Abstract
The in-situ, rapid and non-destructive identification of complex food samples is a long-standing core technical challenge in food chemistry. Herein, vapor assisted desorption chemical ionization mass spectrometry (VADCI-MS) device was independently constructed, which facilitates rapid and non-destructive analysis of complex foods with diverse morphologies. Additionally, an integrated strategy that combines MS fingerprinting, feature selection, and machine learning classification models was established and is particularly effective for complex foods that are challenging to differentiate using conventional methods. As a paradigm, VADCI-MS was employed to rapidly (<1.0 min/sample) and non-destructively obtain the MS fingerprints of 101 batches of Zanthoxylum bungeanum (ZB) samples and 415 feature peaks were selected. Further, using machine learning classification models, a high accuracy differentiation of ZB from different origins was achieved (accuracy rate = 96.88%) and was verified to be substantially improved over traditional GC-MS. Conclusively, VADCI-MS coupled with machine learning holds significant potential for accurate and non-destructive identification of foods.
Similar content being viewed by others
Introduction
A significant challenge in food chemistry is the non-destructive and precise differentiation and authentication of food from various origins. This task is vital not only for aiding consumers in selecting products from their preferred regions but also for assisting regulatory agencies in tracing product origins1. However, it may be challenging to distinguish real geographical sources based on morphological characteristics such as appearance2. This difficulty allows some unscrupulous merchants to deceitfully advertise their product’s variety, geographical origin, and production method in order to maximize profits. Traditional methods struggle to quickly and accurately identify such intentional frauds, thereby posing significant challenges to market regulation and quality control (QC)3,4. Consequently, a pressing issue in the field of food chemistry is the development of a rapid, accurate, and non-destructive analytical technique to achieve quick and precise authentication of the same type of food from different sources5,6.
At present, a variety of detection technologies and methods are utilized in food identification7. The most commonly used non-destructive analysis technologies include spectroscopy and electronic sensor technology8,9. Other frequently employed techniques encompass nuclear magnetic resonance spectroscopy (NMR)10,11, chromatography12, isotope ratio mass spectrometry (IRMS)13, and molecular biology techniques14. Each technology has its own distinct advantages and disadvantages15. While Fourier-transform infrared spectroscopy (FT-IR), hyperspectral imaging technology, and NMR offer rapid detection speed, non-destructive analysis, cost-effectiveness, and environmental benefits16,17,18, they fall short in terms of sensitivity1,19. Recombinant DNA and PCR amplification technologies20 often fail to detect small molecule compounds. Chromatographic techniques, especially chromatography-mass spectrometry, have high accuracy in quality evaluation and discrimination based on the analysis of secondary metabolites in food. However, the indispensable sample pretreatment process is often complicated and time-consuming, and may lead to consumption or loss of key metabolites in samples3,21. Currently, there is no non-destructive and reliable food identification method that meets the requirements of speed, accuracy, no pretreatment, and low cost for precise origin determination to prevent market fraud and adulteration.
Ambient mass spectrometry22,23 has developed rapidly in recent years. It has the advantages of saving sample pretreatment time, enabling direct analysis of complex samples21,24 and environment-friendly25,26,27. Moreover, due to its exceptional capabilities in both qualitative and quantitative analysis, MS enables, to a certain extent, rapid in-situ detection and qualitative discrimination of various food samples when the concentration of the target compounds reaches detectable levels28,29,30. Therefore, this technology has potential applications in non-destructive, rapid mass spectrometry inspection of complex food samples. Machine learning, one of the most prevalent AI technologies today, is increasingly becoming a crucial tool in food research due to its robust feature recognition and extraction capabilities, coupled with its self-learning ability31,32. With the global popularity of Asian cuisine, Zanthoxylum bungeanum Maxim. (ZB) has become an important international condiment with a substantial trade volume. However, its flavor profiles and quality exhibit significant variations across different geographical origins33, which are difficult to distinguish by appearance alone. Coupled with its high economic value, this poses a high risk of origin adulteration and geographical indication fraud2. Therefore, ZB serves as a representative model for studying geographical traceability.
This study initially constructed a vapor-assisted desorption chemical ionization mass spectrometry (VADCI-MS) device to directly acquire mass spectrometric profiles of complex morphology food samples. Compared with the common nitrogen-assisted discharge needle ionization, VADCI technology introduces polar water vapor. The ionization efficiency can be improved by increasing the humidity of the ionization environment34, and its polar vapor may also improve the desorption ability of medium and even high polar compounds with a milder method, thus expanding the detection coverage. The research also aimed to integrate VADCI-MS with machine learning to develop a non-destructive, precise, and rapid inspection strategy for easily confused foods. The study used seven different regions of ZB as a demonstration object. The VADCI-MS allowed for comprehensive characterization of a single sample within 1.0 min. The analysis of comprehensive mass spectrometric fingerprints of 101 batches of ZB, combined with chemometrics and machine learning methods, successfully differentiated and identified the seven different origins. In conclusion, this study innovatively established a comprehensive strategy based on VADCI-MS coupled with machine learning, achieving non-destructive and precise rapid inspection of complex morphological foods. This approach has broad applicability in food production, regulation, and consumption, contributing to the advancement and upgrading of the food industry.
