Abstract
Fatigue is a common and debilitating non-motor symptom in patients with Parkinson’s disease (PD). By integrating the activation likelihood estimation (ALE) meta-analysis and activation network mapping (ANM) technique, the brain network underlying fatigue (referred to as “fatigue-related network”) in PD was identified, comprising widespread brain regions primarily involving the somatomotor and frontoparietal networks. This network was interpreted through transcriptomic patterns, chemoarchitectures, and behavioral relevance. Fatigue-related genes were predominantly enriched in synaptic and actin filament-related biological processes and showed primary expression in neurons and oligodendrocytes. The fatigue-related network closely corresponded to the spatial distribution of acetylcholine, glutamate, and norepinephrine systems and was primarily linked to motor function. This study defines a distinct brain network substrate underlying fatigue in PD, advancing our understanding of the neurobiological mechanisms and facilitating the development of effective therapies for affected patients.
Similar content being viewed by others
Introduction
Parkinson’s disease (PD) is a prevalent neurological disorder, with its incidence doubled over the past three decades1. The pathophysiology of PD is multifaceted, involving aberrant α-synuclein aggregation, mitochondrial dysfunction, lysosomal impairment, vesicle transport disturbances, synaptic transport defects, and neuroinflammation2. Beyond the well-recognized motor symptoms, PD also presents a broad spectrum of non-motor symptoms (NMS) that often go unnoticed during the disease course. Fatigue, one of the most common and debilitating NMS in PD, affects up to 50% of patients and can arise at any stage of the disease, sometimes persisting throughout its course3,4. The 2019 recommendations for managing NMS in PD deem “rasagiline” “possibly useful” for fatigue5, but there remains a lack of updated evidence on the effective treatment for this symptom in PD6. Despite its high prevalence, the pathophysiological mechanisms underlying fatigue in PD remain poorly understood.
Fatigue is a common feature in various neurological conditions, classified as either peripheral fatigue (e.g., muscle fatigability in neuromuscular junction disorders) or central fatigue (e.g., fatigue in PD or multiple sclerosis (MS), affecting the central, peripheral, or autonomic nervous systems)7,8. Central fatigue is hypothesized to arise from dysfunction within the striato-thalamo-cortical loop, which connects the neostriatum to the frontal lobe (including the prefrontal, cingulate, and orbitofrontal cortices)7. Fatigue can also manifest under normal physiological conditions when the cost of an action outweighs its expected benefits, heavily influenced by motivation9. It is suggested that deficits in stimulus evaluation and reward-related decision-making processes, tied to the mesolimbic system’s functional integrity, ultimately lead to central fatigue in PD9,10. Although these circuits are believed to contribute to central fatigue, their neural substrates in PD remain undefined.
Over the past decade, neuroimaging techniques have become integral in investigating the neural substrates of neuropsychiatric disorders. Several studies have reported structural and functional brain abnormalities linked to fatigue in PD11,12,13,14,15,16,17,18,19,20,21. Notably, resting-state functional magnetic resonance imaging (rs-fMRI), a widely used method to indirectly access neuronal activity via blood-oxygen-level-dependent (BOLD) signals, has shown that distressing fatigue in PD is associated with altered regional brain activity and functional connectivity (FC) in various brain areas, including the postcentral gyrus, supplementary motor area (SMA), supramarginal gyrus (SMG), frontal regions, insula, and cingulate cortex11,14,16,17,18,19,20,21. Tessitore et al. observed that fatigue severity correlated with connectivity changes within both the sensorimotor and default mode networks in PD18. However, findings regarding these fatigue-related regions often suffer from poor reproducibility. To address the “reproducibility crisis” in neuroimaging research, activation likelihood estimation (ALE), a widely adopted coordinate-based meta-analysis method22, has been employed to identify consistent patterns of abnormal intrinsic functional activity across studies23,24. While ALE effectively maps convergent activations, it does not capture inter-regional connectivity. Recent approaches emphasize that the pathophysiological basis of neuropsychiatric disorders lies not in isolated brain regions but in disrupted brain networks. The connectome perspective offers a more integrative framework for understanding pathological brain perturbations25,26,27. A novel and well-validated approach, activation network mapping (ANM), enables the mapping of complex behaviors, brain diseases, or clinical manifestations to a shared network. ANM has been applied to localize the network substrates for facial emotion processing28, PD dementia (PDD)29, and suicide attempts in depression30. The diverse functional findings in PD-related fatigue studies could converge into a shared functional brain network, referred to as the “fatigue-related network”. This network represents the core neuroanatomical substrate of fatigue in PD, integrating contributions from multiple brain regions and their functional connections. PD is known to be influenced by a variety of genetic variants and neurotransmitter systems2. Previous clinical cohort and neuroimaging studies have found associations between genetic factors, neurotransmitter levels, and fatigue in PD patients31,32,33,34,35,36. Additionally, the fatigue-related network, as the neurobiological basis of fatigue in PD, may be linked to genetic factors and neurotransmitter systems.
To test these hypotheses, this study aimed to identify and decode the fatigue-related brain network in PD. First, an ALE meta-analysis was conducted on functional activation data from comparisons between PD patients with and without fatigue to integrate findings across published rs-fMRI studies. Second, the ANM method was employed to localize a shared fatigue-related network in PD using a discovery dataset and two independent validation datasets. Third, the identified fatigue-related network was interpreted through transcriptomic patterns (i.e., gene functional features, including the functional enrichment and cell type-specific expression), chemoarchitectures (i.e., neurotransmitter distribution patterns), and behavioral relevance. A schematic overview of the study design is shown in Fig. 1.
First, an ALE meta-analysis was conducted on functional activation data derived from the comparison between PD patients with and without fatigue to integrate findings across published rs-fMRI studies. Second, the significant convergence across these enrolled rs-fMRI findings yielded 12 significant clusters predominantly in the sensorimotor, frontal, and temporal regions. Third, the discovery dataset (referred to as “D1_Old Subjects”) and the two validation datasets (referred to as “D2_1000 Subjects” and “D3_Young Subjects”) were adopted for further ANM procedure. Fourth, the brain network underlying fatigue in PD was localized by the ANM method. Fifth, the identified fatigue-related network in PD was interpreted through transcriptomic patterns (i.e., gene functional features, including the functional enrichment and cell type-specific expression), chemoarchitectures (i.e., neurotransmitter distribution patterns), and behavioral relevance. ALE activation likelihood estimation, ANM activation network mapping, FC functional connectivity, PD Parkinson’s disease, rs-fMRI resting-state functional magnetic resonance imaging.
