Abstract
This study explored how heat treatment (20–80 °C) and surfactants affect the structure and interfacial properties of goat milk fat globule membrane (MFGM) proteins. Heating to 80 °C increased protein particle size but reduced colloidal stability, inducing a molten globule-like state with altered secondary structure. Small-angle X-ray scattering (SAXS) revealed an increase in overall protein size and compressed colloidal calcium phosphate-casein clusters. Ionic surfactants (sodium dodecyl sulfate (SDS), dodecyl trimethylammonium bromide (DTAB)) dissociated aggregates and formed core-shell complexes, enhancing surface activity and thermal stability. The nonionic surfactant polysorbate 20 (PS20) adsorbed onto the MFGM surface with minimal structural disruption due to steric hindrance. Moderate heating and nonionic surfactants are promising for industrial MFGM protein applications. This study provides the first SAXS-based structural insights, recommending moderate heat and nonionic surfactants for optimal industrial MFGM proteins handling.
Similar content being viewed by others
Introduction
The milk fat globule membrane (MFGM) proteins are sophisticated and functionally vital component of mammalian milk, forming a tripartite structure that stabilizes fat globules through its unique composition of proteins, phospholipids, and glycolipids1. This membrane system not only ensures the colloidal stability of fat globules in raw milk but also possesses remarkable bioactive properties, making it a high-value functional ingredient in the food and pharmaceutical industries2,3. While bovine MFGM proteins have been extensively studied4,5, there is growing scientific and commercial interest in the MFGM proteins from goat milk, which serves as an important nutritional alternative due to its superior digestibility and lower allergenic potential6,7.
The functional properties of the MFGM proteins are intrinsically linked to its structural integrity, which is highly susceptible to common dairy processing operations, particularly heat treatment8,9. Previous research has begun to characterize the behavior of caprine MFGM proteins. It is established that heat processing induces compositional changes in goat MFGM proteins, with proteomic analyses revealing selective denaturation of specific protein classes10,11. Furthermore, caprine MFGM proteins exhibits distinct thermal behavior compared to its bovine counterpart, attributed to differences in fat globule size distribution and casein micelle stability12,13. Evidence also suggests that surfactant interactions can modulate MFGM proteins interfacial properties, though mechanistic studies have primarily focused on bovine systems14,15.
Despite these advances, critical knowledge gaps remain regarding the structural and mechanistic responses of goat MFGM proteins16. First, the structural dynamics of how heat-induced modifications propagate through the hierarchical organization of proteins-from secondary structure to supramolecular assemblies-are poorly understood. Second, it is unclear whether thermal and chemical perturbations have interface-specific effects that differ from their impact on the bulk protein structure. Third, the fundamental structure-function relationships-how molecular-scale changes translate to functional alterations in emulsification and stabilization capacity-have not been established for goat MFGM. This gap is exacerbated by a lack of systematic investigation into the role of specific non-covalent interactions (e.g., hydrophobic, electrostatic) in determining stability17.
To address these gaps, the present study was designed to provide a multi-scale analysis of the impacts of heat treatment and non-covalent interactions on the structure and interfacial properties of goat MFGM proteins recovered from a commercially relevant dairy ingredient (goat milk powder), rather than to fully represent the native state of MFGM in freshly secreted milk. We specifically investigated temperature-dependent (20–80 °C) structural transitions and modulated noncovalent interactions using model surfactants: sodium dodecyl sulfate (SDS, hydrophobic/electrostatic), dodecyl trimethylammonium bromide (DTAB, hydrophobic/electrostatic), and polysorbate 20 (PS20, steric stabilization), using a suite of complementary techniques: UHPLC-MS/MS, nuclear magnetic resonance (NMR), rheology, electron microscope (EM) and small-angle x-ray scattering (SAXS) techniques has been implemented18,19.
This work provides, for the first time, a comprehensive view of the solution structure of goat MFGM proteins at the nanoscale and its evolution under thermal and chemical perturbations. The findings offer mechanistic insights into heat- and surfactant-induced reorganization and are expected to establish clear structure-function relationships, providing a scientific basis for optimizing the processing of goat milk-based products.
Results
Identification of MFGM proteins
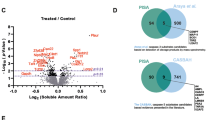
Initially, intact protein mass spectrometry was performed to evaluate the composition of the MFGM proteins using UHPLC-MS/MS. As seen from Fig. 1A, the TIC result presented the several abundances, implying the complexity of identifying peptide fragments. After acquisition and analysis of dada, a total of 462 MFGM proteins were quantitatively identified in MFGM treated at room temperature using Proteome Discoverer 2.5 based proteomic technique with coverage for peptides and proteins over 1%. Although the total protein content in whole milk is relatively low, proteomic profiling demonstrated that (MFGM) contains a remarkably diverse and abundant complement of proteins20.
A Total ion current (TIC) analysis; B RP-HPLC chromatographic spectra analysis; C SDS-PAGE analysis, Lane M: marker, Lane 1: MFGM 80 °C, Lane 2: MFGM 20 °C, Lane 3: MFGM 40 °C, Lane 4: MFGM 60 °C, Lane 5: MFGM with 1 mM SDS, Lane 6: MFGM with 1 mM PS20, Lane 7: MFGM with 1 mM DTAB.
The database retrieval results listed in Table 1 indicated that the top nine proteins with the highest Score Sequest are β-casein, β-lactoglobulin (β-LG), κ-casein, xanthine oxidase/dehydrogenase (XO/XDH), butyrophilin (BTN), αs1-casein, lactotransferrin (LTF), αs2-casein, α-lactalbumin (α-LA) and Ig-like protein. The UHPLC-MS/MS results are, therefore, in accordance with those of a recent study by Han et al.21, who reported that αs1-casein, XO/XDH and LTF played an important role in taxonomic identification in the principal component analysis of bovine and goat milk MFGM proteins.
Caseins were observed to be the highest in the relative abundance value, which is one of the main proteins present in MFGM. The interfacial protein composition of goat milk powder exhibited significant differences compared to previous studies16,22,23, in which, the major bovine MFGM proteins were reported to be BTN, adipophilin (ADPH), lactadherin and XO/XDH24. The primary MFGM proteins in this work, XO/XDH was quantified in fourth highest Score Sequest value, in line with previous studies25. As a membrane-associated protein, XO/XDH plays structural role in the wrapping and secretion of fat globules, and has antimicrobial function20. BTN was observed to be the fifth highest in the relative Score Sequest value, which has been demonstrated to regulate the encephalitogenic T-cell response to myelin oligodendrocyte glycoprotein in experimental autoimmune encephalomyelitis, a model that closely mimics the immunopathological features of human multiple sclerosis (MS)26. Furthermore, the quantification of MFGM proteins via trypsin digestion is limited by two primary factors: (1) membrane proteins exhibit a significantly lower abundance of lysine and arginine cleavage sites compared to cytosolic proteins, thereby reducing proteolytic accessibility; and (2) their inherent hydrophobicity and tendency to form aggregates hinder efficient solubilization, which is essential for complete digestion27.
The HPLC profiles (Fig. 1B) showed a good separation rate. Based on the retention time ranging 5-20 min, the 9 significant common-peaks were labeled accompany with elution time according to top ten proteins with the highest Score Sequest identified in UHPLC-MS/MS (Table 1). The observed phenomenon aligns with the data documented in the literature23. SDS-PAGE electrophoretic separation was used to confirm and to identify the proteins present in MFGM. Figure 1C displays clear and well-resolved protein bands, which were acquired through gel electrophoresis followed by UV irradiation for visualization. In general, higher relative intensities indicate a higher protein solubility8. It needs to explicitly state that peak identification is based on a correlation with SDS-PAGE, HPLC and MS/MS results, supported by literature on MFGM proteins behavior, rather than on comparison with pure standards. Along with the MFGM proteins, significant amounts of casein and whey protein (β-LG and α-LA) can be observed in sample, which was consistent with those obtained from MS/MS and HPLC.
The extremely low levels of MFGM proteins obtained in this work are attributed to the manufacturing treatments, primarily homogenization. Moreover, the MFGM proteins in this work were recovered from a commercially relevant dairy ingredient (goat milk powder), rather than to fully represent the native state of MFGM in freshly secreted milk. This processing decomposed the native MFGM structure, leading to the adsorption of casein and whey proteins from the serum onto the oil-water interface. The resulting novel interfacial layer stabilized the emulsion. Consequently, whey proteins and casein were the main interfacial protein components of MFGM proteins. Similar MFGM proteins profile was observed in infant formulas previously28.
After heat treatment (40 °C, 60 °C and 80 °C), both HPLC (Fig. 1B) and SDS-PAGE results (Fig. 1C) showed the changes of MFGM proteins compositions. The MFGM proteins and whey proteins were significantly reduced with increased temperature. On the other hand, the casein region remained after heat treatment. The heat-induced interactions between MFGM proteins and milk proteins were found by comparison of SDS-PAGE and HPLC which were similar to those of previous studies16,17.
Furthermore, the disappearance and decrease in MFGM proteins were related to heating intensity, and more MFGM proteins were significantly down-regulated at 80 °C treatment than at 40 °C and 60 °C treatment. Previous studies have also shown the decrease in some common MFGM proteins in ultra-pasteurization was greater than that of UHT, suggesting that UHT had a more significant influence on goat MFGM proteins than ultra-pasteurization in terms of the types of MFGM proteins16. Heating may decrease free sulfhydryl (SH) groups in the MFGM and increased disulfide (S-S) groups, suggesting that incorporation of β-LG might be due to association with membrane proteins via disulfide bonds29. In general, the amount of goat MFGM proteins was reduced after heat treatment.
The proteins changed differently in the presence of different surfactants conditions. Upon addition with SDS and DTAB, caseins were significantly reduced as indicated by the decreased elution peak area around 7.8 min (Fig. 1B), and the interfacial composition of MFGM proteins was impacted little. No obvious change in interfacial components when added PS20 in MFGM proteins. This implied that ionic and non-ionic surfactants impacted the MFGM proteins little and can coexist with it. These may be related to the disassociation of casein micelles initiated by SDS and DTAB and the interfacial composition of MFGM protein increased. While PS20 modified on the surface of casein micelles with little alteration of internal casein micelle’s structure, and thus no obvious change interfacial composition of MFGM proteins. This is consistent with the previous studies that charged surfactants, like SDS and DTAB, induced disassociation of casein micelles or caseinate particles, while PS20 formed an adsorbed layer on the casein micelles surface, stabilizing the air/water or oil/water interface with the minimum alteration for the internal structure of casein micelles30,31.
Surfactants (surface-active agents) are widely used in diverse fields, such as (but not limited to) food, pharmaceuticals and nanomaterials and nanotechnology32. The understanding of the formation, structures, and properties of emulsions is essential to the creation and stabilization of structures in food33. The emulsifying characteristics of MFGM proteins are closely associated with its surface composition. The stability of oil-in-water emulsions relies on the surfactants present in the system. Both proteins and polar lipids possess emulsifying properties9. Several studies have reported a synergistic effect with polar lipids when using MFGM proteins as co-emulsifiers, which may lead to different functional properties, for instance, one of the components may form the emulsion while the other improve its stability or physiological functionality2.