Results
GC-MS analysis
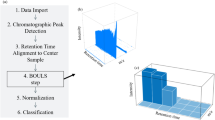
Given the high sensitivity and specificity of MS-based metabolomics, it has become the most widely used method for food identification. This technique is particularly suitable for analyzing ZB, a widely consumed food and condiment known for its unique aroma and abundance of volatile secondary metabolites. The overall workflow of this study is illustrated in Fig. 1. The conventional method for identifying foods with special odors, such as ZB, involves gas chromatography or gas chromatography-mass spectrometry (GC-MS) combined with chemometrics35,36. Thus, the present study firstly employed GC-MS to qualify 101 batches of ZB from 7 origins (the spectra are shown in Fig. S1), and the data were processed using GC Deconvolution 1.5, resulting in a dataset of 148 aligned peaks for multivariate analysis. The results of principal component analysis (PCA), partial least squares discriminant analysis (PLS-DA), and a 200 times permutation test were shown in Fig. 2A–C, respectively. As shown in Fig. 2A, the QC samples clustered closely, indicating that the machine was stable and the data were reliable. However, it can be seen from the figure that the samples from different origins were randomly distributed, which were difficult to distinguish. In addition, R2X = 0.401 and Q2 = 0.12, indicating that the prediction effect of the model is not good and the ability of effective prediction on new data is limited. Moreover, there was 1 sample falling out of the 95% confidence interval. However, in PCA, the samples of SN and GS still showed a tendency to distinguish from other origins.
Schematic diagram of the combination of VADCI-MS and GC-MS, respectively, with multivariate statistical analysis and machine learning for distinguishing the geographical origin of ZB.
A PCA score plot of GC-MS. B PLS-DA score plot of GC-MS. C Permutation test plot of GC-MS.
Given that PCA is an unsupervised technique that often overlooks intragroup variations and random errors when identifying variance components, the supervised pattern recognition method PLS-DA was employed to further distinguish the data. Figure 2B are the results of PLS-DA of ZB samples from different geographical origins with the optimized R2X, R2Y and Q2 of 0.547, 0.447 and 0.277, respectively (Table S1). The results are significantly better than the PCA results. In order to further verify the reliability of this model, a permutation test was conducted 200 times (Fig. 2C), and it can be observed that the model is not overfitting. It can be seen from Fig. 2B that the samples from SN and GS have a certain tendency to distinguish from other producing areas (The m/z of VIP value > 1 in SN and GS can be referred to Table S2.), but the samples produced in HB, SX, SC, YN and SD are difficult to distinguish. In addition, an attempt was made to further utilize machine learning methods to distinguish different origin data obtained by GC-MS (Fig. S2), and it can be found that except for RF (the accuracy rate is 90%), the accuracy rates of the remaining six algorithms are all below 80% in the test set, and through the evaluation of the model (Table S3). To sum up, when using traditional GC-MS combined with chemometric analysis or machine learning classification models, it is difficult to quickly distinguish the different sources of ZB collected in this study. It should be noted that this study did not perform in-depth compound identification, as the primary focus was on the practical application of origin certification. For the accurate identification of ZB components, systematic phytochemical separation followed by high-resolution mass spectrometric characterization is recommended.
Development and optimization of VADCI-MS device
In this study, a novel VADCI-MS device (Fig. 3) was further developed in this research, which can obtain sample spectrum information quickly and without damage. The VADCI device is mainly composed of the following parts: first, the flow rate of nitrogen is controlled by a precision gas flow valve (2.5–40 mL/min). The nitrogen flows through a quartz glass humidifying bottle containing 60 mL of deionized water to ensure that the entire closed gas circuit maintains the desired humidity. The humidified gas enters the sample warehouse through an L-shaped quartz glass tube; the sample is placed at 5.00 mg, and the end of the inlet is inserted into the sample, and the port is 1.5cm away from the bottom of the warehouse. Finally, the volatile components desorbed from the sample in the gas path are purged out by the carrier gas and introduced into the mass spectrometer through the sampling needle for analysis. Compared to traditional GC-MS, VADCI-MS has the following advantages: Firstly, it does not require any pretreatment or separation, greatly saving time and financial costs while causing no damage to the sample, preventing potential changes or loss of key substances that might lead to false positives; Secondly, the collection time is fast. Compared to GC-MS (~20–30 min), VADCI-MS only requires less than 1.0 min to complete the sample collection, and it also has the advantages of simple operation and high-throughput sampling. Thirdly, it does not require any organic solvents, making it environmentally friendly and pollution-free, especially suitable for on-site rapid inspection.
