Abstract
Sustainable food provision for a continuously growing human population is one of the major challenges for the next decades. Cultured meat represents one of the alternatives which is currently extensively explored. Yet, the most appropriate cell type, capable of long-term proliferation and myogenic differentiation, remains to be identified. Bovine mesenchymal stromal cells (MSCs) are considered as a promising cell source. Within the context of cultured meat production, it is mandatory to maximize cell yield per tissue source. Although many enzymatic methods to isolate MSCs from adipose tissue (AT) have been described, cell yield has never been compared. In this study, we evaluate 32 isolation conditions including four enzyme mixtures (Collagenase type I, Collagenase type I + Trypsin, LiberaseTM and Collagenase type IV) at varying concentrations and incubation times, regarding their efficiency to isolate MSCs from bovine subcutaneous AT. The highest cell yield in combination with a low population doubling time was obtained using LiberaseTM at a concentration of 0.1% for 3 h. MSC identity of the cells was confirmed by tri-lineage differentiation potential and cell surface marker expression. Subsequently, isolated cells were myogenically differentiated using 5-aza-2’-deoxycytidine and galectin-1. mRNA levels of the myogenic regulatory factors (MRF) myogenic factor 5 (MYF5), myogenic differentiation 1 (MYOD1), MYF6, and myogenin (MYOG) were increased, while less paired box 3 (PAX3) mRNA expression was observed when compared to undifferentiated MSCs. The presence of desmin (DES), tropomyosin (TM), and myosin heavy chain (MyHC) in myogenically differentiated bovine AT-MSCs was confirmed using immunofluorescence stainings. When considering MSCs from bovine AT as potential cell source to produce cultured meat, it is recommended to use 0.1% LiberaseTM for 3 h to ensure a high cell yield.
Similar content being viewed by others
Introduction
One of the major challenges for the future is providing healthy food to an ever-growing population. This requires a further transition to sustainable food systems. The provision of high-quality proteins and several micronutrients, particularly from animal sources, is essential to human nutrition. However, current food systems and agricultural practices, in particular meat production, are under pressure because of their environmental impact. Therefore, there is an urgent need to develop alternative food production methods to ensure the supply of high-quality proteins in a sustainable manner1.
Cultured meat represents an emerging alternative to traditionally farmed meat. Its production requires a multidisciplinary approach combining biotechnology, tissue engineering, molecular and synthetic biology. Although the first proof of concept of a cultured hamburger was reported a decade ago2, production on a larger scale is not yet within reach. To this end, the main hurdles to overcome are (i) the slow and limited proliferation rate of the currently used satellite cells, (ii) the presence of animal serum or expensive recombinant growth factors and antibiotics in the culture medium, and (iii) the differentiation towards well-structured, mature muscle fibers to provide taste and texture3,4.
Traditionally, satellite cells are used to produce cultured meat. These skeletal muscle stem cells are well described in many species and serve in vivo as a robust cell source for skeletal muscle repair. Their proliferation potential in vitro, however, is limited with expansion through 20 population doublings5, while doubling numbers in the range of 30-50 are required as starting point for downstream differentiation to generate large quantities of cultured meat4. Moreover, expanded satellite cells lose their myogenic differentiation capacity6,7. Therefore, alternative stem cell sources are explored. Due to their virtue of unlimited proliferation potential, embryonic or induced pluripotent stem cells represent interesting alternative cell sources. In 2018, the first stable culture of bovine embryonic stem cells was isolated8. Using these cells, a cell biobank could be established with an unaltered and stable karyotype, as such eliminating further dependence on animals for cell isolation9. Directing these cells towards a muscle cell lineage, however, might be more difficult10, requiring additional bioengineering approaches to achieve fully matured skeletal muscle. Furthermore, it should be mentioned that cultured meat produced from induced pluripotent stem cells should be labeled as ‘genetically modified’, which requires extensive safety testing4.
Only recently, the use of mesenchymal stromal cells (MSCs) is explored in the context of cultured meat production11,12,13. They are considered as another promising cell source due to their abundance, their role during myogenesis and their ability to differentiate into several lineages, such as myocytes and adipocytes14. Additionally, they can proliferate without losing their differentiation potential15. We recently extensively reviewed the advantages and limitations of each cell type currently used for the production of cultured meat16.
Adipose tissue (AT) is commonly used to isolate MSCs from as it is readily available and contains high numbers of MSCs which can be easily harvested17,18. Several approaches to isolate bovine AT-derived MSCs have been described, including mechanical tissue disruption, explant methods or enzymatic digestion. However, cell isolation efficiency is rarely evaluated19. Furthermore, isolation procedures might not only affect total cell yield but also cell viability, proliferation potential, tri-lineage differentiation capacity, and cell surface marker expression20. In the study of Salehinejad et al. (2012), for example, four methods were compared to isolate MSCs from human Wharton’s jelly. The use of a collagenase (Coll)/trypsin enzymatic combination resulted in a higher cell yield and higher expression of pluripotent cell markers such as C-kit and Oct-4, when compared to other isolation methods21.
Within the context of cultured meat production, it is mandatory to maximize cell yield per isolation. As significantly more days are required until first cell harvest using explant cultures22, only enzymatic digestion methods were evaluated in this study. Besides an increased cell viability, other advantages of enzymatic digestion include reproducibility, low-cost, quick sample preparation and potential automation19,22,23,24. Despite these advantages, not all enzymes perform equally well to digest tissues. In fact, the purity and/or the combination of enzymes used affect cell isolation efficiency25.
To identify which enzymes or enzyme combinations, concentrations and incubation times are most frequently used to isolate MSC from AT, a literature search was performed. Across species, Coll type I is most frequently used, while a 1 mg/mL enzyme concentration and a 3 h incubation time are frequently reported across species (Table 1). Other enzymes reported are trypsin, Coll type II and IV, and Liberase (Lib). In two studies, Coll I was combined with trypsin to isolate ovine or bovine MSC from abdominal AT26,27.
Based on this literature search, 32 different conditions to isolate bovine MSCs from AT were selected to identify the most efficient protocol to ensure a maximum cell yield per isolation. When cells were isolated from at least five out of eight donors with an average cell yield exceeding 35 × 106 cells/g tissue at passage (P)1, they were further characterized by determining their proliferation potential, tri-lineage differentiation ability and immunophenotype. Subsequently, cells isolated with the protocol giving the highest yield, were myogenically differentiated.
