trained on FinnGen data. The optimal PGS included 64,518 variants and was associated with an odds ratio of 1.46 (95% CI:1.37–1.54) per standard deviation when validated in the UKBB. This study represents the largest HGSOC GWAS to date – demonstrating that improvements in imputation reference panels and increased sample sizes help to identify HGSOC associated variants that previously went undetected, ultimately improving PGS which can improve personalized HGSOC risk prediction.
Similar content being viewed by others
Introduction
Globally, epithelial ovarian cancer (EOC) is the seventh most common cancer diagnosed in women, with ~314,000 new cases diagnosed each year1. It is the most lethal gynecological cancer, responsible for ~207,000 deaths annually1. EOC is a collection of five major histotypes, namely high-grade serous (HGSOC), endometrioid, clear cell, low-grade serous (LGS) and mucinous, which are thought to have distinct etiology2. HGSOC is the most prevalent accounting for 60–70% of EOC diagnoses2,3, and accounting for most EOCs diagnosed in BRCA1 and BRCA2 pathogenic variant (PV) carriers4,5,6,7,8,9,10,11. Furthermore, HGSOC accounts for the majority of EOC mortality12,13.
To date, 40 genomic regions associated with EOC have been identified through genome-wide association studies (GWAS)14,15,16,17,18,19,20,21,22,23,24,25,26,27,28. For 19 of these regions, HGSOC is the most strongly associated histotype14,15,16,17,18,19,20,21,22,26,27,28. These studies have relied on imputation efforts that used the 1000 Genomes Project29 and Haplotype Reference Consortium30 reference panels, yielding up to ~11 million well-imputed genetic variants. The Trans-Omics for Precision Medicine (TOPMed) reference panel31 and imputation server32 have recently become publicly available. The TOPMed panel consists of ~308 million variants, yielding greater genomic coverage than previously available reference panels, with the added benefit of containing many more low-frequency and rare variants. This prompted us to re-impute genetic variant data from the population-based Ovarian Cancer Association Consortium (OCAC)33, and BRCA1/2 carriers from the Consortium of Investigators of Modifiers of BRCA1 and BRCA2 (CIMBA)34,35 to assess whether the larger coverage of the genome from the TOPMed reference panel leads to detection of novel loci associated with HGSOC risk. We additionally made use of the UK Biobank (UKBB)36,37 to boost the sample size and power to detect associations. We combined these data with summary statistics from FinnGen38,39 and BioBank Japan40,41 to develop and validate polygenic models (PGM) and scores (PGS) for non-mucinous OC.
Results
The genome-wide association analyses for HGSOC were based on up to 398,238 women from OCAC (N = 120,248, 30.2%), UKBB (N = 245,377, 61.6%) and CIMBA (N = 32,613, 8.2%) (Table 1, Supplementary Tables 1, 2). A total of 19,883 (5.0%) women were classified as being diagnosed with HGSOC. The mean (standard deviation, SD) age at diagnosis for women in OCAC and UKBB were 60.2 (10.9) years and 63.5 (10.0) years, respectively. The mean (SD) censoring ages for BRCA1 and BRCA2 PV carriers were 43.7 (SD = 12.0) years and 46.2 (SD = 12.9) years, respectively.
Re-examining previously identified associations with EOC
We looked up the associations for the lead variants previously reported as being associated with EOC in our newly generated results (Supplementary Table 3, Supplementary Figs. 1–41). Most lead variants previously reported to be associated specifically with HGSOC risk replicated in the present meta-analysis of OCAC, UKBB and BRCA1/2 carriers at the significance threshold P < 5 × 10−8. Exceptions were chr2:111138666 (rs17041869)26, chr2:113216387 (rs895412), chr11: 62126500 (rs7937840)26 and chr22:28538325 (rs6005807)22 (Supplementary Table 3). However, the chr2:110525257..111658369 and chr2:112716387..113716387 regions contained other variants that were associated at the genome-wide significance level in the present analysis, whilst the chr11:61626500..62626500 and chr22:28038325..29038325 regions did not contain any variants associated with HGSOC at the genome-wide significance level. It should be noted, however, that the chr2:111138666 (rs17041869) and chr11:62126500 (rs7937840) variants were identified through a cross-cancer (breast, ovarian, and prostate) GWAS26 and were not specifically identified as HGSOC associated variants.
Novel loci associated with HGSOC
Associations with a total of 5786 variants from 44 loci were significant at P < 5 × 10−8. We excluded 5778 variants at 37 loci from further consideration, as they were either near known associated regions (Supplementary Table 3), were not conditionally independent of the lead variant in a nearby known region, or were likely statistical artifacts arising from strata specific effects.
There were eight variants associated (P < 5 × 10−8) with HGSOC, at 5q11, 6p12, 8p21, 9p24-23, 16q22, 17p13 and 19q12 (Table 2, Fig. 1, Supplementary Tables 4, 5).
The dashed line is the genome-wide statistical significance level (P = 5 × 10−8). The eight independent genome-wide statistically significant variants at seven novel loci are shown as red triangles.
The associations at 5q11 (rs528577783-G; RR = 5.15, 95% CI:2.96–8.96), 6p12 (rs1013698558-T; RR = 2.35, 95% CI:1.76–3.14), 8p21 (rs540569242-A; RR = 25.60, 95% CI:9.32–70.31), and 9p24-p23 (rs768719522-T; RR = 10.04, 95% CI:4.38-22.99) were all single rare variants (MAF ≤ 0.15%) associated with large HGSOC effects. The single SNPs associated with HGSOC at 16q22 (rs6979-G; RR = 1.07, 95% CI:1.04-1.09) and 19q12 (rs62107113-A; RR = 1.08, 95% CI:1.05–1.11) were common and conferred modest effects on HGSOC risk. There were two moderately correlated (TOPMed European42 r2 = 0.46, D’ = 0.89) low-frequency (MAF: 1.2% and 2.1%) variants at the 17p13 locus. The TNFS13/TNFSF12-TNFSF13 intronic variant rs143094271-G was associated with a per-allele RR = 1.28 (95% CI:1.18–1.39, P = 7.61 × 10−9); and rs78378222-T, a TP53 3’-UTR variant, with a per-allele RR = 1.44 (95% CI:1.28–1.62, P = 1.76 × 10−9). The association effect size estimates at 16q22, 19q12 and 17p13 were consistent between OCAC/UKBB and BRCA1/2 PV carriers.
Bayesian false-discovery probabilities (BFDPs) indicated that six of these associations are likely to be true, although two rare variants, rs540569242 and rs768719522, had noticeably larger BFDPs (Table 2). Under a model assuming 1:1,000 truly associated variants, the BFDPs were 3.2% for rs540569242 and 11% for rs768719522. The other variants all had BFDP ≤ 0.42%.
Credible causal variants (CCVs)
We defined 52 CCVs across the seven novel regions (Supplementary Table 6, Supplementary Figs. 42–48). Four regions (5q11, 6p12, 8p21, 9p24-23) had only the lead variant as a CCV, whilst the 16q22 (N = 5), 17p13 (N = 3) and 19q12 (N = 40) loci had several CCVs.
Association of the PGS with HGSOC
Of the 1102 PGMs developed using OCAC and CIMBA data, the PGM that performed best in the FinnGen data comprised of 64,518 variants (Supplementary Data 1), denoted PGS64518. In the UKBB validation, the OR per SD of PGS64518 was estimated to be 1.46 (95% CI:1.37–1.54), with discriminatory ability of AUROC = 0.607 (95% CI:0.590–0.623) (Table 3). The association of PGS64518 was strongly attenuated in the BBJ validation (East Asian ancestry women), where the OR per SD was 1.12 (95% CI:1.05–1.20).
When restricting the PGS to include only genotyped variants from the 64,518 genotyped and imputed variants, which may make their implementation easier, PGS with 5957 (all genotyped variants from the 64,518) and 400 variants had similar performance characteristics. Relative to the PGS64518, a PGS considering the 400 most strongly associated genotyped variants, denoted PGS400, resulted in a small decrease in the AUROC to 0.603, and a marginally attenuated OR per SD (OR = 1.43, 95% CI:1.35–1.52) in the UKBB.
Predicted absolute risks for the general population and BRCA1/2 pathogenic variant carriers
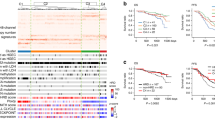
Absolute lifetime risks of developing EOC by age 80 years for a woman from the general population were calculated to be 0.9%, 1.6% and 3.0% at the 5th, 50th and 95th percentiles of the PGS64518 (Table 3, Fig. 2). The absolute lifetime risks (at age 80 years) for BRCA1 PV carriers were predicted to be 25.9%, 42.8% and 64.7% at the 5th, 50th and 95th percentiles of the PGS64518 distribution, respectively (Table 3, Fig. 2). The corresponding risks for PGS64518 for BRCA2 PV carriers at the same PGS percentiles were predicted to be 9.3%, 16.7% and 28.9%, respectively (Table 3, Fig. 2). The range of predicted percentile specific risks for the previously published 36 variant PGS43 was narrower, with risks for the same percentiles of 27.7%, 43.0% and 62.3% for BRCA1 PV carriers, respectively, and 10.1%, 16.9% and 27.5% for BRCA2 PV carriers, respectively. The PGS400 yielded absolute risks which were approximately at the midpoint of the 36 and 64,518 variant PGS absolute risks (BRCA1 PV carriers: 26.7%, 42.9% and 63.6% at the 5th, 50th and 95th percentiles, respectively; BRCA2 PV carriers: 9.7%, 16.8% and 28.3% at the 5th, 50th and 95th percentiles, respectively).
Predicted cumulative risks of developing EOC based on the PGS64518 at various percentiles of the PGS distribution for: a the general population (0.7% for 1st percentile to 3.9% for the 99th percentile), b BRCA1 PV carriers (20.7% for 1st percentile to 74.1% for the 99th percentile), and c BRCA2 PV carriers (7.3% for 1st percentile to 35.7% for the 99th percentile).
There was a total reclassification of BRCA2 PV carriers at the 10% risk threshold when considering PGS400 and PGS64518 of 4.8% and 5.4%, respectively, compared to the 36 variant PGS (Supplementary Table 7).
Discussion
We conducted the largest GWAS to date for HGSOC, both in terms of the sample size and the number of genetic variants assessed. To do this, we made use of large international consortia (OCAC and CIMBA), and harnessed data from the UKBB to bolster the sample size. We also utilized recent advances in imputation reference panels, namely the TOPMed panel, which allowed us to investigate the largest number of genetic variants to date. We identified eight independent variants at seven loci to be genome-wide statistically significant for association with HGSOC risk, including four rare variants (MAF < 1%) and two low-frequency polymorphisms (MAF 1–5%), demonstrating that GWAS with greater genomic coverage for imputation can contribute to identify previously undiscovered rare variant associations. Based on these associations, we defined 52 CCVs that have the strongest statistical evidence for being the likely causal variant for each locus. We also developed a polygenic model that exhibited improved discriminatory ability compared to previously published models.
The four rare variants were associated with large effect sizes, with RR estimates ranging from 2.35 to 25.6. The large effect sizes seem implausible and may reflect a winner’s curse effect with true effect sizes actually being lower than these estimates44. Moreover, they may represent false positive associations, hence we estimated BFDPs45 for the eight novel variants to determine the likelihood of this. Six variants had low BFDPs, indicating they are likely true associations. However, 8p21 rs540569242 and 9p24-p23 rs768719522 had noticeably larger BFDPs and are more likely to represent false positive associations.
Variant rs78378222, in the TP53 3’-UTR, with the major T-allele (AF 98.8%) is associated with an increased risk of HGSOC. The same allele has been associated with an increased risk of triple-negative breast cancer46 with a per-allele relative risk of 1.45, similar to its association with HGSOC (RR = 1.44). In contrast, the minor (G) allele of rs78378222 is associated with increased risks of skin47,48,49,50,51, brain51,52,53,54,55 and prostate cancers47,56. One study also found rs78378222 to be associated with standing height, lean body mass and basal metabolic rate51. The minor allele has been shown to impede TP53 3’-end processing, resulting in downregulated p53 mRNA levels and protein levels, and decreased apoptosis57. Furthermore, germline and somatic variations in TP53 are well established factors in cancer development and progression through its role as a tumor suppressor58,59,60,61 and the TP53 3’-UTR germline variant has been shown to interact with tumor TP53 mutation status62. A previous study by the OCAC found five SNPs in the TP53 region, within ±21 kb of rs78378222, to be associated with invasive EOC63. However, none of these SNPs are in LD with rs78378222.
rs528577783 is a rare intergenic variant between the FST and NDUFS4 genes. FST encodes follistatin, a single-chain gonadal protein that specifically inhibits follicle-stimulating hormone release and is moderately expressed in human reproductive tissues (Supplementary Table 5)64. A linkage study identified FST as a candidate gene for polycystic ovary syndrome (PCOS)65. PCOS may be associated with an increased risk of borderline or postmenopausal ovarian cancer66, although a recent Mendelian Randomization study found that genetically predicted PCOS was associated with decreased risk of ovarian cancer67. A GWAS of African ancestry women found rs37792 near FST to be genome-wide statistically significantly associated with HGSOC in African ancestry women68.
The 6p12 variant rs1013698558, located ~9.7 kb from the GCLC gene, was moderately associated with HGSOC. A study found a synthetic lethality relationship between GCLC and ARID1A-deficient OC cells69, whilst another reports that GCLC inhibition causes apoptosis in ARID1A-deficient cancer cells70. ARID1A has been implicated as a tumor suppressor gene71. It may be that the association we find for rs1013698558 with HGSOC is a result of the interplay between GCLC and ARID1A. The 8p21 variant rs540569242 is 156 kb from CDCA2 and was associated with HGSOC with the largest RR that we report. CDCA2 is involved in the cell division cycle and response to DNA damage. One study found CDCA2 expression is upregulated in ovarian tumor tissue compared with normal tissue72. This study also found that CDCA2 and its 100 most co-expressed genes were primarily involved in cell cycle, oocyte meiosis, progesterone-mediated oocyte maturation, p53 signaling and pyruvate metabolism pathways. We found that the PTPRD intronic variant rs768719522 at the 9p24-p23 locus had a large association RR with HGSOC. PTPRD has been implicated as a tumor suppressor gene73. This gene encodes a protein from the protein tyrosine phosphatase (PTP) family – PTPs are signaling molecules regulating processes such as cell growth, cell differentiation, the mitotic cycle and oncogenic transformation74. The common ACD missense variant rs6979 at 16q22 conferred a small per-allele RR to HGSOC risk. This gene encodes telomere protein TPP1, which is involved in maintenance of telomere length and protecting telomere ends. In addition to the 17p13 TP53 3’-UTR variant association, there was another independent variant associated with HGSOC risk at this locus. The rs143094271 variant is intronic in TNFSF13/TNFSF12-TNFSF13. These genes are members of the tumor necrosis factor family, which are involved in various cellular processes, including survival, proliferation, differentiation, and apoptosis. rs143094271-G has been associated with a decreased risk of having uterine fibroids (OR = 0.70)75. Women with uterine fibroids have been found to be at increased risk of developing OC76. However, despite rs143094271-G being associated with women being less likely to have uterine fibroids, we found that rs143094271-G yielded an increased risk of developing HGSOC. The 19q12 common variant rs62107113 is located ~15 kb from the CCNE1 gene and was associated with a modest increased risk of developing HGSOC. CCNE1 encodes cyclin E1, which regulate cyclin-dependent kinases. Overexpression of CCNE1 has been observed in genomically unstable tumors, notably HGSOC77,78,79,80,81 and triple-negative breast cancer78,82,83,84. Moreover, it has been shown that replication stress in cells overexpressing CCNE1 is likely a consequence of replication initiation, ultimately resulting in DNA damage and genomic instability85. There is evidence indicating CCNE1 amplification is mutually exclusive to BRCA inactivation79. CCNE1 is an exciting novel therapeutic target, Gallo et al. developed an orally bioavailable PKMYT1 inhibitor that activated CDK1 in CCNE1 overexpressed cells, promoting early mitosis in cells undergoing DNA synthesis86.