(1) capillary voltage; (2) the outlet of the sample tube; (3) the inlet of the MS.
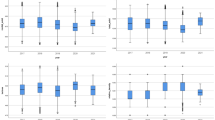
Several key parameters of the device, including the flow rate of nitrogen, capillary voltage, distance between the outlet of sample tube and the inlet of MS, and distance between the discharge needle and the inlet of MS were optimized according to the signal intensity of key components, leading to more effective ionization and detection of samples (Fig. 4). Relying on a closed circuit, through the nitrogen gas flow, the gas can be smoothly pushed into the MS. As shown in Fig. 4A, the responses at 2.5 and 5 mL/min are the lowest (2.67E5 and 4.14E5, respectively), indicating that under this gas flow, the sample can hardly enter the MS. Starting from 10 mL/min, the signal strength has significantly increased, and reached the highest response value (3.14E6) at 25 mL/min. However, it began to decline at 30 mL/min, and reduced to a level similar to that at 5 mL/min (3.66E5) at 35 mL/min. This might be due to the excessively high flow rate of nitrogen, which shortened the time for the sample to enter the mass spectrometer, leading to incomplete or non-ionization of the sample. As illustrated in Fig. 4B, when the capillary voltage is within the range of 3.5–4.5 KV, the sample exhibits a relatively high degree of ionization (3.04E6–3.71E6). However, the optimal ionization effect for the sample is achieved at 4.5KV (3.71E6). Finally, the distance between the sample tube outlet and the MS inlet and the distance between the discharge needle and the MS inlet are also very important parameters. As shown in Fig. 4C~D, the optimal distances are 6 mm and 3 mm, respectively. Especially, the position of the discharge needle is very important. If it is too close to the mass spectrometer (1 mm), it greatly shortens the ionization time of the sample, resulting in a poor response of the sample in MS. The results show that the optimized conditions are a nitrogen flow rate of 25 mL/min, a capillary voltage of 4.5 KV, and the distances between the sample tube outlet and the discharge needle with the MS inlet are 6mm and 3mm, respectively. All other MS parameters in the experiment use the default values of the instrument and do not need further manual optimization.
A N2 flow velocity. B Capillary voltage. C Distance between the outlet of the sample tube and the inlet of MS. D Distance between the discharge needle and the inlet of MS.
ZB from different geographical sources was directly characterized by VADCI-MS
Using the optimized MS conditions, a comprehensive MS characterization was conducted of 101 batches of ZB samples from 7 production areas, with a mass spectrometry information collection time of 1.0 min for each sample. The typical MS fingerprint spectrum of each production area obtained based on VADCI-MS is shown in Fig. 5. Overall, in positive ion mode, although the ZB fingerprints of 7 different origins were basically similar with multiple common peaks (such as m/z153, m/z155, m/z195, m/z213, m/z273, m/z293, m/z307 and m/z351), there were still obvious differences in its response intensity. For instance, m/z155 was the peak with the highest response intensity in ZB from all sources except for the SD production area (in the SD production area, m/z153 had the highest response intensity of 1.19E7), reaching its highest response intensity of 1.15E7 in the GS production area, compared to only 5.98E6 in the SD production area, a difference of ~1.9 times. Notably, m/z351 (response values of 4.60E5 and 4.37E5, respectively) could be clearly observed in the GS and SC production areas, but was almost intuitively undetectable in the remaining production areas. Conversely, m/z307 (response values of 5.83E4 and 4.47E4, respectively), which was difficult to directly observe in the GS and SC production areas, could be distinctly observed in the rest of the production areas, with all response values above E5. A similar situation also occurred for m/z293 in the HB, SX and SN production areas, whose response value was the highest in the SN production area (6.01E5), although this peak was not directly observed in the YN production area, its response also reached 1.09E5, while the rest of the production areas had responses around 4 orders of magnitude. In addition, in areas with relatively low signal intensity, differences in compounds can be seen more obviously, such as m/z195 in the SN production area, with a relative intensity equivalent to 10–15% of the highest peak, while this peak is hard to see in the fingerprint of the SD production area. To better interpret the fingerprints, the secondary fragmentation information of some compounds was further obtained through MS2 mode (Fig. S3) and further determined the possible structure of some compounds by literature research and online database (Table S4).