Results
Isolation of bovine AT-MSCs using different conditions
In preliminary experiments, MSCs isolated using the culture explant method were passaged for the first time after approximately 10 days in combination with a substantially lower cell yield (data not shown). Therefore, only enzymatic digestion methods were evaluated in this study. To identify the most efficient method to isolate bovine MSCs from AT, the following conditions were evaluated: four enzymes (Coll IA, Coll IA + Trypsin (T), LibTM and Coll IV), two concentrations (0.1% and 0.04%) and four incubation times (3 h, 6 h, overnight (ON) and 24 h).
As a high isolation efficiency and cell viability are mandatory for further applications, we considered an isolation condition successful when (i) cells of P0 were observed within 21 days, (ii) cell yield was minimally 1 × 106 cells/g AT at P1 (based on preliminary experiments), and (iii) cell viability was higher than 95%. Spindle-shaped plastic-adherent cells were observed for all conditions within 2 days of culture, except when using Coll IV at 0.04% for 6 h, ON and 24 h (Fig. 1A). For the latter conditions, no plastic-adherent cells were observed even after 21 days of culture. Cells were passaged when at least 5 colony forming units (CFU) were observed, which was within four to seven days after start of the culture. Cell viability exceeded 95% for all isolation conditions. The average cell yield ranged from 4.38 × 106 to 130 × 106 cells/g AT (Fig. 1A).
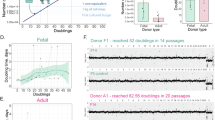
A Quantitative data of total cell yield (cells/g AT) per isolation condition. The dotted line represents one of the selection criteria for further characterization (i.e. average cell yield ≥35×106 cells/g AT). The colored lines indicate the mean total cell yield per isolation condition. B The effect of the enzyme on cell yield was statistically compared to Coll IA (i.e. the most frequently reported enzyme for AT-MSC isolation), while enzyme concentration (0.1%, highest cell yield) and incubation time (3 h, lowest incubation time) were fixed. AT adipose tissue, IQR interquartile range, ON overnight, Coll collagenase, LibTM liberaseTM. *Cells isolated using these conditions were further characterized to confirm their MSC identity.
Results were statistically analyzed to compare the conditions in terms of cell yield. For each enzyme, the effect of enzyme concentration and incubation time on cell yield was assessed. Varying the enzyme concentration or incubation time did not result in significant differences in cell yield (Fig. 1A). Nevertheless, higher cell yields were observed when the 0.1% enzyme concentration was combined with low incubation times (3 h and 6 h) (Fig. 1A).
Subsequently, the effect of the enzyme on cell yield was assessed in comparison to Coll IA, which is the most frequently reported enzyme for AT-MSC isolation. For this analysis, we selected the 0.1% enzyme concentration (highest cell yield) and 3 h (lowest incubation time). Only LibTM 0.1% at 3 h performed significantly better when compared to Coll IA 0.1% at 3 h (Fig. 1B). Isolation with LibTM resulted in an average cell yield ranging from 30.48 × 106 to 67.1 × 106 cells/g AT.
To confirm the MSC identity of the isolated cells, following criteria were used to select the most efficient methods: (i) cells were obtained from at least five out of eight donors, and (ii) average total cell yield exceeding 35 × 106 cells/g AT (based on preliminary results, dotted line, Fig. 1A). As such, cells isolated using LibTM at 0.1% (3 h, 6 h and ON), Coll IA at 0.04% (6 h) and Coll IA at 0.1% (6 h) were further characterized by determining their proliferation potential, tri-lineage differentiation ability and immunophenotype.
Proliferation potential of bovine AT-MSCs
To evaluate the effect of the selected isolation conditions on cell proliferation, days in culture (starting from isolation until > 5 CFU were observed) was compared between the selected isolation conditions using a time-to-event analysis (Fig. 2A). As early as 5 days after start of the culture, > 5 CFU were reached with cells isolated using LibTM at 0.1% for 3 and 6 h and Coll IA at 0.04% for 6 h. For the isolation conditions LibTM at 0.1% for ON and Coll IA at 0.1% for 6 h, > 5 CFU were reached at 6 and 7 days after start of culture, respectively (Fig. 2A). The time to reach > 5 CFU did not significantly differ between the selected isolation conditions. It must be mentioned however that only when using LibTM at 0.1% for 3 h, cells were obtained from all donors (Fig. 2A). No cells were passaged within 21 days for 1 donor with the LibTM 0.1% 6 h and ON, and Coll IA 0.1% 6 h condition, and for 2 donors with the Coll IA 0.04% 6 h condition (Fig. 2A). Furthermore, the in vitro proliferation potential was evaluated during eight consecutive passages. Cells isolated with LibTM at 0.1% for 3 and 6 h showed the lowest population doubling time. In all other selected conditions, the population doubling time increased with population doublings (Fig. 2B).
A Time-to-event analysis of bovine AT-MSCs isolated using the selected isolation conditions. An offset was set to visualize all selected conditions on the graph. B The in vitro proliferation potential of bovine AT-MSCs was evaluated during eight consecutive passages. The dots show the median of the population doubling time in hours (h) while the bars show the interquartile range (IQR). IQR interquartile range, ON overnight, Coll collagenase, LibTM liberaseTM.
Tri-lineage differentiation potential of bovine AT-MSCs
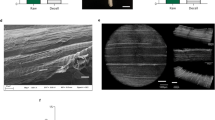
To confirm the MSC identity of the isolated cells of the selected conditions, their tri-lineage differentiation potential was quantified by a differentiation ratio. The latter is obtained using an image-based analysis by dividing the area % of the differentiation signal by the area % of the nuclear signal or cell culture area. As illustrated in Fig. 3, all isolated cells were able to differentiate towards adipocytes, osteocytes and chondrocytes, respectively (Fig. 3A–C). No significant differences in tri-lineage differentiation potential were observed between the selected isolation conditions.
Isolation methods are linked in an interaction plot and the mean differentiation ratio ± standard deviation is shown for (A) adipogenic, (B) osteogenic, and (C) chondrogenic differentiation. Donors were linked in an interaction plot and the percentage variability related to inter-donor variation is shown. Each color represents a selected isolation condition. Representative images for each lineage are showed below the control or differentiation condition, respectively. Scale bar = 200 µm. GAG glycosaminoglycans, ON overnight, Coll collagenase, LibTM liberaseTM.
Immunophenotypic profile of bovine AT-MSCs
Following the criteria formulated for human MSCs, MSCs must express CD73, CD90 and CD105 but lack expression of the hematopoietic or endothelial markers CD11b or CD14, CD34, CD45, CD79α or CD19 and MHC class II28,29. However, MSCs represent a heterogeneous cell population as illustrated by differences in marker expression between species and sources30,31,32,33. For example, our previous research showed that bovine AT-MSCs express CD29 and CD44 and lacked expression of CD14, CD45, CD73, CD79α and MHC class II30. Figure 4 shows that the AT-MSCs isolated with the selected protocols also expressed CD29 and CD44 and lacked expression of CD14, CD45, CD73, CD79α and MHC class II (Fig. 4). No significant differences in immunophenotypic profile were observed between the selected isolation conditions.