Results
Convergence of rs-fMRI findings in fatigue of PD
Five studies reporting abnormal activations in PD patients with fatigue (total n = 88) compared to those without fatigue (total n = 120) were identified (Table S1). Convergence analysis of these rs-fMRI findings revealed 12 significant clusters, primarily located in sensorimotor, frontal, and temporal regions (Table S2). All clusters exhibited 80% reproducibility in the jackknife sensitivity analysis, with each cluster remaining significant in 80% of leave-one-study-out iterations.
Fatigue-related network of PD
The fatigue-related network in PD encompassed a distributed set of brain regions, predominantly the SMA, precentral and postcentral gyri, SMG, inferior frontal gyrus, and superior temporal gyrus. This network primarily overlapped with the somatomotor (37.95%) and frontoparietal (26.42%) networks. Validation analyses using two additional datasets (D2_1000 Subjects and D3_Young Subjects) showed that the derived networks closely resembled those from the discovery dataset (D1_Old Subjects). Specifically, the network from the D2_1000 Subjects dataset was predominantly linked to the somatomotor (28.12%) and frontoparietal (19.89%) networks, while the network from the D3_Young Subjects dataset was mainly associated with the somatomotor (29.57%) and frontoparietal (20.81%) networks (Figs. 2 and 3; and Table S3).
ALE activation likelihood estimation, ANM activation network mapping, D1_Old Subjects the discovery dataset from the First People’s Hospital of Yunnan Province, D2_1000 Subjects the first validation dataset from Neurosynth (https://www.neurosynth.org/locations/); D3_Young Subjects, the second validation dataset from Kunming University of Science and Technology, L left, R right, PD Parkinson’s disease.
Polar plots show the proportion of overlapping voxels between the fatigue-related network and a canonical network to all voxels within the fatigue-related network in PD. A The network from the D1_Old Subjects dataset primarily overlapped with the somatomotor (37.95%) and frontoparietal (26.42%) networks; B The network from the D2_1000 Subjects dataset was predominantly linked to the somatomotor (28.12%) and frontoparietal (19.89%) networks; C The network from the D3_Young Subjects dataset was mainly associated with the somatomotor (29.57%) and frontoparietal (20.81%) networks. D1_Old Subjects the discovery dataset from the First People’s Hospital of Yunnan Province, D2_1000 Subjects the first validation dataset from Neurosynth (https://www.neurosynth.org/locations/), D3_Young Subjects the second validation dataset from Kunming University of Science and Technology, PD Parkinson’s disease.
Fatigue-related transcriptomic profiles of PD
A total of 406 overlapping genes were identified between the D1_Old Subjects and D2_1000 Subjects datasets and combined into a single gene set (referred to as “D1&2”) for subsequent analysis (Fig. 4A). These gene sets were primarily enriched in actin filament-based processes and synapse-related biological processes, such as modulation of chemical synaptic transmission, regulation of synapse structure or activity, trans-synaptic signaling, and synapse organization (Fig. 4B, Fig. S1, and Table S4). Further analysis of these genes as cell-type markers revealed consistent distribution across gene sets. Specifically, genes with positive weights in the fatigue-related network were predominantly enriched in oligodendrocyte markers, while negatively weighted genes were primarily associated with markers of excitatory and inhibitory neurons (Fig. 4C and Table S5).
A Genes associated with the fatigue-related network were identified from D1_Old Subjects and D2_1000 Subjects gene sets respectively, and a total of 406 overlapping genes were identified between them (referred to as “D1&2”); B The D1&2 gene set showed significantly enriched biological processes (PFDR < 0.05); C These genes as cell-type markers revealed consistent distribution across gene sets, predominantly enriched in excitatory neurons, inhibitory neurons, and oligodendrocytes (***PFDR < 0.001). Astro astrocytes, D1_Old Subjects the discovery dataset from the First People’s Hospital of Yunnan Province, D2_1000 Subjects the first validation dataset from Neurosynth (https://www.neurosynth.org/locations/), D3_Young Subjects the second validation dataset from Kunming University of Science and Technology, Endo endothelial cells, FDR false discovery rate, Micro microglia, N-Ex excitatory neurons, N-In inhibitory neurons, Oligo oligodendrocytes, OPC oligodendrocyte precursor cells, PD Parkinson’s disease.
Fatigue-related chemoarchitectures of PD
The relationship between neurotransmitter receptor distribution and the fatigue-related network was investigated using multiple linear regression (Fig. 5, Figs. S2 and Fig. S3, and Table S6). The model accounted for 36% of the variance (F = 64.83, adjusted R2 = 0.36). The acetylcholine, glutamate, and norepinephrine systems significantly contributed to the fatigue-related network in PD, with acetylcholine accounting for 29.15%, glutamate for 22.88%, and norepinephrine for 18.30% in the discovery dataset (D1_Old Subjects). Similar contributions were observed in validation datasets: D2_1000 Subjects (acetylcholine = 33.51%, glutamate = 18.90%, norepinephrine = 18.46%) and D3_Young Subjects (acetylcholine = 23.43%, glutamate = 22.52%, norepinephrine = 17.96%). Robustness was assessed through multiple validations: (1) significant Pspin values were obtained across datasets (D1_Old Subjects = 0.00019, D2_1000 Subjects = 0.00179, D3_Young Subjects = 0.01779), mitigating spatial autocorrelation biases and verifying result reliability; (2) correlation coefficients between glutamate and dopamine (r = 0.735), and between glutamate and serotonin (r = 0.787), were relatively high (exceeding the common threshold of 0.7), while all other neurotransmitter pairs exhibited correlation coefficients below 0.7 (Table S7). Additionally, the variance inflation factors (VIF) values for glutamate (VIF = 6.89) and dopamine (VIF = 6.01) exceeded the conventional caution threshold of 5, suggesting moderate multicollinearity, whereas all other neurotransmitters had VIF values below 5 (Table S8); (3) similar contributions from acetylcholine, glutamate, and norepinephrine were observed when the analysis was repeated using a 100-parcel atlas (Fig. S4); and (4) false discovery rate (FDR) correction at P < 0.001 yielded near-zero P-values for the three neurotransmitters (Table S6).