A GS; B HB; C SC; D SD; E YN; F SN; G SX. (RA: relative abundance).
VADCI-MS combined with multivariate statistical analysis, can intuitively and accurately distinguish ZB from different sources
The number of initial chromatographic peaks obtained in this study was 450. After filtering according to the screening criteria established in part 4.4, 415 peaks were retained. Subsequently, in order to better and more intuitively demonstrate the superiority of VADCI-MS in distinguishing between different producing areas compared with GC-MS, chemometric multivariate statistical analysis was used to visualize the results (Fig. 6). Figure 6A is the PCA score plot of ZB from seven producing areas, with R2X = 0.998 and Q2 = 0.989, indicating that the model has stable classification effects and high prediction efficiency. Moreover, compared to GC-MS, the samples from each producing area show a clear separation trend. In order to better promote the distinction between samples from different producing areas, PLS-DA was further utilized to differentiate the data (Fig. 6B). After optimization, the values of R2X, R2Y and Q2 are 0.996, 0.866 and 0.798 respectively (Table S5), both R2Y and Q2 values exceed 0.5, and no sample shows abnormal deviation within the 95% confidence space, indicating the reliability of the PLS-DA model, strong explanatory power, and high data processing efficiency. In addition, to verify the effectiveness of the model, 200 permutation tests were also conducted (Fig. 6C), indicating that the model has not been overfit and has good predictive ability. On the basis that the PLS-DA model can distinguish different producing areas of ZB, we performed VIP screening (Fig. S4) to demonstrate the efficacy of VADCI-MS in detecting differences in ZB metabolites from different producing areas. The characteristics that met the standard of VIP > 1.5, p < 0.05 were considered to be relevant features for distinguishing between groups (Table S6). As a result, a total of 38 ZB metabolites with statistical significance were screened. It can be clearly seen from Table S6. that the peaks mentioned in the previous context, which can be clearly distinguished from the graph, all have high VIP values, among which the value of m/z155 is the highest (5.2651), indicating that it may be the key metabolite for distinguishing different producing areas of ZB. In summary, based on the MS fingerprints obtained from VADCI-MS combined with chemometric models, the differentiation and identification of ZB sources with high similarity can be basically achieved.
A PCA score plot of VADCI-MS. B PLS-DA score plot of VADCI-MS. C Permutation test plot of VADCI-MS.
VADCI-MS combined with machine learning, accurately identifies different production areas of ZB
Based on the results above, PLS-DA was used to conduct a preliminary classification. In order to make a better distinction of different producing areas, a variety of machine learning models were employed for further analysis, including Random Forest (RF), Long Short-Term Memory (LSTM), Gated Recurrent Unit (GRU), Bidirectional Long Short-Term Memory (BiLSTM), Convolutional Neural Network (CNN), Artificial Neural Network (ANN) and Back Propagation (BP) (Fig. 7). Figure 7A is the ROC curve of the test set of these 7 types of classification models. It is worth noting that the AUC of the three models of GRU, BP and RF reached more than 0.95 in the test set, which showed excellent performance in distinguishing the ZB producing areas (training set data can be seen in Fig. S5). To further evaluate the performance, generalization ability, and prevent model overfitting, the model was assessed based on metrics such as Accuracy, Precision, Recall, F1_score, and Table S7 lists the detailed parameters of the seven types of models. It was found that the GRU model performs better overall in distinguishing production areas, with a test set AUC score of 0.9821 and a classification accuracy of 96.88%. Figure 7B represents its test set confusion matrix, showing that only one GS sample in the test set was incorrectly classified as SD, and the rest were all correctly classified. The results show that combining VADCI-MS with machine learning classification models enables more accurate and faster classification.
A ROC curve results of the test set for seven machine learning models. B~H Confusion matrix results of the test set for the seven machine learning models. (B GRU, C BP, D RF, E BiLSTM, F LSTM, G ANN, H CNN).