Quantitative data is presented as the median of the % positive cells, while the bars show the interquartile range (IQR). Each color represents a selected isolation condition. IQR interquartile range, ON overnight, Coll collagenase, LibTM LiberaseTM.
Myogenic differentiation potential of bovine AT-MSCs
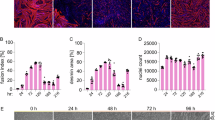
Subsequently, AT-MSCs, isolated with LibTM 0.1% for 3 h, were myogenically differentiated for 14 days using 10 µM of the DNA methyltransferase inhibitor 5-aza-2’-deoxycytidine and 100 nM of the myoblast-secreted factor galectin-1. Myogenically differentiated AT-MSCs showed cell protrusions, cell alignment and fusion of nuclei, while undifferentiated MSCs maintained their spindle shaped morphology (Fig. 5A).
A Morphology of myogenically differentiated AT-MSCs changed as visualized by bright-field microscopy. B Expression of myogenic markers PAX3, MYF5, MYOD1, MRF4, and MYOG at mRNA level. Quantitative data is presented as the median of the log2 fold change, while the bars show the interquartile range (IQR). **: significant difference compared to undifferentiated bovine AT-MSCs (p = 0.0078). C Expression of myogenic markers DES, (D) TM, and (E) MyHC at protein level. Quantitative data is presented as the median of the log10 FITC/DAPI ratio, while the bars show the IQR. Scale bar = 100 µm. DES desmin, TM tropomyosine, MyHC myosin heavy chain, Diff differentiated, PAX3 paired box 3, MYF5/6 myogenic factor 5/6, MYOD1 myogenic determination 1, MYOG myogenin.
Following 14 days of myogenic differentiation, AT-MSCs expressed decreasing mRNA levels of paired box 3 (PAX3) in combination with elevated mRNA levels of myogenic factor 5 (MYF5), myogenic differentiation 1 (MYOD1), MYF6, and myogenin (MYOG) when compared to undifferentiated MSCs (Fig. 5B). Using immunofluorescence stainings, the presence of desmin (DES), tropomyosin (TM), and myosin heavy chain (MyHC) was confirmed in differentiated AT-MSCs at day 14 (Fig. 5C–E). Interestingly, MyHC expression was also observed in undifferentiated AT-MSCs at day 14 (Fig. 5E).
Discussion
As maximizing both isolation efficiency as well as cell yield per isolation are mandatory when considering cell sources to produce cultured meat, different protocols to isolate bovine AT-MSCs were evaluated. Although all enzymes or enzyme combinations used in this study were already described in literature to isolate AT-MSCs in different species, there was only one isolation condition combination that ensured the isolation of MSCs from every donor with a high cell yield, namely LibTM 0.1% for 3 h. No significant differences were observed between the selected isolation conditions regarding proliferation potential, tri-lineage differentiation capacity or immunophenotype of the bovine AT-MSC.
To the best of our knowledge, LibTM has not yet been used to isolate bovine AT-MSCs. It consists of a mixture of highly purified Coll I and II together with the neutral protease thermolysin, that breaks peptide bonds in fibronectin, collagen I, II and non-specific cell adhesion proteins34. In our study, this enzyme combination resulted in a higher cell yield, which might indicate that bovine AT is more efficiently degraded when compared to the other enzymes or enzyme combinations. Although already described for the isolation of AT-MSCs, Coll IV resulted in significantly less successful isolations, which can be explained by the fact that collagen type IV is less present in subcutaneous bovine AT35. Indeed, the extracellular matrix of bovine subcutaneous AT consists mainly of collagen types I, III, V and VI, while collagen types II and IV are less abundant35,36.
For each enzyme, the effect of enzyme concentration (0.1% and 0.04%) and incubation time (3 h, 6 h, ON and 24 h) on cell yield and successful isolation was evaluated. Although 0.1% is frequently used, a variation of concentrations is described in literature, ranging from 0.001% to 0.2% (Table 1)37. A higher concentration might accelerate the enzymatic reaction as long as there is substrate available38. In our study, no significant effect of enzyme concentration on cell yield was observed. However, a lower concentration of Coll IV clearly resulted in less successful isolations and a lower cell yield (Fig. 1A).
Longer incubation times may enhance cell release39 but might be associated with an extensive degradation of extracellular matrix and cell membrane proteins, resulting in less plastic adherence and thus a lower cell yield40. Additionally, an increased collagenase digestion time (more than 50 min) might result in a significantly decreased number of viable adipocytes41. In our study, no significant differences were observed between incubation times when enzyme and enzyme concentration were fixed. It should be mentioned however that the cell yield decreased when AT was incubated longer (ON and 24 h) with Coll IA combined with trypsin. This might be explained by the rather aggressive action of trypsin21.
Different protocols are described to isolate bovine MSCs of which Coll IA 0.1% is frequently used42,43,44. In our study, LibTM at 0.1% for 3 h is identified as the preferred method, as bovine AT-MSCs were isolated from every donor with a high cell yield and low population doubling time, when compared to the other isolation conditions. To generate large quantities of cultured meat, doubling numbers in the range of 30-50 are necessary4. In our study, the population doubling time of MSCs after isolation with LibTM at 0.1% for 3 and 6 h did not increase with population doublings, indicating that their proliferation potential is preserved during eight consecutive passages. As 2–3 population doublings per passage were observed, resulting in a population doubling range of 15.77-17.61 at P8 (data not shown), the above-mentioned criterium is within reach. After repeated passaging, MSCs may enter a state of replicative senescence, also referred to as the Hayflick limit45. In our hands, however, senescence of bovine AT-MSCs is only observed after 15–16 passages (data not shown).
While meat products are predominantly composed of muscle, the presence of fat and its lipid composition significantly influences meat quality, contributing to the perception of flavors, mouthfeel sensation, appearance, texture, and nutritional values13. In this study, bovine AT-MSC were successfully differentiated adipogenically and myogenically. Downregulation of PAX3 was observed in combination with increased MRF expression, which is indicative for MSC transition into the myogenic lineage. Indeed, the PAX3 protein level of myotomal cells will decline once MRF expression increases during embryogenesis46. Although we and others demonstrated that bovine MSCs are able to differentiate myogenically, it must be mentioned that only immature myotubes are formed, with limited fusion capacity47,48. In future studies, myogenic differentiation efficiency towards well-structured, mature myotubes, being 10-100 µm in diameter and predominantly containing contractile proteins, such as actin and myosin4, should be optimized.