A The multivariate linear regression analysis was performed to investigate the relationship between the fatigue-related network and neurotransmitter systems; B The relative contributions of each neurotransmitter system to the network were assessed (*P < 0.001); C The scatterplot was generated to display the similarity between the neurotransmitter predicted density map and the network. Blue dots represent individual brain regions; the red line indicates linear regression fit; light blue shading shows 95% confidence interval. The overall correspondence is quantified by an adjusted R² = 0.36 (Pspin = 0.00019, spin-based permutation test). The gray histogram (lower right) shows residual distribution, with the red dashed line indicating zero residual error. D1_Old Subjects the discovery dataset from the First People’s Hospital of Yunnan Province, GABA γ-aminobutyric acid, PD Parkinson’s disease.
Behavioral relevance related to fatigue in PD
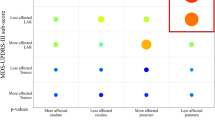
The fatigue-related network in PD showed significant correlations with multiple behavioral terms across datasets (Fig. 6 and Fig. S5). The most prominent functional association was with “motor” (D1_Old Subjects: z = 20.85; D2_1000 Subjects: z = 17.01; D3_Young Subjects: z = 17.53).
The terms shown here were significantly associated with the network. D1_Old Subjects the discovery dataset from the First People’s Hospital of Yunnan Province, PD Parkinson’s disease.
Discussion
By integrating the ALE meta-analysis and ANM technique, the brain network underlying fatigue in PD was localized based on resting-state functional abnormalities identified in PD patients with fatigue from previous neuroimaging studies. Our findings delineated a fatigue-related network composed of extensive brain regions, primarily involving the somatomotor and frontoparietal networks. A series of decoding analyses revealed that (1) genes significantly associated with the fatigue-related network were predominantly enriched in synaptic and actin filament-related biological processes; (2) these genes showed primary expression in excitatory neurons, inhibitory neurons, and oligodendrocytes; (3) the fatigue-related network closely corresponded to the spatial distribution of acetylcholine, glutamate, and norepinephrine systems; and (4) the behavioral relevance of the fatigue-related network was primarily linked to motor function. These results provide empirical evidence for a distinct brain network substrate underlying fatigue in PD, advancing our integrated understanding of the neurobiological mechanisms contributing to this debilitating symptom.
Despite the widespread use of neuroimaging techniques in human neuroscience to localize diseases, symptoms, or brain functions, concerns about the reproducibility crisis have increasingly surfaced. The lack of reproducibility can be attributed to factors such as small sample sizes, suboptimal study designs, analytic procedures, and limited statistical power. Furthermore, studies with poor reproducibility may identify different subregions within a shared connected brain network, suggesting that heterogeneous findings could be localized within connected brain networks to enhance the precision of disease or symptom localization25,37,38. To address these concerns, an ALE meta-analysis was performed to provide a comprehensive view of distributed neuroimaging results and establish consensus on the spatial locations of regional alterations in PD patients with fatigue across studies. To further elucidate the functional architecture of fatigue in PD, ANM was employed to combine the identified regions of interest with rs-fMRI connectome data from a healthy cohort, constructing the brain network of fatigue in PD. Given that PD is an age-related neurodegenerative disease2, elderly individuals were recruited as the discovery cohort (D1_Old Subjects). Additional validation analyses were conducted using a large healthy cohort of 1,000 subjects (D2_1000 Subjects) and a separate cohort of young individuals (D3_Young Subjects). These analyses collectively revealed a complex and interconnected brain network associated with fatigue in PD.
Within the pathophysiological framework of fatigue, the symptom is described as a difficulty in initiating or sustaining voluntary activities. The extent of voluntary activities depends on the applied efforts, which are influenced by both internal and external motivational inputs and feedback from motor, sensory, and cognitive systems. Chaudhuri and Behan proposed that pathological fatigue, an amplified sense of physiological fatigue, may arise from alterations in one or more variables regulating the applied efforts8. Kuppuswamy further suggested that the core mechanism of fatigue involves abnormal interactions between motor and sensory systems39. It is well-established that brain regions exhibiting abnormal activation or connectivity, primarily within the somatomotor network, such as the SMA, precentral and postcentral gyri, and paracentral lobule, are implicated in the fatigue process in PD14,16,17,18,20. These observations are consistent with our finding that the fatigue-related network in PD predominantly involves the somatomotor network. These regions may represent a core neural substrate underlying the pathogenesis of fatigue in PD. Furthermore, our results suggest that the fatigue-related network also includes the frontoparietal network, a distinct cognitive control network involved in flexibly modulating and interacting with other functional brain networks40. Previous resting-state functional neuroimaging studies in PD have consistently shown abnormal activation, connectivity, or functional coordination within the frontoparietal network, including the superior frontal gyrus (dorsolateral part), SMG, midcingulate cortex, and insula, correlating with fatigue symptom severity14,19,20. Additionally, Liu et al. combined arterial spin labeling (ASL) perfusion fMRI with a simple reaction time task to explore the neural correlates of fatigue, further supporting the pivotal role of the frontoparietal network in mediating fatigue in PD41.
Transcriptomic analyses revealed an association between the fatigue-related network in PD and gene expression enriched in biological processes that regulate synaptic structure and transmission, particularly those dependent on or modulating the actin cytoskeleton. The actin cytoskeleton, a dynamic network of actin filaments, plays a pivotal role in maintaining the structural integrity of pre- and post-synaptic elements and in regulating neurotransmission, including vesicle release and endocytosis in the pre-synaptic compartment, as well as neurotransmitter receptor localization and trafficking in the post-synaptic compartment42,43. Growing evidence suggests that both synaptic dysfunction and disruptions in the actin networks are key contributors to neurodevelopmental and neurodegenerative disorders, including PD44,45,46,47. In PD, previous studies have proposed that synaptic dysfunction in specific brain regions (e.g., hippocampus) or neurotransmitter systems (e.g., acetylcholine, norepinephrine, serotonin, and systems) may underlie the complex PD phenotype, encompassing cognitive impairment, depression, fatigue, and psychotic symptoms35,36,48,49,50. Moreover, alpha-synuclein (α-Syn) has been shown to inhibit cofilin, an actin-destabilizing factor, leading to abnormal actin overstabilization, which may impair synaptic signaling46. Genes involved in actin filament-related processes may therefore influence neurotransmission by modulating synaptic actin dynamics. Cellular gene set analyses revealed that fatigue-related genes in PD were significantly expressed in neurons and oligodendrocytes. Neurodegenerative diseases are characterized by the progressive loss of selectively vulnerable neuronal populations. However, increasing evidence suggests that glial cells also play a critical role in disease onset and progression through various physiological and pathological mechanisms51. A recent study integrating genome-wide association study (GWAS) data with single-cell transcriptomic profiles from the mouse nervous system showed that PD was genetically associated not only with cholinergic and monoaminergic neurons (e.g., dopaminergic neurons) but also with oligodendrocytes52. Oligodendrocytes are essential for myelin formation and repair, and the degree of myelination has been proposed as a key determinant of neuronal vulnerability to Lewy pathology in PD51,53. Alterations in white matter tracts have also been linked to fatigue severity in PD, suggesting that white matter abnormalities may serve as potential biomarkers for fatigue12.