Analysis heat-treatment induced-change in spectra
The UV absorption spectra of proteins and amino acids in aqueous solution exhibit characteristic electronic transitions across the 185–320 nm range. The most prominent absorption bands (255–280 nm) originate from the aromatic side chains of three amino acids: tryptophan (Trp, λmax ≈ 280 nm), tyrosine (Tyr, λmax ≈ 275 nm), and phenylalanine (Phe, λmax ≈ 257 nm). The peptide backbone demonstrates strong absorption near 190 nm with a weaker secondary peak between 210 and 220 nm34. The MFGM proteins presents two main peaks, the major one locates at ~210 nm contributed by the π → π* transition in the peptide bond absorbs light35, the minor one is at ~280 nm originated from Trp and Tyr residues. With increased treating temperature (40, 60, and 80 °C), the absorption intensity enhanced at ~210 nm, while conversely, the absorption intensity decreased at ~280 nm and an iso-absorptive point was found at ~220 nm. This demonstrated that the heat treatment prompted the interactions between peptides and prompted the energy transfer of aromatic amino acids residues, leading to an increased absorbance in 210 nm and a decrease in absorption band around 280 nm.
Heat treatment is conventionally employed to enhance product safety and extend shelf-life. Denatured proteins may undergo structural unfolding, adsorb at oil-water interfaces, form aggregates, or interact competitively with emulsifiers. In dairy systems, heat treatments between 60–95 °C specifically modify MFGM interfaces, predominantly through covalent binding of whey proteins (β-LG, α-LA) to MFGM-associated proteins36. Previous research has established that thermal processing significantly reduces β-LG and α-LA concentrations37. This reduction initiates above 60 °C through disulfide bond cleavage, generating reactive free sulfhydryl (-SH) groups. At elevated temperatures (>70 °C), structural denaturation facilitates whey protein aggregation via intermolecular disulfide bridging. These protein complexes subsequently associate with casein micelle surfaces through hydrophobic interactions and covalent cross-linking38.
In the MFGM fraction, disulfide-mediated binding occurs between β-LG, α-LA and MFGM proteins (XDH/XO, BTN and LTF)17, consistent with findings in this work. As heat intensity increases (60–80 °C), β-LG, α-LA progressively associates with MFGM proteins through two established pathways16: (1) covalent conjugation via disulfide linkages between exposed free thiols in β-LG, α-LA and cysteine residues of MFGM proteins39; and (2) adsorption of casein micelles onto milk fat globules, followed by β-LG-caseins complex formation. Upon heating, the heat-induced β-Lg/α-La-casein complex became incorporated into the structure of the MFGM proteins40. On the other hand, contemporary studies further reveal that non-covalent interactions-particularly hydrophobic forces, electrostatic attractions, and hydrogen bonding-significantly contribute to β-LG-MFGM proteins associations alongside covalent binding, while disulfide bond formation is not crucial for the heat-induced interaction between β-LG and MFGM proteins40.
With addition of SDS, the decrease in absorption intensity at 210 nm and the increasement in absorbance at 280 nm were observed (Fig. 2B), suggesting that SDS changed the interfacial characteristics of MFGM proteins and reduced the interactions between peptides, which resulting in the exposure of the originally embedded aromatic amino acids, thereby improving the UV absorption ability. Upon adding DTAB, the absorption band at 210 nm and 240 nm increased, accompany with a decreased absorbance at 280 nm (Fig. 2C), indicating that peptide chain interactions and Trp residues embedding occurred simultaneously. It was found that two iso-absorptive points at 220 nm and 260 nm, suggesting that the dissociation and aggregation of MFGM proteins may coexist. Figure 2D showed that PS20 significantly decreased the absorption at 280 nm no change in the band around 210 nm, which clearly demonstrated that PS20 only formed an absorption layer on the surface of MFGM proteins, which may stabilize the air/water or oil/water interface with the minimum alteration for the internal structure of MFGM proteins.
A–D UV-Vis absorption spectra analysis. E–H fluorescence spectra analysis. I–J FT-IR, circular dichroism spectra (CD) and NMR analysis, respectively.
The fluorescence properties of MFGM proteins upon heat treatment and surfactants were further investigated. The intrinsic fluorescence of MFGM proteins is mainly presented by Trp residues with a maximum emission at 330 nm (Fig. 2E and Table 2), meaning that the Trp residues are embedded in the interior of MFGM proteins with a considerable hydrophobic environment. With increasing treating temperature, the intrinsic fluorescence intensity of MFGM proteins increased with a maximum around 331 nm, suggesting that upon increasing temperature, the hydrophobic environment of Trp residue was changed little, the indole ring lone pair electron transfer was restrained and self-quenching of Trp residues was weakened, which may enhance the fluorescence intensity of MFGM proteins.
As shown in Fig. 2F–H and Table 2, the fluorescence intensity of MFGM proteins underwent a two-step transition, upon adding 1 mM SDS, increased firstly and then decreased with a blue-shift in emission from 330 nm to 324 nm. With addition of DTAB and PS20, emission of MFGM proteins red-shifted from 330 nm to 333 nm with an increased intensity. These findings indicated that the intermolecular quenching of Trp residues of MFGM proteins, such as excited-state electron transfer, excited-state proton transfer41, NH…π interaction quenching and self-energy transfer was also prompted when bound by SDS, DTAB and PS2018,30. The hydrophobic environment of Trp residues was increased in the presence of SDS while it decreased upon addition of DTAB and PS-20.
FTIR was further utilized to characterize the heat treatment and surfactants induced change in structure of MFGM proteins. As seen from Fig. 2I, the spectral variation of the amide II band (1500–1600 cm−1) is mainly due to N–H bending and C–N stretching vibrations. The peaks at 2800–3100 cm−1 were mainly related to C−H stretching and hydrophobic interactions42. The absorption peak was generated near 1750 cm−1, which corresponded to the stretching vibration of the ester chain C=O43. With increased treatment temperature, the new absorption peaks were generated around 1600 cm−1 and 3300 cm−1, and the vibration near 2850 cm−1 was strengthened. This indicated that the hydrophobic interaction dominated the heat treatment-induced interactions of MFGM proteins. Upon adding SDS, DTAB and PS20, the vibrations around 1300 cm−1, 1600 cm−1 and 2850 cm−1 were intensified, and interestingly, the absorption peak shifted from 3300 cm−1 to 3500 cm−1, possibly due to the interaction between surfactants and MFGM proteins that exposed aliphatic amino acids to dodecyl chain of surfactant, resulting in increased hydrophobicity.
Far-UV circular dichroism (CD) was utilized to detect the change in secondary structure of MFGM proteins upon heat treatment and surfactants. As seen from Fig. 2J, MFGM proteins exhibit showed a typical CD curve with two minimum ellipticity (θmin) at 206 nm and a shoulder at 225 nm indicating a disordered structure of casein micelles, which are the main components of MFGM proteins in this work and rich in the proline-glutamine rich (P, Q-rich) sequences44. Compared to CD spectra of goat casein micelles (GCM), MFGM proteins showed a more significant ellipticity at 225 nm, due to the contribution from whey proteins, XDH/XO and BTN. The secondary structure content of MFGM proteins, helix, antiparallel, turn and unordered conformation has been calculated as 2.3%, 38.1%, 14.9% and 44.6%, respectively (Table 2), suggesting the MFGM proteins partially unordered characteristics. With increased treating temperature, the negative ellipticity at 206 nm enhanced with remarkable blue-shift, and the ellipticity at 225 nm was degraded. When treating temperature came to 80 °C, the helical content and unordered structure increased to 11.3% and 47.3%, while the antiparallel and turn content reduced to 24.8% and 16.6%.
This observed shift in secondary structure composition indicates that heat treatment induced a significant conformational rearrangement within the MFGM proteins, characterized by a relative increase in α-helix and unordered content accompanied by a reduction in antiparallel structures. This specific pattern of structural changes signifies a reorganization of the MFGM proteins into a molten globule state-an intermediate unfolding state characterized by the enhancement of secondary structure elements (particularly α-helices) while maintaining a dynamic and flexible three-dimensional conformation45. This phenomenon is not unique to MFGM proteins; it has been documented in previous reports that heat treatment can induce the formation of a molten globule state in several other proteins, such as β-LG, duck liver protein, and rabbit meat protein46. For the complex MFGM protein system, heat treatment may not induce a simple, direct unfolding pathway. Instead, it promotes a structural reorganization that leads to a molten globule-like state, in which secondary structure elements (particularly α-helices) are stabilized or become more pronounced in the CD spectrum due to the compaction of the overall structure, despite the loss of native three-dimensional structure45. It is crucial to emphasize that the apparent increase in helical content induced by heat treatment does not reflect a net gain of native-like ordering, but rather a repartitioning of structural elements within a partially folded intermediate.
The addition of SDS and DTAB decreased the negative ellipticity at 206 nm significant blue-shift. And the calculated secondary structure content given in Table 2 shows that MFGM proteins exhibit an enhancement in helix, turn and unordered conformation, but reduced antiparallel in the presence 1 mM SDS and DTAB. Despite the behavior of SDS and DTAB modified CD curve of MFGM proteins is similar to that of heat treatment, the mechanism of increasing ordered structure was different. Our previous study indicated that SDS or DTAB not only induced the dissociation of casein micelles or caseinate particles resulting a decreased turbidity reduction, but also converted β-strand into α-helix and unordered conformation18,30. The phosphorylated caseins in micelles primarily associated with colloidal calcium phosphate (CCP) to form micellar structure, and CCP failed to provided adequate hydrophobic force, which may lead to increase unordered structure of casein micelles by associating with SDS dodecyl chains18. The β-domain and unstructured region(s) were involved in the α-LA molten globule state modulated by CnTAB, and the long-chain C16TAB increased secondary structure content of α-LA47. This supports that the hydrophobic interaction of surfactant alkyl chains plays a decisive role in increasing the helical ordered structure of proteins.
In the presence of PS20, the CD curve of MFGM proteins showed little modification in secondary structure content with a gentle blue-shift of ellipticity at 206 nm (Table 2 and Fig. 2J). It is consistent with previous findings that PS20 mainly bound and formed a layer on the protein’s surface due to the steric hindrance of polysorbate, rather than interrupted the internal structure of protein18,31.