In addition to the 40 loci previously found to be associated with EOC14,15,16,17,18,19,20,21,22,23,24,25,26,27, we have identified genetic variants at a further seven loci associated with HGSOC, taking the number of loci associated with EOC to 47, 26 specifically with HGSOC. The previous known loci (52 variants at 40 loci) explained 8.5% of the polygenic variance of OC, assuming a total polygenic variance of 2.004 from Lee et al.87. The six variants that we report to be associated with HGSOC with low BFDP explain an additional 2.8%, making the total variance explained by GWAS identified variants to be 11.3%.
For previously identified variants, the estimated associations from general population data (combined OCAC and UKBB) ORs were broadly consistent with the estimated HRs for BRCA1 and BRCA2 PV carriers (Supplementary Table 3). Furthermore, testing for heterogeneity of effects resulted in a small number of variants exhibiting differences in effect sizes between population-based ORs and carrier HRs. None of the eight novel associations exhibited any heterogeneous effects between population-based ORs and BRCA1/2 PV carrier HRs (Supplementary Table 4).
Using the S4 method, we developed a 64,518 variant PGM, whose PGS was associated with a per SD OR of 1.46 (95% CI:1.37–1.54) and had discriminatory ability of AUROC = 0.607 (95% CI:0.590–0.623). There was a small improvement over the previous best performing PGM developed by Dareng et al.43 (18,007 variants; OR per unit SD = 1.42; AUROC = 0.596) developed using similar methodology. The PGS400 could be more practical to implement in a clinical setting, since it contains fewer variants, all of which are known to successfully genotype from the OncoArray experiment, compared to the best performing PGS64518 which requires imputation. We found that the PGS400 had similar performance to the optimal PGS64518. This also suggests that most of the predictive ability of the PGS may derive from genotyped SNPs with the largest variance contributions.
When each of the PGS were tested in individuals of East Asian ancestry from BBJ, the PGS associations were strongly attenuated, each to a similar degree. For women of East Asian ancestry, the PGS consisting of genotyped variants with 5957 and 3448 variants performed marginally better, with a slightly larger OR per PGS SD. The observed attenuations for East Asian women compared with European women are likely a result of the PGM derivation data being strongly weighted towards Europeans, as the majority of OCAC and CIMBA samples came from this ancestry group.
We calculated lifetime risks of developing EOC by PGS percentiles for the optimal 64,518 variant PGS for BRCA1/2 carriers. The risks ranged from 25.9% to 64.7% for BRCA1 carriers, and 9.3% to 28.9% for BRCA2 carriers, at the 5th and 95th percentiles, respectively. The range of risks for the 36 variant PGS that is currently implemented in the CanRisk ovarian cancer risk prediction algorithm43,87 were narrower than those for the PGS64518. The lifetime risks based on the PGS400 that used a subset of 400 reliably genotyped variants included in the PGS64518 at these percentiles sat approximately at the midpoints of the risks from the 64,518 and 36 variant PGS. We compared what risk reclassification (lower risk, <10%, or higher risk, ≥10% lifetime risk) would occur when using the PGS64518 or PGS400 versus the 36 variant PGS for BRCA2 PV carriers. We were unable to assess reclassification for BRCA1 carriers as the lifetime absolute risks at the lowest percentiles of the PGS distributions were always above 10%. We found that the PGS64518 and PGS400 led to total reclassification of risk groups of around 5% versus the 36 variant PGS. Most reclassification shifted women from lower risk (<10%) using the 36 variant PGS to higher risk (≥10%) using the alternative PGS64518 or PGS400. Taken together, these estimated lifetime risks and risk reclassifications will help to more accurately determine a carriers’ risk and inform clinical management of risk, such as the timing of risk reducing surgery or the initiation of cancer surveillance. In practice, PGS should be used in combination with pathogenic variants in ovarian cancer susceptibility genes and other risk lifestyle/hormonal factors in validated multifactorial cancer risk prediction models, such as CanRisk87,88,89, to improve comprehensive risk assessment. The clinical implementation of PGMs is mainly in the context of risk management for women with a family history of ovarian cancer. The CanRisk breast/ovarian cancer risk prediction model87,88,89 currently incorporates polygenic scores for both breast and ovarian cancer, together with lifestyle risk factors, family history and moderate/high penetrance risk alleles. The familial risk used in the model is the risk after exclusion of the contribution of polygenic risk as the polygenic risk explains ~11.3% of the excess familial risk. A key assumption that underlies the model is that the effects of the various risk factors are multiplicative (log-additive). There is a large body of evidence supporting this assumption90,91,92,93,94,95,96,97,98,99.
Strengths of this study include using the TOPMed imputation reference panel, enabling us to assess a larger number of low-frequency and rare variants than previous studies. A further strength was its power to detect low-frequency and rare variant associations, as well as previously unidentified common variant associations. This was facilitated by additional genotyped samples included in both OCAC and CIMBA and using population-based data from UKBB, resulting in the largest sample size analyzed for assessing genetic variant associations with HGSOC risk.
Limitations include the fact that the GWAS discovery data available were primarily of European ancestry; the associations of these variants are likely to differ for women of non-European ancestries, as they are likely to have different frequencies and LD patterns. A limitation of the PGM was that the derivation data differed from the GWAS discovery data presented here. Ideally, all the discovery GWAS data would have been used for PGM development. However, it was essential to validate the PGM on independent data, hence the OCAC and CIMBA data were used for development, whilst the UKBB data were reserved for validation. Lastly, the PGM training data (FinnGen) did not have specific histotypes available, meaning we were only able to consider overall EOC in the PGM training. As we were investigating non-mucinous OC, we would ideally have had specific EOC histotypes available at each stage of the PGM development, training and validation. However, given that HGSOC is the most prevalent EOC histotype, it is unlikely to have a major impact on the PGM hyperparameter fine-tuning.
Future research may aim to fine-map the novel loci identified here, to refine the candidate causal variants associated with HGSOC risk; and in-silico analyses may identify candidate target genes or pathways for further experimental studies100. Additionally, future research could aim to identify novel variants associated with other OC histotypes and to discover novel associations for other ancestries.
We have shown that improvements in imputation reference panels that have larger genomic coverage and increased sample sizes can assist in identification of novel HGSOC associated variants that previously went undetected, either from absence from genotyping or imputation reference panels, or from lack of power to detect associations. Furthermore, these associations can be used to develop PGM that outperform previous best efforts that can be incorporated into cancer risk prediction algorithms to improve personalized risk prediction for HGSOC.
Methods
Study samples
OCAC participants were enrolled in 65 studies from 16 countries and a large European multinational nested case-control study (Supplementary Table 1). OCAC individual participant data were used for GWAS discovery analyses and developing polygenic models (PGMs).
CIMBA study participants were enrolled in 64 studies from 28 countries (Supplementary Table 2). Eligibility was restricted to women aged at least 18 years at the time of recruitment who carried a PV in either BRCA1 or BRCA2. Data collected included year of birth, PV description, age at recruitment, age at last follow-up, and age at breast and ovarian cancer (invasive, fallopian tube and peritoneal) diagnoses, and age or date of prophylactic surgeries (bilateral mastectomy and bilateral oophorectomy). Most participants were recruited through cancer genetics clinics and enrolled in regional/national research studies. CIMBA individual participant data were used in the GWAS discovery and in PGM development.
The UK Biobank (UKBB) is a large-scale biomedical research resource, with detailed genetic and health data on half a million UK participants36,37. For the purposes of these analyses, data from 245,377 female participants of European ancestry were used. UKBB individual participant data were used in the GWAS discovery analyses and to independently validate PGS.
FinnGen is a large collection of newly recruited and legacy samples from Finnish biobanks, research institutes, universities, university hospitals, international pharmaceutical partners, the Finnish Blood Service, the Finnish Biobank Cooperative, and the Finnish Institute for Health and Welfare, utilizing Finnish longitudinal health register data collected on every resident of Finland since 196938,39. FinnGen summary statistics (data freeze 8) based on 150,658 women (149,394 controls, 1264 EOC cases of any histotype) were used to train PGM hyperparameters.
BioBank Japan (BBJ) is a large biobank resource containing clinical and genetic data on over 300,000 participants40,41. BBJ summary statistics based on 61,457 women (60,614 controls, 843 EOC cases) were used for assessing PGS associations for women of East Asian ancestry101.
Genotyping and re-imputation using the TOPMed reference panel
Genotyping of OCAC and CIMBA samples were performed on one of two custom single nucleotide polymorphism (SNP) genotyping arrays, the iCOGS23,102 array or OncoArray22,103,104. The iCOGS array included ~210,000 SNPs that were selected for previous evidence of association with breast, ovarian and prostate cancer. The OncoArray is a custom genotyping chip consisting of ~533,000 SNPs, approximately half of which is a GWAS backbone that tags common SNPs. A standard quality control (QC) process was applied, including assessment of SNP call rate, allele frequency, genotyping intensity clustering, Hardy-Weinberg equilibrium, and SNP concordance from duplicated samples104. OCAC had additional samples genotyped on GWAS arrays14,15,16. These data were imputed to the TOPMed reference panel (version R2 on GRCh38, with 97,256 samples)31 using the Michigan Imputation Server32,105. Phasing was performed with Eagle2106 and imputation with Minimac107,108. Prior to imputation, variants were excluded from imputation genotype files if they: (1) were not in Hardy-Weinberg equilibrium (P < 1 × 10−7 in controls, or P < 1 × 10−12 in cases); (2) had poor cluster plots; or (3) had a call rate <95% (common variants), or a call rate <98% (rare variants, MAF < 1%). We used https://www.well.ox.ac.uk/~wrayner/tools/ script HRC-1000G-check-bim-v4.3.0.pl to remove variants not on the TOPMed reference panel or align them to the correct strand. This tool excluded variants where the genotyped frequency differed from the panel by more than 0.20. Samples were randomized into batches of <25,000 to meet the maximum sample requirement of the imputation server31,32,105. The same list of variants was included for each genotype batch. Details of the UKBB genotyping and imputation to a combined UK10K109,110 and HRC30 reference panel have been described elsewhere36,37. The OCAC, CIMBA and UKBB analyses were based on 142 million, 104 million and 60 million well-imputed (imputation r2 > 0.30) variants, respectively (Table 1). Downstream meta-analyses were restricted to variants that had minor allele counts (MACs) of MAC > 5 and did not have heterogeneous effects (Cochran Q-statistic, Phet > 1 × 10−8) in the meta-analysis of OCAC studies. Variants were aligned to the Genome Reference Consortium Human Build 38 (GRCh38/hg38).
Analyses including FinnGen and BBJ data made use of summary statistic data only. Details of FinnGen and BBJ genotyping and imputation have been described elsewhere38,39,40,41,101.
Statistical analyses of OCAC and UKBB data
We examined the associations between genotypes and HGSOC risk in the OCAC data using logistic regression (using custom software). Analyses were conducted separately for OncoArray, iCOGS, and five GWAS datasets14,15,16,111 and were combined by fixed-effects inverse-variance weighted meta-analysis (Fig. 3). We included project-specific principal components (PCs) as covariates in the model with the number of PCs based on the inflection point observed in a scree plot (Supplementary Fig. 49). PCs for OncoArray data were calculated using 33,661 uncorrelated (r2 < 0.10) common (MAF > 0.05) variants. Calculations were performed using a custom program (PCAcalc), available at https://github.com/CCGE-Cambridge/OCAC_CIMBA_HGSOC. Details of PC calculations for the other genotype data has been previously described elsewhere19,23. All women were of European ancestry and unrelated, determined using genetic data22,103. Ancestry was evaluated using the FastPop software112. Women with >80% European ancestry were retained for statistical analyses. Relatedness was checked by genetic concordance statistics. This process has been described previously22. Briefly, women with concordance statistics between 0.74 and 0.86 were considered to be related (values > 0.86 considered duplicates). In instances of case-control pairs, the case was retained for statistical analyses and the control was excluded. For instances of case-case and control-control pairs, the sample with the lower call rate was excluded.
Schema describing the analytical process for the GWAS analyses from OCAC, UKBB and CIMBA, and subsequent meta-analyses.
The UKBB analysis was restricted to women with European ancestry, selected based on their PCs. EOC histotypes were classified using diagnostic codes provided by UKBB, with “serous”, “undifferentiated”, or “other” classified as HGSOC, a methodology similar to that used in OCAC22. Association analyses between genotypes and HGSOC risk were assessed by logistic regression (Fig. 3). We adjusted for the top four PCs and age at recruitment.
Statistical analyses of BRCA1 and BRCA2 pathogenic variant carriers
Analyses of CIMBA data were limited to carriers of European ancestry, determined by genetic data and multidimensional scaling22,103. We used 33,661 common uncorrelated variants (the same set used to calculate PCs) to calculate kinship coefficients between all CIMBA participants and 267 HapMap samples (CHD, JPT, YRI and CEU ancestries). These kinship coefficients were converted to distances and then underwent multidimensional scaling. Using the top two PCs, the proportion of European ancestry for each participant was calculated. Women with >27% non-European ancestry were excluded, ensuring that women with Ashkenazi Jewish ancestry were retained for statistical analyses. Association analyses were performed separately by genotyping array (iCOGS or OncoArray), and separately for BRCA1 and BRCA2 PV carriers (Fig. 3). iCOGS and OncoArray associations were combined by fixed-effects inverse-variance weighted meta-analysis to estimate BRCA1 and BRCA2 PV carrier specific associations. The association analysis was carried out within a survival analysis framework, by modeling the retrospective likelihood of observing the genotypes conditional on the disease phenotypes to adjust for the non-random ascertainment with respect to disease phenotypes113,114. The censoring process followed carriers from birth until the first occurrence of: EOC (including fallopian tube and peritoneal cancers) diagnosis, risk-reducing salpingo-oophorectomy, or study entry. Breast cancer diagnoses were not considered to be a censoring event and EOC was the endpoint of interest. Associations were then assessed using the score test statistic based on the retrospective likelihood113,114 assuming BRCA1 and BRCA2 PV carrier specific and age-cohort specific EOC incidences88. Analyses were stratified by country and Ashkenazi Jewish ancestry, and to account for relatedness between individuals we calculated kinship adjusted variances115. Only variants that were available through OncoArray genotyping and imputation were considered, as the majority of samples were available from this genotyping platform (Table 1). As HGSOC is the predominant histotype for both BRCA1 and BRCA2 PV carriers, the associations were combined by fixed-effects and inverse variance weighted meta-analysis using the METAL software116.