To make the introduced machine learning classification strategy more applicable, with the LSTM classification model serving as an example, the analysis strategy was further improved, offering a new potential optimization strategy for subsequent scholars. It can be seen from Table S7 that the model has a precision, Recall, F1 score, and AUC of 0.9524, 0.9286, 0.9238, and 0.9643 on the training set, respectively. But these parameters drop to 0.9000, 0.8571, 0.8651, and 0.9018 on the test set, respectively, which indicates that the model may have overfitted or poor generalization problems. To address such problems, two strategies are proposed to optimize the algorithm: one is RMBO-LSTM, and the other is Bayes-CNN-LSTM. The former, RMBO, is a novel meta-heuristic algorithm (intelligent optimization algorithm)37, which can be combined with LSTM to comprehensively optimize key parameters such as regularization parameters, the number of hidden layer neurons, and learning rate, aiming to achieve the best optimization model effect and prevent overfitted. The latter, on the one hand, combines the function of the Bayes optimizer to automatically adjust the hyperparameters in the LSTM model, such as learning rate, hidden layer size, batch size, etc., to find the optimal parameter combination. On the other hand, it uses CNN as an auxiliary feature extraction tool. Specifically, it inputs the feature values extracted by CNN into LSTM, helping LSTM better understand the data and make more accurate predictions. In addition, it reduces the amount of information that LSTM needs to process, thereby improving the efficiency of LSTM. As can be seen from Fig. S6, after optimization, the adaptability and accuracy of the model have significantly improved. Subsequently, by performing performance evaluations on the three models (Table S8), it can be found that the joint application of RMBO and Bayes-CNN with the LSTM model reduces the risk of overfitting of the LSTM, and improves the accuracy, precision, and other performance metrics of the model. In summary, through the application of 7 classification models, it was found that the GRU classification model is the best model in terms of prediction accuracy and reliability. At the same time, LSTM was selected as an example and proposed two new optimization model strategies, namely RMBO-LSTM and Bayes-CNN-LSTM, which can be used to optimize models with poor performance, thereby establishing a more comprehensive classification model suitable for more foods.
To sum up, although the potential of this research strategy has been verified in complex ZB samples, its extension to other foods still needs to be systematically evaluated to determine the influence of environmental factors (such as temperature, humidity) and sample types on the detection stability in future work, so as to comprehensively demonstrate its broad applicability. For example, in terms of different types of samples, the gas circulation design of the VADCI closure contributes to stabilizing the local environment around high-moisture samples. Its solvent-free extraction and analysis characteristics at normal temperature and pressure can prevent the extra oxidation of high-fat samples during processing. For surface heterogeneous samples, the original samples can be subjected to rapid in-situ detection to obtain data with overall representativeness.
Discussion
In conclusion, this study successfully developed a VADCI-MS device based on ambient MS. It overcomes the inherent disadvantages of traditional GC-MS analysis, such as complex pretreatment and slow analysis speed, and is a more efficient and environmentally friendly method for batch processing of complex samples. More significantly, VADCI-MS shows a great advantage in the number of compounds detected. This enhanced capability may be attributed to the fact that VADCI is using water vapor to impinge on the surface of the sample, which makes it not only possible to send in volatile compounds, but also possible to desorb even moderately polar or even highly polar compounds. In addition, the increased ionization environment humidity by water vapor can further improve the ionization efficiency. At the same time, this device was combined with machine learning to establish a set of non-destructive and rapid coherent strategies for identifying and distinguishing the origin of complex matrix foods. In this study, 101 batches of ZB samples from seven different production areas were first characterized by VADCI-MS to obtain fingerprint maps. Subsequently, interference values were screened out, and finally, different production areas of ZB were successfully distinguished using machine learning. The GRU classification model achieved 96.88% accuracy, 97.14% accuracy, 96.43% recall, 96.37 F1 score and 98.21% AUC on the test set. In addition, two algorithm optimization strategies were proposed to enhance the applicability of machine learning in the food origin classification prediction model. In this study, the analytical strategy based on VADCI-MS successfully achieved rapid and non-destructive identification of ZB from different geographical origins with highly similar appearances. It should be noted that the VADCI device in this study is built on a laboratory mass spectrometry platform and is not suitable for field applications such as markets or production sites. The key to subsequent research is to combine it with miniaturized and portable mass spectrometers to fully tap its potential in the field of food authentication, which is of great significance for improving the level of food QC and promoting the standardization process.
Methods
Sample collection
A total of 101 batches of ZB were collected from seven major producing provinces in China. All samples were identified as dried mature pericarps of ZB. Prior to analysis, all samples were packaged in sealed bags and stored at 4 °C in a refrigerator at the Traditional Chinese Medicine Resource Center of the Chinese Academy of Traditional Chinese Medicine, with assigned codes ZB20240001–ZB20240101. The images and geographical locations of the ZB samples are shown in Fig. S7. Table S9 shows detailed information on different ZB production areas. An MS104S analytical balance (Mettler-Toledo, Switzerland) was used for weighing. Deionized water (18.2 MΩ/cm) purified by a Mill-Q water purification system (Billerica, MA, USA) in the laboratory was used throughout the experiment.