The 1H-NMR chemical shifts in proteins serve as sensitive probes for local conformational dynamics. These spectral perturbations arise from the collective contributions of: (i) backbone (φ, ψ) and sidechain (χ) torsion angles, (ii) hydrogen-bonding networks, (iii) vectorial electric fields, (iv) paramagnetic ring current effects, and (v) steric constraints in the local environment48. In this work, the variations in chemical shift distribution in four regions of stack spectra including methyl -CH3- (0.5–1.5 ppm), aliphatic -CH2- (1.5–3.5 ppm), Hα amide region (3.5–5 ppm), and aromatic/HN-amino region (5.5–9 ppm) (Fig. 2K–M). With heat treatment, the deshielding was observed in the methyl and aromatic/HN-amino regions with a downfield shift of the methyl and Hα amide region around 1.2 ppm and 3.6 ppm (Fig. 2N). As added surfactants, the reduced signal intensity can be seen clearly in the aliphatic region around 3.3–3.5 ppm. The lower peak intensity for methyl groups can be due losses of signal from the methyl and amide side chains of aliphatic amino acids, Asn and Gln. The sensitivity of ¹H chemical shifts to local electronic environments originates from their modulation by multiple structural and physical factors, including: (i) protein conformational states, (ii) hydrogen-bonding networks, (iii) magnetic anisotropy from aromatic ring currents, and (iv) local electrostatic field gradients49. In addition, the downfield shielding of the 1H chemical shift can result from changes in the different components in the secondary structure50. Due to the wide variety of MFGM proteins types, it is indeed tough to obtain accurate information on the primary structure using NMR. However, in this work, the downfield shift for the amide region was observed, which can relate to a higher presence of helix structures at higher temperature and bound by surfactant.
Analysis of interfacial behavior and thermal stability
We also monitored the sizes, zeta potential and interfacial behavior of the MFGM proteins upon heat treatment and interacted with surfactants. Figure 3A, B show a bimodal pattern distribution from 40-400 nm with an average size of 323 nm for MFGM proteins at 20 °C (p < 0.05). It has been reported that the goat casein micelles have a unimodal distribution with a mean particle size between ~180 nm to ~250 nm18,51,52. Therefore, this peak corresponds to the part of contribution from casein micelles. The average size of the MFGM proteins in raw milk ranges between 0.1 and 20 µm in size2,53, which can significantly change during processing8. The change in particle size can, for instance, indicate of MFGM disruption11 or aggregation54. The MFGM from infant formulas showed unimodal distribution pattern with a mean particle size from 360 nm to 390 nm28. Thus, the smaller size of the MFGM proteins was due to the homogenization process during manufacturing goat milk powder. To ensure the physical stability of the powder during long-term storage, this process forms an oil-in-water emulsion. Milk proteins, when combined with carbohydrates, further enhance stability by creating functional emulsifiers that also produce smaller MFGM. The fat globule particle size is significant related to the lipid digestibility and bioavailability55.
A–B size distribution. C–E zeta potential, contact angle and surface tension analysis. F differential scanning calorimetry analysis.
Additionally, a minor peak between 40 and 100 nm was obtained. Holzmüller et al. assumed this the small fragments also related to MFGM fragments56. Michalski et al. identified that appearance of a new population of small particles with a size of 40–200 nm upon mechanical treatments (homogenization, ultrasonication and shear) as a small fat globule population57. The pattern distribution of heat treated MFGM proteins was bimodal and the heat treatment at temperature exceeding 60 °C resulted in a significant increase in mean particle size, indicating the heat-induced aggregation of MFGM proteins. Ma et al. reported increased αs1-casein and β-casein levels in the MFGM fraction of both bovine and caprine milk following holder pasteurization17. This shift is primarily attributed to heat-induced casein dissociation from micelles and subsequent diffusion into the serum phase, where caseins aggregate with whey proteins via electrostatic interactions and disulfide bonding58. Heating up to 60 °C unfolds and denatures whey proteins, exposing reactive groups and facilitating these interactions59. Thus, the change of proteins on the interface of MFGM, or the attachment of casein or whey proteins on MFGM was responsible for the heat-induced aggregation.
MFGM proteins show a unimodal pattern distribution ranging 100 nm to 380 nm with a mean size of ~250 nm and ~260 nm after added SDS and DTAB, respectively. This suggests that SDS and DTAB led to the dissociation of MFGM proteins and thus resulting in a reduced and homogeneous size of system. Interestingly, MFGM proteins with DTAB added has a larger particle size and distribution than that with SDS added, implying the neutralization of positively charged DTAB with oppositely charged protein may improve the size of the system. No significant change of 1 mM PS20 in the size distribution of MFGM proteins was observed, demonstrating that noncovalent bonds derived from PS20 may be insufficient to disrupt the interactions between peptides to lead dissociation of MFGM proteins.
The stability of MFGM proteins has been associated with zeta potential value, as illustrated in Fig. 3C, all the samples showed a negative zeta potential (p < 0.05). This is because the initial solution had a neutral pH (pH = 7.0), which is higher than the isoelectric points of whey protein (pI = 5.1) and casein (average pI = 4.6), the major components of MFGM proteins in this work, leading to a greater dissociation of acidic groups in the proteins, resulting in an overall negative charge60. The zeta potential of MFGM proteins with pH 7.0, when measured at 20 °C, was −37.70 mV, which was much higher than the previous studies showed that the zeta potential value of whole milk at physiological pH (6.60–7.0) ranged from −13.7 mV to −22 mV61,62. Sun et al. investigated the changes in Milk fat globules and MFGM proteins after cream separation from different pH bovine raw milk, in which, zeta potential value for raw milk between pH 6.74 and 5.30 at 25 °C is −33.60 mV63, which are quite close to our results.
The increasing treated temperature increased the zeta potential of MFGM proteins to −26.73 mV at 80 °C. This might be a result of a modification to the MFGM proteins at molecular level upon heat treatment, which leads to an reduction in the exposure of negatively charged and/or hydrophobic segments64. The denatured whey proteins interact with κ-casein through hydrophobic bonding and disulfide exchanges to form the whey protein/κ-casein complexes whereby change in the ionic strength of the continuous phase increases65. This decreases the thickness of the electrical double layers and their potential for overlapping, and thus resulting in the reduction in zeta potential of MFGM proteins62.
The zeta potential of MFGM proteins is closely related to the type of surfactant added, meaning that anionic surfactants (SDS) increase the zeta potential of MFGM proteins, while cationic surfactants (DTAB) decrease it. The zeta potential of MFGM proteins was found to decrease when adding non-ionic surfactant (PS20), suggesting that the modification of PS20 on the surface of MFGM proteins reduced the exposure of negatively charged segments and/or electrical double layers and downregulated the zeta potential of system.
The hydrophilic and hydrophobic groups of proteins interact with the water and oil phases, impacting interfacial protein. We further implemented the contact angle (θ) and surface tension measurement to investigate the hydrophilic and hydrophobic nature of MFGM proteins. As shown in Fig. 3D, the θ values of all samples are less than 90° (p < 0.05), indicating that they are hydrophilic property66. The MFGM proteins showed a θ value as ~25 ° at 20 °C, lower than that of pure goat casein micelles with a value of ~55° reported by Silva et al.67. This suggests that interfacial composition of MFGM proteins changed the hydrophobic property of casein micelles. The heat treatment increased the θ value of MFGM proteins, demonstrating that the heat-induced aggregation enhanced the size of particle and thus increased the θ value. While, the θ value of MFGM proteins reduced significantly upon adding surfactant, although that of value in the presence of PS20 did not decrease as much as the other two.
Figure 3E indicates the surface tension value of MFGM proteins as 50.65 mN/m (p < 0.05), which is comparable the previous results of casein micelles and MFGM materials68. Upon heat treatment and surfactants, the surface tension of MFGM protein showed a significant reduction in surface tension. However, the mechanisms by which heat treatment and surfactants increase surface activity may be different, the former being due to denatured whey proteins interact with κ-casein through hydrophobic bonding and disulfide exchanges to form the whey protein/κ-casein complexes whereby contributed to the surface activity of MFGM proteins. The latter was due to surfactant reduced surface tension and increased the surface activity.
A reduced contact angle measurement typically indicates decreased interfacial tension within the MFGM proteins-surfactant system, which aligns with Gibbs adsorption theory, which establishes that surfactant accumulation at the air-water interface directly correlates with solution surface tension reduction in equilibrium conditions69. These results suggest that the heat treatment induced aggregation of MFGM proteins, which fortify the absorption of protein at the air-water interface and decrease surface tension and contact angle of the solution. In the presence of surfactant, a modification by surrounded surfactant molecules and complex of surfactant-MFGM proteins absorbed at air–water interface leads to a reduction in the surface tension and contact angle of a solution. The difference in interfacial behavior of heat treated MFGM proteins from protein bound by surfactants was related to the aggregation and dissociation of MFGM protein induced by heat and surfactants. A similar result was reported by Chen et al., who demonstrated that MFGM-stabilized interfaces seem to have a complex heterogeneous structure consisting of MFGM proteins particles, free phospholipids and proteins. The phospholipids can adsorb reversibly at the interface, the MFGM particles and proteins typically have a much higher desorption energy68.
Differential scanning calorimetry (DSC) thermogram of MFGM proteins upon heat treatment and surfactants are presented in Fig. 4B and corresponding parameters, such as denaturation temperature and enthalpy change are listed in Table 2. The MFGM protein shows an endothermic peak around 94.04 °C with a height of 0.516 mW/mg and −124 mJ/mg of ΔH. The denaturation temperature of heat treated MFGM protein (40, 60, and 80 °C) increased to 105.65, 108.92, and 111.81 °C with a value of ΔH as −136.1, −138.6 and −159.8 mJ/mg, respectively. This clearly demonstrates that the energy required to unfold the protein molecules increased, signifying enhanced thermal stability at the molecular level, which could be due to the interactions among proteins requiring higher temperature to degrade its molecular structure and thus resulted in a higher denatured temperature and enthalpy change, as reported previously70. It has been reported that heat treatment results in casein micellar depolymerization, which can significantly decrease the heat stability of the milk71. We obtained the opposite results, which was related to the structural rearrangement induced by heat treatment as a transition towards a more ordered molten globule-like state. In this partially folded intermediate, the protein backbone gained conformational rigidity, thereby increasing its resistance to heat denaturation while maintaining a compact overall structure.
A–G Heat treatment (20, 40, 60 and 80 ℃), SDS, DTAB and PS-20 on the colloidal stability of MFGM proteins. H–J The corresponding TSI and apparent viscosity analysis.
The addition of surfactants significantly decreased height of MFGM proteins thermogram, meaning that the dissociation of protein induced by surfactants decreased the endothermic peak intensity. Interestingly, MFGM proteins bound by SDS, DTAB and PS20 presented a higher denaturation temperature of 107.47, 99.43 and 94.37 °C. This suggests that the incorporation of surfactants significantly enhanced the thermal stability of MFGM proteins. Specifically, the interaction of SDS or DTAB with the proteins led to the formation of a more compact and ordered structure, thereby raising the denaturation temperature. Although SDS and DTAB initially caused dissociation of protein aggregates, the resultant complexes maintained thermal stability. This phenomenon can be attributed to the surfactant-induced increase in α-helix content, as evidenced by CD results. In contrast, the PS20 primarily adsorbed onto the surface of MFGM proteins, forming a stabilizing layer with minimal perturbation to the internal structure, consequently exerting negligible impact on the protein’s intrinsic thermal stability.