Meta-analyses
We pooled the combined OCAC and UKBB summary association data (per-allele odds ratios, ORs) with the combined BRCA1 and BRCA2 PV carrier summary association data (per-allele hazard ratios, HRs) by fixed-effects inverse-variance weighted meta-analysis using METAL116 to give per-allele relative risks (RRs, a combination of population-based ORs and BRCA1/2 carrier HRs) (Fig. 3).
Eliminating likely statistical artifacts
The associations of all variants with genome-wide significant associations and falling outside known regions were re-evaluated to eliminate likely spurious associations potentially due to unstable effect estimates from strata with small numbers. For this purpose, the associations were re-analyzed, pooling individual level data from OCAC and UKBB, incorporating 29 PCs derived across all genotyping projects (PCs set to 0 for women not in specific studies/projects). The UKBB data were also adjusted for age (set to 0 for the OCAC samples). Any variant with imputation accuracy r2 < 0.30 within a panel were considered missing for that particular panel. The associations for BRCA1/2 PV carriers were reassessed assuming all study participants came from a single stratum.
We also re-evaluated potentially novel associations with variants in regions proximal to known regions by performing approximate conditional analyses117. This approach utilized summary statistics from the combined OCAC, UKBB and BRCA1/2 PV carrier meta-analysis and the linkage disequilibrium (LD) structure from 111,304 women genotyped on the OncoArray from OCAC and CIMBA.
For variants passing these checks, we calculated Bayesian false-discovery probabilities (BFDPs)45. BFDPs are approximate Bayes factors, statistical measures of the strength of evidence in favor of a given statistical model over another. They are akin to false-positive reporting probabilities, aiming to control the number of false positive associations whilst not dismissing too many associations that may merit scientific interest, by including Bayesian priors on an effect or association. BFDPs are especially useful in contexts where many statistical tests are performed, such as GWAS. We assumed prior probabilities of 1:1000 and 1:10,000 variants being truly associated. BFDPs were calculated using the BFDP function from the R gap package118.
Genomic database searches
Variants found to be associated with HGSOC at the genome-wide statistical significance level that also passed the additional checks detailed above, and the nearest genes to these variants, were looked up in genomic resource databases. We accessed dbSNP74, Phenoscanner (version 2)119,120,121, PheWeb (version 1.3.15)122, GTEx64 and eQTLGen123,124 (accessed on 15th May 2023).
Defining credible causal variants
The lead variant (variant with the smallest P-value) at each novel region may not be causal. Therefore, we identified lists of credible causal variants (CCVs) that are likely to contain the genetic variant responsible for altering HGSOC risk defined as the set of variants within ±500 kb of the lead variant whose P-value was within two orders of magnitude of the lead variant’s P-value125.
Development of polygenic risk models
Previous analyses of PGMs specific to HGS, LGS, and mucinous histotypes showed that all histotypes, except mucinous, were associated with either the HGSOC PGS or LGSOC PGS28. Genetic correlations between mucinous ovarian cancer and other ovarian cancer histotypes have been estimated to be 0.24, 0.21, 0.23 and −0.21 with high-grade serous ovarian cancer, low-grade serous ovarian cancer, endometrioid ovarian cancer and clear cell, respectively28. Whereas the genetic correlations are stronger for pairs of the other non-mucinous histotypes28. Thus, the non-mucinous histotypes were combined here126. The PGMs were developed on summary statistics using associations obtained from a meta-analysis of the OCAC and CIMBA data (Fig. 4). To maximize sample size and genetic diversity, the OCAC summary statistics came from a meta-analysis of European (133,369 controls, 25,707 cases), East Asian (3871 controls, 2736 cases) and African (1099 controls, 556 cases) ancestry women, considering the associations with non-mucinous OC. Hence, these OCAC summary statistics differed from those obtained from the analyses described in ‘Statistical analyses of OCAC and UKBB data’, as a result of analyzing additional cases from non-mucinous histotypes (and further controls from studies previously excluded because they had no HGSOC cases), and additional women of East Asian and African ancestry. The CIMBA summary statistics were identical to those generated in the analyses described in ‘Statistical analyses of BRCA1 and BRCA2 pathogenic variant carriers’.
S4 select and shrink using summary statistics, PGM polygenic model, NMOC non-mucinous ovarian cancer, LD linkage disequilibrium.
We developed 1102 PGMs (sets of variants and their weights) for non-mucinous OC using the Select and Shrink with Summary Statistics (S4) method43,126. We used two P-value to LD r2 ratios to select variants: (i) P:r2 < 0.02 that resulted in ~64k variants (562 PGMs tested) and (ii) P:r2 < 0.15 resulting in ~394k variants (540 models tested). The LD structure was weighted to reflect the average effects from each ancestry based on a subset of OCAC OncoArray data. Each model had different combinations of model hyperparameters (shrinkage parameters controlling small and large variant effect sizes, and an overall shrinkage parameter) that were trained using summary statistics based on 150,658 women (149,394 controls, 1264 EOC cases) from FinnGen (data freeze 8)38,39. EOC histotype data was not available for FinnGen, hence we used the associations with overall EOC for PGM training.
Polygenic scores (PGS) are PGMs applied to observed or imputed genotypes. We used the resultant PGM to calculate PGS on individual-level data in the UKBB36,37 to test its association with HGSOC and calculated its discriminatory ability for HGSOC by estimating the area under the ROC (AUROC) curve. AUROCs were calculated using the R pROC package auc function127.
We assessed the performance of the PGS for women of East Asian ancestry using BBJ data. As individual-level data were unavailable, we used association summary statistics from BBJ101 and assessed the PGS performance using a previously described method for evaluating PGS on summary statistic data126. The reference panel used consisted of individuals of East Asian ancestry from the 1000 Genomes Project29.
We also created candidate PGMs consisting of genotyped variants that could be more easily applied in clinical settings by selecting variants known to reliably genotype from the OncoArray103, many of which had been chosen for their relevance to ovarian cancer. We selected a subset of genotyped variants, based on “relative importance”, where the importance of each variant is approximately proportional to p*(1-p)*β2, where β is the log-RR and p is the minor allele frequency for that variant. We ranked each variant based on relative importance and selected the top N desired variants.
Absolute risks of EOC by PGS percentile
We calculated predicted lifetime risks (to age 80 years) of developing EOC for women in the general population (unselected for PV status), BRCA1 and BRCA2 PV carriers at the 1st, 5th, 20th, 50th (median), 80th, 95th and 99th percentiles of the various PGS distributions, following previously published methodology128. To ensure consistency with known EOC risks for the general population, BRCA1 and BRCA2 PV carriers, average age-specific EOC incidence rates were constrained over PGS percentiles to agree with external EOC incidence rates for the general population129 and BRCA1/2 carriers130.
We examined the number of BRCA2 PV carriers genotyped on the OncoArray that would transition between risk groups (low (<10%) or high (≥10%) lifetime risk) of developing EOC, based on their observed PGS percentile. These risk reclassification analyses were limited to BRCA2 carriers as their lifetime risks transition over the 10% lifetime risk threshold, whereas a BRCA1 carrier is already at substantially increased lifetime risk.
Ethics statement
All study participants provided written informed consent and participated in research studies at the host institute under ethically approved protocols. This study was conducted in accordance with the Declaration of Helsinki.
All study participants provided written informed consent and participated in research or clinical studies at the host institute under ethically approved protocols. The studies and their approving institutes are: Australian site of the Breast Cancer Family Registry (BCFR-AU) - The University of Melbourne Health Sciences Human Ethics Sub-Committee; Northern California site of the Breast Cancer Family Registry (BCFR-NC) - Northern California Cancer Center Institutional Review Board; New York site of the Breast Cancer Family Registry (BCFR-NY) - Columbia University Medical Center Institutional Review Board; Ontario site of the Breast Cancer Family Registry (BCFR-ON) - Mount Sinai Hospital Research Ethics Board; Philadelphia site of the Breast Cancer Family Registry (BCFR-PA) - Institutional Review Board Fox Chase Cancer Center; Utah site of the Breast Cancer Family Registry (BCFR-UT) - Institutional Review Board University of Utah; Baltic Familial Breast and Ovarian Cancer Consortium (BFBOCC) - Centrālā medicīnas ētikas Komiteja; Lietuvos Bioetikos Komitetas; BRCA-gene mutations and breast cancer in South African women (BMBSA) - University of Pretoria and Pretoria Academic Hospitals Ethics Committee; Beckman Research Institute of the City of Hope (BRICOH) - City of Hope Medical Center Institutional Review Board; Copenhagen Breast Cancer Study (CBCS) - De Videnskabsetiske Komiteer I Region Hovedsladen; Spanish National Cancer Centre (CNIO) - Instituto de Salud Carlos III Comité de Bioética y Bienestar Animal; City of Hope Cancer Center (COH) - City of Hope Institutional Review Board; CONsorzio Studi ITaliani sui Tumori Ereditari Alla Mammella (CONSIT TEAM) - Comitato Etico Indipendente della Fondazione IRCCS “Istituto Nazionale dei Tumori”; National Centre for Scientific Research Demokritos (DEMOKRITOS) - Bioethics committee of NCSR “Demokritos”, 240/EHΔ/11.3; National Centre for Scientific Research Demokritos (DEMOKRITOS) - Papageorgiou Hospital Ethics Committee; Dana Farber Cancer Institute (DFCI) - Dana Farber Cancer Institute Institutional Review Board; Deutsches Krebsforschungszentrum (DKFZ) - Ethik-Kommission des Klinikums der Universität; Deutsches Krebsforschungszentrum (DKFZ) - Hospital Universitario de San Ignacio Comité de Investigaciones y Etica; Deutsches Krebsforschungszentrum (DKFZ) - Shaukat Khanum Memorial Cancer Hospital and Research Centre Institutional Review Board; Epidemiological study of BRCA1 and BRCA2 mutation carriers (EMBRACE) - Anglia & Oxford MREC; Fox Chase Cancer Center (FCCC) - Institutional Review Board Fox Chase Cancer Center; Fundación Pública Galega de Medicina Xenómica - Comite Autonomico de Etica da Investigacion de Galicia; German Consortium of Hereditary Breast and Ovarian Cancer (GC-HBOC) - Ethik-Kommission der Medizinischen Fakultät der Universät zu Köln; Genetic Modifiers of cancer risk in BRCA1/2 mutation carriers (GEMO) - Comité consultatif sur le traitement de I’information en matière de recherche dans le domaine de la santé; Georgetown University (GEORGETOWN) - MedStar Research Institute - Georgetown University Oncology Institutional Review Board; Ghent University Hospital (G-FAST) - Universitair Ziekenhuis Gent - Ethics Committee; Hospital Clinico San Carlos (HCSC) - Comité Ético de Investigación Clínia Hospital Clínico San Carlos; Helsinki Breast Cancer Study (HEBCS) - Helsingin ja uudenmaan sairaanhoitopiiri (Helsinki University Central Hospital ethics committee); HEreditary Breast and Ovarian study Netherlands (HEBON) - Protocol Toetsingscommissie van het Nederlands Kanker Instituut/Antoni van Leeuwenhoek Ziekenhuis; Molecular Genetic Studies of Breast- and Ovarian Cancer in Hungary (HUNBOCS) - Institutional Review Board of the Hungarian National Institute of Oncology; University Hospital Vall d’Hebron (HVH) - The Hospital Universitario Vall d’Hebron Clinical Research Ethics Committee; Institut Català d’Oncologia (ICO) - Catalan Institute of Oncology Institutional Review Board; International Hereditary Cancer Centre (IHCC) - Komisji Bioetycznej Pomorskiej Akademii Medycznej (Pomeranian Medical University Bioethics Committee); Iceland Landspitali - University Hospital (ILUH) - Vísindasiđanefnd National Boethics Committee; Interdisciplinary Health Research International Team Breast Cancer Susceptibility (INHERIT) - Comité d'éthique de la recherche du Centre Hospitalier Universitaire de Québec; Istituto Oncologico Veneto Hereditary Breast and Ovarian Cancer Study (IOVHBOCS) - Centro Oncologico Regionale Azienda Ospedale Di Padova Comitato Etico; Portuguese Oncology Institute-Porto Breast Cancer Study - COMISSÃO DE ÉTICA PARA A SAÚDE (CES); Kathleen Cuningham Foundation Consortium for Research into Familial Breast Cancer (KCONFAB) - Queensland Institute of Medical Research - Human Research Ethics Committee; Kathleen Cuningham Foundation Consortium for Research into Familial Breast Cancer (KCONFAB) - Peter MacCallum Cancer Centre Ethics Committee; University of Kansas Medical Center(KUMC) - The University of Kansas Medical Center Human Research Protection Program; Mayo Clinic (MAYO) - Mayo Clinic Institutional Review Boards; McGill University (MCGILL) - McGill Faculty of Medicine Institutional Review Board; Modifier Study of Quantitative Effects on Disease (MOD-SQUAD) - Mayo Clinic Institutional Review Boards; Memorial Sloane Kettering Cancer Center (MSKCC) - Human Biospecimen Utilization Committee; Memorial Sloane Kettering Cancer Center (MSKCC) - Memorial Sloan-Kettering Cancer Center IRB; General Hospital Vienna (MUV) - Ethikkommission der Medizinischen Universität Wien; Women’s College Research Institute Hereditary Breast and Ovarian Cancer Study - University of Toronto Health Sciences Review Ethics Board; National Cancer Institute (NCI) - NIH Ethics Office; National Israeli Cancer Control Center (NICCC) - Carmel Medical Center Institutional Review Board (Helsinki Committee); N.N. Petrov Institute of Oncology (NNPIO) - N.N. Petrov Institional Ethical Committee; NorthShore University HealthSystem (NORTHSHORE) - Institutional Review Board of NorthShore University HealthSystem; NRG Oncology (NRG_ONCOLOGY) - Cancer Prevention and Control Protocol Review Committee; Ontario Cancer Genetics Network (OCGN) - University Health Network Research Ethics Board; The Ohio State University Comprehensive Cancer Center (MACBRCA) - The Ohio State University Cancer Institutional Review Board; Odense University Hospital (OUH) - Den Videnskabsetiske Komité for Region Syddanmark; Pisa Breast Cancer Study (PBCS) - Azienda Ospedaliera Pisana Comitato Etico per lo studio del farmaco sull’uomo; Sheba Medical Centre - Chaim Sheba Medical Center IRB; Swedish Breast Cancer Study (SWE-BRCA) - Regionala Etikprövningsnämnden Stockholm; University of Chicago (UCHICAGO) - The University of Chicago Biological Sciences Division. Institutional Review Board (BSD IRB); University of California Los Angeles (UCLA) - UCLA Institutional Review Board (UCLA IRB); University of California San Francisco (UCSF) - Human Research Protection Program Institutional Review Board (IRB); UK and Gilda Radner Familial Ovarian Cancer Registries (UKGRFOCR) - Roswell Park Cancer Institute IRB; UK and Gilda Radner Familial Ovarian Cancer Registries (UKGRFOCR) - Cambridge Local Research Ethics Committee; University of Pennsylvania (UPENN) - University of Pennsylvania Institutional Review Board; Cancer Family Registry University of Pittsburg (UPITT) - University of Pittsburgh Institutional Review Board; University of Texas MD Anderson Cancer Center (UTMDACC) - University of Texas MD Anderson Cancer Center Office of Protocol Research Institutional Review Board; Victorian Familial Cancer Trials Group (VFCTG) - Peter MacCallum Cancer Centre Ethics Committee; Women’s Cancer Program at Cedars-Sinai Medical Center (WCP) - (Cedars-Sinai Medical Center) CSMC Institutional Review Board. GynBiobank (WMH) – Western Sydney Local Health District Human Research Ethics Committee.