Our analysis of neurotransmitter systems revealed significant correlations between the acetylcholine, glutamate, and norepinephrine systems and the fatigue-related network in PD. The central cholinergic system plays a key role in regulating a wide range of human behaviors by enhancing adaptive responses to environmental stimuli while suppressing responses to non-urgent inputs54. In vivo imaging studies have identified a “cholinergic phenotype” in PD, in which cholinergic dysfunction contributes to specific clinical manifestations, such as cognitive decline and neuropsychiatric symptoms55. While direct investigations into the relationship between cholinergic dysfunction and fatigue in PD are limited, cholinergic impairment has been implicated in chronic fatigue syndrome (CFS)56,57,58, suggesting a potential pathophysiological link between cholinergic dysfunction and fatigue in PD. Glutamate, the major excitatory neurotransmitter in the central nervous system, is associated with neurodegeneration and excitotoxicity when overactive. Evidence indicates that glutamate overactivity and excitotoxicity significantly contribute to the progression of neuroinflammation in PD59,60, potentially disrupting effort perception and homeostatic regulation—both processes closely linked to fatigue. Norepinephrine, involved in arousal regulation and neuroinflammatory modulation61,62, is another promising candidate for investigation in PD-related fatigue. Recent work by Di Vico et al. provided direct support for a non-dopaminergic (glutamatergic and noradrenergic) contribution to fatigue in PD, highlighting that decreased glutamate-enriched FC was particularly localized in the SMA, which is also a key region in our fatigue-related network63. Similarly, previous studies have suggested that the dysregulation of glutamatergic/noradrenergic neurotransmission is involved in mediating central fatigue in MS64,65. The robustness of our key findings was supported by multiple validations, including significant Pspin values, different atlas parcellations, and FDR correction. Notably, glutamate exhibited relatively high correlations with dopamine and serotonin, and both glutamate and dopamine showed moderate multicollinearity. While the multiple linear regression model indicated a substantial contribution of the glutamate system to the fatigue-related network, this finding should be interpreted with caution, as its contribution may be partially confounded by the effects of dopamine and serotonin. An electrophysiological study suggested that an altered dopamine/serotonin balance is involved in the genesis of central fatigue in PD66. Previous positron emission tomography (PET) studies have demonstrated the relevance of monoamine neurotransmitters (dopamine and serotonin) in PD-related fatigue, with nigrostriatal and insular dopaminergic degeneration and serotonergic dysfunction in the basal ganglia (BG) and limbic circuits associated with fatigue35,36. These monoamine-associated brain regions are predominantly subcortical. While earlier studies focused on individual neurotransmitter systems, our findings emphasize the integrated roles of multiple neurotransmitters within the fatigue-related network. Beyond these micro-level associations, the fatigue-related network was also significantly related to motor function. Previous meta-analyses have shown that motor dysfunction is more severe in PD patients with fatigue3.
The strengths of this study include the application of a novel network mapping approach (i.e., ANM), validation across independent datasets (including a large sample (D2_1000 Subjects) and a younger cohort (D3_Young Subjects)), and comprehensive multi-dimensional decoding analyses. However, several limitations should be considered. First, the small number of studies and their cross-sectional design limit the ability to draw definitive conclusions regarding the precise localization of the fatigue-related network in PD. Our study focused on a subset of neural correlates of fatigue detectable via rs-fMRI meta-analysis, rather than providing a comprehensive mapping of all known neural correlates. Second, only one meta-analysis method (i.e., ALE) was applied, and alternative methods may yield different results, as seen in previous studies on PDD29. Third, the presence of moderate multicollinearity in neurotransmitter systems (e.g., glutamate) may affect the interpretation of their individual contributions to the fatigue-related network. Finally, the associations between the fatigue-related network and specific biological mechanisms (e.g., synaptic processes, actin filament dynamics, and oligodendrocyte involvement) were presented in a deterministic manner, based on indirect spatial correlations with transcriptomic datasets, rather than patient-derived molecular data. While biologically plausible, these associations should be considered hypotheses rather than proven mechanisms. Similarly, the links between this network and neurotransmitter systems derived from atlas-based PET receptor density maps in healthy populations (not PD-specific imaging or biochemical assays) suggest potential vulnerabilities, rather than confirmed neurochemical alterations.
In conclusion, this study localized the heterogeneous neural activations related to fatigue in PD into the fatigue-related network by integrating the ALE meta-analysis and ANM approach. This network was closely associated with synaptic and actin filament-related biological processes, neurons and oligodendrocytes, acetylcholine, glutamate, and norepinephrine neurotransmitter systems, and motor function. These findings may represent the neurobiological basis of fatigue in PD and provide valuable insights into potential therapeutic targets for affected patients.