Analysis of colloidal stability and apparent viscosity
Turbiscan Lab stability analyzer uses backscattering measurements to analyze colloidal stability mechanisms. The MFGM proteins samples were scanned continuously within 24 h, the variation of the transmitted light (T) and the backscattered light (BS) with measuring time as a function of the height of measuring cell could be obtained, which reflected the movement trend of the particles and then predicted the stability of MFGM proteins72. The ΔBS curves (ΔBS = BSt − BSt = 0) exhibited characteristic patterns over time, reflecting colloidal destabilization, which allows for precise quantification of the instability process73. The ΔBS intensity decreases with increasing particle size or decreasing particle concentration when particle diameter exceeds the wavelength. Consequently, a declining ΔBS in the bottom layer signals reduced particle concentration, while an increasing ΔBS in the top layer indicates elevated particle concentration74. A smaller change of ΔBS suggests a solution system with higher stability. As shown in Fig. 4A, the ΔBS profiles of MFGM proteins exhibited with stability phenomena with less change, the backscattered light intensity on the center of the scan increased slightly 1% at the middle of the sample cell (4–36 mm). This behavior is primarily attributed to Brownian motion and gravity, which drive particle movement and collisions, causing the MFGM proteins particles to aggregate. Upon heat treated (Fig. 4B–D), MFGM proteins showed the reduced stability, and ΔBS variation amplitude increased to 2% with more change compared to MFGM proteins treated at 20 °C, indicating an increase in particle concentration and thermal-induced instability. This confirmed that the thermal treatment induced aggregation of MFGM proteins. After addition of surfactant (Fig. 4E–G), ΔBS profiles of MFGM proteins presented more change in variation amplitude as 3%, suggesting that surfactant can accelerate the migration rate of particles and improve the instability of the MFGM protein. This may explain by the movement and collisions of dissociated MFGM protein by surfactants.
The physical stabilization kinetics of the emulsion can be evaluated by measuring the turbiscan stability index (TSI)75. The higher is the TSI value, the lower the stability of the colloidal system. As shown in Fig. 4H, during the 22 h testing period, the TSI for all the samples kept increasing, indicating an unstable emulsion system. Compared to other groups, the MFGM proteins at 20 °C showed highest emulsion stability, of which the TSI value even reached ~3 at the end of 22 h. The MFGM proteins treated at 40, 60, and 80 °C exhibited TSI value of ~4.5, ~5.8 and ~6.2, suggesting the heat treatment decreased the colloidal stability of MFGM proteins.
In contrast to the enhanced molecular thermal stability, the heated MFGM protein samples exhibited a significant decline in colloidal stability. This was evidenced by a notable decrease in the absolute zeta potential value, indicating a substantial reduction in the electrostatic repulsion between MFGM proteins particles. Consequently, the transmission and backscattering profiles from turbiscan analysis confirmed increased particle aggregation over time, underscoring the compromised integrity of the protein solution at the colloidal level.
The MFGM proteins with 1 mM SDS exhibited the lowest emulsion stability (TSI∼12). According to Fig. 4I, the MFGM proteins with 1 mM PS20 exhibited higher TSI ~ 4.4 before 4.4 h, after that, MFGM proteins adding SDS showed highest TSI ~ 12 at the end of 22 h, meaning the lowest emulsion stability. This was probably caused by the competitive adsorption between surfactants and MFGM proteins at the oil/water interface improving the migration particles. Despite the MFGM proteins were unstable to creaming, but no significant flocculation or coalescence occurred shortly after addition of surfactant.
Figure 4J shows the apparent viscosity curves of the MFGM proteins. All samples exhibited typical shear-thinning flow behavior, with a linear decrease in viscosity with the increased shear rate ranging from 0.1 s−1 to 1000 s−1. The observed viscosity reduction stems from a shear-induced decrease in flow resistance, caused by the disentanglement of macromolecular chains that were previously entangled76. Moreover, the viscosity of MFGM proteins treated at 20, 40, 60, and 80 °C in the initial measurement period exhibited as 0.04197, 0.06163, 0.1416, and 0.3575 showing a viscosity-dependent correlation with treated temperature. This suggests that the heat treatment induced aggregation and form a more compact and stable structure of MFGM proteins, enhancing its shear resistance and increasing the apparent viscosity. Additionally, the PS20 polyoxyethylene sorbitan structures contributed to the highest viscosity and thickening effect, while it showed faster descent rate compared to SDS or DTAB group, suggesting PS2 promoted associated MFGM proteins separate, resulting in a reduced flow resistance and viscosity with faster speed.
The colloidal stability and apparent viscosity analysis reveal the thermal treatment induced aggregation of MFGM proteins and a more compact structure but less stability of colloidal. Surfactant leads to dissociation of MFGM proteins and promotes the migration and collisions, and thus resulting in a less stable colloidal system compared to native MFGM proteins. Rearrangement of MFGM proteins bound by surfactant also induces a more compact and stable structure and improves its shear resistance and apparent viscosity. Therefore, thermal treatment and surfactant could be an option for improving the mechanical stress and viscosity of nanomaterials based on MFGM proteins.
Analysis of solution structure using SAXS
SAXS offers a way to examine the overall shape and oligomerization state of biological macromolecules under quasi native conditions in solution77. We used SAXS to explore the solution structure, distinguish oligomeric state of MFGM proteins, define global structural parameters (gyration of radius, particle volume, etc.) and identify shapes. Further investigation of impacts of thermal treatment and surfactants.
As seen from Fig. 5A, MFGM proteins can be summarized into three structural characteristics based on the SAXS scattering q range: (1) the global size of MFGM protein reflected by the q range lower than 0.01 Å−1 78, (2) the ‘incompressible regions’ or CCP-casein nanoclusters in the casein micelles contributed the scattering of the middle-q range from 0.01 to 0.1 Å−1 79, (3) the high-q range larger than 0.1 Å−1 corresponds to the heterogenous protein (caseins, whey, BTN, XO, etc.) assemblies19. The Guinier plot (Fig. 5D, q = 0.00999-0.01568 Å−1) shows good linear, suggesting that further structural analysis of SAXS data is reliable. The p(r) curve shows a maximum dimension (Dmax) as 756 Å with a maximum frequency of 230 Å (Fig. 5E). The structural parameters listed in Table 2, gyration of radius (Rg) was calculated as 172.2 Å with an aspect ratio of 2.2 (the ratio of Dmax to twice of Rg), indicating an ellipsoidal shape of MFGM proteins in solution. The molecular weight (MW) was obtained as 1.99 × 103 kDa with a volume (Vp) of 2.39 × 103 nm3, suggesting a rather large complex of MFGM protein in solution. The Kratky plot shows a bell-shaped scattering profile around middle qRg region, indicating a compact structure of MFGM proteins (Fig. 5F). The scattering at ~7 qRg region was contributed by the colloidal scattering from the CCP. The structural parameters of MFGM proteins, Rg, MW, and Vp obtained in this work are higher than previously reported in the literature for goat casein micelles investigated by SAXS18, which attributes to the association of MFGM proteins donating the structural property. Despite that the polydispersity and structural heterogeneity of MFGM proteins obscuring the linearity of the Guinier region and making it difficult to obtain its accurate dimension, the overall size of MFGM proteins revealed by SAXS in this work is comparable with the previous results28,57.
A–O Effects of thermal treatment (20, 40, 60 and 80 ℃), SDS, DTAB and PS-20 on the SAXS curve, pair-distance distribution, p(r) plot and Kratky plot of MFGM proteins (1 mg/mL), respectively.
Upon thermal treatment, the SAXS profile changes in three major regions as follows: (1) increased I(0) and slope in Guinier region (Figs. 5B, D) indicates the increasing whole size of MFGM proteins, (2) reduced intensity around q = 0.01 Å−1 means the space among CCP-caseins clusters compressed, (3) increased intensity around q = 0.08 Å−1 suggests the increasing size of CCP-clusters, (4) maintained intensity of high-q range larger than 0.1 Å−1 suggests the little change in heterogenous protein upon increasing temperature. The p(r) curve shows shift of maximum frequency to longer pair-distance shown in Fig. 5E. The Kratky plot maintained similar shape but the peak shifted to the lower qRg region (Fig. 5F). The structural parameters listed in Table 2 exhibits that the values of Rg, MW, Vp and aspect ratio increased to 178.08 Å, 2.6 × 103 kDa (1.3 times that of control group), 3.13 × 103 nm3 (1.3 times that of control group) and 2.3. These results clearly suggest that thermal treatment induced aggregation of MFGM protein and the molecular weight and volume increased by 1.3 times as the temperature up to 80 °C.
As shown in Fig. 5G and Table 2, with adding SDS, the scattering intensity at low-q region (Guinier region) reduced significantly with a value Rg of 157.06 Å, meaning a reduced size of MFGM proteins bound by SDS. Three scattering “bumps” at q = 0.08, 0.15 and 0.3 Å−1 with increasing scattering intensity can been seen, suggesting a promoted interaction of CCP-caseins and formation of SDS-nanocluster complex with a three-layer core-shell structure. Additionally, the pronounced “bump” in the high-q region (q = 0.3 Å−1) contributed by the electron density contrast between the hydrophilic headgroups and hydrophobic alkyl chains of surfactant, forming a core-shell structure80. Figure 5H showed a Dmax of 729 Å in p(r) curve with a downgraded oscillation, suggested the reduced size of MFGM proteins upon SDS bound. Prominent features at qRg = 10, 30, and 50 in Kratky plots supported the three-layer core-shell structure with an aspect ratio of 2.3. The value of MW and Vp was calculated as 640 kD and 770 nm3, one-third of the control group, meaning the SDS-induced dissociation of larger MFGM proteins aggregates. Interestingly, the typical core-shell scattering characteristic was observed for MFGM proteins when bound with 0.1 mM SDS and a negative oscillation can be observed adding 1 mM SDS (significantly lower than its CMC of ~4 mM at neutral condition81), indicating that the MFGM proteins facilitated the formation of a remarkably core–shell structure with SDS.
After addition of DTAB (Fig. 5J), the intensity of SAXS profile at Guiner region reduced with a value Rg of 152.95 Å, and a typical scattering “bump” for core-shell structure was observed at q = 0.2 Å−1, which suggests that DTAB degraded the size of MFGM proteins forming a core-shell like complex and degraded the interactions among CCP-casein nanoclusters. The p(r) curve of DTAB showed a reduced oscillation and shifted to the lower pair-distance significantly with a Dmax of 703 Å and an aspect ratio of 2.3 (Fig. 5K), demonstrating a dissociation of MFGM proteins induced by DTAB. The value of MW and Vp was calculated as 710 kD and 860 nm3 (Table 2), larger a bit than that of SDS group. The scattering in Kratky plots shown in Fig. 5L went up when bound with DTAB, implying DTAB-MFGM proteins complex modified the interaction between CCP and caseins little. Typically, the core-shell scattering was found after adding low DTAB amount (0.1 mM), lower that the CMC of ~6 mM reported previously30, which indicated that DTAB cut off the interactions among CCP-casein nanoclusters, dissociated MFGM proteins, and then covered the surface of nanoclusters forming core-shell like structure.