GC-MS analysis
ZB from different geographical sources was detected according to the established method38. Specifically, the GC-MS system comprises the Thermo Scientific AI/AS 1310 liquid sampling automation, the Thermo Scientific™ TRACE™ 1310 gas chromatograph analysis system, and the Thermo Scientific™ TSQ™ 8000 Gas Chromatograph/Triple Quadrupole Mass Spectrometer (Thermo Fisher Scientific, Waltham, MA, USA). For each ZB sample, precisely measure 150.00 mg and transfer it into a 2 mL centrifuge tube. Add 500 μL of n-hexane solvent (Merck, Darmstadt, Germany) and incubate at 50 °C for 30 min. After incubation, collect the supernatant and filter it through an Acrodisc Syringe Filter measuring 13 mm with a 0.2 μm GHP membrane (Waters, USA). Transfer the filtrate into a 1.5 mL sample vial and organize them sequentially in the sample tray. In addition, to ensure the reliability of the analysis results, 10 μL was sucked in turn from the prepared samples and combined and mixed as QC samples. An autosampler was used to draw 1 μL of sample into the GC-MS system for subsequent analysis.
The chromatographic conditions were as follows: a TG-5 MS capillary column with an inner diameter of 30 m × 0.25 mm, DF of 0.25 μm was used, with He as the carrier gas (flow rate 0.8 mL/min). The injection port temperature was 190 °C, with a split ratio of 10:1. The initial column temperature was 50 °C, maintained for 3 min, then increased to 95 °C at a rate of 15 °C/min and held for 1 min; it was then raised to 110 °C at a rate of 2 °C/min and held for 1 min; subsequently, it was increased to 170 °C at a rate of 4 °C/min and held for 1 min; finally, it was raised to 200 °C at a rate of 15 °C/min and maintained for 3 min, with a solvent delay time of 3.5 min. The mass spectrometry conditions were as follows: an electron ionization (EI) source type with an energy of 70 eV, transfer line temperature of 280 °C; ion source temperature of 250 °C, mass scan range of 50–500 m/z, scan rate of 0.200 scans per second. Data analysis was performed using the specialized software Xcalibur (Thermo Scientific) and deconvolution of the GC-MS results using GC Deconvolution 1.5, as well as peak alignment.
Optimization of VADCI-MS parameters and direct analysis of ZB samples
All the experiments were executed utilizing a novel VADCI ion source, which was installed on the Thermo Scientific™ LTQ XL™ linear ion trap mass spectrometer (Thermo Fisher Scientific, Waltham, MA, USA). Specifically, the device can be divided into three parts. Firstly, a gas flow valve controls the nitrogen flow rate. Secondly, there is a humidification bottle (deionized water, ~60 mL) that allows the nitrogen to carry water vapor. This water vapor assists in the desorption and ionization of volatile components in the sample, enhancing the number and intensity of compounds detected by mass spectrometry34. The parameters of the MS are crucial in determining the signal response of target compounds. Therefore, optimization was carried out for several factors, including the capillary voltage, N2 flow velocity, and the distances between the outlet of the sample tube and the MS ion transmission tube, as well as the discharge needle and the MS ion transmission tube. The optimization ranges were set as follows: capillary voltage (2.5~5 KV), N2 flow velocity (2.5~35 mL/min), distance between the sample tube outlet and the MS ion transmission tube (4~16 mm), and distance between the discharge needle and the MS ion transmission tube (1~5 mm). Based on the results of the pre-experiment, the sample responded better in positive ion mode, so the mass spectrometer was set to operate in positive ion mode, with a mass range of m/z 50–500. To demonstrate the device’s feasibility and operational simplicity, all ZB samples used in this experiment were ZB particles (5.00 g), without any pretreatment or pre-separation. Each optimized parameter was repeatedly collected 3 times to ensure the repeatability of the method. The entire data acquisition process for ZB can be completed within 1.0 min.
Multivariate statistical analysis based on VADCI-MS fingerprint spectrum
VADCI-MS measured MS fingerprint spectra were processed using Xcalibur’s ‘subtract spectra’ command to remove blank signals, yielding pure sample spectra. The data was exported to Excel to extract m/z values and signal intensities, followed by filtering out peaks with overall response values below 1.0E2 to minimize low-response interference. Finally, PCA and PLS-DA (SIMCA 14.1) were used to develop a differentiation model for ZB from various sources.
Machine-learning model construction and evaluation
The implementation of various machine learning algorithms was carried out using MATLAB Online (https://matlab.mathworks.com/). These algorithms included RF, LSTM, GRU, BiLSTM, CNN, ANN, BP, Red-billed Blue Magpie Optimizer-LSTM (RMBO-LSTM), and Bayes-CNN-LSTM.