Methods
Search strategy and study selection
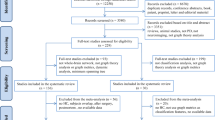
The meta-analysis was conducted following the guidelines of the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA). Neuroimaging studies that examined functional alterations in PD patients with fatigue compared to those without fatigue were identified through a search of two databases (PubMed and Web of Science) conducted in July 2025. The search terms included: (human) AND ((Parkinson’s disease) OR (Parkinson disease)) AND ((fatigue) OR (tiredness)) AND ((MRI) OR (magnetic resonance imaging) OR (resting state fMRI) OR (rs fMRI) OR (resting state functional magnetic resonance imaging) OR (functional magnetic resonance imaging) OR (fMRI)). Manual searches were also performed within the reference lists of relevant reviews and meta-analysis articles. Inclusion and exclusion criteria were established a priori to ensure systematic and unbiased study selection. The inclusion criteria were: (i) studies that compared whole-brain activations between PD patients with and without fatigue based on rs-fMRI data; (ii) studies that provided Montreal Neurological Institute (MNI) or Talairach coordinates for significant brain regions. Exclusion criteria were: (i) studies in which experimental tasks were performed during fMRI scanning; (ii) studies that were meta-analyses, reviews, case reports, letters, editorials, or meeting abstracts; (iii) studies not published in English. Study selection was conducted independently by two investigators (FY and YH), with discrepancies resolved through consensus with a third investigator (JW). The flow diagram of the study screening and selection process is provided in Fig. S6, and a list of excluded studies is presented in Table S9. Since individual studies may report multiple contrasts (e.g., PD patients with fatigue versus healthy controls (HC) and PD patients without fatigue versus HC), our analysis focused on the contrast between PD patients with and without fatigue, rather than on individual studies. The risk of bias (RoB) was assessed using the Newcastle-Ottawa Scale (NOS). Detailed star ratings for each included study across all NOS domains are provided in Table S10. Relevant information on sample characteristics and brain coordinates of peak voxels in significant clusters was extracted from each study (Table S1). Talairach coordinates were converted to MNI coordinates.
ALE meta-analysis
The GingerALE software package (https://www.brainmap.org/ale) was used to identify the significant convergence of activation foci from different resting-state fMRI experiments (e.g., PD patients with fatigue >PD patients without fatigue, PD patients with fatigue <PD patients without fatigue) relative to a null distribution of random spatial association. Statistical significance was set at P < 0.005, and the volume threshold was set at >300 mm³. Reported peak coordinates were in MNI space. The Jackknife sensitivity analysis was performed to validate robustness.
Discovery and validation datasets
The discovery dataset (referred to as “D1_Old Subjects”) obtained from the First People’s Hospital of Yunnan Province comprised 41 normal subjects (20 males, mean age = 64.27 ± 3.76 years), and the validation dataset (referred to as “D3_Young Subjects”) obtained from Kunming University of Science and Technology consisted of 62 normal subjects (24 males, mean age = 22.02 ± 2.26 years). All participants were recruited from local communities through poster advertisements. Exclusion criteria included MRI contraindications, severe somatic or neuropsychiatric diseases, and a history of head injury. Demographic information is shown in Table S11. Ethical approval was obtained from the local ethical institutional review boards (D1_Old Subjects: KHLL2024-KY026; D3_Young Subjects: KMUST-MEC-207), and written informed consent was provided by all participants. The additional validation dataset (referred to as “D2_1000 Subjects”) from Neurosynth (https://www.neurosynth.org/locations/) comprised 1,000 healthy adults, and the fMRI data were acquired using the consistent MRI sequences and hardware (3.0-Tesla field strength, 12-channel receive coil array)67. Notably, the age ranges of participants were as follows: 60–75 years for the D1_Old Subjects dataset, 18–35 years for the D2_1000 Subjects dataset, and 18-30 years for the D3_Young Subjects dataset.
Magnetic resonance imaging acquisition and preprocessing
The rs-fMRI data of D1_Old Subjects and D3_Young Subjects datasets were obtained on a 3.0-Tesla Siemens Prisma scanner equipped with 20-channel and 64-channel head coils, respectively. All subjects were instructed to lie comfortably in a supine position with their eyes closed. A foam pad was used to reduce head movement, and earplugs were used to minimize noise interference. Scans used the following parameters: repetition time/echo time = 1500 ms/30 ms, voxel size = 3.0 × 3.0 × 3.0 mm3, acquisition matrix size = 64 × 64, field of view (FOV) = 192 × 192 mm2, 250 volumes with a slice gap of 0.75 mm. Data were processed using the Statistical Parametric Mapping software (SPM12, https://www.fil.ion.ucl.ac.uk/spm-statistical-parametric-mapping/) and the Data Processing & Analysis for Brain Imaging toolkit (DPABI, http://rfmri.org/DPABI). The rs-fMRI data were processed using the following steps: removal of the first 10 time points, slice timing corrections, spatial realignment to the first volume, spatial normalization into the standard MNI space, resampling to 3 × 3 × 3 mm3 for each voxel, spatial smoothing with 8-mm full-width at half-maximum (FWHM) Gaussian kernel, and nuisance signal regression (including the Friston 24-parameters, white matter (WM), and cerebrospinal fluid (CSF) signals). The head motion parameters for all participants were <2.5 mm maximum displacement in the x, y, or z plane and <2.5° angular rotation about each axis. A bandpass filter (0.01–0.10 Hz) was applied to extract low-frequency signals for each voxel. Scrubbing with linear interpolation was further used to remove the bad images exceeding the predefined motion threshold (frame displacement (FD) > 0.5).
ANM analysis and relation to brain networks
The ANM approach was used to construct the fatigue-related network of PD. First, three millimeter-radius spheres were created at each of the above peak coordinates to yield seed masks. Second, based on the preprocessed rs-fMRI data, the seed-to-whole brain FC map for each subject was calculated by computing Pearson’s correlation coefficients between time courses of the seed and each voxel within the whole brain, followed by Fisher’s z-transformation. Third, the subject-level z-maps were entered into a voxel-wise one-sample t-test to generate t-maps. Fourth, the group-level t-map was thresholded at P < 0.05 with FDR correction, and regions with cluster size less than 30 were removed, on which the obtained t-map was binarized (t > 0 = 1, t < 0 = 0). Finally, all binarized maps were overlapped and thresholded at 50% to produce a group-level network, that is, the fatigue-related network in PD. For the sake of interpretability, the spatial relations between our fatigue-related network and each of eight established brain networks, including seven cortical networks (i.e., default, dorsal attention, frontoparietal, limbic, somatomotor, ventral attention, and visual networks67) and one subcortical network (including the amygdala, hippocampus, BG, and thalamus68), were assessed by calculating the proportion of overlapping voxels to all voxels within the fatigue-related network in PD.