In conclusion, our study showed that only the LibTM 0.1% 3 h isolation condition resulted in a high MSC yield from all donors, of which the adipogenic and myogenic differentiation potential was confirmed. This study paves the way to consider MSCs as attractive alternative cell source to produce cultured meat.
Materials and methods
Cell isolation methods
Bovine MSCs were isolated from subcutaneous AT (at the back region, near shoulder) from eight healthy, adult (2–4 years) female Belgian White Blue cows, collected in the local abattoir. Briefly, AT samples were transported within 1–2 h to the lab in phosphate buffered saline (PBS) without Ca2+/Mg2+ (Gibco) containing 0.05 mg/mL gentamycin (Gibco). Tissue samples were extensively washed to remove blood, cut in small pieces, weighed, transferred into a 4-well plate and minced with a sterile pipet tip for 1–2 min in an enzymatic solution at 38.5 °C in a humidified atmosphere containing 5% CO2. The following isolation conditions were evaluated: the enzymes Coll IA (1262 units/mg, Sigma, C9722), LibTM (13 units/mg, Sigma, LIBTM-RO) and Coll IV (Sigma, 1267 units/mg, C5138) as well as a combination of Coll IA with trypsin 0.25% (Sigma, 1609 units/mg, T4799) were tested at two concentrations (0.1% and 0.04%) and four incubation times (3 h, 6 h, ON ( ± 15 h) and 24 h) (Fig. 6). Subsequently, the enzymatic reaction was neutralized with an equal amount of cold culture medium, consisting of low glucose Dulbecco’s Modified Eagle Medium (DMEM-LG; Invitrogen) supplemented with 30% fetal bovine serum (FBS; Sigma), 10–11 M dexamethasone (Sigma), 1% antibiotic-antimycotic solution (Sigma) and 1% L-glutamine (Invitrogen)49. After 1 min, the non-buoyant fraction was filtered using a 70 µm cell strainer. Cells were collected after centrifugation for 5 min at 400 × g at room temperature (RT). After two washing steps, cells were resuspended into culture medium, seeded in a 25 cm² culture flask and cultured at 38.5 °C in a humidified atmosphere containing 5% CO2. After 24 h, non-adherent cells were removed by replacing the culture medium. Culture medium was replaced 2–3 times/week and when at least five CFU were observed, cells were passaged using 2.5 mg/mL trypsin-0.2 mg/mL EDTA (Sigma). Cell viability and concentration was determined by trypan blue exclusion using a Neubauer hemocytometer. Cells were seeded at a density of 5000 cells/cm2 and cultured in expansion medium (culture medium with the omission of dexamethasone) from P1 onwards50. An overview of the experimental design is provided in Fig. 6.
For each AT sample (n = 8), bovine MSCs were isolated using 32 conditions consisting of four different enzymes (Coll IA, Coll IA + T, LibTM, Coll IV), two concentrations (0.1% and 0.04%) and four incubations times (3 h, 6 h, ON and 24 h). Isolation conditions were considered as successful when (i) cells were observed within 21 days after isolation (P0), (ii) cell yield was minimally 1 × 106 cells/g AT at P1, and (iii) cell viability was higher than 95%. Subsequently, bovine MSCs were further characterized, i.e. proliferation potential, tri-lineage differentiation ability (at P4) and immunophenotype (at P4), when (i) cells were obtained in at least five out of eight donors and (ii) the average total cell yield was higher than 35 × 106 cells/g AT. Finally, cells isolated with the most efficient isolation protocol, were myogenically differentiated. Figure created with BioRender. ON overnight, MSCs mesenchymal stromal cells, P passage, IF immunofluorescence.
Proliferation potential of bovine AT-MSCs
During eight consecutive passages, cells were seeded at a density of 5000 cells/cm² in six-well culture plates and sub-cultured at 70–80% confluency with expansion medium. Population doublings were calculated based on the following formula: population doublings = ln(Nf/Ni)/ln2 (Ni, the initial cell number; Nf: the harvest cell number). The population doubling time was calculated from the population doublings and cell culture time in hours for each passage by the following formula: population doubling time = (cell culture time/population doublings) x 2449.
Tri-lineage differentiation potential of bovine AT-MSCs
Following the guidelines formulated by the International Society for Cellular Therapy for human MSCs28, undifferentiated bovine MSCs were induced after three passages towards the adipogenic, chondrogenic, and osteogenic lineage, respectively, as routinely performed in our lab30,51. Non-induced MSCs cultured in expansion medium were used as appropriate negative controls.
Briefly, adipogenic differentiation was performed in 24-well culture dishes with 21,000 cells/cm² cultured until 90-100% confluency. Adipogenic differentiation was induced by subsequent cycles of 72 h culturing in adipogenic induction medium (DMEM-LG supplemented with 1 µM dexamethasone, 0.5 mM 3-isobutyl-1-methylxanthine (Sigma), 10 µg/mL rh-insuline (Sigma), 0.2 mM indomethacin (Sigma), 15% rabbit serum (Sigma), 50 µg/mL gentamycin and 1% antibiotic-antimycotic solution) followed by 24 h of culturing in the adipogenic maintenance medium (identical to the adipogenic induction medium except for the omission of dexamethasone, indomethacin and 3-isobutyl-1-methylxanthine). After 14 days, adipogenic differentiation was assessed using Oil Red O histological staining with a Mayer’s modified hematoxylin (Abcam) counterstaining.
Osteogenic differentiation was performed in 24-well culture dishes with 10,000 cells/cm² cultured in expansion medium until 90-100% confluency. Osteogenic differentiation was induced with osteogenic medium (DMEM-LG supplemented with 10% FBS, 0.05 mM L-ascorbic acid-2-phosphate (Sigma), 100 nM dexamethasone, 10 mM β-glycerophosphate (Sigma), 50 µg/mL gentamycin and 1% antibiotic-antimycotic solution). After 21 days, osteogenic differentiation was evaluated using the Alizarin Red S histological staining (Sigma) according to the manufacturer’s instructions.
Chondrogenic differentiation was induced using a micromass culture system, i.e. 250,000 cells were centrifuged in 15 mL Falcon tubes for 5 min at 300 g at RT. Subsequently, 0.5 mL chondrogenic medium (basal differentiation medium (Lonza), supplemented with 10 ng/mL transforming growth factor-β3 (Lonza)) was added to each tube without disturbing the cell pellet. After 21 days, cell pellets were fixed overnight with 4% formaldehyde (Carl Roth), and further processed for paraffin sectioning. Chondrogenic differentiation was evaluated by Alcian blue (Sigma) histological staining, with 0.1% Nuclear Fast Red (Sigma) counterstaining.
