Abstract
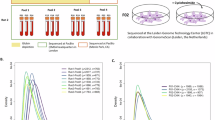
Paralogous genes challenge short-read sequencing (SRS) due to high sequence similarity. Although long-read sequencing (LRS) improves resolution, the extent to which it resolves paralogous genes remains unclear. This study evaluates the capability of LRS by integrating in silico mappability-based predictions with clinical data to generate SRS- and LRS-unresolved gene lists, and by assessing whether a paralog-specific phasing, Paraphase, can overcome remaining limitations. Mappability was simulated across read lengths (250 bp to 14 kb) to predict unresolved regions and validated against mapping quality (MQ) from 66 high-fidelity LRS samples. Paraphase was applied to 79 paralog groups. Among 645 medically relevant (MR) genes unresolved by SRS, 419 (65.0%) were predicted to be resolved by LRS, while 226 (35.0%) remained unresolved. These predictions correlated with clinical MQ (χ² = 92.43, p < 2.2 × 10−16; κ = 0.37), with significant differences between LRS-resolved and LRS-unresolved MR genes (W = 63,656, p < 2.2 × 10−16; r = 0.36). Paraphase resolved 61 groups (77.2%), providing additional resolution beyond LRS. LRS improves paralogous gene resolution but cannot fully eliminate paralog blind spots. Curated gene lists define boundaries of LRS utility for clinical interpretation, while Paraphase adds complementary resolution, supporting an integrated framework combining predictive modeling with algorithmic strategies.
Similar content being viewed by others
Data availability
The GRCh38 reference genome FASTA file used for mappability analysis is available from the UCSC Genome Browser (https://hgdownload.soe.ucsc.edu/goldenPath/hg38/bigZips/). The GRCh38 version with excluded alternate contigs was downloaded from the Pacific Biosciences GitHub repository (https://github.com/PacificBiosciences/reference_genomes). The GEM library tool was downloaded from SourceForge (https://sourceforge.net/projects/gemlibrary/files/). The code for Paraphase is available on GitHub (https://github.com/PacificBiosciences/paraphase). Owing to ethical constraints and the sensitive nature of clinical genomic data, full BAM files from patients cannot be made publicly available. The data are available following review and approval by the corresponding author’s institution and under appropriate data use agreements upon request.
References
Koonin, E. V. Orthologs, paralogs, and evolutionary genomics. Annu. Rev. Genet. 39, 309–338 (2005).
Kuzmin, E., Taylor, J. S. & Boone, C. Retention of duplicated genes in evolution. Trends Genet. 38, 59–72 (2022).
Drobek, M. Paralogous genes involved in embryonic development: lessons from the eye and other tissues. Genes 13, 2082 (2022).
Ebbert, M. T. W. et al. Systematic analysis of dark and camouflaged genes reveals disease-relevant genes hiding in plain sight. Genome Biol. 20, 97 (2019).
Olivucci, G. et al. Long read sequencing on its way to the routine diagnostics of genetic diseases. Front. Genet. 15, 1374860 (2024).
Derrien, T. et al. Fast computation and applications of genome mappability. PLoS ONE 7, e30377 (2012).
Mandelker, D. et al. Navigating highly homologous genes in a molecular diagnostic setting: a resource for clinical next-generation sequencing. Genet. Med. 18, 1282–1289 (2016).
Eid, J. et al. Real-time DNA sequencing from single polymerase molecules. Science 323, 133–138 (2009).
Mikheyev, A. S. & Tin, M. M. A first look at the Oxford Nanopore MinION sequencer. Mol. Ecol. Resour. 14, 1097–1102 (2014).
Wenger, A. M. et al. Accurate circular consensus long-read sequencing improves variant detection and assembly of a human genome. Nat. Biotechnol. 37, 1155–1162 (2019).
Logsdon, G. A., Vollger, M. R. & Eichler, E. E. Long-read human genome sequencing and its applications. Nat. Rev. Genet. 21, 597–614 (2020).
Stephens, Z. D. & Iyer, R. K. Measuring the mappability spectrum of reference genome assemblies. In Proc. 2018 ACM International Conference on Bioinformatics, Computational Biology, and Health Informatics 47–52 (ACM, 2018).
Sanford Kobayashi, E. et al. Approaches to long-read sequencing in a clinical setting to improve diagnostic rate. Sci. Rep. 12, 16945 (2022).
Chen, X. et al. Genome-wide profiling of highly similar paralogous genes using HiFi sequencing. Nat. Commun. 16, 2340 (2025).
Chen, X. et al. Comprehensive SMN1 and SMN2 profiling for spinal muscular atrophy analysis using long-read PacBio HiFi sequencing. Am. J. Hum. Genet 110, 240–250 (2023).
Hops, W. et al. HiFi long-read genomes for difficult-to-detect, clinically relevant variants. Am. J. Hum. Genet. 112, 450–456 (2025).
Wagner, J. et al. Benchmarking challenging small variants with linked and long reads. Cell Genom. 2, 100128 (2022).
Prodanov, T. & Bansal, V. Sensitive alignment using paralogous sequence variants improves long-read mapping and variant calling in segmental duplications. Nucleic Acids Res. 48, e114 (2020).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Sahlin, K., Baudeau, T., Cazaux, B. & Marchet, C. A survey of mapping algorithms in the long-reads era. Genome Biol. 24, 133 (2023).
Li, W. & Freudenberg, J. Mappability and read length. Front. Genet. 5, 381 (2014).
Neph, S. et al. BEDOPS: high-performance genomic feature operations. Bioinformatics 28, 1919–1920 (2012).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Acknowledgements
This work was supported by the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea (grant number: RS-2025-02263889).
Author information
Authors and Affiliations
Contributions
Conceptualization: S.K., J.L, and M.S.; Data curation: H.S., H.L., H.I., and S.C.; Formal analysis: S.K. and J.J.; Methodology: S.K., J.J., Y.K., J.L., and M.S.; Supervision: J.L. and M.S.; Visualization: S.K.; Writing—original draft: S.K.; Writing—review and editing: all authors. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kim, S.K., Jang, J., Kim, Y. et al. Integrative analysis of in silico predictions and clinical evidence to delineate the capability of HiFi long-read sequencing in paralogous genes. npj Genom. Med. (2026). https://doi.org/10.1038/s41525-026-00555-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41525-026-00555-2