Before model training, the GC-MS and VADCI-MS data are first imported into Excel, and corresponding numerical labels are assigned to samples from different origins. The processed data is then imported into machine learning, where the ‘mapminmax’ function is used for normalization, unifying the scale of various features to the same range to enhance the model’s efficiency in learning relationships between different features. To further eliminate the influence of the original data order on training, the ‘randperm’ function is used to randomly shuffle all 101 ZB samples, and they are divided into training and testing sets at a ratio of 7:3. Model performance was evaluated using Accuracy, Precision, Recall, F1_score, and AUC, with formulas detailed in Table S10. Finally, confusion matrices visualized the predictive performance of each algorithm.
Data availability
All relevant data supporting this study are included within the manuscript and supplementary information. Additional raw data will be made available from the corresponding author upon reasonable request.
Code availability
The codes used to generate the results in this study are available on request from the corresponding author.
References
Xu, Y., Zhang, J. & Wang, Y. Recent trends of multi-source and non-destructive information for quality authentication of herbs and spices. Food Chem. 398, 133939 (2023).
Zhang, D. et al. Rapid determination of geographical authenticity and pungency intensity of the red Sichuan pepper (Zanthoxylum bungeanum) using differential pulse voltammetry and machine learning algorithms. Food Chem. 439, 137978 (2024).
Zhong, D. et al. Development of sequential online extraction electrospray ionization mass spectrometry for accurate authentication of highly-similar Atractylodis Macrocephalae. Food Res. Int. 175, 113681 (2024).
Gan, Y., et al. Using HS-GC-MS and flash GC e-nose in combination with chemometric analysis and machine learning algorithms to identify the varieties, geographical origins and production modes of Atractylodes lancea. Ind. Crops Prod. 209, 117955 (2024).
Modupalli, N.et al. Emerging non-destructive methods for quality and safety monitoring of spices. Trends Food Sci. Technol. 108, 133–147 (2021).
Valletta, M. et al. Mass spectrometry-based protein and peptide profiling for food frauds, traceability and authenticity assessment. Food Chem. 365, 130456 (2021).
Liu, J. et al. Application and prospect of metabolomics-related technologies in food inspection. Food Res. Int. 171, 113071 (2023).
Chen, Q. et al. Recent advances in emerging imaging techniques for non-destructive detection of food quality and safety. Trends Anal. Chem. 52, 261–274 (2013).
Kiani, S., Minaei, S. & Ghasemi-Varnamkhasti, M. Integration of computer vision and electronic nose as non-destructive systems for saffron adulteration detection. Comput. Electron. Agric. 141, 46–53 (2017).
Hatzakis, E. Nuclear magnetic resonance (NMR) spectroscopy in food science: a comprehensive review. Compr. Rev. Food Sci. Food Saf. 18, 189–220 (2019).
Schmitt, C. et al. Detection of peanut adulteration in food samples by nuclear magnetic resonance spectroscopy. J. Agric. Food Chem. 68(49), 14364–14373 (2020).
Campmajó, G. et al. Assessment of paprika geographical origin fraud by high-performance liquid chromatography with fluorescence detection (HPLC-FLD) fingerprinting. Food Chem. 352, 129397 (2021).
Giannioti, Z. et al. Isotope ratio mass spectrometry (IRMS) methods for distinguishing organic from conventional food products: a review. TrAC, Trends Anal. Chem. 170, 117476 (2024).
El Sheikha, A. F. & Montet, D. How to determine the geographical origin of seafood?. Crit. Rev. Food Sci. Nutr. 56, 306–317 (2016).
Yang, H. et al. Advanced analytical techniques for authenticity identification and quality evaluation in essential oils: a review. Food Chem. 451, 139340 (2024).
Wang, J. et al. Adulteration detection of multi-species vegetable oils in camellia oil using Raman spectroscopy: Comparison of chemometrics and deep learning methods. Food Chem. 141314. https://doi.org/10.1016/j.foodchem.2024.141314 (2024).
Damto, T., Zewdu, A. & Birhanu, T. Application of Fourier transform infrared (FT-IR) spectroscopy and multivariate analysis for detection of adulteration in honey markets in Ethiopia. Curr. Res. Food Sci. 7, 100565 (2023).
Özdoğan, G., Lin, X. & Sun, D.-W. Rapid and noninvasive sensory analyses of food products by hyperspectral imaging: Recent application developments. Trends Food Sci. Technol. 111, 151–165 (2021).
Angeli, L. et al. 1H NMR spectroscopy combined with chemometrics for detection of turmeric adulteration in Italian saffron (Crocus sativus L.). Food Control 179, 111560 (2026).
Barbosa, A. J., Sampaio, I. & Santos, S. Re-visiting the occurrence of mislabeling in frozen “pescada-branca” (Cynoscion leiarchus and Plagioscion squamosissimus – Sciaenidae) sold in Brazil using DNA barcoding and octaplex PCR assay. Food Res. Int. 143, 110308 (2021).