Images of adipogenic and osteogenic differentiation were captured using an inverted microscope (DMi1, Leica Biosystems, Nussloch, Germany) at a magnification of 200x and 100x, respectively. Images of chondrogenic differentiation were captured at 200x using a bright-field microscope (DM LB2, Leica Biosystems, Nussloch, Germany). Tri-lineage differentiation potential of bovine MSCs was quantified as previously described, using an image acquisition protocol based on color deconvolution51.
Multi-color flow cytometry
Following the guidelines formulated by the International Society for Cellular Therapy for human MSCs28, undifferentiated bovine MSCs from the fourth passage were immunophenotyped using multi-color flow cytometry, as previously described30. For panel 1, approximately 500,000 cells were incubated for 30 min at 4°C in the dark with a CD73-specific antibody (Ab) (Bioss Antibodies, polyclonal, 1:50). After blocking and washing, cells were incubated with an APC-Cy7 secondary Ab (AAT Bioquest, 1:100) for 20 min at 4°C in the dark. The cells were washed and subsequently blocked for 15 min using 10% rabbit serum (Sigma). Next, cells were incubated with a biotinylated CD90-specific Ab (Bioss Antibodies, polyclonal, 1:50). After washing, cells were incubated with a secondary Streptavidin PerCP-Cy5.5 (Invitrogen, 1:100) together with the directly labeled primary monoclonal Abs CD29-APC (BioLegend, TS2/16, 1:50), CD45-FITC (BioRad, CC1, 1:10) and MHC II-(R)PE (BioRad, CC158, 1:20). After washing, cell pellets were finally resuspended in 100 µL PBS with 0.001 mM Sytox Blue (Invitrogen). For panel 2, approximately 500,000 cells were stained with a fixable live/dead violet stain (Invitrogen). After one washing step, cells were incubated with the CD34-specific Ab (Bioss Antibodies, polyclonal, 1:100). After blocking and washing, cells were incubated with a PE-Cy5 secondary Ab (ThermoFisher, 1:100) together with the directly labeled primary monoclonal Abs CD14-(R)PE (BioRad, CC-G33, 1:100) and CD44-FITC (BioRad, IL-A118, 1:10). Subsequently, cells were fixed and permeabilized using BD Cytofix/CytopermTM (BD Biosciences) for 20 min at 4°C. After blocking and washing with 1x Perm/Wash washing buffer (BD Biosciences), cells were incubated with the CD79α-AF700 Ab (BioRad, HM57, 1:50) for 20 min at 4°C in the dark. After a washing step, cell pellets were finally resuspended in 100 µL PBS. For both panels, at least 10,000 viable single cells were analyzed using a CytoFLEX flow cytometer (Beckman Coulter), and data was subsequently analyzed in the CytExpert software. All data were corrected for autofluorescence, compensated and compared to specific fluorescence minus one controls. Compensation for spectral overlap between fluorophores was performed using an automatic calibration technique and subsequently evaluated individually with a compensation matrix.
Myogenic differentiation
To assess the myogenic differentiation potential, bovine AT-MSCs of P4 were seeded at a density of 5,000 cells/cm² in 6-well (for RT-qPCR) and 24-well culture plates (immunofluorescence) and cultured to 70–80% confluency. Subsequently, myogenic differentiation was induced for 14 days using myogenic differentiation medium consisting of high glucose Dulbecco’s Modified Eagle Medium (DMEM-HG; ThermoFisher) supplemented with 10% FBS, 1% antibiotic-antimycotic solution, 10 µM 5-aza-2’-deoxycytidine (Sigma) and 100 nM galectin-1 (R&D Systems), as described by Okamura et al. (2018)47.
For RT-qPCR analysis, 500,000 cells were trypsinized, washed with PBS and stored as cell pellets at -80°C until RNA extraction. After thawing in 1 mL TRIR (ABgene) at RT for 5 min, total RNA was isolated in 30 µL using the Aurum Total RNA Mini Kit (Bio-Rad), according to the manufacturer’s instructions and including an on-column DNase treatment of 20 min at RT. RNA purity and concentration was measured via Nanodrop analysis (Thermo Scientific) and DNA contamination was checked via minus-RT PCR with the TBP assay52. Up to 1 µg of DNA-free RNA was converted into cDNA with random hexamers and oligo-dT using the ImProm-II Reverse Transcription System (Promega). The integrity and the PCR amplificability of the cDNA (1/10 dilution) was confirmed via the UBC integrity assay53.
qPCR reactions were performed on 2 µL 1/10 diluted cDNA with the KAPA SYBR FAST qPCR Master Mix (KAPA Bio-systems) and 500 nM primers (Table S1) in a final volume of 10 μL. The reactions were performed on the CFX96 Touch Real-Time PCR Detection System (Bio-Rad). Initial denaturation at 95 °C for 3 min to activate the DNA polymerase was followed by 40 cycles of denaturation at 95 °C for 20 s and a combined annealing/extension/signal detection step at the primer annealing temperature for 40 s (Table S1). Finally, a melt curve analysis was performed (from 70 °C to 95 °C with 0.5 °C increments of 5 s) to confirm that the detected signals came from the intended amplicons. All reactions were performed in duplicate and a no template control was included in each run. A four-fold dilution series of one reference cDNA sample was additionally included for each gene to acquire PCR efficiency based on relative standard curves. Calculation of the Cq-values (quantification cycle), PCR efficiencies, correlation coefficients and analysis of the melting curves was performed by means of the CFX Maestro Software.
Additionally, inter-run calibration was performed. When inter-sample variability exceeded a Cq-value of 0.5, a third run was performed. The mean Cq-value of the replicates was used for further data processing. The expression stability of 5 commonly used reference genes (HPRT1, TBP, ACTB, SDHA and YWHAZ) was determined in the samples. NormFinder for Microsoft Excel was applied to identify the most efficient reference genes54. Applying this software program, HPRT1 and YWHAZ were indicated as the most stable genes in the samples, since the average stability values of these genes was 0.127. Relative quantification was calculated based on the delta Cq-method, with HPRT1 and YWHAZ used as reference genes and undifferentiated MSCs as a negative control.