Data availability
GWAS summary statistics from OCAC and CIMBA are publicly available from the GWAS Catalog under study accession identifiers GCST90455658 (OCAC), GCST90455659 (CIMBA BRCA1 PV carriers), GCST90455660 (CIMBA BRCA2 PV carriers). The summary statistics from the meta-analysis of OCAC, UKBB and CIMBA associations are also available under study accession identifier GCST90455661. Variants included in the polygenic scores and their corresponding weights are publicly available from the PGS Catalog under identifiers PGS005086 (PGS64518), PGS005087 (PGS5957), PGS005088 (PGS3448) and PGS005089 (PGS400). The FinnGen summary statistics used were from data freeze 8. The summary statistics across all data freezes are available at https://www.finngen.fi/en/access_results and the data freeze 8 summary statistics are available at https://r8.finngen.fi/. The Biobank Japan summary statistics used were from Sakaue et al.101. The summary statistic data for all traits are available at https://pheweb.jp/ and the ovarian cancer summary statistic data can be accessed here https://pheweb.jp/pheno/OvC.
Code availability
Code and software used to implement analyses is publicly available at https://github.com/CCGE-Cambridge/OCAC_CIMBA_HGSOC.
References
Sung, H. et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Prat, J. New insights into ovarian cancer pathology. Ann. Oncol. 23, x111–x117 (2012).
Peres, L. C. et al. Invasive epithelial ovarian cancer survival by histotype and disease stage. J. Natl Cancer Inst. 111, 60–68 (2019).
Risch, H. A. et al. Population BRCA1 and BRCA2 mutation frequencies and cancer penetrances: a kin-cohort study in Ontario, Canada. J. Natl Cancer Inst. 98, 1694–1706 (2006).
Risch, H. A. et al. Prevalence and penetrance of germline BRCA1 and BRCA2 mutations in a population series of 649 women with ovarian cancer. Am. J. Hum. Genet. 68, 700–710 (2001).
Shaw, P. A. et al. Histopathologic features of genetically determined ovarian cancer. Int. J. Gynecol. Pathol. 21, 407–411 (2002).
Zhang, S. et al. Frequencies of BRCA1 and BRCA2 mutations among 1,342 unselected patients with invasive ovarian cancer. Gynecol. Oncol. 121, 353–357 (2011).
Schrader, K. A. et al. Germline BRCA1 and BRCA2 mutations in ovarian cancer: utility of a histology-based referral strategy. Obstet. Gynecol. 120, 235–240 (2012).
Wang, Y. K. et al. Genomic consequences of aberrant DNA repair mechanisms stratify ovarian cancer histotypes. Nat. Genet. 49, 856–865 (2017).
Lakhani, S. R. et al. Pathology of ovarian cancers in BRCA1 and BRCA2 carriers. Clin. Cancer Res. 10, 2473–2481 (2004).
O’Mahony, D. G. et al. Ovarian cancer pathology characteristics as predictors of variant pathogenicity in BRCA1 and BRCA2. Br. J. Cancer 128, 2283–2294 (2023).
Singh, N., McCluggage, W. G. & Gilks, C. B. High-grade serous carcinoma of tubo-ovarian origin: recent developments. Histopathology 71, 339–356 (2017).
Kobel, M. et al. Differences in tumor type in low-stage versus high-stage ovarian carcinomas. Int. J. Gynecol. Pathol. 29, 203–211 (2010).
Song, H. et al. A genome-wide association study identifies a new ovarian cancer susceptibility locus on 9p22.2. Nat. Genet. 41, 996–1000 (2009).
Bolton, K. L. et al. Common variants at 19p13 are associated with susceptibility to ovarian cancer. Nat. Genet. 42, 880–884 (2010).
Goode, E. L. et al. A genome-wide association study identifies susceptibility loci for ovarian cancer at 2q31 and 8q24. Nat. Genet. 42, 874–879 (2010).
Bojesen, S. E. et al. Multiple independent variants at the TERT locus are associated with telomere length and risks of breast and ovarian cancer. Nat. Genet. 45, 371–384 (2013).
Permuth-Wey, J. et al. Identification and molecular characterization of a new ovarian cancer susceptibility locus at 17q21.31. Nat. Commun. 4, 1627 (2013).
Pharoah, P. D. et al. GWAS meta-analysis and replication identifies three new susceptibility loci for ovarian cancer. Nat. Genet 45, 362–370 (2013).
Shen, H. et al. Epigenetic analysis leads to identification of HNF1B as a subtype-specific susceptibility gene for ovarian cancer. Nat. Commun. 4, 1628 (2013).
Kuchenbaecker, K. B. et al. Identification of six new susceptibility loci for invasive epithelial ovarian cancer. Nat. Genet. 47, 164–171 (2015).
Phelan, C. M. et al. Identification of 12 new susceptibility loci for different histotypes of epithelial ovarian cancer. Nat. Genet. 49, 680–691 (2017).
Couch, F. J. et al. Genome-wide association study in BRCA1 mutation carriers identifies novel loci associated with breast and ovarian cancer risk. PLoS Genet. 9, e1003212 (2013).
Chen, K. et al. Genome-wide association study identifies new susceptibility loci for epithelial ovarian cancer in Han Chinese women. Nat. Commun. 5, 4682 (2014).
Kelemen, L. E. et al. Genome-wide significant risk associations for mucinous ovarian carcinoma. Nat. Genet. 47, 888–897 (2015).
Kar, S. P. et al. Genome-wide meta-analyses of breast, ovarian, and prostate cancer association studies identify multiple new susceptibility loci shared by at least two cancer types. Cancer Discov. 6, 1052–1067 (2016).
Lawrenson, K. et al. Functional mechanisms underlying pleiotropic risk alleles at the 19p13.1 breast-ovarian cancer susceptibility locus. Nat. Commun. 7, 12675 (2016).
Dareng, E. O. et al. Integrative multi-omics analyses to identify the genetic and functional mechanisms underlying ovarian cancer risk regions. Am. J. Hum. Genet. 111, 1061–1083 (2024).
Genomes Project Consortium et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
McCarthy, S. et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat. Genet. 48, 1279–1283 (2016).
Taliun, D. et al. Sequencing of 53,831 diverse genomes from the NHLBI TOPMed Program. Nature 590, 290–299 (2021).
Das, S. et al. Next-generation genotype imputation service and methods. Nat. Genet. 48, 1284–1287 (2016).
OCAC. Ovarian Cancer Association Consortium, https://ocac.ccge.medschl.cam.ac.uk/ (2023).
Chenevix-Trench, G. et al. An international initiative to identify genetic modifiers of cancer risk in BRCA1 and BRCA2 mutation carriers: the Consortium of Investigators of Modifiers of BRCA1 and BRCA2 (CIMBA). Breast Cancer Res. 9, 104 (2007).
CIMBA. Consortium of Investigators of Modifiers of BRCA1 & BRCA2, https://cimba.ccge.medschl.cam.ac.uk/ (2023).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Biobank, U. UK Biobank, https://www.ukbiobank.ac.uk/ (2023).
Kurki, M. I. et al. FinnGen provides genetic insights from a well-phenotyped isolated population. Nature 613, 508–518 (2023).
FinnGen. FinnGen, https://www.finngen.fi/en (2023).
Nagai, A. et al. Overview of the BioBank Japan project: study design and profile. J. Epidemiol. 27, S2–S8 (2017).
Japan, B. BioBank Japan, https://biobankjp.org/en/ (2023).
Huang, L. et al. TOP-LD: a tool to explore linkage disequilibrium with TOPMed whole-genome sequence data. Am. J. Hum. Genet. 109, 1175–1181 (2022).
Dareng, E. O. et al. Polygenic risk modeling for prediction of epithelial ovarian cancer risk. Eur. J. Hum. Genet. 30, 349–362 (2022).
Xiao, R. & Boehnke, M. Quantifying and correcting for the winner’s curse in genetic association studies. Genet. Epidemiol. 33, 453–462 (2009).
Wakefield, J. A Bayesian measure of the probability of false discovery in genetic epidemiology studies. Am. J. Hum. Genet. 81, 208–227 (2007).
Zhang, H. et al. Genome-wide association study identifies 32 novel breast cancer susceptibility loci from overall and subtype-specific analyses. Nat. Genet. 52, 572–581 (2020).
Stacey, S. N. et al. A germline variant in the TP53 polyadenylation signal confers cancer susceptibility. Nat. Genet. 43, 1098–1103 (2011).
Stacey, S. N. et al. Germline sequence variants in TGM3 and RGS22 confer risk of basal cell carcinoma. Hum. Mol. Genet. 23, 3045–3053 (2014).
Stacey, S. N. et al. New basal cell carcinoma susceptibility loci. Nat. Commun. 6, 6825 (2015).
Chahal, H. S. et al. Genome-wide association study identifies 14 novel risk alleles associated with basal cell carcinoma. Nat. Commun. 7, 12510 (2016).
Di Giovannantonio, M. et al. Heritable genetic variants in key cancer genes link cancer risk with anthropometric traits. J. Med. Genet. 58, 392–399 (2021).
Melin, B. S. et al. Genome-wide association study of glioma subtypes identifies specific differences in genetic susceptibility to glioblastoma and non-glioblastoma tumors. Nat. Genet. 49, 789–794 (2017).
Egan, K. M. et al. Rare TP53 genetic variant associated with glioma risk and outcome. J. Med. Genet. 49, 420–421 (2012).
Enciso-Mora, V. et al. Low penetrance susceptibility to glioma is caused by the TP53 variant rs78378222. Br. J. Cancer 108, 2178–2185 (2013).
Wang, Z. et al. Further confirmation of germline glioma risk variant rs78378222 in TP53 and its implication in tumor tissues via integrative analysis of TCGA data. Hum. Mutat. 36, 684–688 (2015).
Conti, D. V. et al. Trans-ancestry genome-wide association meta-analysis of prostate cancer identifies new susceptibility loci and informs genetic risk prediction. Nat. Genet. 53, 65–75 (2021).
Li, Y. et al. Single nucleotide variation in the TP53 3’ untranslated region in diffuse large B-cell lymphoma treated with rituximab-CHOP: a report from the International DLBCL Rituximab-CHOP Consortium Program. Blood 121, 4529–4540 (2013).
Bode, A. M. & Dong, Z. Post-translational modification of p53 in tumorigenesis. Nat. Rev. Cancer 4, 793–805 (2004).
Olivier, M., Hussain, S. P., Caron de Fromentel, C., Hainaut, P. & Harris, C. C. TP53 Mutation Spectra and Load: A Tool for Generating Hypotheses on the Etiology of Cancer, 247–270 (IARC Science Publication, 2004).
Levine, A. J. p53, the cellular gatekeeper for growth and division. Cell 88, 323–331 (1997).
Vogelstein, B., Lane, D. & Levine, A. J. Surfing the p53 network. Nature 408, 307–310 (2000).
Zhang, P. et al. Germline and somatic genetic variants in the p53 pathway interact to affect cancer risk, progression, and drug response. Cancer Res. 81, 1667–1680 (2021).
Schildkraut, J. M. et al. Single nucleotide polymorphisms in the TP53 region and susceptibility to invasive epithelial ovarian cancer. Cancer Res. 69, 2349–2357 (2009).
GTEx. Genotype-Tissue Expression project, https://gtexportal.org/home/ (2023).
Urbanek, M. et al. Thirty-seven candidate genes for polycystic ovary syndrome: strongest evidence for linkage is with follistatin. Proc. Natl Acad. Sci. USA 96, 8573–8578 (1999).
Frandsen, C. L. B. et al. Risk of epithelial ovarian tumors among women with polycystic ovary syndrome: a nationwide population-based cohort study. Int. J. Cancer 153, 958–968 (2023).
Harris, H. R. et al. Association between genetically predicted polycystic ovary syndrome and ovarian cancer: a Mendelian randomization study. Int J. Epidemiol. 48, 822–830 (2019).
Manichaikul, A. et al. Identification of novel epithelial ovarian cancer loci in women of African ancestry. Int J. Cancer 146, 2987–2998 (2020).
Lin, C. H. et al. Glutamate-cysteine ligase catalytic subunit as a therapeutic target in acute myeloid leukemia and solid tumors. Am. J. Cancer Res. 11, 2911–2927 (2021).
Ogiwara, H. et al. Targeting the vulnerability of glutathione metabolism in ARID1A-deficient cancers. Cancer Cell 35, 177–190.e178 (2019).
Wu, R. C., Wang, T. L. & Shih Ie, M. The emerging roles of ARID1A in tumor suppression. Cancer Biol. Ther. 15, 655–664 (2014).
Li, W. et al. A pan-cancer analysis reveals the diagnostic and prognostic role of CDCA2 in low-grade glioma. PLOS ONE 18, e0291024 (2023).
Veeriah, S. et al. The tyrosine phosphatase PTPRD is a tumor suppressor that is frequently inactivated and mutated in glioblastoma and other human cancers. Proc. Natl Acad. Sci. USA 106, 9435–9440 (2009).
Sherry, S. T., Ward, M. & Sirotkin, K. dbSNP-database for single nucleotide polymorphisms and other classes of minor genetic variation. Genome Res. 9, 677–679 (1999).
Rafnar, T. et al. Variants associating with uterine leiomyoma highlight genetic background shared by various cancers and hormone-related traits. Nat. Commun. 9, 3636 (2018).
Harris, H. R. et al. Racial differences in the association of endometriosis and uterine leiomyomas with the risk of ovarian cancer. Obstet. Gynecol. 141, 1124–1138 (2023).
Broggini, M. et al. Cell cycle-related phosphatases CDC25A and B expression correlates with survival in ovarian cancer patients. Anticancer Res. 20, 4835–4840 (2000).
Schraml, P. et al. Cyclin E overexpression and amplification in human tumours. J. Pathol. 200, 375–382 (2003).
Cancer Genome Atlas Research, N Integrated genomic analyses of ovarian carcinoma. Nature 474, 609–615 (2011).
Goundiam, O. et al. Histo-genomic stratification reveals the frequent amplification/overexpression of CCNE1 and BRD4 genes in non-BRCAness high grade ovarian carcinoma. Int. J. Cancer 137, 1890–1900 (2015).
Kang, E. Y. et al. CCNE1 and survival of patients with tubo-ovarian high-grade serous carcinoma: an Ovarian Tumor Tissue Analysis consortium study. Cancer 129, 697–713 (2023).
Cancer Genome Atlas, N Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012).
Curtis, C. et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486, 346–352 (2012).