Transcriptomic analysis
The development of brain-wide gene expression atlases, such as the Allen Human Brain Atlas (AHBA), has enabled the integration of neuroimaging phenotypes with gene expression patterns69, bridging the gap between connectome and transcriptome. Following the standard neuroimaging-transcriptome analysis pipeline, this study constructed a gene expression matrix (152 regions × 10,027 genes) from the AHBA dataset (http://www.brain-map.org)70. Due to gene expression data being available for only two subjects in the right hemisphere, the analysis in this study focused on the left hemisphere. Partial least squares (PLS) regression, well-suited for high-dimensional gene expression data71, was applied to examine the relationship between the fatigue-related network and all 10,027 genes. The normalized gene expression matrix served as the predictor variable, while the unthresholded overlap map for the fatigue-related network was treated as the response variable. The first PLS component (PLS1), a linear combination of gene expression values, showed a strong correlation with regions within the fatigue-related network. The alignment test was used to test the statistical significance of PLS1, and bootstrapping (1000 iterations) was performed to assess the weighting coefficients of individual genes. The corrected weight was determined by the ratio of the raw weight to its bootstrap standard error. Genes were ranked according to the corrected weights, with those showing significant positive weights (all P < 4.98 × 10⁻⁶) being selected.
To understand the biological processes associated with the fatigue-related network in PD, an enrichment analysis was conducted. Genes with significant PLS1 weights were input into the Metascape (https://metascape.org/) for Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis, with a significance threshold of FDR-corrected P < 0.05. These genes were further assigned to cell types (excitatory and inhibitory neurons, astrocytes, microglia, endothelial cells, oligodendrocytes, and oligodendrocyte precursor cells) based on bulk-tissue AHBA expression data72. The overlap of genes within each cell type was evaluated through permutation testing (5000 iterations), and FDR correction was applied to determine statistical significance.
Neurotransmitter analysis
Neurotransmitter density maps for nine different neurotransmitter systems (acetylcholine, cannabinoid, dopamine, γ-aminobutyric acid (GABA), glutamate, histamine, norepinephrine, opioid, and serotonin), derived from PET data, were used to examine the spatial distribution of neurotransmitter receptors and identify specific receptor-disorder associations73. The fatigue-related whole brain was parcellated into 1000 subregions, and multivariate linear regression was applied to assess the contribution of each neurotransmitter system30.
To ensure robustness, this study controlled for spatial autocorrelation, assessed collinearity among neurotransmitter maps (e.g., Pearson correlation coefficients and VIF), and evaluated result consistency across different atlas parcellations (e.g., 100 subregions) and statistical thresholds (e.g., FDR-corrected P < 0.001). To control for spatial autocorrelation, this study employed spatial permutation null models74,75. Specifically, activation maps were projected onto the fsLR32k surface space to generate surface-based parcels. Parcel centroids were identified as the vertices closest to the center of each parcel, and the parcel coordinates were then randomly rotated across the spherical surface. The original parcel values were reassigned to the nearest vertices in the rotated set (5000 iterations), and the regression model was refitted for each rotation. Pspin values were derived from the null distribution generated by these rotations, representing the proportion of permutations yielding a test statistic as extreme as or more extreme than the observed value. Pspin serves as a spatially informed P-value that accounts for the inherent spatial autocorrelation, thereby reducing the risk of false positives. Statistical significance was typically assessed at Pspin < 0.05. To assess potential multicollinearity, this study calculated Pearson correlation coefficients and VIF values. Specifically, the neurotransmitter density maps derived from PET data were registered to a standard brain space. Neurotransmitter density values were extracted from each of the 1000 brain subregions, and Pearson correlation coefficients were computed to assess spatial correlations among the nine neurotransmitter systems. Moreover, this study computed the VIF values for each neurotransmitter system, using the formula:
where R²ᵢ is the coefficient of determination from a linear regression model with the i-th neurotransmitter as the dependent variable, and all other neurotransmitters as independent variables.
Behavioral relevance analysis
The behavioral relevance of the fatigue-related network was assessed through a NeuroSynth meta-analysis based on 24 topic terms from Margulies et al.76. The unthresholded overlap maps were divided into 20 binary masks (from 0-5% to 95-100% overlap in 5% increments) and used as inputs for the meta-analysis. The output provided z-statistics for each topic term (PFDR < 0.05), with significant topic terms ranked by the weighted mean of their z-scores for visualization.
Data availability
The data supporting the findings of this study are available on request from the corresponding author.
Code availability
The code supporting the findings of this study is available on request from the corresponding author.
References
Collaborators, G. B. D. P.sD. Global, regional, and national burden of Parkinson’s disease, 1990-2016: a systematic analysis for the Global Burden of Disease Study 2016. Lancet Neurol. 17, 939–953 (2018).
Bloem, B. R., Okun, M. S. & Klein, C. Parkinson’s disease. Lancet 397, 2284–2303 (2021).
Siciliano, M. et al. Fatigue in Parkinson’s disease: a systematic review and meta-analysis. Mov. Disord. 33, 1712–1723 (2018).
Pont-Sunyer, C. et al. The onset of nonmotor symptoms in Parkinson’s disease (the ONSET PD study). Mov. Disord. 30, 229–237 (2015).
Seppi, K. et al. Update on treatments for nonmotor symptoms of Parkinson’s disease-an evidence-based medicine review. Mov. Disord. 34, 180–198 (2019).
Foltynie, T. et al. Medical, surgical, and physical treatments for Parkinson’s disease. Lancet 403, 305–324 (2024).
Chaudhuri, A. & Behan, P. O. Fatigue and basal ganglia. J. Neurol. Sci. 179, 34–42 (2000).
Chaudhuri, A. & Behan, P. O. Fatigue in neurological disorders. Lancet 363, 978–988 (2004).
Boksem, M. A. & Tops, M. Mental fatigue: costs and benefits. Brain Res Rev. 59, 125–139 (2008).
Pauletti, C. et al. Central fatigue and attentional processing in Parkinson’s disease: An event-related potentials study. Clin. Neurophysiol. 130, 692–700 (2019).
Hou, Y. et al. Resting-state fMRI study on drug-naive early-stage patients with Parkinson’s disease and with fatigue. Parkinsonism Relat. Disord. 105, 75–82 (2022).
Kang, S. Y. et al. Neural and dopaminergic correlates of fatigue in Parkinson’s disease. J. Neural Transm.127, 301–309 (2020).
Kluger, B. M. et al. Structural brain correlates of fatigue in older adults with and without Parkinson’s disease. Neuroimage Clin. 22, 101730 (2019).
Li, J. et al. Alterations in regional homogeneity of resting-state brain activity in fatigue of Parkinson’s disease. J. Neural Transm. 124, 1187–1195 (2017).