For immunofluorescence analysis, cells were fixed in 4% formaldehyde for 20 min at RT. After two washing steps in PBS, cells were permeabilized with 0.1% TritonTM X-100 (Sigma) in PBS for 5 min at RT. The cells were washed again twice in PBS and blocked with 1% bovine serum albumin in PBS for 30 min at RT. Subsequently, cells were incubated with mouse anti-DES (1:100, Sigma, D1033), TM (1:100, Sigma, T9283) and MyHC (1:100, Sigma, M4276) primary Abs at 4°C ON in the dark. The cells were washed twice with PBS followed by 1 h incubation with goat anti-mouse IgG fluorescent isothiocyanate (FITC, 1:100, BioRad) secondary Ab at RT in the dark. Cells were washed twice with PBS and nuclei were visualized using 4,6-diamidino-2-phenylindole dihydrochloride (DAPI, 1:1000, ThermoFisher). Following three washing steps, stained cells were imaged using fluorescence microscopy (Leica DMi8, 10x magnification). For image analysis, two different wells per donor and four images per well were analysed.
Data analysis
The effect of incubation time and enzyme concentration on cell yield (cells/g AT) was assessed within each enzyme using a Kruskal-Wallis test with Dunn’s correction for multiple comparisons. The effect of the enzyme on cell yield was assessed compared to Coll IA (most frequently reported enzyme for AT-MSC isolation), with fixed parameters of time (3 h) and concentration (0.1%), using a Kruskal-Wallis test with Dunn’s post hoc test and Bonferroni correction for multiple comparisons. A Cox proportional hazards frailty model was fit to study the association between the growth of cells (1 = 70–80% confluency at P1, 0 = no plastic-adherent cells past 21 days) and the selected isolation conditions. A Kaplan-Meier graph was generated for visualizing the time in culture of P1 cells. The analysis was performed using R Studio (Version 1.2.1335, RStudio, Boston, MA, USA), using the package “survival”. Quantitative data of the population doubling time, multi-color flow cytometry, RT-qPCR and immunofluorescence is presented as median (interquartile range, IQR) from eight replicates. Data of population doubling time and multi-color flow cytometry were statistically analyzed using a Kruskal-Wallis test with Dunn’s post hoc test and Bonferroni correction for multiple comparisons. Data of RT-qPCR and immunofluorescence were statistically analyzed using a Wilcoxon matched-pairs signed rank t-test.
Data analysis for quantification of tri-lineage differentiation was performed using R Studio, using the package “nlme”. Normality was evaluated by visual examination of Q-Q plots and equality of variances was assessed by plotting the residuals against the fitted values. The effect of the condition, i.e. differentiated vs non-induced cells, was evaluated using repeated measures ANOVA (a univariate approach, condition = fixed factor and donor = random factor).
All data were visualized using GraphPad Prism 8.0.2.
Data availability
Data files presented in this study are available upon request, please contact the corresponding author Catharina De Schauwer.
References
Warner, R. D. Review: analysis of the process and drivers for cellular meat production. Animal 13, 3041–3058 (2019).
Post, M. J. Cultured meat from stem cells: Challenges and prospects. Meat Sci. 92, 297–301 (2012).
Olenic, M. & Thorrez, L. Cultured meat production: what we know, what we don’t know and what we should know. Ital. J. Anim. Sci. 22, 749–753 (2023).
Thorrez, L. & Vandenburgh, H. Challenges in the quest for ‘clean meat.’. Nat. Biotechnol. 37, 215–216 (2019).
Kadim, I. T., Mahgoub, O., Baqir, S., Faye, B. & Purchas, R. Cultured meat from muscle stem cells: A review of challenges and prospects. J. Integr. Agric. 14, 222–233 (2015).
Choi, K. H. et al. Muscle stem cell isolation and in vitro culture for meat production: A methodological review. Compr. Rev. Food Sci. Food Saf. 20, 429–457 (2021).
Ding S. et al. Maintaining bovine satellite cells stemness through p38 pathway. Sci. Rep. 8 https://doi.org/10.1038/s41598-018-28746-7 (2018).
Bogliotti, Y. S. et al. Efficient derivation of stable primed pluripotent embryonic stem cells from bovine blastocysts. Proc. Natl. Acad. Sci. USA 115, 2090–2095 (2018).
Ben-Arye, T. & Levenberg, S. Tissue engineering for clean meat production. Front Sustain Food Syst. 3, 46 (2019).
Stephens, N. et al. Bringing cultured meat to market: Technical, socio-political, and regulatory challenges in cellular agriculture. Trends Food Sci. Technol. 78, 155–166 (2018).
Hanga, M. P. et al. Bioprocess development for scalable production of cultivated meat. Biotechnol. Bioeng. 117, 3029–3039 (2020).
Naraoka, Y. et al. Isolation and characterization of tissue resident CD29-positive progenitor cells in livestock to generate a three-dimensional meat bud. Cells 10, 2499 (2021).
Zagury, Y. et al. Engineered marble-like bovine fat tissue for cultured meat. Commun. Biol. 5, 927 (2022).
Nawaz, S. et al. Molecular characterization of bovine amniotic fluid derived stem cells with an underlying focus on their comparative neuronal potential at different passages. Ann. Anat. 228, (2020). https://doi.org/10.1016/j.aanat.2019.151452 (2020).
Kim, G. Y. et al. Comparative analysis of porcine adipose- and Wharton’s jelly-derived mesenchymal stem cells. Animals 13, 2947 (2023).
Olenic, M. et al. Livestock cell types with myogenic differentiation potential: considerations for the development of cultured meat. Animal https://doi.org/10.1016/j.animal.2024.101242 (2024).
Fei, W. et al. Synergistic effect of hydrogen and 5-Aza on myogenic differentiation through the p38 MAPK signaling pathway in adipose-derived mesenchymal stem cells. Int. J. Stem Cells 16, 78–92 (2022).
Helms, F. et al. Complete myogenic differentiation of adipogenic stem cells requires both biochemical and mechanical stimulation. Ann. Biomed. Eng. 48, 913–926 (2020).
Gugjoo, M. B., Amarpal, Fazili, M. R., Shah, R. A. & Sharma, G. T. Mesenchymal stem cell: Basic research and potential applications in cattle and buffalo. J. Cell. Physiol. 234, 8618–8635 (2019).
Bajek, A. et al. Does the harvesting technique affect the properties of adipose-derived stem cells? The comparative biological characterization. J. Cell. Biochem. 118, 1097–1107 (2017).
Salehinejad, P. et al. Comparison of different methods for the isolation of mesenchymal stem cells from human umbilical cord Wharton’s jelly. Vitr. Cell. Dev. Biol. Anim. 48, 75–83 (2012).
Gittel, C. et al. Isolation of equine multipotent mesenchymal stromal cells by enzymatic tissue digestion or explant technique: Comparison of cellular properties. BMC Vet. Res. 9, 221 (2013).