Jiang, Y. Z. et al. Genomic and transcriptomic landscape of triple-negative breast cancers: subtypes and treatment strategies. Cancer Cell 35, 428–440.e425 (2019).
Jones, R. M. et al. Increased replication initiation and conflicts with transcription underlie Cyclin E-induced replication stress. Oncogene 32, 3744–3753 (2013).
Gallo, D. et al. CCNE1 amplification is synthetic lethal with PKMYT1 kinase inhibition. Nature 604, 749–756 (2022).
Lee, A. et al. Comprehensive epithelial tubo-ovarian cancer risk prediction model incorporating genetic and epidemiological risk factors. J. Med. Genet. 59, 632–643 (2022).
Lee, A. et al. Enhancing the BOADICEA cancer risk prediction model to incorporate new data on RAD51C, RAD51D, BARD1 updates to tumour pathology and cancer incidence. J. Med. Genet. 59, 1206–1218 (2022).
Yang, X. et al. Validation of the BOADICEA model for epithelial tubo-ovarian cancer risk prediction in UK Biobank. Br J Cancer 131, 1473–1479 (2024).
Milne, R. L. et al. Assessing interactions between the associations of common genetic susceptibility variants, reproductive history and body mass index with breast cancer risk in the breast cancer association consortium: a combined case-control study. Breast Cancer Res. 12, R110 (2010).
Nickels, S. et al. Evidence of gene-environment interactions between common breast cancer susceptibility loci and established environmental risk factors. PLoS Genet. 9, e1003284 (2013).
Rudolph, A. et al. Investigation of gene-environment interactions between 47 newly identified breast cancer susceptibility loci and environmental risk factors. Int. J. Cancer 136, E685–E696 (2015).
Clyde, M. A. et al. Risk prediction for epithelial ovarian cancer in 11 United States-based case-control studies: incorporation of epidemiologic risk factors and 17 confirmed genetic loci. Am. J. Epidemiol. 184, 579–589 (2016).
Rudolph, A. et al. Joint associations of a polygenic risk score and environmental risk factors for breast cancer in the Breast Cancer Association Consortium. Int. J. Epidemiol. 47, 526–536 (2018).
Vachon, C. M. et al. Joint association of mammographic density adjusted for age and body mass index and polygenic risk score with breast cancer risk. Breast Cancer Res. 21, 68 (2019).
Kachuri, L. et al. Pan-cancer analysis demonstrates that integrating polygenic risk scores with modifiable risk factors improves risk prediction. Nat. Commun. 11, 6084 (2020).
Kapoor, P. M. et al. Assessment of interactions between 205 breast cancer susceptibility loci and 13 established risk factors in relation to breast cancer risk, in the Breast Cancer Association Consortium. Int. J. Epidemiol. 49, 216–232 (2020).
Archambault, A. N. et al. Risk stratification for early-onset colorectal cancer using a combination of genetic and environmental risk scores: an international multi-center study. J. Natl Cancer Inst. 114, 528–539 (2022).
Wang, X. et al. Genome-wide interaction analysis of menopausal hormone therapy use and breast cancer risk among 62,370 women. Sci. Rep. 12, 6199 (2022).
Brnich, S. E. et al. Recommendations for application of the functional evidence PS3/BS3 criterion using the ACMG/AMP sequence variant interpretation framework. Genome Med. 12, 3 (2019).
Sakaue, S. et al. A cross-population atlas of genetic associations for 220 human phenotypes. Nat. Genet. 53, 1415–1424 (2021).
Gaudet, M. M. et al. Identification of a BRCA2-specific modifier locus at 6p24 related to breast cancer risk. PLoS Genet. 9, e1003173 (2013).
Amos, C. I. et al. The OncoArray Consortium: a network for understanding the genetic architecture of common cancers. Cancer Epidemiol. Biomark. Prev. 26, 126–135 (2017).
Milne, R. L. et al. Identification of ten variants associated with risk of estrogen-receptor-negative breast cancer. Nat. Genet. 49, 1767–1778 (2017).
TOPMed. TOPMed Imputation Server, https://imputation.biodatacatalyst.nhlbi.nih.gov/ (2023).
Loh, P. R. et al. Reference-based phasing using the Haplotype Reference Consortium panel. Nat. Genet 48, 1443–1448 (2016).
Howie, B., Fuchsberger, C., Stephens, M., Marchini, J. & Abecasis, G. R. Fast and accurate genotype imputation in genome-wide association studies through pre-phasing. Nat. Genet 44, 955–959 (2012).
Fuchsberger, C., Abecasis, G. R. & Hinds, D. A. minimac2: faster genotype imputation. Bioinformatics 31, 782–784 (2015).
Consortium, U. K. et al. The UK10K project identifies rare variants in health and disease. Nature 526, 82–90 (2015).
UK10K. UK10K Project, https://www.uk10k.org/ (2014).
Permuth-Wey, J. et al. LIN28B polymorphisms influence susceptibility to epithelial ovarian cancer. Cancer Res 71, 3896–3903 (2011).
Li, Y. et al. FastPop: a rapid principal component derived method to infer intercontinental ancestry using genetic data. BMC Bioinforma. 17, 122 (2016).
Antoniou, A. C. et al. RAD51 135G->C modifies breast cancer risk among BRCA2 mutation carriers: results from a combined analysis of 19 studies. Am. J. Hum. Genet. 81, 1186–1200 (2007).
Barnes, D. R. et al. Evaluation of association methods for analysing modifiers of disease risk in carriers of high-risk mutations. Genet. Epidemiol. 36, 274–291 (2012).
Antoniou, A. C. et al. A locus on 19p13 modifies risk of breast cancer in BRCA1 mutation carriers and is associated with hormone receptor-negative breast cancer in the general population. Nat. Genet. 42, 885–892 (2010).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Yang, J. et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, 369–375 (2012).
Zhao, J. H. gap: genetic analysis package. J. Stat. Softw. 23, 1–18 (2007).
Staley, J. R. et al. PhenoScanner: a database of human genotype-phenotype associations. Bioinformatics 32, 3207–3209 (2016).
Kamat, M. A. et al. PhenoScanner V2: an expanded tool for searching human genotype-phenotype associations. Bioinformatics 35, 4851–4853 (2019).
PhenoScanner. PhenoScanner V2: A database of human genotype-phenotype associations, http://www.phenoscanner.medschl.cam.ac.uk/ (2023).
PheWeb. PheWeb, https://pheweb.org/UKB-TOPMed/ (2023).
Vosa, U. et al. Large-scale cis- and trans-eQTL analyses identify thousands of genetic loci and polygenic scores that regulate blood gene expression. Nat. Genet. 53, 1300–1310 (2021).
eQTLGen. eQTLGen, https://www.eqtlgen.org/ (2023).
Udler, M. S., Tyrer, J. & Easton, D. F. Evaluating the power to discriminate between highly correlated SNPs in genetic association studies. Genet Epidemiol. 34, 463–468 (2010).
Tyrer, J. P. et al. Improving on polygenic scores across complex traits using select and shrink with summary statistics (S4) and LDpred2. BMC Genomics. 25, 878 (2024).
Robin, X. et al. pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinforma. 12, 77 (2011).
Antoniou, A. C. et al. Common breast cancer susceptibility alleles and the risk of breast cancer for BRCA1 and BRCA2 mutation carriers: implications for risk prediction. Cancer Res. 70, 9742–9754 (2010).
UK, C. R. Cancer Research UK: Ovarian Cancer incidence by age, https://www.cancerresearchuk.org/health-professional/cancer-statistics/statistics-by-cancer-type/ovarian-cancer/incidence#heading-One (2024).
Kuchenbaecker, K. B. et al. Risks of breast, ovarian, and contralateral breast cancer for BRCA1 and BRCA2 mutation carriers. JAMA 317, 2402–2416 (2017).
Acknowledgements
This work was supported by Cancer Research UK grant: PPRPGM-Nov20\100002 and PRCPJT-May21\100006; by core funding from the NIHR Cambridge Biomedical Research Centre (NIHR203312) [*]. *The views expressed are those of the author(s) and not necessarily those of the NIHR or the Department of Health and Social Care. The Ovarian Cancer Association Consortium is supported by a grant from the Ovarian Cancer Research Fund thanks to donations by the family and friends of Kathryn Sladek Smith (PPD/RPCI.07). The scientific development and funding for this project were in part supported by the US National Cancer Institute GAME-ON Post-GWAS Initiative (U19-CA148112). This study made use of data generated by the Wellcome Trust Case Control consortium that was funded by the Wellcome Trust under award 076113. The OCAC OncoArray genotyping project was funded through grants from the U.S. National Institutes of Health (CA1X01HG007491-01 (C.I.A.), U19-CA148112 (T.A.S.), R01-CA149429 (C.M.P.) and R01-CA058598 (M.T.G.); Canadian Institutes of Health Research (MOP-86727 (L.E.K.) and the Ovarian Cancer Research Fund (A.B.). The COGS project was funded through a European Commission’s Seventh Framework Programme grant (agreement number 223175 - HEALTH-F2-2009-223175). Funding for individual studies: A.A.S.: National Institutes of Health (RO1-CA142081); A.O.V.: The Canadian Institutes for Health Research (MOP-86727); A.U.S.: The Australian Ovarian Cancer Study Group was supported by the U.S. Army Medical Research and Materiel Command (DAMD17-01-1-0729), National Health & Medical Research Council of Australia (199600, 400413 and 400281), Cancer Councils of New South Wales, Victoria, Queensland, South Australia and Tasmania and Cancer Foundation of Western Australia (Multi-State Applications 191, 211 and 182). The Australian Ovarian Cancer Study gratefully acknowledges additional support from Ovarian Cancer Australia and the Peter MacCallum Foundation; B.A.V.: ELAN Funds of the University of Erlangen-Nuremberg; B.E.L.: National Kankerplan; B.G.S.: Breast Cancer Now and the UK NIHR Biomedical Research Centre at the Institute of Cancer Research. B.V.U.: National Institutes of Health/National Center for Advancing Translational Sciences (ULTR000445 and 1S10RR025141-01); C.A.M.: National Institutes of Health Research Cambridge Biomedical Research Centre and Cancer Research UK Cambridge Cancer Centre; C.H.A.: Innovative Research Team in University (PCSIRT) in China (IRT1076); C.N.I.: Instituto de Salud Carlos III (PI 12/01319); Ministerio de Economía y Competitividad (SAF2012); C.O.E.: Department of Defense (W81XWH-11-2-0131); C.O.N.: National Institutes of Health (R01-CA063678, R01-CA074850; R01-CA080742); D.K.E.: Ovarian Cancer Research Fund; D.O.V.: National Institutes of Health (R01-CA112523 and R01-CA87538); E.M.C.: Dutch Cancer Society (EMC 2014-6699); E.P.C.: The coordination of EPIC is financially supported by the European Commission (DG-SANCO) and the International Agency for Research on Cancer. The national cohorts are supported by Danish Cancer Society (Denmark); Ligue Contre le Cancer, Institut Gustave Roussy, Mutuelle Générale de l’Education Nationale, Institut National de la Santé et de la Recherche Médicale (INSERM) (France); German Cancer Aid, German Cancer Research Center (DKFZ), Federal Ministry of Education and Research (BMBF) (Germany); the Hellenic Health Foundation (Greece); Associazione Italiana per la Ricerca sul Cancro-AIRC-Italy and National Research Council (Italy); Dutch Ministry of Public Health, Welfare and Sports (VWS), Netherlands Cancer Registry (NKR), LK Research Funds, Dutch Prevention Funds, Dutch ZON (Zorg Onderzoek Nederland), World Cancer Research Fund (WCRF), Statistics Netherlands (The Netherlands); ERC-2009-AdG 232997 and Nordforsk, Nordic Centre of Excellence programme on Food, Nutrition and Health (Norway); Health Research Fund (FIS), PI13/00061 to Granada, PI13/01162 to EPIC-Murcia, Regional Governments of Andalucía, Asturias, Basque Country, Murcia and Navarra, ISCIII RETIC (RD06/0020) (Spain); Swedish Cancer Society, Swedish Research Council and County Councils of Skåne and Västerbotten (Sweden); Cancer Research UK (14136, C570/A16491 and C8221/A19170), UK Medical Research Council (1000143 and MR/M012190/1); G.E.R.: German Federal Ministry of Education and Research, Programme of Clinical Biomedical Research (01 GB 9401) and the German Cancer Research Center; G.R.C.: the European Union (European Social Fund - ESF) and Greek national funds through the Operational Program “Education and Lifelong Learning” of the National Strategic Reference Framework - Research Funding Program of the General Secretariat for Research & Technology (SYN11_10_19); G.R.R.: Roswell Park Cancer Institute Alliance Foundation (P30 CA016056); H.A.W.: U.S. National Institutes of Health (R01-CA58598, N01-CN-55424 and N01-PC-67001); H.J.O.: German Research Foundation (Do 761/15-1); Rudolf-Bartling Foundation; H.M.O.: Intramural funding; Rudolf-Bartling Foundation; H.O.C.: Helsinki University Research Fund; H.O.P.: Department of Defense (DAMD17-02-1-0669) and NCI (K07-CA080668, R01-CA95023, P50-CA159981 MO1-RR000056 R01-CA126841); H.U.O.: German Research Foundation (Do 761/15-1); Rudolf-Bartling Foundation; J.G.O.: JSPS KAKENHI grant; J.P.N.: Grant-in-Aid for the Third Term Comprehensive 10-Year Strategy for Cancer Control from the Ministry of Health, Labour and Welfare; K.R.A.: Korea Health Technology R&D Project through the Korea Health Industry Development Institute, and the National R&D Program for Cancer Control, Ministry of Health & Welfare, Republic of Korea (HI16C1127; 0920010); L.A.X.: American Cancer Society Early Detection Professorship (SIOP-06-258-01-COUN) and the National Center for Advancing Translational Sciences (UL1TR000124); L.U.N.: European Research Council (ERC-2011-AdG 294576), Swedish Cancer Society, Swedish Research Council, Beta Kamprad Foundation; M.A.C.: National Institutes of Health (R01-CA122443, P30-CA15083, P50-CA136393); Mayo Foundation; Minnesota Ovarian Cancer Alliance; Fred C. and Katherine B. Andersen Foundation; Fraternal Order of Eagles; M.A.L.: National Institutes for Health (R01- CA61107) Danish Cancer Society (94 222 52) from the, Copenhagen, Denmark; and the Mermaid I project; M.A.S.: Malaysian Ministry of Higher Education (UM.C/HlR/MOHE/06) and Cancer Research Initiatives Foundation; M.A.Y.: National Institutes of Health (R01-CA122443, P30-CA15083, P50-CA136393); Mayo Foundation; Minnesota Ovarian Cancer Alliance; Fred C. and Katherine B. Andersen Foundation; M.C.C.: Cancer Council Victoria, National Health and Medical Research Council