Qiu, Z. et al. Sensitive quantitation of ultra-trace toxic aconitines in complex matrices by perfusion nano-electrospray ionization mass spectrometry combined with gas-liquid microextraction. Talanta 269, 125402 (2024).
Takáts, Z. et al. Mass spectrometry sampling under ambient conditions with desorption electrospray ionization. Science 306, 471–473 (2004).
Cooks, R. G. et al. Ambient mass spectrometry. Science 311, 1566–1570 (2006).
Qiu, Z. et al. Rapid authentication of different herbal medicines by heating online extraction electrospray ionization mass spectrometry. J. Pharm. Anal. 13, 296–304 (2023).
Molina-Díaz, A. et al. Ambient mass spectrometry from the point of view of green analytical chemistry. Curr. Opin. Green. Sustain. Chem. 19, 50–60 (2019).
Guo, Y. et al. Integrating ambient ionization with miniature mass spectrometry to advance green analytical chemistry: an overview. Green. Anal. Chem. 13, 100257 (2025).
Zou, Y., Tang, W. & Li, B. Mass spectrometry in the age of green analytical chemistry. Green. Chem. 26, 4975–4986 (2024).
Qiu, Z.-D. et al. Real-time toxicity prediction of Aconitum stewing system using extractive electrospray ionization mass spectrometry. Acta Pharm. Sin. B 10, 903–912 (2020).
Qiu, Z.-D. et al. Limitation standard of toxic aconitines in Aconitum proprietary Chinese medicines using on-line extraction electrospray ionization mass spectrometry. Acta Pharm. Sin. B 10, 1511–1520 (2020).
WeiC. et al. Rapid toxicity characterization of Aconitum herbal medicines using perfusion nano-electrospray ionization mass spectrometry. J. Pharm. Anal. 14, 101016 (2024).
Li, X. et al. Application of novel hybrid deep leaning model for cleaner production in a paper industrial wastewater treatment system. J. Clean. Prod. 294, 126343 (2021).
Veloso de Melo, V. & Banzhaf, W. Automatic feature engineering for regression models with machine learning: An evolutionary computation and statistics hybrid. Inf. Sci. 430-431, 287–313 (2018).
Ke, J. et al. Application of HPLC fingerprint based on acid amide components in Chinese prickly ash (Zanthoxylum). Ind. Crops Prod. 119, 267–276 (2018).
Chen, H. et al. Surface desorption atmospheric pressure chemical ionization mass spectrometry for direct ambient sample analysis without toxic chemical contamination. J. Mass Spectrom. 42, 1045–1056 (2007).
Feng, X. et al. Discrimination and characterization of the volatile organic compounds in eight kinds of huajiao with geographical indication of China using electronic nose, HS-GC-IMS and HS-SPME-GC–MS. Food Chem. 375, 131671 (2022).
Shao, Y. et al. Comparison and discrimination of the terpenoids in 48 species of huajiao according to variety and geographical origin by E-nose coupled with HS-SPME-GC-MS. Food Res. Int. 167, 112629 (2023).
Fu, S. et al. Red-billed blue magpie optimizer: a novel metaheuristic algorithm for 2D/3D UAV path planning and engineering design problems. Artif. Intell. Rev. 57, 134 (2024).
Yang, Y. et al. GC-MS analysis of volatile components in Zanthoxylum armatum by double solvent micro cell headspace extraction. Food Ferment. Ind. 46, 214–219 (2020).
Acknowledgements
This work was supported by the Beijing Natural Science Foundation (No. 7252249), the Fundamental Research Funds for the Central Public Welfare Research Institutes (No. ZZ14-YQ-047), the Scientific and technological innovation project of China Academy of Chinese Medical Sciences (No. CI2023E002, CI2023C039YGL) and CACMS Innovation Fund (No. CI2021A04118, CI2021B014).
Author information
Authors and Affiliations
Contributions
Y.W.: writing—original draft, data curation, formal analysis, investigation, methodology, visualization. S.G.: formal analysis, software. J.W.: writing—original draft, data curation, methodology. X.L.: data curation, methodology. Y.Z.: data curation. L.Z.: formal analysis. Y.H.: investigation. L.K.: writing—review & editing. Z.Q.: Writing—review & editing, supervision, conceptualization. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, Y., Gao, S., Wu, J. et al. Non-destructive food authentication by vapor-assisted desorption chemical ionization mass spectrometry paired with machine learning. npj Sci Food 9, 271 (2025). https://doi.org/10.1038/s41538-025-00650-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41538-025-00650-1