Hornick, J. E., Duncan, F. E., Shea, L. D. & Woodruff, T. K. Multiple follicle culture supports primary follicle growth through paracrine-acting signals. Reproduction 145, 19–32 (2013).
Schmidt, V. et al. Comparison of the enzymatic efficiency of Liberase TM and tumor dissociation enzyme: Effect on the viability of cells digested from fresh and cryopreserved human ovarian cortex. Reprod. Biol. Endocrinol. 16, 57 (2018).
Aronowitz, J. A., Lockhart, R. A. & Hakakian, C. S. Mechanical versus enzymatic isolation of stromal vascular fraction cells from adipose tissue. SpringerPlus 23, 713 (2015).
Grzesiak, J., Krzysztof, M., Karol, W. & Joanna, C. Isolation and morphological characterization of ovine adipose-derived mesenchymal stem cells in culture. Int. J. Stem Cells 4, 99–104 (2011).
Lu, T. et al. Isolation and characterization of adipose-derived mesenchymal stem cells (ADSCs) from cattle. Appl. Biochem. Biotechnol. 174, 719–728 (2014).
Dominici, M. et al. Minimal criteria for defining multipotent mesenchymal stromal cells. The International Society for Cellular Therapy position statement. Cytotherapy 8, 315–317 (2006).
Viswanathan, S. et al. Mesenchymal stem versus stromal cells: International Society for Cell & Gene Therapy (ISCT®) Mesenchymal Stromal Cell committee position statement on nomenclature. Cytotherapy 21, 1019–1024 (2019).
Heyman, E. et al. Validation of multiparametric panels for bovine mesenchymal stromal cell phenotyping. Cytometry 2023, 1–12 (2023).
De Schauwer, C. et al. In search for cross-reactivity to immunophenotype equine mesenchymal stromal cells by multicolor flow cytometry. Cytometry 81A, 312–323 (2012).
Suga, H. et al. Functional implications of CD34 expression in human adipose-derived stem/progenitor cells. Stem Cells Dev. 18, 1201–1210 (2019).
Mishra, S. et al. Umbilical cord tissue is a robust source for mesenchymal stem cells with enhanced myogenic differentiation potential compared to cord blood. Sci. Rep. 10, 18978 (2020).
Schmidt, V. M. et al. Comparison of the enzymatic efficiency of Liberase TM and tumor dissociation enzyme: Effect on the viability of cells digested from fresh and cryopreserved human ovarian cortex. Reprod. Biol. Endocrinol. 16, 1–14 (2018).
Sheng, H. et al. Weighted gene co-expression network analysis identifies key modules and central genes associated with bovine subcutaneous adipose tissue. Front. Vet. Sci. 9, 1–12 (2022).
Nakajima, I., Yamaguchi, T., Ozutsumi, K. & Aso, H. Adipose tissue extracellular matrix: Newly organized by adipocytes during differentiation. Differentiation 63, 193–200 (1998).
Bukowska, J. et al. Adipose-derived stromal/stem cells from large animal models: From basic to applied science. Stem Cell Rev. 17, 719–738 (2021).
Liu, S. Enzymes (ed. Liu, S.) 297-373.
Burja, B. et al. An optimized tissue dissociation protocol for single-cell RNA sequencing analysis of fresh and cultured human skin biopsies. Front. Cell. Dev. Biol. 10, 1–17 (2022).
Skog, M. et al. The effect of enzymatic digestion on cultured epithelial autografts. Cell Transplant. 28, 638–644 (2019).
Seaman, S. A., Tannan, S. T., Cao, Y., Peirce, S. M. & Lin, K. Y. Differential effects of processing time and duration of collagenase digestion on human and murine fat grafts. Plast. Reconstr. Surg. 136, 1–11 (2015).
Lara, E. et al. Endometritis and in vitro PGE2 challenge modify properties of cattle endometrial mesenchymal stem cells and their transcriptomic profile. Stem Cells Int. 2017, 1–16 (2017).
Campos, L. L. et al. Isolation, culture, characterization and cryopreservation of stem cells derived from amniotic mesenchymal layer and umbilical cord tissue of bovine fetuses. Pesq. Vet. Bras. 37, 278–286 (2017).
Yang, J. et al. Isolation and biological characterization of tendon-derived stem cells from fetal bovine. Vitr. Cell. Dev. Biol. 52, 846–856 (2016).
Liu, J. et al. Senescence in mesenchymal stem cells: functional alterations, molecular mechanisms, and rejuvenation strategies. Front. Cell. Dev. Biol. 8, 1–17 (2020).
Kassar-Duchossoy, L. et al. Pax3/Pax7 mark a novel population of primitive myogenic cells during development. Genes Dev. 19, 1426–1431 (2005).
Okamura, L. H. et al. Myogenic differentiation potential of mesenchymal stem cells derived from fetal bovine bone marrow. Anim. Biol. 29, 1–11 (2018).
Ramírez-Espinosa, J. J. et al. Bovine (Bos taurus) bone marrow mesenchymal cell differentiation to adipogenic and myogenic lineages. Cells Tissues Organs 16, 51–64 (2015).
De Schauwer, C. et al. Characterization and profiling of immunomodulatory genes of equine mesenchymal stromal cells from non-invasive sources. Stem Cell Res. Ther. 5, 1–13 (2014).
De Schauwer, C. et al. Optimization of the isolation, culture, and characterization of equine umbilical cord blood mesenchymal stromal cells. Tissue Eng. Part C. 17, 220 (2011).
Heyman, E. et al. Validation of a color deconvolution method to quantify MSC tri-lineage differentiation across species. Front. Vet. Sci. 9, 1–15 (2022).
Bustin, S. et al. A-Z of Quantitative PCR. IUL Biotechnology Series (2004), International University Line, La Jolla, California.
Van Poucke, M., Peelman, L. J. Flexible, multi-use, PCR-based nucleic acid integrity assays based on the ubiquitin C gene. BioRxiv 168195 https://doi.org/10.1101/168195 (2017).
Andersen Lindbjerg, C. et al. Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 64, 5245–5250 (2004).
Sampaio, R. V. et al. Generation of bovine (Bos indicus) and buffalo (Bubalus bubalis) adipose tissue derived stem cells: isolation, characterization, and multipotentiality. Genet. Mol. Res. 14, 53–62 (2015).
Hepsibha, P. et al. Multipotent differentiation potential of buffalo adipose tissue-derived mesenchymal stem cells. Asian J. Anim. Vet. Adv. 6, 772–788 (2011).