Surfactant molecules (SDS/DTAB) initially interact with the MFGM protein aggregates through electrostatic and hydrophobic forces. At this stage, the primary effect is the disruption of protein-protein interactions within existing aggregates. This leads to the breakdown of larger aggregates into smaller subunits or individual protein molecules, which is observed as a reduction in apparent molecular weight and hydrodynamic size.
Following the initial dissociation, the surplus surfactant molecules continue to bind to the now-separated protein subunits. The hydrophobic tails of the surfactants associate with the hydrophobic regions of the proteins, while the hydrophilic heads face outward toward the aqueous environment. This rearrangement can lead to the formation of a core-shell-like complex, where the protein constitutes a core, surrounded by a surfactant shell. PS20 modified the SAXS profile of MFGM proteins slightly, the scattering intensity in low-q region decreased and the intensity in high-q region raised (Fig. 5M) with a value of Rg 176.5 Å. A faint scattering “bump” for core-shell structure was observed at q = 0.15 Å−1. The value of MW and Vp was calculated as 1.79 × 103 kD and 2.16 × 103 nm3 with an aspect ratio of 2.2 (Table 2), comparable with that of control group. The p(r) plot exhibited a Dmax of 775 Å with a higher r shift to 281 Å (Fig. 5N). Figure 5O showed the noticeable characteristics of CCP-casein nanocluster in Kratky plots of MFGM proteins, suggesting the CCP nanoclusters maintained. A pronounced scattering at q = 0.15 Å−1 was observed afterwards adding 0.1 mM PS-20, contributed by the positive/negative electron density contrast between the micellar shell and core of PS20 in solution, which was comparable with the CMC of PS20 reported in literature as 60 μM82. These findings demonstrated that no significant change in overall size of MFGM proteins was observed and the fluctuation in structural parameters of MFGM proteins could be due to the extra electron density of PS-20 micelles coated on the surface of MFGM proteins forming layer-like shell.
Polyoxyethylene sorbitan, a nonionic surfactant, is widely employed as food and pharmaceuticals filed as emulsifier, hydrocolloids and nanomaterial. PS20 primarily stabilizes interfaces (air/water, oil/water) by forming solid-like adsorption layers that reduce surface tension to near zero. PS20 prevents protein aggregation induced by interfaces or hydrophobic interactions, achieved by competing with proteins for the oil-water interface or binding to their exposed hydrophobic segments31. Interaction with small proteinaceous particles at interfaces can generate a surface “skin,” linked to the “stagnant surface” mechanism. While lower in affinity than skins formed by ionic surfactants, this PS-20-derived skin offers greater stability83. PS20 interacts with MFGM proteins through a surface-adsorption mechanism, forming a protective layer that does not substantially alter the protein’s intrinsic interfacial activity (e.g., its capacity to reduce equilibrium surface tension). However, this surface-bound PS20 layer significantly enhances the bulk colloidal stability of the protein dispersion by providing steric hindrance that prevents aggregation. Consequently, the system exhibits increased viscosity profile, while the foundational interfacial behavior remains governed by the MFGM proteins themselves.
In order to obtain the overall shape of MFGM proteins in solution, the ab initio DAMMIN models and DENSS reconstructions were further obtained. As seen from Fig. 6, the fitting of all of samples showed good agreement with the SAXS data structure. It was intuitively presented the MFGM proteins solution structure and the impacts of thermal treatment and surfactants on the structure The resolution of the DENSS reconstruction is estimated to be 80 Å. The excluded volume regions between DAMMIN and DENSS may account for the extra volume from the water holding capacity and electron donated by surfactants.
Top: Fitting result of experimental SAXS data (dark circle) with theoretical calculation of DAMMIN (red line) and DENSS (blue line) program. Bottom: ab initio model envelope (dummy dots) calculated by DAMMIN overlays with electron density reconstructed by DENSS (gray surface).
It must be pointed out that the FTIR and CD spectra indicated an increase in local secondary structure (e.g., α-helix content) upon treatment, SAXS revealed concomitant changes in the global supramolecular organization, such as heat-induced aggregation or surfactant-induced dissociation. These findings are reconciled by a model in which the treatments induce a partial unfolding of the native structure, leading to a reorganized state. In this state, the polypeptide chain refolds into more stable secondary structures locally, while the disruption of tight three-dimensional packing promotes self-association or interaction with surfactants at a larger scale, thereby altering the solution’s colloidal architecture.
Morphological analysis
The morphological characterization of MFGM proteins was further implemented using scanning electron microscopy (SEM) and transmission electron microscopy (TEM). As shown in Fig. 7A, MFGM protein at 20 °C distributed homogeneously and exhibited a nearly spherical shape with an average diameter of ~50 nm. The larger spherical shape particles with an average diameter of ~100 nm corresponded to casein micelles, which was., which was consistent with the TEM results of goat casein micelles reported previously18. Upon thermal treatment (Fig. 7B–D), the amount of MFGM proteins decreased and size increased with an amorphous shape, confirming the thermal treatment induced aggregation of MFGM proteins. The electron microscopy results of MFGM proteins were in line with the previous results of heat treatment on the behavior of protein and fat globules84
A–G Effects of 20, 40, 60, 80 ℃, SDS, DTAB and PS-20 on the SEM (left pane) and TEM (right pane) micrographs of MFGM proteins, respectively.
After addition of SDS, casein micelles could not be observed and MFGM proteins further lost its initial morphology and was dissociated into smaller particles with a good homogeneity with an average dimeter of ~20 nm (Fig. 7D) This suggested that SDS initiated the dissociation of MFGM proteins, especially for the casein micelles. Upon adding DTAB (Fig. 7E), MFGM proteins dissociated into particles with an average diameter of ~50 nm particles observed by SEM and TEM images. As shown in Fig. 7F, PS20 formed a layer on the surface of MFGM proteins, and MFGM proteins maintained a similar size and shape, suggestion no significant modification in MFGM proteins upon bound by PS20. The major role of PS20 could form adsorbed layer on the surface of MFGM proteins to stabilize the air/water or oil/water interface, which was in line with the SAXS results.
It should be noted that the SEM/TEM imaging revealed that heat-treated MFGM proteins formed large, irregular aggregates at the micrometer scale, SAXS analysis indicated the presence of compact, ellipsoidal-like structures at the nanometer scale within the hydrated solution. This apparent discrepancy is reconciled by a hierarchical assembly model wherein proteins first form dense, nanoscale clusters in solution (detected by SAXS), which subsequently coalesce into larger, amorphous super-structures upon sample preparation for microscopy. The ellipsoidal model from SAXS thus describes the local packing and overall shape of the primary building blocks of the aggregate, rather than the final, complex macroscopic morphology observed by electron microscopy.
Discussion
In this work, the MFGM proteins were extracted and identified from goat reconstituted milk powder (GRMP). Caseins, XO/XDH, BTN and whey were major components of goat MFGM proteins. The heat treatment (20, 40, 60, and 80 °C) and different charged surfactants (SDS, DTAB and PS20) impacted the interfacial characteristics and structure of goat MFGM protein significantly. The heat treatment decreased the zeta potential, thermal stability and contact angle, but increased size and surface tension of MFGM proteins. With increasing treated temperature, MFGM proteins exhibited a trend of aggregation due to the reduction of free sulfhydryl (SH) groups and increased disulfide (S-S) groups in the MFGM proteins, suggesting that incorporation of whey might be due to association with MFGM proteins and caseins via disulfide bonds. The observed heat-induced increase in α-helix and unordered structures, concomitant with a decrease in antiparallel β-sheet content, signifies a structural reorganization of the MFGM proteins into a molten globule state. Complementary SAXS analysis confirmed an increase in the overall size of the MFGM proteins assemblies upon heating, consistent with a partial unfolding and aggregation driven by the exposure of hydrophobic regions in the molten globule state, while revealing that the fundamental architecture of the heterogeneous protein assemblies remained largely intact at the nanoscale.
The interaction of SDS and DTAB with MFGM proteins follows a sequential two-stage mechanism. In the initial stage, both surfactants disrupt intermolecular interactions within native protein aggregates, leading to their dissociation into smaller subunits—a process reflected in the observed two-thirds reduction in molecular weight and volume. In the subsequent stage, these liberated subunits reorganize with surfactant molecules to form structurally defined core–shell complexes, wherein the protein core is stabilized by a surfactant shell. This structural rearrangement yields three key functional outcomes: (1) enhanced colloidal stability and compactness compared to native aggregates; (2) increased surface activity manifested by reduced surface tension and accelerated adsorption at the air–water interface; and (3) increased thermal stability, attributable to the surfactant-induced stabilization of the MFGM proteins in a molten globule-like state characterized by heightened structural order and dynamic flexibility. .PS20 interacts MFGM proteins primarily through a surface-dominated mechanism, forming an adsorbed layer that stabilizes interfaces like the air-water or oil-water boundary without significantly penetrating or altering the protein’s internal structure. This behavior, attributed to the steric hindrance and modest binding affinity of PS20, explains why its binding does not fundamentally change the intrinsic interfacial properties of the MFGM proteins (e.g., its capacity to reduce equilibrium surface tension), as the pre-formed surfactant layer modulates interfacial energy while preserving the protein’s core architecture. However, this surface-bound PS20 layer significantly enhances the bulk colloidal stability of the protein dispersion by providing steric hindrance that prevents aggregation. Consequently, the system exhibits increased viscosity profile, while the foundational interfacial behavior remains governed by the MFGM proteins themselves. In conclusion, moderate heat treatment and PS20 could be an option for manufacture and application of MFGM proteins.
As far as we know, it is the first time to report the impacts of heat and noncovalent interactions on the solution structure of goat MFGM proteins using SAXS. These findings could contribute to a better understanding impact of thermal treatment and additives on the interfacial properties and structure of goat MFGM proteins. This work also provides new insight for structural modification of MFGM proteins-based food and nanomaterials modulated by heat and noncovalent bonding.
Methods
Materials
GRMP used in this study was sourced from Longteng Biological Dairy Co., Ltd. (Yunnan, China). The product complies with the Chinese national standard for milk powder (GB 19644) and holds the Food Production License Number SC10553012207005. Its nutritional composition, expressed on a weight/weight basis, is as follows: 30% protein (comprising 24% casein and 6% whey protein), 28.7% fat, and 35.8% lactose. SDS, DTAB and PS20 were purchased from Macklin Biochemical Technology Co., Ltd. (Shanghai, China) and used without further purification. Ultra-high-purity water was prepared using a Millipore SAS-67120 (Molsheim, Cedex, France).