Lu, Z. H. et al. Global cortical atrophy may be associated with fatigue in patients with parkinson’s disease. J. Clin. Neurosci. 79, 172–177 (2020).
Shan, A. et al. Aberrant voxel-based degree centrality and functional connectivity in Parkinson’s disease patients with fatigue. Cns Neurosci. Therapeutics 29, 2680–2689 (2023).
Siciliano, M. et al. Supplementary motor area functional connectivity in “drug-naive” Parkinson’s disease patients with fatigue. J. Neural Transm. 127, 1133–1142 (2020).
Tessitore, A. et al. Functional connectivity underpinnings of fatigue in “Drug-Naïve” patients with Parkinson’s disease. Mov. Disord. 31, 1497–1505 (2016).
Yuan, Y. S. et al. Impaired Interhemispheric Synchrony in Parkinson’s Disease with Fatigue. J. Pers. Med. 12, https://doi.org/10.3390/jpm12060884 (2022).
Zhang, J. J. et al. Abnormal resting-state neural activity and connectivity of fatigue in Parkinson’s disease. CNS Neurosci. Ther. 23, 241–247 (2017).
Zabala-Gomez, O. et al. Fatigue brain network functional connectivity in Parkinson’s disease. Neuroscience 579, 122–128 (2025).
Eickhoff, S. B., Bzdok, D., Laird, A. R., Kurth, F. & Fox, P. T. Activation likelihood estimation meta-analysis revisited. Neuroimage 59, 2349–2361 (2012).
Giehl, K., Tahmasian, M., Eickhoff, S. B. & van Eimeren, T. Imaging executive functions in Parkinson’s disease: an activation likelihood estimation meta-analysis. Parkinsonism Relat. Disord. 63, 137–142 (2019).
Tahmasian, M. et al. Resting-state functional reorganization in Parkinson’s disease: an activation likelihood estimation meta-analysis. Cortex 92, 119–138 (2017).
Darby, R. R., Joutsa, J. & Fox, M. D. Network localization of heterogeneous neuroimaging findings. Brain 142, 70–79 (2019).
Fornito, A., Zalesky, A. & Breakspear, M. The connectomics of brain disorders. Nat. Rev. Neurosci. 16, 159–172 (2015).
Fox, M. D. Mapping Symptoms to Brain Networks with the Human Connectome. N. Engl. J. Med. 379, 2237–2245 (2018).
Peng, S., Xu, P., Jiang, Y. & Gong, G. Activation network mapping for integration of heterogeneous fMRI findings. Nat. Hum. Behav. 6, 1417–1429 (2022).
Weil, R. S., Hsu, J. K., Darby, R. R., Soussand, L. & Fox, M. D. Neuroimaging in Parkinson’s disease dementia: connecting the dots. Brain Commun. 1, fcz006 (2019).
Li, J. et al. Divergent suicidal symptomatic activations converge on somato-cognitive action network in depression. Mol Psychiatry, https://doi.org/10.1038/s41380-024-02450-7 (2024).
Hou, Y. et al. Disrupted topological organization of resting-state functional brain networks in Parkinson’s disease patients with glucocerebrosidase gene mutations. Neuroradiology 65, 361–370 (2023).
De Michele, G. et al. Motor and non-motor features in Parkinson’s Disease patients carrying GBA gene mutations. Acta Neurol. Belg. 123, 221–226 (2023).
Pan, H. X. et al. GCH1 variants contribute to the risk and earlier age-at-onset of Parkinson’s disease: a two-cohort case-control study. Transl. Neurodegener. 9, 31 (2020).
Jiang, Y. et al. Parkin is the most common causative gene in a cohort of mainland Chinese patients with sporadic early-onset Parkinson’s disease. Brain Behav. 10, e01765 (2020).
Chou, K. L., Kotagal, V. & Bohnen, N. I. Neuroimaging and clinical predictors of fatigue in Parkinson disease. Parkinsonism Relat. Disord. 23, 45–49 (2016).
Pavese, N., Metta, V., Bose, S. K., Chaudhuri, K. R. & Brooks, D. J. Fatigue in Parkinson’s disease is linked to striatal and limbic serotonergic dysfunction. Brain 133, 3434–3443 (2010).
Poldrack, R. A. et al. Scanning the horizon: towards transparent and reproducible neuroimaging research. Nat. Rev. Neurosci. 18, 115–126 (2017).
Cheng, Y. et al. Brain network localization of gray matter atrophy, neurocognitive and social cognitive dysfunction in schizophrenia. Biol Psychiatry, https://doi.org/10.1016/j.biopsych.2024.07.021 (2024).
Kuppuswamy, A. The fatigue conundrum. Brain 140, 2240–2245 (2017).
Marek, S. & Dosenbach, N. U. F. The frontoparietal network: function, electrophysiology, and importance of individual precision mapping. Dialogues Clin. Neurosci. 20, 133–140 (2018).
Liu, W. et al. Perfusion imaging of fatigue and time-on-task effects in patients with Parkinson’s disease. Front Aging Neurosci. 14, 901203 (2022).
Cingolani, L. A. & Goda, Y. Actin in action: the interplay between the actin cytoskeleton and synaptic efficacy. Nat. Rev. Neurosci. 9, 344–356 (2008).
Gentile, J. E., Carrizales, M. G. & Koleske, A. J. Control of synapse structure and function by actin and its regulators. Cells 11, https://doi.org/10.3390/cells11040603 (2022).
Lepeta, K. et al. Synaptopathies: synaptic dysfunction in neurological disorders—a review from students to students. J. Neurochem 138, 785–805 (2016).
Bernstein, B. W., Maloney, M. T. & Bamburg, J. R. Actin and diseases of the nervous system. Adv. Neurobiol. 5, 201–234 (2011).
Kounakis, K. & Tavernarakis, N. The cytoskeleton as a modulator of aging and neurodegeneration. Adv. Exp. Med. Biol. 1178, 227–245 (2019).
Munoz-Lasso, D. C., Roma-Mateo, C., Pallardo, F. V. & Gonzalez-Cabo, P. Much more than a scaffold: cytoskeletal proteins in neurological disorders. Cells 9, https://doi.org/10.3390/cells9020358 (2020).