Ayala-Cuellar, A. P. et al. Characterization of canine adipose tissue-derived mesenchymal stem cells immortalized by SV40-T retrovirus for therapeutic use. J. Cell. Physiol. 234, 16630–16642 (2019).
Sánchez, R. et al. Canine adipose-derived mesenchymal stromal cells enhance neuro regeneration in a rat model of sciatic nerve crush injury. Cell Transplant. 28, 47–54 (2019).
Murakami, M. et al. Trophic effects and regenerative potential of mobilized mesenchymal stem cells from bone marrow and adipose tissue as alternative cell sources for pulp/dentin regeneration. Cell Transplant. 24, 1753–1765 (2015).
Bearden, R. N. et al. In-vitro characterization of canine multipotent stromal cells isolated from synovium, bone marrow, and adipose tissue: a donor-matched comparative study. Stem Cell Res. Ther. 8, 218 (2017).
Sasaki, A. et al. Canine mesenchymal stem cells from synovium have a higher chondrogenic potential than those from infrapatellar fat pad, adipose tissue, and bone marrow. PLOS ONE 13, 1–20 (2018).
Mellado-López, M. et al. Plasma rich in growth factors induces cell proliferation, migration, differentiation, and cell survival of adipose-derived stem cells. Stem Cells Int. 2017, 1–11 (2017).
Russell, K. A., Gibson, T. W. G., Chong, A., Co, C. & Koch, T. G. Canine platelet lysate is inferior to fetal bovine serum for the isolation and propagation of canine adipose tissue- and bone marrow-derived mesenchymal stromal cells. PLOS ONE 10, 1–14 (2015).
Neupane, M., Chang, C. C., Kiupel, M. & Yuzbasiyan-Gurkan, V. Isolation and characterization of canine adipose-derived mesenchymal stem cells. Tissue Eng. Part A 14, 1007–1015 (2008).
Guercio, A. et al. Production of canine mesenchymal stem cells from adipose tissue and their application in dogs with chronic osteoarthritis of the humeroradial joints. Cell. Biol. Int. 36, 189–194 (2012).
Devireddy, L. R., Myers, M., Screven, R., Liu, Z. & Boxer, L. A serum-free medium formulation efficiently supports isolation and propagation of canine adipose-derived mesenchymal stem/ stromal cells. PLOS ONE 14, 1–21 (2019).
Ahn, J. et al. Anti-tumor effect of adipose tissue derived-mesenchymal stem cells expressing interferon-β and treatment with cisplatin in a xenograft mouse model for canine melanoma. PLOS ONE 8, 1–11 (2013).
Frisbie, D. D., Kisiday, J. D., Kawcak, C. E., Werpy, N. M. & McIlwraith, C. W. Evaluation of adipose-derived stromal vascular fraction or bone marrow-derived mesenchymal stem cells for treatment of osteoarthritis. J. Orthop. Res. 27, 1675–1680 (2009).
de Oliveira, S. et al. Triiodothyronine has no enhancement effect on the osteogenic or chondrogenic differentiation of equine adipose tissue stem cells. J. Equine Vet. 86, 1–7 (2020).
Raabe, O. et al. Hydrolyzed fish collagen induced chondrogenic differentiation of equine adipose tissue-derived stromal cells. Histochem. Cell. Biol. 134, 545–554 (2010).
Adamič, N. et al. Effect of intrabronchial administration of autologous adipose-derived mesenchymal stem cells on severe equine asthma. Stem Cell Res. Ther. 13, 23 (2022).
Clarke, L. E. et al. Growth differentiation factor 6 and transforming growth factor-beta differentially mediate mesenchymal stem cell differentiation, composition, and micromechanical properties of nucleus pulposus constructs. Arthritis Res. Ther. 16, 1–13 (2014).
Alstrup, T., Eijken, M., Bohn, A. B., MØller, B., & Damsgaard, T. E. Isolation of adipose tissue-derived stem cells: enzymatic digestion in combination with mechanical distortion to increase adipose tissue-derived stem cell yield from human aspirated fat. Curr. Protoc. Stem Cell Biol. 48 https://doi.org/10.1002/cpsc.68 (2019).
Cao, Y. et al. Human adipose tissue-derived stem cells differentiate into endothelial cells in vitro and improve postnatal neovascularization in vivo. Biochem. Biophys. Res. Commun. 332, 370–379 (2005).
Vahedi, P. et al. Advantages of sheep infrapatellar fat pad adipose tissue derived stem cells in tissue engineering. Adv. Pharm. Bull. 6, 105–110 (2016).
Godoy, R. F., Alves, A. L. G., Gibson, A. J., Lima, E. M. M. & Goodship, A. E. Do progenitor cells from different tissue have the same phenotype? Res. Vet. Sci. 96, 454–459 (2014).
Heidari, B. et al. Comparison of proliferative and multilineage differentiation potential of sheep mesenchymal stem cells derived from bone marrow, liver, and adipose tissue. Avicenna J. Med. Biotechnol. 5, 104–117 (2013).
Acknowledgements
This study was funded by the CUSTOMEAT project (FWO-SBO project S002821N). Additionally, we gratefully acknowledge Joachim Christiaens and Delphine Ameye (Department of Pathobiology, Pharmacology and Zoological Medicine, Faculty of Veterinary Medicine, Ghent University) for their excellent technical assistance.
Author information
Authors and Affiliations
Contributions
E.H. was involved in conception and design, execution of experiments, data analysis and manuscript writing. E.D.V. was involved in data analysis and manuscript editing. K.G. performed the statistical analysis. M.V.P. and L.P. were involved in RT-qPCR experiments and manuscript writing. C.D.S. and B.D. were involved in conception and design, data analysis and manuscript editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Heyman, E., Devriendt, B., De Vlieghere, E. et al. Evaluation of enzymatic protocols to optimize efficiency of bovine adipose tissue-derived mesenchymal stromal cell isolation. npj Sci Food 8, 70 (2024). https://doi.org/10.1038/s41538-024-00313-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41538-024-00313-7
This article is cited by
-
Establishment and characterization of a continuous adipose-derived stem cell line from Epinephelus fuscoguttatus and its application in cell-cultured fish meat
Marine Life Science & Technology (2026)
-
Establishing protocols for the efficient expansion of canine and feline adipose tissue-derived mesenchymal stromal cells following cell isolation
BMC Veterinary Research (2025)
-
Donor age and breed determine mesenchymal stromal cell characteristics
Stem Cell Research & Therapy (2025)
-
Pooled CRISPR screens identifies key regulators of bovine stem cell expansion for cultured meat
Communications Biology (2025)
-
Development of a sustainable strategy for cultured fat production based on serum-free 3D culture of bovine adipose stem cells
Scientific Reports (2025)