Preparation of goat MFGM proteins
Goat reconstituted milk was prepared by dispersing the milk powder in Milli-Q water to a final solids content of 25% (w/v, equivalent to 25 g of powder in 100 mL of water). The dispersion was centrifuged at 5570 rpm for 15 min at 4 °C. The upper milk fat globule layer, enriched with MFGM proteins, was carefully collected. The collected milk fat was resuspended in 10 mM phosphate-buffered saline (PBS), pH 7.4 (10 mL per gram of fat). The suspension was mixed thoroughly by shaking in a 37 °C water bath for 30 min to facilitate the removal of loosely associated proteins. The sample was then centrifuged at 7200 rpm for 15 min at 4 °C to recover the washed milk fat layer. This PBS washing procedure was repeated three times to ensure thorough purification. A final wash was performed using an equivalent volume of distilled water to remove residual salts from the PBS buffer. The purified milk fat globules were resuspended in 100 mM Tris-HCl buffer, pH 7.4 (5 mL per gram of fat). The mixture was incubated in a 95 °C water bath for 5 min to solubilize MFGM proteins through heat-induced release, followed by vigorous vortexing for 1 min to ensure complete mixing. The sample was centrifuged at 10,300 rpm for 20 min at 4 °C. The supernatant, containing the solubilized MFGM proteins, was collected as the crude MFGM proteins fraction. For further purification and concentration, 200 μL of the MFGM proteins supernatant was mixed with 1 mL of pre-chilled acetone (–20 °C) in a 1:5 (v/v) ratio. The mixture was stored at –20 °C for 20 h to facilitate complete protein precipitation. The precipitate was collected by centrifugation at 10,300 rpm for 40 min at 4 °C. The resulting protein pellet was freeze-dried to obtain the purified MFGM protein powder, which was stored at –80 °C for subsequent analysis.
Heat treatment and noncovalent interactions
Heat treatment was performed using a precision water incubator (HH-2A, Ronghua Instrument Co., Ltd, China) at temperatures of 20, 40, 60, and 80 °C for durations of 30 min. Immediately after heating, the samples were cooled in an ice-water bath for 5 min to terminate the heat treatment before subsequent analysis. The interaction of MFGM with surfactants (SDS, DTAB and PS20) was processed as follows. Stock solutions of SDS, DTAB and PS20 (100 mM) were prepared in Milli-Q water. The MFGM samples were mixed with the surfactant solutions to achieve final surfactant concentrations of 1 mM with constant gentle agitation of 1 min to ensure equilibrium.
UHPLC-MS/MS analysis
Aliquots containing 150 μg of total proteins were subjected to enzymatic digestion (trypsin) at 37 °C for overnight. Following desalting, the supernatant was collected for nanoflow liquid chromatography separation prior to analysis using a using a Nano upgraded HPLC (EASY-nLC 1200, Thermo Fisher) coupled with a high-resolution mass spectrometry Q-Exactive HF (Thermo Fisher) in Standard Sci-tech Innovation (Qingdao, China).
Chromatographic separation was achieved using an Acclaim PepMap™ RSLC C18 nanoLC column (150 mm × 50 μm, 2 μm; Thermo Fisher Scientific). The mobile phases consisted of 0.1% (v/v) formic acid in water (Eluent A) and 0.1% (v/v) formic acid in 80% (v/v) acetonitrile (Eluent B). A sample volume of 1 μL was injected, and the peptides were eluted at a constant nanoflow rate of 300 nL/min with the following gradient program: 0–98 min, 8% to 28% B; 98–113 min, 28% to 37% B; 113–117 min, 37% to 100% B; 117–120 min, 100% B. The mass spectrometer was operated in positive/negative polarity mode with spray voltage of 2.4 kV, capillary temperature of 275 °C, sheath gas flow rate of 0 arb and aux gas flow rate of 0 arb. The raw MS/MS data were processed using Proteome Discoverer software (version 2.5, Thermo Fisher Scientific). Protein identification was conducted by searching the fragmentation spectra against a relevant species-specific protein database. The false discovery rate for both peptide and protein identification was set to ≤1% to ensure high confidence in the results.
Polyacrylamide gel electrophoresis analysis
The elution samples (2 mg/mL) were collected and their composition was checked by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE). SDS-PAGE was conducted using 15% (v/w) acrylamide separating gel and 4% acrylamide stacking gel. Samples were prepared in 10 mM Tris-HCl buffer (pH 6.8) containing 2% SDS and 0.2% β-mercaptoethanol. Electrophoresis was performed at 15 and 20 mA in the stacking and separating gels, respectively. The gel sheets were stained with Coomassie brilliant blue R-250 in 50% trichloroacetic acid (C2HCl3O2), and destained in 7% acetic acid (methanol: acetic acid: water, 227:37:236 [v/v/v])19.
HPLC analysis
In this study, an Agilent 1100 Series chromatograph (Agilent Technologies, Santa Clara, CA) equipped with a binary pump, autosampler and UV detector was used. Separation of the proteins was performed on a Zorbax 300 SB reverse-phase C18 column with a silica-based packing (3.5 μm, 300 Å 150 × 4.6 mm I.D.; Agilent Technologies). The flow rate was set at 1.0 mL/min, and the column temperature was controlled between 25–30 °C. The sample solution described in section “Analysis of interfacial behavior and thermal stability” with a concentration of 1 mg/mL was prepared. d. The mobile phase consisted of solvent A (0.1% trifluoroacetic acid in water) and solvent B (0.1% trifluoroacetic acid in acetonitrile). The gradient elution program was as follows: initial conditions were maintained at 5% B, followed by a linear increase to 60% B over 10 min, then a further ramp to 95% B within 5 min. Detection wavelength was set up 280 nm to assess the chromatographic profile of the MFGM proteins samples following different treatments (heat, interaction with surfactants) described in section “Analysis of interfacial behavior and thermal stability”.
UV-vis absorbance, fluorescence and far-UV circular dichroism (CD) measurements
UV-vis absorbance spectra of MFGM protein samples, following different treatments described in section “Analysis of interfacial behavior and thermal stability”, were recorded from 200 to 400 nm. Steady-state fluorescence measurements were performed using Dual-FL (Horiba Jobin-Yvon) with an excitation wavelength of 280 nm and the emission range between 290 and 500 nm at room temperature and quartz cuvette (volume = 1.5 mL, optical path length = 1.0 cm). Far-UV CD spectra were recorded in a 1 cm-quartz cuvette on the Chirascan plus CD spectrometer (Applied Photophysics Ltd). Wavelength scans were recorded in the range of 190–260 nm (bandwidth 1.0 nm and path 0.1 cm), and the scanning speed is 10 nm/min with four accumulations averaged to obtain the final spectrum. The results are presented as mean residue molar ellipticity θ (deg cm2 dmol−1) v.s. wavelength (nm). The secondary content was analyzed using Beta Structure Selection (BeStSel) (https://bestsel.elte.hu)85. Background contribution from the buffer was subtracted. All measurement were carried out at room temperature.
Fourier transform infrared measurement
The Fourier-transform infrared (FTIR) spectra of the MFGM protein samples following various treatments described in section “Analysis of interfacial behavior and thermal stability”, were recorded at room temperature (~25 °C) using a Bruker Equinox 55 spectrometer (Bruker Optik GmbH, Ettlingen, Germany). The instrument was configured with a liquid nitrogen-cooled mercury–cadmium–telluride detector to enhance sensitivity in the mid-infrared region. Spectral data were acquired in the range of 4000–400 cm⁻¹ with a high resolution of 2 cm⁻¹. To ensure an optimal signal-to-noise ratio, each spectrum was obtained by averaging 512 consecutive scans. The sample (60 μg) was spread on a germanium ATR plate (50 mm × 20 mm × 2 mm, Bruker Optik) with an aperture of 45° yielding 25 internal reflections. Reference spectra of the germanium plate were automatically recorded after purging with dry air for 15 min. These reference spectra were then rationed against the recently run sample spectra. The sample was slowly dried under a stream of N2. After that, the plate was sealed in a universal sample holder and rehydrated by flushing the holder with N2 saturated with D2O for 4 h at room temperature.
Nuclear magnetic resonance (NMR) spectrum
NMR measurements for MFGM proteins samples, following different treatments described in section “Analysis of interfacial behavior and thermal stability”, were performed at 800 MHz Bruker spectrometer equipped with a field gradient probe. NMR samples had the typical volume of 600 μL with 10% D2O content. A total of 4 scans were collected with a recycling delay of 1 s. In each run, 8 dummy scans were first applied to the sample before the actual experiment carried out. For protein self-diffusion measurements, the spectral width was 20 ppm and 16 scans were collected with a recycling delay of 2 s. NMR tubes (5-mm) were used and all the measurement were performed at 20 °C. All 1H NMR data were Fourier transformed with 0.3- to 1.0-Hz exponential line broadening.
Size and interfacial characteristics measurements
The size of MFGM proteins samples, following different treatments described in section “Analysis of interfacial behavior and thermal stability”, was detected using a laser nanoparticle analyzer (Winner802, Weina, China) based on the dynamic light scattering theory with a scattering angle of 90° in the size range of 1–1000 nm at room temperature.
The surface wettability of MFGM protein sampleswas measured according to Li et al. method86. The sessile drop method in which a droplet of the tested liquid was placed on a horizontal film surface with a BDA25S (Krüss GmbH, Hamburg, Germany) contact angle meter, equipped with a high-speed camera and an image analysis software. Specifically, using a precision syringe to deposit 3.0 μL of deionized water droplets on the air side of the film surface. To decrease the possibility of human intervention that may arise from droplet volume loss, the contact angle at 2 s after deposition of the droplets on substrate chosen to investigate. The measurements were implemented at least five parallels for each sample.
Differential scanning calorimetry (DSC) measurement
Thermal stability properties of MFGM proteins samples powder (5 mg), following different treatments described in section “Analysis of interfacial behavior and thermal stability”, were determined using differential scanning calorimeters (DSC) (NETZSCH 3500, Germany) with a Concavus AI crucible under the scanning rate of 10 K/min within the temperature range from 5 to 200 °C. For the curve analysis, buffer-buffer tracings were recorded under the same conditions and subtracted from the sample endotherms. Data analyses consisted of manual peak integration of the DSC thermograms, followed by calculation of the enthalpy change (∆H) and denaturation temperature (Td) were used the DSC supplementary software.
Small-angle X-ray scattering (SAXS) measurement
The SAXS measurements of MFGM proteins samples following different treatments described in section “Analysis of interfacial behavior and thermal stability” were performed on BioSAXS-BL19U2 beamline of National Facility for Protein Science Shanghai (NFPS) at the Shanghai Synchrotron Radiation Facility (SSRF)87. The wavelength of the monochromatic X-ray beam was set to λ = 1.033 Å, with a q-range from 0.006 to 0.467 Å−1 (q = 4πsin(θ)/λ, where 2θ is the scattering angle). All measurements were performed at 25 oC. A conventional SAXS setup was equipped with a robotic sample changer with an exposure time of 1 s per frame, collected by a Pilatus 2 M photon counting detector (DECTRIS Ltd., Switzerland). Buffer subtraction, data reduction and normalization were performed using BioXTAS RAW software (Version 1.2.1)88. SAXS data analysis was carried out using the ATSAS software program suite (Version 3.0.0)89. The radius of gyration (Rg) and forward scattering I(0) were obtained by the Guinier analysis “Autorg” function, the distribution of pair distances P(r) and the maximum interatomic distance (Dmax) within molecules were determined using GNOM integrated in PRIMUSqt. Low-resolution ab initio bead models were built up with the program DAMMIN using scattering data within a q range of 0.009–0.467 Å−1, which generates models represented by an ensemble of densely packed beads90. Thirty-two independent runs were performed, and the resulting models were averaged by DAMAVER program and were filtered using DAMFILT program to generate the final model. The alignment of the optimal predicted structure with the ab initio model was superimposed using SUPCOMB program based on the normalized spatial discrepancy (NSD) criteria. Data quality was evaluated based on χ2 value between the theoretical scattering curve and experimental SAXS data. Electron density map generation using program DENSS91. The same output pair distribution functions and Dmax values from GNOM as used for DAMMIN was used. Twenty independent models were generated for each dataset. The best principal axis of the ellipsoid describing the molecular dimensions to which to apply the symmetry was determined by applying it to each axis in turn in independent simulations. The bead models generated by the programs DAMMIN and DENSS were transformed into volumetric maps using Situs and displayed using UCSF Chimera.