Sweet, E. S., Saunier-Rebori, B., Yue, Z. & Blitzer, R. D. The Parkinson’s disease-associated mutation LRRK2-G2019S impairs synaptic plasticity in mouse hippocampus. J. Neurosci. 35, 11190–11195 (2015).
Ignatavicius, A., Matar, E. & Lewis, S. J. G. Visual hallucinations in Parkinson’s disease: spotlight on central cholinergic dysfunction. Brain, https://doi.org/10.1093/brain/awae289 (2024).
Buddhala, C. et al. Dopaminergic, serotonergic, and noradrenergic deficits in Parkinson disease. Ann. Clin. Transl. Neurol. 2, 949–959 (2015).
Han, S., Gim, Y., Jang, E. H. & Hur, E. M. Functions and dysfunctions of oligodendrocytes in neurodegenerative diseases. Front Cell Neurosci. 16, 1083159 (2022).
Bryois, J. et al. Genetic identification of cell types underlying brain complex traits yields insights into the etiology of Parkinson’s disease. Nat. Genet 52, 482–493 (2020).
Orimo, S. et al. Unmyelinated axons are more vulnerable to degeneration than myelinated axons of the cardiac nerve in Parkinson’s disease. Neuropathol. Appl. Neurobiol. 37, 791–802 (2011).
Picciotto, M. R., Higley, M. J. & Mineur, Y. S. Acetylcholine as a neuromodulator: cholinergic signaling shapes nervous system function and behavior. Neuron 76, 116–129 (2012).
Bohnen, N. I. et al. Cholinergic system changes in Parkinson’s disease: emerging therapeutic approaches. Lancet Neurol. 21, 381–392 (2022).
Loebel, M. et al. Antibodies to beta adrenergic and muscarinic cholinergic receptors in patients with chronic fatigue syndrome. Brain Behav. Immun. 52, 32–39 (2016).
Madill, P. V. Chronic fatigue syndrome and the cholinergic hypothesis. JAMA 292, 2723; author reply 2723 (2004).
Yamamoto, S. et al. Reduction of [11C](+)3-MPB binding in brain of chronic fatigue syndrome with serum autoantibody against muscarinic cholinergic receptor. PLoS One 7, e51515 (2012).
Almohmadi, N. H. et al. Glutamatergic dysfunction in neurodegenerative diseases focusing on Parkinson’s disease: role of glutamate modulators. Brain Res Bull. 225, 111349 (2025).
Wang, J., Wang, F., Mai, D. & Qu, S. Molecular mechanisms of glutamate toxicity in Parkinson’s disease. Front. Neurosci. 14, 585584 (2020).
Berridge, C. W. Noradrenergic modulation of arousal. Brain Res Rev. 58, 1–17 (2008).
Ghasemi, M. & Mehranfard, N. Neuroprotective actions of norepinephrine in neurological diseases. Pflug. Arch. 476, 1703–1725 (2024).
Di Vico, I. A. et al. Molecular-informed network analysis unveils fatigue-related functional connectivity in Parkinson’s disease. Mov. Disord. https://doi.org/10.1002/mds.30214 (2025).
Cercignani, M. et al. Cognitive fatigue in multiple sclerosis is associated with alterations in the functional connectivity of monoamine circuits. Brain Commun. 3, fcab023 (2021).
Arm, J. et al. Altered in vivo brain GABA and glutamate levels are associated with multiple sclerosis central fatigue. Eur. J. Radiol. 137, 109610 (2021).
Pauletti, C. et al. Serotonergic central tone in Parkinson’s disease with fatigue: Evidence from the loudness dependence of auditory evoked potentials (LDAEP). Neurosci. Lett. 764, 136242 (2021).
Yeo, B. T. et al. The organization of the human cerebral cortex estimated by intrinsic functional connectivity. J. Neurophysiol. 106, 1125–1165 (2011).
Fan, L. et al. The Human Brainnetome Atlas: a new brain atlas based on connectional architecture. Cereb. Cortex 26, 3508–3526 (2016).
Fornito, A., Arnatkeviciute, A. & Fulcher, B. D. Bridging the gap between connectome and transcriptome. Trends Cogn. Sci. 23, 34–50 (2019).
Hawrylycz, M. J. et al. An anatomically comprehensive atlas of the adult human brain transcriptome. Nature 489, 391–399 (2012).
Li, W. et al. Disentangling the switching behavior in functional connectivity dynamics in autism spectrum disorder: insights from developmental cohort analysis and molecular-cellular associations. Adv. Sci. 12, e2403801 (2025).
Seidlitz, J. et al. Transcriptomic and cellular decoding of regional brain vulnerability to neurogenetic disorders. Nat. Commun. 11, 3358 (2020).
Hansen, J. Y. et al. Mapping neurotransmitter systems to the structural and functional organization of the human neocortex. Nat. Neurosci. 25, 1569–1581 (2022).
Alexander-Bloch, A. F. et al. On testing for spatial correspondence between maps of human brain structure and function. Neuroimage 178, 540–551 (2018).
Markello, R. D. & Misic, B. Comparing spatial null models for brain maps. Neuroimage 236, 118052 (2021).
Margulies, D. S. et al. Situating the default-mode network along a principal gradient of macroscale cortical organization. Proc. Natl Acad. Sci. USA 113, 12574–12579 (2016).
Acknowledgements
The present study was supported by Yunnan Fundamental Research Projects (No. 202501AV070005 and 202201BE070001-004), the National Natural Science Foundation of China (No. 82301427 and 82160275), the Sichuan Science and Technology Program (No. 2023NSFSC1576), joint Medical Project of Kunming University of Science and Technology and the First People's Hospital of Yunnan Province (KUST-KH2023006Y), Dehong Prefecture Academician & Expert Workstations: Talent Program for Vitalizing Border Areas (No. 2025RC013).
Author information
Authors and Affiliations
Contributions
J.W. and Y.H. contributed to the conception and design of the study; F.Y. and Y.H. contributed to the acquisition and data analysis; F.Y., J.S., Z.S., L.T., R.Y., L.Z., Y.M., H.S., and J.H. contributed to the drafting of the manuscript or part of it; J.W., Y.H., Y.G., and Z.S. contributed to the critical review of the manuscript. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, F., Shen, J., Sun, Z. et al. Mapping the neural and molecular basis underlying fatigue in Parkinson’s disease. npj Parkinsons Dis. 12, 3 (2026). https://doi.org/10.1038/s41531-025-01216-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41531-025-01216-4