of Australia (NHMRC) grants number 209057, 251533, 396414, and 504715; M.D.A.: DOD Ovarian Cancer Research Program (W81XWH-07-0449); M.E.C.: NIH (CA54281, CA164973, CA63464); M.O.F.: Moffitt Cancer Center, Merck Pharmaceuticals, the state of Florida, Hillsborough County, and the city of Tampa; N.C.O.: National Institutes of Health (R01-CA76016) and the Department of Defense (DAMD17-02-1-0666); N.E.C.: National Institutes of Health R01-CA54419 and P50-CA105009 and Department of Defense W81XWH-10-1-02802; N.H.S.: UM1 CA186107, P01 CA87969, R01 CA49449, R01-CA67262, UM1 CA176726; N.J.O.: National Cancer Institute (NIH-K07 CA095666, R01-CA83918, NIH-K22-CA138563, and P30-CA072720) and the Cancer Institute of New Jersey; If Sara Olson and/or Irene Orlow is a co-author, please add NCI CCSG award (P30-CA008748) to the funding sources; N.O.R.: Helse Vest, The Norwegian Cancer Society, The Research Council of Norway; N.T.H.: Radboud University Medical Centre; O.P.L.: National Health and Medical Research Council (NHMRC) of Australia (APP1025142) and Brisbane Women’s Club; O.R.E.: OHSU Foundation; O.V.A.: This work was supported by Canadian Institutes of Health Research grant (MOP-86727) and by NIH/NCI 1 R01CA160669-01A1; P.L.C.: Intramural Research Program of the National Cancer Institute; P.O.C.: Pomeranian Medical University; P.O.L.: Intramural Research Program of the National Cancer Institute; P.V.D.: Canadian Cancer Society and Cancer Research Society GRePEC Program; R.B.H.: National Health and Medical Research Council of Australia; R.M.H.: Cancer Research UK, Royal Marsden Hospital; R.P.C.: National Institute of Health (P50 CA159981, R01CA126841); S.E.A.: Cancer Research UK (C490/A10119 C490/A10124 C490/A16561); UK National Institute for Health Research Biomedical Research Centres at the University of Cambridge; S.I.S.: NIH, National Institute of Environmental Health Sciences, Z01 ES044005 and Z01-ES049033; S.M.C.: The Swedish Research Council; S.O.N.: National Health Research and Development Program, Health Canada, grant 6613-1415-53; S.R.O.: Cancer Research UK (C536/A13086, C536/A6689) and Imperial Experimental Cancer Research Centre (C1312/A15589); S.T.A.: NIH grants U01 CA71966 and U01 CA69417; S.W.E.: Swedish Cancer foundation, WeCanCureCancer and årKampMotCancer foundation; S.W.H.: NIH (NCI) grant R37-CA070867; T.B.O.: National Institutes of Health (R01-CA106414-A2), American Cancer Society (CRTG-00-196-01-CCE), Department of Defense (DAMD17-98-1-8659), Celma Mastery Ovarian Cancer Foundation; T.O.R.: NIH grants R01 CA063678 and R01 CA063682; U.C.I.: NIH R01-CA058860 and the Lon V Smith Foundation grant LVS-39420; U.H.N.: Princess Margaret Cancer Centre Foundation-Bridge for the Cure; U.K.O.: The UKOPS study was funded by The Eve Appeal (The Oak Foundation) and supported by the National Institute for Health Research University College London Hospitals Biomedical Research Centre; U.K.R.: Cancer Research UK (C490/A6187), UK National Institute for Health Research Biomedical Research Centres at the University of Cambridge; U.S.C.: P01CA17054, P30CA14089, R01CA61132, N01PC67010, R03CA113148, R03CA115195, N01CN025403, and California Cancer Research Program (00-01389V-20170, 2II0200); V.A.N.: BC Cancer Foundation, VGH & UBC Hospital Foundation; V.T.L.: NIH K05-CA154337; W.M.H.: National Health and Medical Research Council of Australia, Enabling Grants ID 310670 & ID 628903. Cancer Institute NSW Grants 12/RIG/1-17 & 15/RIG/1-16; W.O.C.: National Science Centren (N N301 5645 40) The Maria Sklodowska-Curie Memorial Cancer Center and Institute of Oncology, Warsaw, Poland. This work was supported by the Gray Foundation. GCT and ABS are NHMRC Research Fellows. iCOGS: the European Community’s Seventh Framework Programme under grant agreement n° 223175 (HEALTH-F2-2009-223175) (COGS), Cancer Research UK (C1287/A10118, C1287/A 10710, C12292/A11174, C1281/A12014, C5047/A8384, C5047/A15007, C5047/A10692, C8197/A16565), the National Institutes of Health (CA128978) and Post-Cancer GWAS initiative (1U19 CA148537, 1U19 CA148065 and 1U19 CA148112 - the GAME-ON initiative), the Department of Defence (W81XWH-10-1-0341), the Canadian Institutes of Health Research (CIHR) for the CIHR Team in Familial Risks of Breast Cancer (CRN-87521), and the Ministry of Economic Development, Innovation and Export Trade (PSR-SIIRI-701), Komen Foundation for the Cure, the Breast Cancer Research Foundation, and the Ovarian Cancer Research Fund. OncoArray: the PERSPECTIVE and PERSPECTIVE I&I projects funded by the Government of Canada through Genome Canada and the Canadian Institutes of Health Research, the ‘Ministère de l’Économie, de la Science et de l’Innovation du Québec’ through Genome Québec, and the Quebec Breast Cancer Foundation; the NCI Genetic Associations and Mechanisms in Oncology (GAME-ON) initiative and Discovery, Biology and Risk of Inherited Variants in Breast Cancer (DRIVE) project (NIH Grants U19 CA148065 and X01HG007492); and Cancer Research UK (C1287/A10118 and C1287/A16563). ACBRCA: FAPESP/Fundação de Amparo à Pesquisa do Estado de São Paulo (2014/50943-1) São Paulo, Brazil, CNPq/Conselho Nacional de Desenvolvimento Científico e Tecnológico (465682/2014-6), Brazil. BCFR: U01 CA164920 from the National Cancer Institute. The content of this manuscript does not necessarily reflect the views or policies of the National Cancer Institute or any of the collaborating centers in the Breast Cancer Family Registry (BCFR), nor does mention of trade names, commercial products, or organizations imply endorsement by the US Government or the BCFR. BFBOCC: Lithuania (BFBOCC-LT): Research Council of Lithuania grant SEN-18/2015. BIDMC: Breast Cancer Research Foundation. BMBSA: Cancer Association of South Africa (PI Elizabeth J. van Rensburg). BRICOH: SLN was partially supported by the Morris and Horowitz Families Endowed Professorship. CNIO: CNIO study is partially funded by Instituto de Salud Carlos III, project reference PI19/00640, cofunded by the European Regional Development Fund (ERDF), “A way to make Europe” and the Spanish Network on Rare Diseases (CIBERER). CCGCRN: Research reported in this publication was supported by the Breast Cancer Research Foundation (project 20-172), National Cancer Institute of the National Institutes of Health under grant number R25CA112486, and RC4CA153828 (PI: J. Weitzel) from the National Cancer Institute and the Office of the Director, National Institutes of Health. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. CONSIT TEAM: Associazione Italiana Ricerca sul Cancro (AIRC; IG2015 no.16732) to P. Peterlongo. This work was partially supported by the Italian Ministry of Health with Ricerca Corrente and 5×1000 Funds (European Institute of Oncology). CZECANCA: Charles University/VFN projects Cooperatio - Laboratory Diagnostics/Medical Genetics, NU20-03-00285, NU20-09-00355, NU20-03-00016/ RVO-VFN 64165. DEMOKRITOS: European Union (European Social Fund – ESF) and Greek national funds through the Operational Program “Education and Lifelong Learning” of the National Strategic Reference Framework (NSRF) - Research Funding Program of the General Secretariat for Research & Technology: SYN11_10_19 NBCA. Investing in knowledge society through the European Social Fund. DFKZ: German Cancer Research Center. EMBRACE: Cancer Research UK Grants PRCPJT-Nov21\100004, C1287/A23382 and C1287/A26886. D. Gareth Evans and Fiona Lalloo are supported by an NIHR grant to the Biomedical Research Centre, Manchester. The Investigators at The Institute of Cancer Research and The Royal Marsden NHS Foundation Trust are supported by an NIHR grant to the Biomedical Research Centre at The Institute of Cancer Research and The Royal Marsden NHS Foundation Trust. Ros Eeles and Elizabeth Bancroft are supported by Cancer Research UK Grant C5047/A8385. Ros Eeles is also supported by NIHR support to the Biomedical Research Centre at The Institute of Cancer Research and The Royal Marsden NHS Foundation Trust. The FCCC and KUMC cohorts: The University of Kansas Cancer Center (P30 CA168524), the Kansas Institute for Precision Medicine (P20GM130423) and the Kansas Bioscience Authority Eminent Scholar Program. A.K.G. was funded by R01CA140323, R01CA214545, R01CA260132, 5U10CA180888, and by the Chancellors Distinguished Chair in Biomedical Sciences Professorship. FPGMX: A.Vega is supported by the Spanish Health Research Foundation, Instituto de Salud Carlos III (ISCIII), partially supported by FEDER funds through Research Activity Intensification Program (contract grant numbers: INT15/00070, INT16/00154, INT17/00133, INT20/00071), and through Centro de Investigación Biomédica en Red de Enferemdades Raras CIBERER (ACCI 2016: ER17P1AC7112/2018); Autonomous Government of Galicia (Consolidation and structuring program: IN607B), and by the Fundación Mutua Madrileña and Asociación Española Contra el Cáncer (AECC). GC-HBOC: German Cancer Aid (grant no 110837 and 113049, Rita K. Schmutzler), Federal Ministry of Education and Research, Germany (grant no 01GY1901), and the European Regional Development Fund and Free State of Saxony, Germany (LIFE - Leipzig Research Centre for Civilization Diseases, project numbers 713-241202, 713-241202, 14505/2470, 14575/2470). GEMO, a study from the National Cancer Genetics Network UNICANCER Genetic Group, France.: Ligue Nationale Contre le Cancer; the Association “Le cancer du sein, parlons-en!” Award, the Canadian Institutes of Health Research for the “CIHR Team in Familial Risks of Breast Cancer” program, the Fondation ARC pour la recherche sur le cancer (grant PJA 20151203365) and the French National Institute of Cancer (INCa grants AOR 01 082, 2001-2003, 2013-1-BCB-01-ICH-1 and SHS-E-SP 18-015). GEORGETOWN: the Non-Therapeutic Subject Registry Shared Resource at Georgetown University (NIH/NCI grant P30-CA051008), the Fisher Center for Hereditary Cancer and Clinical Genomics Research, and Swing Fore the Cure. HCSC: Spanish Ministry of Science and Innovation, ISCIII (Hayley) co-funded by FEDER Regional Development European Funds (EU). HEBCS: Helsinki University Hospital Research Fund, the Finnish Cancer Society and the Sigrid Juselius Foundation. The Hereditary Breast and Ovarian Cancer Research Group Netherlands (HEBON) consists of the following Collaborating Centers: Netherlands Cancer Institute; Erasmus Medical Center, Rotterdam; Leiden University Medical Center; Radboud University Nijmegen Medical Center; University Medical Center Utrecht; Amsterdam UMC, Univ of Amsterdam; Amsterdam UMC, Vrije Universiteit Amsterdam; Maastricht University Medical Center; University of Groningen; and acknowledges contributions of the Netherlands Comprehensive Cancer Organisation (IKNL), the nationwide network and registry of histo- and cytopathology in The Netherlands (PALGA), the Dutch Cancer Society grants NKI1998-1854, NKI2004-3088, NKI2007-3756, the Netherlands Organization of Scientific Research grant NWO 91109024, the Pink Ribbon grants 110005 and 2014-187.WO76, the BBMRI grant NWO 184.021.007/CP46 and the Transcan grant JTC 2012 Cancer 12-054. HEBCS: The Helsinki University Hospital Research Fund, The Sigrid Jusélius Foundation, The Cancer Foundation Finland. HRBCP: Hong Kong Sanatorium and Hospital, Dr Ellen Li Charitable Foundation, The Kerry Group Kuok Foundation, National Institute of Health1R 03CA130065, and North California Cancer Center. HUNBOCS: Hungarian Research Grants KTIA-OTKA CK-80745, NKFI_OTKA K-112228, and NKFI-FK-135065. HVH/VHIO: Orland Díez, Sara Gutiérrez-Enríquez and Judith Balmaña were funded by the Spanish Instituto de Salud Carlos III (ISCIII), an initiative of the Spanish Ministry of Economy and Innovation, partially supported by European Regional Development FEDER Funds, grant numbers PI16/01218, PI16/11363, PI19/01195 and PI19/01303 ICO: The authors would like to particularly acknowledge the support of the Asociación Española Contra el Cáncer (AECC), the Instituto de Salud Carlos III (organismo adscrito al Ministerio de Economía y Competitividad) and “Fondo Europeo de Desarrollo Regional (FEDER), una manera de hacer Europa” (PI10/01422, PI13/00285, PIE13/00022, PI15/00854, PI16/00563 and CIBERONC) and the Institut Català de la Salut and Autonomous Government of Catalonia (2009SGR290, 2014SGR338 and PERIS Project MedPerCan). ICARE: Inherited Cancer Registry (ICARE) based at the Vanderbilt-Ingram Cancer Center is funded, in part, through the Ingram Professorship. IFBCRC (Iranian Familial Breast Cancer Research Consortium) is funded by MCI (Motamed Cancer Institute) of ACECR. IHCC: PBZ_KBN_122/P05/2004 and the program of the Minister of Science and Higher Education under the name “Regional Initiative of Excellence” in 2019–2022 project number 002/RID /2018/19 amount of financing 12 000 000 PLN. INHERIT: Canadian Institutes of Health Research for the “CIHR Team in Familial Risks of Breast Cancer” program – grant # CRN-87521 and the Ministry of Economic Development, Innovation and Export Trade – grant # PSR-SIIRI-701. IOVHBOCS: Ministero della Salute and “5x1000” Istituto Oncologico Veneto grant. IPOBCS: Liga Portuguesa Contra o Cancro. kConFab: The National Breast Cancer Foundation, and previously by the National Health and Medical Research Council (NHMRC), the Queensland Cancer Fund, the Cancer Councils of New South Wales, Victoria, Tasmania and South Australia, and the Cancer Foundation of Western Australia. KOHBRA: the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), and the National R&D Program for Cancer Control, Ministry of Health & Welfare, Republic of Korea (HI16C1127; 1020350; 1420190). MAYO: NIH grants CA116167, CA192393, CA176785, and CA253187 an NCI Specialized Program of Research Excellence (SPORE) in Breast Cancer (CA116201),and a grant from the Breast Cancer Research Foundation. The MACBRCA study is funded by the Research Centre for Genetic Engineering and Biotechnology (RCGEB) “Georgi D. Efremov”, Macedonian