Electron microscopy measurements
SEM measurements of MFGM proteins samples following different treatments described in section “Analysis of interfacial behavior and thermal stability” were performed recording to the method reported by Sun et al.81. One drop of sample (1 mg mL−1) was dipped on a polyvinyl difluoride organic membrane (PVDF, Tech Sep, Miribel, France). The membranes with samples attached to them were directly dehydrated with a supercritical carbon dioxide (Balzer CPD 020, Blazer Union, Furstentum, Liechtenstein). The dried samples were sputter coated with 10 nm layer of gold/platinum and imaged with an SEM (ZEISS, Sigma 300, Germany) at 15 kV.
TEM measurements of MFGM proteins samples following different treatments described in section “Analysis of interfacial behavior and thermal stability” were performed according to method of Hu et al.31 that ∼5 μL uranyl acetate (0.5%) negative-stained sample (0.1 mg mL−1) was placed onto carbon coated grid and drying in air. TEM (JEOL, JEM-2100Plus, Japan) operating at 100 kV was used for imaging equipped with an UltraScan 4000 4 × 4k digital camera using the image acquisition software Digital Micrograph (Gatan Software Team Inc., Pleasanton).
Statistical processing
The statistical analyses were performed using Graphpad prism 6.0 statistical software. All experimental data are presented as mean ± standard deviation (mean ± SD). One-way analysis of variance (ANOVA) with Tukey’s multiple comparisons was used to compare more than two groups.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Wang, Y. et al. Fractionation of milk fat globule membrane enriched materials from caprine milk cream. Int. Dairy J. 144, 105698 (2023).
Jukkola, A., Partanen, R., Xiang, W., Heino, A. & Rojas, O. J. Food emulsifiers based on milk fat globule membranes and their interactions with calcium and casein phosphoproteins. Food Hydrocoll. 94, 30–37 (2019).
Arranz, E. & Corredig, M. Invited review: Milk phospholipid vesicles, their colloidal properties, and potential as delivery vehicles for bioactive molecules. J. Dairy Sci. 100, 4213–4222 (2017).
Livney, Y. D., Ruimy, E., Ye, A. M., Zhu, X. & Singh, H. A milkfat globule membrane-inspired approach for encapsulation of emulsion oil droplets. Food Hydrocoll. 65, 121–129 (2017).
Gallier, S., Acton, D., Garg, M. & Singh, H. Natural and processed milk and oil body emulsions: bioavailability, bioaccessibility and functionality. Food Struct. 13, 13–23 (2017).
Rashidinejad, A., Birch, E. J. & Everett, D. W. Interactions between milk fat globules and green tea catechins. Food Chem. 199, 347–355 (2016).
Holzmüller, W. & Kulozik, U. Technical difficulties and future challenges in isolating membrane material from milk fat globules in industrial settings—a critical review. Int. Dairy J. 61, 51–66 (2016).
Jukkola, A. & Rojas, O. J. Milk fat globules and associated membranes: colloidal properties and processing effects. Adv. Colloid Interface Sci. 245, 92–101 (2017).
Nie, C. et al. Structure, Biological functions, separation, properties, and potential applications of milk fat globule membrane (MFGM): a review. Nutrients 16, 587 (2024).
Ye, A., Singh, H., Taylor, M. W. & Anema, S. Characterization of protein components of natural and heat-treated milk fat globule membranes. Int. Dairy J. 12, 393–402 (2002).
Huang, Y. et al. Effects of various thermal treatments on interfacial composition and physical properties of bovine milk fat globules. Food Res. Int. 167, 112580 (2023).
Wiking, L., Gregersen, S. B., Hansen, S. F. & Hammershøj, M. Heat-induced changes in milk fat and milk fat globules and its derived effects on acid dairy gelation—a review. Int. Dairy J. 127, 105213 (2022).
Bermúdez-Aguirre, D., Mawson, R. & Barbosa-Cánovas, G. V. Microstructure of fat globules in whole milk after thermosonication treatment. J. Food Sci. 73, E325–E332 (2008).
Huppertz, T., Uniacke-Lowe, T. & Kelly, A. L. in Advanced Dairy Chemistry, Volume 2: Lipids (eds Paul L. H. McSweeney, Patrick F. Fox, & James A. O’Mahony) 133–167 (Springer International Publishing, 2020).
Verruck, S., Dantas, A. & Prudencio, E. S. Functionality of the components from goat’s milk, recent advances for functional dairy products development and its implications on human health. J. Funct. Foods 52, 243–257 (2019).
Yan, D. et al. Changes in caprine milk fat globule membrane proteins after heat treatment using a label-free proteomics technique. Foods 11, 2705 (2022).
Ma, Y., Zhang, L., Wu, Y. & Zhou, P. Changes in milk fat globule membrane proteome after pasteurization in human, bovine and caprine species. Food Chem. 279, 209–215 (2019).
Yang, L. et al. Surfactant charge tuning alters casein micelle structure and complexation behavior. Food Hydrocoll. 164, 111145 (2025).
Du, Z. et al. Study on internal structure of casein micelles in reconstituted skim milk powder. Int. J. Biol. Macromol. 224, 437–452 (2023).
Wang, C. et al. Proteomic characterization and comparison of milk fat globule membrane proteins of Saanen goat milk from 3 habitats in China using SWATH-MS technique. J. Dairy Sci. 106, 2289–2302 (2023).
Han, B., Zhang, L. & Zhou, P. Comparison of milk fat globule membrane protein profile among bovine, goat and camel milk based on label free proteomic techniques. Food Res. Int. 162, 112097 (2022).
Pisanu, S. et al. The sheep milk fat globule membrane proteome. J. Proteom. 74, 350–358 (2011).
Holzmüller, W. & Kulozik, U. Quantification of MFGM proteins in buttermilk and butter serum by means of a stain free SDS-PAGE method. J. Food Compos. Anal. 49, 102–109 (2016).
Affolter, M., Grass, L., Vanrobaeys, F., Casado, B. & Kussmann, M. Qualitative and quantitative profiling of the bovine milk fat globule membrane proteome. J. Proteom. 73, 1079–1088 (2010).
Sun, Y., Wang, C., Sun, X. & Guo, M. Comparative proteomics of whey and milk fat globule membrane proteins of Guanzhong goat and Holstein cow mature milk. J. Food Sci. 84, 244–253 (2019).
Stefferl, A. et al. Butyrophilin, a milk protein, modulates the encephalitogenic T cell response to myelin oligodendrocyte glycoprotein in experimental autoimmune encephalomyelitis. J. Immunol. 165, 2859–2865 (2000).
Fong, B. Y. & Norris, C. S. Quantification of milk fat globule membrane proteins using selected reaction monitoring mass spectrometry. J. Agric. Food Chem. 57, 6021–6028 (2009).
Pan, Y. et al. Comparative analysis of interfacial composition and structure of fat globules in human milk and infant formulas. Food Hydrocoll. 124, 107290 (2022).
Lee, S. J. & Sherbon, J. W. Chemical changes in bovine milk fat globule membrane caused by heat treatment and homogenization of whole milk. J. Dairy Res. 69, 555–567 (2002).
Cao, F. et al. Insights on the structure of caseinate particles based on surfactants-induced dissociation. Food Hydrocoll. 104, 105766 (2020).
Hu, X. et al. How much can we trust polysorbates as food protein stabilizers—the case of bovine casein. Food Hydrocoll. 96, 81–92 (2019).
Miyazawa, T., Itaya, M., Burdeos, G. C., Nakagawa, K. & Miyazawa, T. A critical review of the use of surfactant-coated nanoparticles in nanomedicine and food nanotechnology. Int. J. Nanomed. 16, 3937–3999 (2021).
Kralova, I. & Sjöblom, J. Surfactants used in food industry: a review. J. Dispers. Sci. Technol. 30, 1363–1383 (2009).
Prasad, S. et al. Near UV-Visible electronic absorption originating from charged amino acids in a monomeric protein. Chem. Sci. 8, 5416–5433 (2017).
Kuipers, B. J. H. & Gruppen, H. Prediction of molar extinction coefficients of proteins and peptides using UV absorption of the constituent amino acids at 214 nm to enable quantitative reverse phase high-performance liquid chromatography−mass spectrometry analysis. J. Agric. Food Chem. 55, 5445–5451 (2007).
Chandra Roy, M., Zhang, L., Liu, X. & Zhou, P. Investigation of caprine milk serum proteome and glycated proteome changes during heat treatment using robust ion mobility time-of-flight proteomic techniques. Int. Dairy J. 110, 104798 (2020).
Zhang, L., Zhou, R., Zhang, J. & Zhou, P. Heat-induced denaturation and bioactivity changes of whey proteins. Int. Dairy J. 123, 105175 (2021).
Anema, S. G. Heat-induced changes in caseins and casein micelles, including interactions with denatured whey proteins. Int. Dairy J. 122, 105136 (2021).
Ye, A., Singh, H., James Oldfield, D. & Anema, S. Kinetics of heat-induced association of β-lactoglobulin and α-lactalbumin with milk fat globule membrane in whole milk. Int. Dairy J. 14, 389–398 (2004).
Hansen, S. F., Nielsen, S. D., Rasmusen, J. T., Larsen, L. B. & Wiking, L. Disulfide bond formation is not crucial for the heat-induced interaction between β-lactoglobulin and milk fat globule membrane proteins. J. Dairy Sci. 103, 5874–5881 (2020).
Sun, Y., Oseliero, P. L. & Oliveira, C. L. P. alpha-Lactalbumin and sodium dodecyl sulfate aggregates: denaturation, complex formation and time stability. Food Hydrocoll. 62, 10–20 (2017).
Zhao, Y. et al. Milk fat globule membrane (MFGM) phospholipid-Whey protein interaction characterization and its effect on physicochemical, interfacial properties and evaluation of in vitro digestion of emulsions - Inspired by the MFGM. Food Hydrocoll. 155, 110173 (2024).
Ma, Q. et al. Interaction between whey protein and soy lecithin and its influence on physicochemical properties and in vitro digestibility of emulsion: A consideration for mimicking milk fat globule. Food Res. Int. 163, 112181 (2023).