Academy of Sciences and Arts (MASA). MCGILL: Jewish General Hospital Weekend to End Breast Cancer, Quebec Ministry of Economic Development, Innovation and Export Trade. Marc Tischkowitz is supported by an NIHR grant to the Biomedical Research Centre, Cambridge. MODSQUAD: MH CZ - DRO (MMCI, 00209805), and by Charles University in Prague project UNCE204024 (MZ). MSKCC: the Breast Cancer Research Foundation, the Robert and Kate Niehaus Clinical Cancer Genetics Initiative, the Andrew Sabin Research Fund and a Cancer Center Support Grant/Core Grant (P30 CA008748). NAROD: 1R01 CA149429-01. NCCS: NMRC Clinician Scientist Award (CSA), Terry Fox Foundation, Lee Foundation, PRECISE, Ministry of Education and NCCS Cancer Fund. NCI: the Intramural Research Program of the US National Cancer Institute, NIH, and by support services contracts NO2-CP-11019-50, N02-CP-21013-63 and N02-CP-65504 with Westat, Inc, Rockville, MD. NICCC: Clalit Health Services in Israel, the Israel Cancer Association and the Breast Cancer Research Foundation (BCRF), NY. NNPIO: Russian Science Foundation (grant 21-75-30015). NRG Oncology: U10 CA180868, NRG SDMC grant U10 CA180822, NRG Administrative Office and the NRG Tissue Bank (CA 27469), the NRG Statistical and Data Center (CA 37517) and the Intramural Research Program, NCI. OSUCCG: Ohio State University Comprehensive Cancer Center. OBRCA: funding from the Norwegian Cancer Society, contract 194751-2017. PBCS: Italian Association of Cancer Research (AIRC) [IG 2013 N.14477] and Tuscany Institute for Tumors (ITT) grant 2014-2015-2016. PCCM-CTCR is supported by the Corporate Social Responsibility Fund from Bajaj Auto Ltd, Grant GC-2528. SEABASS: Ministry of Science, Technology and Innovation, Ministry of Higher Education (UM.C/HlR/MOHE/06) and Cancer Research Initiatives Foundation. SGBCC was supported by the National Research Foundation Singapore (NRF-NRFF2017-02), NUS start-up Grant, National University Cancer Institute Singapore (NCIS) Centre Grant [NMRC/CG/NCIS/2010, NMRC/CG/012/2013, CGAug16M005], Breast Cancer Prevention Programme (BCPP), Asian Breast Cancer Research Fund, and the NMRC Clinician Scientist Award (SI Category) [NMRC/CSA-SI/0015/2017]. SMC: the Israeli Cancer Association. SWE-BRCA: the Swedish Cancer Society. UCHICAGO: NCI Specialized Program of Research Excellence (SPORE) in Breast Cancer (CA125183), R01 CA142996, 1U01CA161032 and by the Ralph and Marion Falk Medical Research Trust, the Entertainment Industry Fund National Women’s Cancer Research Alliance and the Breast Cancer research Foundation. OIO is an ACS Clinical Research Professor. UCLA: Jonsson Comprehensive Cancer Center Foundation; Breast Cancer Research Foundation. UCSF: UCSF Cancer Risk Program and Helen Diller Family Comprehensive Cancer Center. UKFOCR: Cancer Research UK. UPENN: Breast Cancer Research Foundation; Susan G. Komen Foundation for the cure, Basser Research Center for BRCA, NCI P30 CA016520. UPITT/MWH: Hackers for Hope Pittsburgh. VFCTG: Victorian Cancer Agency, Cancer Australia, National Breast Cancer Foundation. WCP: Dr Karlan is funded by the American Cancer Society Early Detection Professorship (SIOP-06-258-01-COUN) and the National Center for Advancing Translational Sciences (NCATS), Grant UL1TR000124. We are grateful to the family and friends of Kathryn Sladek Smith for their generous support of the Ovarian Cancer Association Consortium through their donations to the Ovarian Cancer Research Fund. The OncoArray and COGS genotyping projects would not have been possible without the contributions of the following: Per Hall (COGS); Douglas F. Easton, Kyriaki Michailidou, Manjeet K. Bolla, Qin Wang (BCAC), Marjorie J. Riggan (OCAC), Rosalind A. Eeles, Ali Amin Al Olama, Zsofia Kote-Jarai, Sara Benlloch (PRACTICAL), Joe Dennis, Alison M. Dunning, Andrew Lee, Ed Dicks, Craig Luccarini and the staff of the Centre for Genetic Epidemiology Laboratory, Javier Benitez, Anna Gonzalez-Neira and the staff of the CNIO genotyping unit, Jacques Simard and Daniel C. Tessier, Francois Bacot, Daniel Vincent, Sylvie LaBoissière and Frederic Robidoux and the staff of the McGill University and Génome Québec Innovation Centre, Stig E. Bojesen, Sune F. Nielsen, Borge G. Nordestgaard, and the staff of the Copenhagen DNA laboratory, and Julie M. Cunningham, Sharon A. Windebank, Christopher A. Hilker, Jeffrey Meyer and the staff of Mayo Clinic Genotyping Core Facility. We thank the study participants, doctors, nurses, clinical and scientific collaborators, health care providers and health information sources who have contributed to the many studies contributing to this manuscript. We thank all the women who have taken part in the contributing studies and we acknowledge the many study nurses, research assistants and all clinical and scientific collaborators. We thank the following individuals for their contribution to individual studies: A.O.V.: Jennifer Koziak, Mie Konno, Michelle Darago, Faye Chambers and the Tom Baker Cancer Centre Translational Laboratories; A.U.S.: The complete AOCS Study Group can be found at www.aocstudy.org B.E.L.: Gilian Peuteman, Thomas Van Brussel, Annick Van den Broeck and Joke De Roover; B.V.U.: The dataset(s) used for the analyses described were obtained from; C.A.M.: This work was supported by Cancer Research UK; the University of Cambridge; National Institute for Health Research Cambridge Biomedical Research Centre; C.H.A.: Innovative Research Team in University (PCSIRT) in China (IRT1076); C.H.N.: To thank all members of Department of Obstetrics and Gynaecology, Hebei Medical University, Fourth Hospital and Department of Molecular Biology, Hebei Medical University, Fourth Hospital; C.O.E.: Gynecologic Cancer Center of Excellence (W81XWH-11-2-0131); C.O.N.: The cooperation of the 32 Connecticut hospitals, including Stamford Hospital, in allowing patient access, is gratefully acknowledged. This study was approved by the State of Connecticut Department of Public Health Human Investigation Committee. Certain data used in this study were obtained from the Connecticut Tumor Registry in the Connecticut Department of Public Health. The authors assume full responsibility for analyses and interpretation of these data; D.K.E.: OCRF; E.P.C.: To thank all members and investigators of the Rotterdam Ovarian Cancer Study. Dutch Cancer Society (EMC 2014-6699); G.E.R.: The German Ovarian Cancer Study (GER) thank Ursula Eilber for competent technical assistance; H.O.C.: The study was supported by the Helsinki University Research Fund; J.G.O.: JSPS KAKENHI grant; K.R.A.: This study (Ko-EVE) was supported by a grant from the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), and the National R&D Program for Cancer Control, Ministry of Health & Welfare, Republic of Korea (HI16C1127; 0920010); L.U.N.: ERC -2011-AdG, Swedish Cancer Society, Swedish Research Council; M.A.S.: We would like to thank Famida Zulkifli and Ms Moey for assistance in patient recruitment, data collection and sample preparation. The Malaysian Ovarian Cancer Genetic Study is funded by research grants from the Malaysian Ministry of Higher Education (UM.C/HIR/MOHE/06) and charitable funding from Cancer Research Initiatives Foundation (MAY: P30-CA015083, P50-CA1363936, R01-CA248288); M.C.C.: MCCS cohort recruitment was funded by VicHealth and Cancer Council Victoria. The MCCS was further supported by Australian NHMRC grants 209057, 251553 and 504711 and by infrastructure provided by Cancer Council Victoria. Cases and their vital status were ascertained through the Victorian Cancer Registry (VCR) and the Australian Institute of Health and Welfare (AIHW), including the National Death Index and the Australian Cancer Database; M.O.F.: the Total Cancer Care™ Protocol and the Collaborative Data Services and Tissue Core Facilities at the H. Lee Moffitt Cancer Center & Research Institute, an NCI designated Comprehensive Cancer Center (P30-CA076292), Merck Pharmaceuticals and the state of Florida; N.H.S.: The NHS/NHSII studies thank the following state cancer registries for their help: AL, AZ, AR, CA, CO, CT, DE, FL, GA, ID, IL, IN, IA, KY, LA, ME, MD, MA, MI, NE, NH, NJ, NY, NC, ND, OH, OK, OR, PA, RI, SC, TN, TX, VA, WA, and WY; N.J.O.: contact Elisa Bandera; O.P.L.: Members of the OPAL Study Group (http://opalstudy.qimrberghofer.edu.au/); R.P.C.: National Institute of Health (P50 CA159981, R01CA126841); S.E.A.: SEARCH team, Craig Luccarini, Caroline Baynes, Don Conroy; S.I.S.: The Sister Study (SISTER) is supported by the Intramural Research Program of the NIH, National Institute of Environmental Health Sciences (Z01-ES044005 and Z01-ES049033); S.O.N.: National Health Research and Development Program, Health Canada, grant 6613-1415-53; S.R.O.: To thank all members of Scottish Gynaecological Clinical Trails group and SCOTROC1 investigators; S.W.E.: Swedish Cancer foundation, WeCanCureCancer and årKampMotCancer foundation; S.W.H.: The SWHS is supported primarily by NIH grant R37-CA070867. We thank the participants and the research staff of the Shanghai Women’s Health Study for making this study possible; U.C.I.: The UCI Ovarian cancer study is supported by the National Institutes of Health, National Cancer Institute grants CA58860, and the Lon V Smith Foundation grant LVS-39420; U.H.N.: Princess Margaret Cancer Centre Foundation-Bridge for the Cure; U.K.O.: We particularly thank I. Jacobs, M.Widschwendter, E. Wozniak, A. Ryan, J. Ford and N. Balogun for their contribution to the study; U.K.R.: Carole Pye; V.A.N.: BC Cancer Foundation, VGH & UBC Hospital Foundation; W.M.H.: We thank the Gynaecological Oncology Biobank at Westmead, a member of the Australasian Biospecimen Network-Oncology group, which is funded by the National Health and Medical Research Council Enabling Grants ID 310670 & ID 628903 and the Cancer Institute NSW Grants 12/RIG/1-17 & 15/RIG/1-16 with financial support from the Sydney West Translational Cancer Research Centre, funded by the Cancer Institute NSW (15/TRC/1-01). All the families and clinicians who contribute to the studies; Catherine M. Phelan for her contribution to CIMBA until she passed away on 22 September 2017; Sue Healey, in particular taking on the task of mutation classification with the late Olga Sinilnikova; clinicians, patients, researchers, technicians and nurses of A.C. Camargo Cancer Center for their contribution to this study; Oncogenetic Department, Clinical and Functional Genomics Group, Center of Genomic Diagnostics, Biobank and other International Research Center-CIPE’ facilities at AC. Camargo Cancer Center, especially Karina Miranda Santiago, Giovana Tardin Torrezan, José Claudio Casali, Nirvana Formiga and Fabiana Baroni Makdissi; Maggie Angelakos, Judi Maskiell, Gillian Dite, Helen Tsimiklis; members and participants in the New York site of the Breast Cancer Family Registry; members and participants in the Ontario Familial Breast Cancer Registry; Vilius Rudaitis and Laimonas Griškevičius; Drs Janis Eglitis, Anna Krilova and Aivars Stengrevics; Yuan Chun Ding and Linda Steele for their work in participant enrollment and biospecimen and data management; Bent Ejlertsen and Anne-Marie Gerdes for the recruitment and genetic counseling of participants; Alicia Barroso, Rosario Alonso and Guillermo Pita; all the individuals and the researchers who took part in CONSIT TEAM (Consorzio Italiano Tumori Ereditari Alla Mammella), in particular: Jacopo Azzollini, Bernard Peissel, Daniela Zaffaroni, Laura Ottini, Giuseppe Giannini, Gabriele Lorenzo Capone, Liliana Varesco, Viviana Gismondi, Maria Grazia Tibiletti, Daniela Furlan, Antonella Savarese, Aline Martayan, Stefania Tommasi, Brunella Pilato, Bernardo Bonanni, Maria Rosaria Calvello, Irene Feroce, Monica Marabelli, Matilde Risti, Cristina Zanzottera, Loris Bernard, Elena Marino and the personnel of the Cogentech Cancer Genetic Test Laboratory, Milan, Italy. The FCCC cohort (Godwin) acknowledges Ms. JoEllen Weaver and Dr. Betsy Bove, and the KUMC cohort (Sharma and Godwin) acknowledge the support of Michele Park, Lauren DiMartino, Alex Webster and the current and past members of the Biospecimen Repository Core Facility (BRCF) at KUMC; all participants, clinicians, family doctors, researchers, and technicians for their contributions and commitment to the DKFZ study and the collaborating groups in Lahore, Pakistan (Noor Muhammad, Sidra Gull, Seerat Bajwa, Faiz Ali Khan, Humaira Naeemi, Saima Faisal, Asif Loya, Mohammed Aasim Yusuf) and Bogota, Colombia (Diana Torres, Ignacio Briceno, Fabian Gil). FPGMX: members of the Cancer Genetics group (IDIS): Ana Blanco, Miguel Aguado, Uxía Esperón and Belinda Rodríguez; the GIIS025 research nurses and staff for their contributions to this resource, and the many families who contribute to GIIS025; IFE - Leipzig Research Centre for Civilization Diseases (Markus Loeffler, Joachim Thiery, Matthias Nüchter, Ronny Baber); Genetic Modifiers of Cancer Risk in BRCA1/2 Mutation Carriers (GEMO) study is a study from the National Cancer Genetics Network UNICANCER Genetic Group, France. We wish to pay a tribute to Olga M. Sinilnikova, who with Dominique Stoppa-Lyonnet initiated and coordinated GEMO until she sadly passed away on the 30th June 2014. The team in Lyon (Olga Sinilnikova, Mélanie Léoné, Laure Barjhoux, Carole Verny-Pierre, Sylvie Mazoyer, Francesca Damiola, Valérie Sornin) managed the GEMO samples until the biological resource centre was transferred to Paris in December 2015 (Noura Mebirouk, Fabienne Lesueur, Dominique Stoppa-Lyonnet). We want to thank all the GEMO collaborating groups for their contribution to this study: Coordinating Centre, Service de Génétique, Institut Curie, Paris, France: Muriel Belotti, Ophélie Bertrand, Anne-Marie Birot, Bruno Buecher, Sandrine Caputo, Chrystelle Colas, Emmanuelle Fourme, Marion Gauthier-Villars, Lisa Golmard, Marine Le Mentec, Virginie Moncoutier, Antoine de Pauw, Claire Saule, Dominique Stoppa-Lyonnet, and Inserm U900, Institut Curie, Paris, France: Fabienne Lesueur, Noura Mebirouk, Yue Jiao. Contributing Centres : Unité Mixte de Génétique Constitutionnelle des Cancers Fréquents, Hospices Civils de Lyon - Centre Léon Bérard, Lyon, France: Nadia Boutry-Kryza, Alain Calender, Sophie Giraud, Mélanie Léone. Institut Gustave Roussy, Villejuif, France: Brigitte Bressac-de-Paillerets, Odile Cabaret, Olivier Caron, Marine Guillaud-Bataille, Etienne Rouleau. Centre Jean Perrin, Clermont–Ferrand, France: Yves-Jean Bignon, Nancy Uhrhammer. Centre Léon Bérard, Lyon, France: Valérie Bonadona, Sophie Dussart, Christine Lasset, Pauline Rochefort. Centre François Baclesse, Caen, France: Pascaline Berthet, Laurent Castera, Dominique Vaur. Institut Paoli Calmettes, Marseille, France: Violaine Bourdon, Catherine Noguès, Tetsuro Noguchi, Cornel Popovici, Audrey Remenieras, Hagay Sobol. CHU Arnaud-de-Villeneuve, Montpellier, France: Isabelle Coupier, Pascal Pujol. Centre Oscar Lambret, Lille, France: Claude Adenis, Aurélie Dumont, Françoise Révillion. Centre Paul Strauss, Strasbourg, France: Danièle Muller. Institut Bergonié, Bordeaux, France: Emmanuelle Barouk-Simonet, Françoise Bonnet, Virginie Bubien, Anaïs Dupré, Anne Floquet, Michel Longy, Marie Louty, Cécile Maninna, Nicolas Sevenet, Institut Claudius Regaud, Toulouse, France: Laurence Gladieff, Rosine Guimbaud, Viviane Feillel, Christine Toulas. CHU Grenoble, France: Hélène Dreyfus, Dominique Leroux, Clémentine Legrand, Christine Rebischung. CHU Dijon, France: Amandine Baurand, Geoffrey Bertolone, Fanny Coron, Laurence Faivre, Caroline Jacquot, Sarab Lizard, Sophie Nambot. CHU St-Etienne, France: Caroline Kientz, Marine Lebrun, Fabienne Prieur. Hôtel Dieu Centre Hospitalier, Chambéry, France: Sandra Fert Ferrer. Centre Antoine Lacassagne, Nice, France: Véronique Mari. CHU Limoges, France: Laurence Vénat-Bouvet. CHU Nantes, France: Stéphane Bézieau, Capucine Delnatte. CHU Bretonneau, Tours and Centre Hospitalier de Bourges France: Isabelle Mortemousque. Groupe Hospitalier Pitié-Salpétrière, Paris, France: Florence Coulet, Mathilde Warcoin. CHU Vandoeuvre-les-Nancy, France: Myriam Bronner, Johanna Sokolowska. CHU Besançon, France: Marie-Agnès Collonge-Rame. CHU Poitiers, Centre Hospitalier d’Angoulême and Centre Hospitalier de Niort, France: Stéphanie Chieze-Valero, Paul Gesta, Brigitte Gilbert-Dussardier. Centre Hospitalier de La Rochelle : Hakima Lallaoui. CHU Nîmes Carémeau, France : Jean Chiesa. CHI Poissy, France: Denise Molina-Gomes. CHU Angers, France : Olivier Ingster; CHU de Martinique, France: Odile Bera; Mickaelle Rose; Drs. Taru A. Muranen and Carl Blomqvist, RN Outi Malkavaara; The Hereditary Breast and Ovarian Cancer Research Group Netherlands (HEBON) consists of the following Collaborating Centers: Netherlands Cancer Institute (coordinating center), Amsterdam, NL: M.A. Rookus, F.B.L. Hogervorst, F.E. van Leeuwen, M.A. Adank, M.K. Schmidt, D.J. Jenner; Erasmus Medical Center, Rotterdam, NL: J.M. Collée, A.M.W. van den Ouweland, M.J. Hooning, I.A. Boere; Leiden University Medical Center, NL: C.J. van Asperen, P. Devilee, R.B. van der Luijt, T.C.T.E.F. van Cronenburg; Radboud University Nijmegen Medical Center, NL: M.R. Wevers, A.R. Mensenkamp; University Medical Center Utrecht, NL: M.G.E.M. Ausems, M.J. Koudijs; Amsterdam UMC, Univ of Amsterdam, NL: I. van de Beek; Amsterdam UMC, Vrije Universiteit Amsterdam, NL: K. van Engelen, J.J.P. Gille; Maastricht University Medical Center, NL: E.B. Gómez García, M.J. Blok, M. de Boer; University of Groningen, NL: L.P.V. Berger, A.H. van der Hout, M.J.E. Mourits, G.H. de Bock; The Netherlands Comprehensive Cancer Organisation (IKNL): S. Siesling, J. Verloop; The nationwide network and registry of histo- and cytopathology in The Netherlands (PALGA): E.C. van den Broek; the study participants and the registration teams of IKNL and PALGA for part of the HEBON data collection; Hong Kong Sanatorium and Hospital; the Hungarian Breast and Ovarian Cancer Study Group members (Attila Patócs, János Papp, Anikó Bozsik, Timea Pócza, Zoltán Mátrai, Lajos Géczi, National Institute of Oncology, Budapest, Hungary) and the clinicians and patients for their contributions to this study; Fatemeh Yadegari, Shiva Zarinfam and Rezvan Esmaeili for their role in participant enrollment and biospecimen and data management; the study participants and registration teams of the Hereditary Cancer Genetics Group of the Valld’Hebron Institute of Oncolgy (VHIO) and the Clinical and Molecular Genetics Department of the University Hospital Vall d’Hebron (HVH), the Cellex Foundation for providing research facilities, and CERCA Programme/Generalitat de Catalunya for institutional support; members and participants of the Inherited Cancer Registry (ICARE); the ICO Hereditary Cancer Program team led by Dr. Gabriel Capella; the ICO Hereditary Cancer Program team led by Dr. Gabriel Capella; Dr Martine Dumont for sample management and skillful assistance; Catarina Santos and Pedro Pinto; members of the Center of Molecular Diagnosis, Oncogenetics Department and Molecular Oncology Research Center of Barretos Cancer Hospital; Heather Thorne, Eveline Niedermayr, all the kConFab research nurses and staff, the heads and staff of the Family Cancer Clinics, and the Clinical Follow Up Study (which has received funding from the NHMRC, the National Breast Cancer Foundation, Cancer Australia, and the National Institute of Health (USA)) for their contributions to this resource, and the many families who contribute to kConFab; the KOBRA Study Group; all participants and the collaborators from RCGEB “Georgi D. Efremov”, MASA (Ivana Maleva Kostovska, Simona Jakovcevska, Sanja Kiprijanovska), University Clinic of Radiotherapy and Oncology (Snezhana Smichkoska, Emilija Lazarova, Marina Iljovska), Adzibadem-Sistina Hospital (Katerina Kubelka-Sabit, Dzengis Jasar, Mitko Karadjozov), and Re-Medika Hospital (Andrej Arsovski and Liljana Stojanovska) for their contributions and commitment to the MACBRCA study; Csilla Szabo (National Human Genome Research Institute, National Institutes of Health, Bethesda, MD, USA); Eva Machackova (Department of Cancer Epidemiology and Genetics, Masaryk Memorial Cancer Institute and MF MU, Brno, Czech Republic); Petra Kleiblova, Marketa Janatova, Jana Soukupova (Institute of Medical Biochemistry and Laboratory Diagnostics, 1st Faculty of Medicine, Charles University and General University Hospital in Prague (VFN), Czechia), Petra Zemankova, Petr Nehasil (Institute of Pathological Physiology, 1st Faculty of Medicine, Charles University, Czechia), Michal Vocka (Department of Oncology, General University Hospital in Prague (VFN), Czechia), Anne Lincoln, Lauren Jacobs; the participants in Hereditary Breast/Ovarian Cancer Study and Breast Imaging Study for their selfless contributions to our research; the NICCC National Familial Cancer Consultation Service team led by Sara Dishon, the lab team led by Dr. Flavio Lejbkowicz, and the research field operations team led by Dr. Mila Pinchev; the staff of Genetic Health Service NZ and the families who have contributed; members and participants in the Ontario Cancer Genetics Network; Hayley Cassingham. Leigha Senter, Kevin Sweet, Julia Cooper, and Amber Aielts; research nurses and staff of Breast Unit, Pauls Stradins Clinical University Hopsital, RSUIO and the many families who contribute to the CIMBA registry of RSUIO; Yip Cheng Har, Nur Aishah Mohd Taib, Phuah Sze Yee, Norhashimah Hassan and all the research nurses, research assistants and doctors involved in the MyBrCa Study for assistance in patient recruitment, data collection and sample preparation, Philip Iau, Sng Jen-Hwei and Sharifah Nor Akmal for contributing samples from the Singapore Breast Cancer Study and the HUKM-HKL Study respectively; the National Cancer Centre Singapore Cancer Genetics Service (NCCS) for patient recruitement; the Meirav Comprehensive breast cancer center team at the Sheba Medical Center; Christina Selkirk; Åke Borg, Håkan Olsson, Helena Jernström, Karin Henriksson, Katja Harbst, Maria Soller, Ulf Kristoffersson; from Gothenburg Sahlgrenska University Hospital: Anna Öfverholm, Margareta Nordling, Per Karlsson, Zakaria Einbeigi; from Stockholm and Karolinska University Hospital: Anna von Wachenfeldt, Annelie Liljegren, Annika Lindblom, Brita Arver, Gisela Barbany Bustinza, Johanna Rantala; from Umeå University Hospital: Beatrice Melin, Christina Edwinsdotter Ardnor, Monica Emanuelsson; from Uppsala University: Hans Ehrencrona, Maritta Hellström Pigg, Richard Rosenquist; from Linköping University Hospital: Marie Stenmark-Askmalm, Sigrun Liedgren; Cecilia Zvocec, Qun Niu; Joyce Seldon and Lorna Kwan; Dr. Robert Nussbaum, Beth Crawford, Kate Loranger, Julie Mak, Nicola Stewart, Robin Lee, Amie Blanco and Peggy Conrad and Salina Chan; Patricia Harrington; Geoffrey Lindeman, Marion Harris, Joanne McKinley, Simone McInerny, and Ella Thompson for performing all DNA amplification. This research has been conducted using the UK Biobank Resource under Application Number 28126. We want to acknowledge the participants and investigators of the FinnGen study. The BioBank Japan Project was supported by the Tailor-Made Medical Treatment program of the Ministry of Education, Culture, Sports, Science, and Technology (MEXT), the Japan Agency for Medical Research and Development (AMED). The Genotype-Tissue Expression (GTEx) Project was supported by the Common Fund of the Office of the Director of the National Institutes of Health, and by NCI, NHGRI, NHLBI, NIDA, NIMH, and NINDS. The data used for the analyses described in this manuscript were obtained from: https://gtexportal.org/home/gene/ the GTEx Portal on 03/31/2023.
Author information
Authors and Affiliations
Contributions
Writing group: D.R.B., J.P.T., A.C.A., P.D.P.P. Statistical analyses of OCAC and UK Biobank data: J.P.T., D.R.B., P.D.P.P. Statistical analyses of CIMBA data: D.R.B., J.P.T., A.C.A. Meta-analyses: D.R.B., J.P.T., A.C.A., P.D.P.P. OCAC database management: A.B. CIMBA database management: G.L., M.K.B. Supervision of OCAC statistical analyses: P.D.P.P. Supervision of CIMBA statistical analyses: A.C.A. Provision of DNA samples and/or phenotypic data: A.M.A., K.A., N.A., I.L.A., H.A.-C., A.A., B.K.A., J.B., E.V.B., R.B.B., L.P.V.B., A.B.d.G., P.B., K.B., L.B., A.M.B., M.J.B., K.A.B., N.V.B., J.D.B., H.B., S.S.B., M.A.C., I.C., C.C., K.B.M.C., S.V.C., L.S.C., M.B.D., A.D.-M., M.d.l.H., A.d., A.D., Y.D., J.A.D., S.M.D., T.D., Z.E., C.E., D.G.E, L.F., R.T.F., F.F., M.F., E.F., M.N.F., P.A.G., A.G.-M., G.G., A.K.G., A.G.-N., M.H.G., J.G., A.G.-G., U.H., T.v.O.H., H.R.H., J.H., F.H., F.B.L.H., M.J.H., J.L.H., C.D.H., D.G.H., E.N.I., L.I., A.J., P.A.J., R.J., E.M.J., S.K., B.Y.K., C.J.K., L.A.L.M.K., I.K., J.K., Y.L., O.L., K.L., J.L., F.L., C.Lo., P.L.M., S.M., T.M., I.A.M., U.M., R.L.M., F.M., J.M.M., M.M., K.B.M., S.L.N., F.C.N., C.N., E.O., O.I.O., A.O., L.P., H.P., C.L.Pe., I.S.P., A.P., T.P., P-C.P., B.N.P., P.P., C.B.P, D.P., M.P., P.R., M.U.R., G.Re., G.Ri., D.P.S., N.S., V.W.S., P.Sh., W.S., C.F.S., K.S., A.P.S., P.So., M.C.S., D.S.-L., R.S., C.S., Y.Y.T., M.R.T., K.L.T., L.V.T., M.Ti., A.E.T., T.V.G., A.V., D.R.V.E., P.M.W., J.N.W., N.W., A.S.W., S.J.W., A.H.W., S.Y., Y.Y., A.Z., A.B., F.J.C., E.L.G., M.T.G., A.N.M., K.O., S.J.R., H.A.R., J.M.S., M.Th., J.S., D.F.E., M.R.J., G.C.-T., S.A.G., A.C.A., P.D.P.P. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
L.B. has received honoraria for lectures from GlaxoSmithKline, Merck Sharp and Dohme, has received a research grant from AstraZeneca for a researcher-initiated trial, and has had leadership roles in Onkologisk Forum between 2018 and 2022 and in the Nordic Society of Gynaecological Oncology (NSGO) and NSGO-Clinical Trial Unit since 2021. She is faculty member the Gynaecological Cancers group, European Society of Medical Oncology (ESMO) external expert in gynecologic oncology for the Norwegian National Expert Panel for Persons with Serious Life-Shortening Diseases and Member of the national specialist group in oncology, Sykehusinnkjøp HF as well as board member for KinN Therapeutics Bergen - a biopharmaceutical company focused on tailored preclinical oncology services for the development of novel anticancer compounds. A.d. has received research support from AstraZeneca and Illumina, and speakers fee from AstraZeneca. A.G.-M. has received research funding for my role, has been funded by grants from the Medical Research Council (MRC), Cancer Research UK (CRUK), National Institute for Health Research (NIHR) and The Eve Appeal, has funded research collaborations with industry - iLOF (intelligent Lab on Fiber), RNA Guardian, Micronoma and Mercy BioAnalytics, has research collaborations with academic institutions: University of Cambridge, QIMR Berghofer Medical Research Institute, Imperial College London, University of Innsbruck and Dana Farber USA, is a Co-Director Research Domain Trials, and has consultancy arrangement with Mercy BioAnalytics. F.H. has received personal fees and is on advisory boards of NovoCure, PharmaMar, AstraZeneca, Roche, Tesaro, GSK, Zailabs, Abbvie, MSD. A.E.T. has received grant funding through the National Institutes of Health and has study section membership for the National Institutes of Health. P.M.W. has received grant funding from AstraZeneca (06/2017-03/2021) for an unrelated study of ovarian cancer. D.F.E. and A.C.A. are listed as creators of the BOADICEA model which has been licensed by Cambridge Enterprise (University of Cambridge) for commercial purposes.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Barnes, D.R., Tyrer, J.P., Dennis, J. et al. Genome-wide association study of 398,238 women unveils seven loci associated with high-grade serous ovarian cancer. npj Genom. Med. 10, 73 (2025). https://doi.org/10.1038/s41525-025-00529-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41525-025-00529-w