Holt, C., Raynes, J. K. & Carver, J. A. Sequence characteristics responsible for protein-protein interactions in the intrinsically disordered regions of caseins, amelogenins, and small heat-shock proteins. Biopolymers 110, e23319 (2019).
Rico-Pasto, M., Zaltron, A., Davis, S. J., Frutos, S. & Ritort, F. Molten globule–like transition state of protein barnase measured with calorimetric force spectroscopy. Proc. Natl. Acad. Sci. 119, e2112382119 (2022).
Lou, K. et al. Molten globule-state protein structure: Perspectives from food processing applications. Food Res. Int. 198, 115318 (2024).
Sun, Y. et al. The role of hydrophobic interactions in the molten globule state of globular protein modulated by surfactants. Colloids Surf. B Biointerfaces 230, 113490 (2023).
Shen, Y. & Bax, A. Protein backbone chemical shifts predicted from searching a database for torsion angle and sequence homology. J. Biomol. NMR 38, 289–302 (2007).
Mulder, F. A. A. & Filatov, M. NMR chemical shift data and ab initio shielding calculations: emerging tools for protein structure determination. Chem. Soc. Rev. 39, 578–590 (2010).
Wang, Y. & Jardetzky, O. Investigation of the neighboring residue effects on protein chemical shifts. J. Am. Chem. Soc. 124, 14075–14084 (2002).
Hovjecki, M., Miloradovic, Z., Rac, V., Pudja, P. & Miocinovic, J. Influence of heat treatment of goat milk on casein micelle size, rheological and textural properties of acid gels and set type yoghurts. J. Texture Stud. 51, 680–687 (2020).
Minić, D. A. P. et al. Goat milk proteins enriched with Agaricus blazei Murrill ss. Heinem extracts: electrophoretic, FTIR, DLS and microstructure characterization. Food Chem. 402, 134299 (2023).
Postelmans, A., Aernouts, B., Jordens, J., Van Gerven, T. & Saeys, W. Milk homogenization monitoring: fat globule size estimation from scattering spectra of milk. Innov. Food Sci. Emerg. Technol. 60, 102311 (2020).
Luo, J., Wang, Y., Guo, H. & Ren, F. Effects of size and stability of native fat globules on the formation of milk gel induced by rennet. J. Food Sci. 82, 670–678 (2017).
Liang, L., Zhang, X., Wang, X., Jin, Q. & McClements, D. J. Influence of dairy emulsifier type and lipid droplet size on gastrointestinal fate of model emulsions: in vitro digestion study. J. Agric. Food Chem. 66, 9761–9769 (2018).
Holzmüller, W., Gmach, O., Griebel, A. & Kulozik, U. Casein precipitation by acid and rennet coagulation of buttermilk: Impact of pH and temperature on the isolation of milk fat globule membrane proteins. Int. Dairy J. 63, 115–123 (2016).
Michalski, M.-C., Michel, F. & Geneste, C. Appearance of submicronic particles in the milk fat globule size distribution upon mechanical treatments. Le. Lait. 82, 193–208 (2002).
Corredig, M., Nair, P. K., Li, Y., Eshpari, H. & Zhao, Z. Invited review: understanding the behavior of caseins in milk concentrates. J. Dairy Sci. 102, 4772–4782 (2019).
Anema, S. G. & Li, Y. Effect of pH on the association of denatured whey proteins with casein micelles in heated reconstituted skim milk. J. Agric Food Chem. 51, 1640–1646 (2003).
Liu, Y. et al. Impact of fatty acid carbon chain length and protein composition on physicochemical and digestive properties of MFGM contained emulsions. Food Sci. Nutr. 13, e70220 (2025).
Michalski, M.-C., Michel, F., Sainmont, D. & Briard, V. Apparent ζ-potential as a tool to assess mechanical damages to the milk fat globule membrane. Colloids Surf. B Biointerfaces 23, 23–30 (2002).
Gülseren, İ, Alexander, M. & Corredig, M. Probing the colloidal properties of skim milk using acoustic and electroacoustic spectroscopy. Effect of concentration, heating and acidification. J. Colloid Interface Sci. 351, 493–500 (2010).
Sun, Y., Roos, Y. H. & Miao, S. Changes in milk fat globules and membrane proteins prepared from pH-adjusted bovine raw milk. Foods 11, 4107 (2022).
Tholstrup Sejersen, M. et al. Zeta potential of pectin-stabilised casein aggregates in acidified milk drinks. Int. Dairy J. 17, 302–307 (2007).
Han, R. et al. Distribution and variation in proteins of casein micellar fractions response to heat-treatment from five dairy species. Food Chem. 365, 130640 (2021).
Binks, B. P. Particles as surfactants—similarities and differences. Curr. Opin. Colloid Interface Sci. 7, 21–41 (2002).
Silva, N. N., Bahri, A., Guyomarc’h, F., Beaucher, E. & Gaucheron, F. AFM study of casein micelles cross-linked by genipin: effects of acid pH and citrate. Dairy Sci. Technol. 95, 75–86 (2015).
Chen, M., Sagis, L. M. C. & Sun, Q. Emulsification and dilatational surface rheology of ultrasonicated milk fat globule membrane (MFGM) materials. LWT 133, 110094 (2020).
Miller, R., Aksenenko, E. V. & Fainerman, V. B. Dynamic interfacial tension of surfactant solutions. Adv. Colloid Interface Sci. 247, 115–129 (2017).
Zarif, B., Shabbir, S., Shahid, R., Noor, T. & Imran, M. Proteosomes based on milk phospholipids and proteins to enhance the stability and bioaccessibility of β-carotene. Food Chem. 429, 136841 (2023).
Qin, Y. et al. Comparative study on the microstructure and functional properties of casein in goat milk processed by different methods. Int. J. Food Sci. Technol. 56, 1682–1689 (2021).
Yue, M., Huang, M., Zhu, Z., Huang, T. & Huang, M. Effect of ultrasound assisted emulsification in the production of Pickering emulsion formulated with chitosan self-assembled particles: stability, macro, and micro rheological properties. LWT 154, 112595 (2022).
Ma, Y. et al. Structural modification of whey protein isolate via electrostatic complexation with Tremella polysaccharides and its effect on emulsion stability at pH 4.5. Int. J. Biol. Macromol. 297, 139870 (2025).
Zhao, S. et al. The stability of three different citrus oil-in-water emulsions fabricated by spontaneous emulsification. Food Chem. 269, 577–587 (2018).
Liu, J. et al. Investigation into the influence of droplet size on the stability of diesel emulsions based on multiple light scattering. J. Mol. Liq. 390, 123182 (2023).
Yu, J., Wang, X. -y, Li, D., Wang, L. -j & Wang, Y. Development of soy protein isolate emulsion gels as extrusion-based 3D food printing inks: effect of polysaccharides incorporation. Food Hydrocoll. 131, 107824 (2022).
Gräwert, T. W. & Svergun, D. I. Structural modeling using solution small-angle X-ray scattering (SAXS). J. Mol. Biol. 432, 3078–3092 (2020).
Ingham, B. et al. Revisiting the interpretation of casein micelle SAXS data. Soft Matter 12, 6937–6953 (2016).
Pedersen, J. S., Møller, T. L., Raak, N. & Corredig, M. A model on an absolute scale for the small-angle X-ray scattering from bovine casein micelles. Soft Matter 18, 8613–8625 (2022).
Chen, R. et al. Developments in small-angle X-ray scattering (SAXS) for characterizing the structure of surfactant-macromolecule interactions and their complex. Int. J. Biol. Macromol. 251, 126288 (2023).
Sun, Y. et al. Sulfate dodecyl sodium-induced stability of a model intrinsically disordered protein, bovine casein. Food Hydrocoll. 82, 19–28 (2018).
Ruiz-Peña, M., Oropesa-Nuñez, R., Pons, T., Louro, S. R. W. & Pérez-Gramatges, A. Physico-chemical studies of molecular interactions between non-ionic surfactants and bovine serum albumin. Colloids Surf. B Biointerfaces 75, 282–289 (2010).
Mustan, F. et al. Interplay between bulk aggregates, surface properties and foam stability of nonionic surfactants. Adv. Colloid Interface Sci. 302, 102618 (2022).
Ye, A., Cui, J., Dalgleish, D. & Singh, H. Effect of homogenization and heat treatment on the behavior of protein and fat globules during gastric digestion of milk. J. Dairy Sci. 100, 36–47 (2017).
Micsonai, A. et al. BeStSel: webserver for secondary structure and fold prediction for protein CD spectroscopy. Nucleic Acids Res. 50, W90–W98 (2022).
Li, Y. et al. Exploration of structure-activity relationship between IgG1 and IgE binding ability and spatial conformation in ovomucoid with pulsed electric field treatment. LWT 141, 110891 (2021).
Liu, G. et al. Upgraded SSRF BL19U2 beamline for small-angle X-ray scattering of biological macromolecules in solution. J. Appl.Crystallogr. 51, https://doi.org/10.1107/S160057671801316X (2018).
Hopkins, J. B., Gillilan, R. E. & Skou, S. BioXTAS RAW: improvements to a free open-source program for small-angle X-ray scattering data reduction and analysis. J. Appl. Crystallogr. 50, 1545–1553 (2017).
Franke, D. et al. ATSAS 2.8: a comprehensive data analysis suite for small-angle scattering from macromolecular solutions. J. Appl. Crystallogr. 50, 1212–1225 (2017).
Svergun, D. I. Restoring low resolution structure of biological macromolecules from solution scattering using simulated annealing. Biophys. J. 76, 2879–2886 (1999).
Grant, T. D. Ab initio electron density determination directly from solution scattering data. Nat. Methods 15, 191–193 (2018).
Acknowledgements
We thank staff members at BL19U2 beamline of NFPS at SSRF for assistance with SAXS data collection. C.C. discloses support for the research of this work from Hohhot City's “Unveiling and Commanding” Science and Technology Plan Project (2023-Unveiling and Commanding-Agriculture-2). Z.Y.D. discloses support for publication of this work from the Basic Research Program of Yunnan Province (202501AT070020) and Yunnan Provincial Department of Education (2025J0136). Y. Sun discloses support for publication of this work from the Basic Research Program of Yunnan Province (202201AT070031) and Yunnan Xingdian Talent (XDYC-QNRC-2022-0739).
Author information
Authors and Affiliations
Contributions
Y.Y.L, Y. Song, L.B.S., and Y.Y.Z. collected the data, validated the data, prepared Figs. 1–7, and prepared Tables 1, 2. C.C., Z.Y.D., and Y. Sun conceptualized the study, developed the methodology, acquired funding, administered the project, supervised the research and wrote the original draft. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Luo, Y., Song, Y., Su, L. et al. Impacts of heat and noncovalent bonding on interfacial property and structure of goat milk fat globule membrane proteins. npj Sci Food 10, 21 (2026). https://doi.org/10.1038/s41538-025-00668-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41538-025-00668-5