Abstract
Lactic acid bacteria (LAB) are commonly used to degrade nitrite in dry fermented sausages. However, the impact of nitrite-degrading LAB on flavor formation and its regulatory effects on the microbial community, free amino acid (FAA), and lipid metabolism associated with flavor development are not yet fully understood. This study investigated the effects of Lacticaseibacillus rhamnosus H7—a nitrite-reducing strain—on the microbial community, flavor compounds, and metabolites using multi-omics and FAAs analysis. Results showed that the inoculation of Lcb. rhamnosus H7 became dominant, promoted the release of FAAs, and enhanced the accumulation and transformation of specific lipids (e.g., PE, TG, and PC). Correlation analysis revealed that Lcb. rhamnosus H7 was positively associated with the abundance of Weissella hellenica and Weissella minor. This might be further linked to enhanced amino acid and lipid metabolism, which coincided with the increased formation of key flavor compounds such as hexanoic acid and ethyl 2,4-hexadienoate. Overall, this study demonstrates that the inoculation of Lcb. rhamnosus H7 can modulate microbial structure and promote amino acid and lipid metabolism, improving the content of characteristic flavor compounds. This study also provides a reasonable hypothesis for the potential mechanism by which LAB regulate the flavor of low-nitrite dry fermented sausages.

Similar content being viewed by others
Introduction
As an essential additive in dry fermented sausage processing, nitrite plays an irreplaceable role in developing the characteristic color, inhibiting spoilage microorganisms, and contributing to flavor formation1. However, nitrite can react with amines to form carcinogenic nitrosamines, posing a potential risk to human health2. Therefore, developing safe and effective technologies to reduce nitrite residues has become a research priority. Lactic acid bacteria (LAB) are considered an ideal strategy for nitrite degradation due to their acid-producing capacity, generally recognized safety status, and positive contribution to the flavor of fermented meat products3. Although numerous LAB strains with high nitrite degradation efficiency have been screened, most studies focus solely on degradation performance. There is a notable lack of in-depth investigation into how these nitrite-degrading strains influence flavor formation, particularly in low-nitrite environments where flavor deterioration is a common challenge.
Flavor formation in dry fermented sausages is a complex biochemical process driven by the metabolic activities of the microbial community. Specifically, lipid and amino acid metabolism are the two core pathways determining the flavor profile4. During fermentation, microbial-derived enzymes (such as lipases and proteases) catalyze the breakdown of lipids and proteins into free fatty acids and amino acids, which serve as precursors for volatile flavor compounds (VFCs) like aldehydes, ketones, and alcohols5. Previous research has established that starter cultures can modulate the structure of the microbial community, thereby influencing these metabolic pathways6,7. However, the role of nitrite-degrading LAB in reshaping the microbial network and metabolic fluxes (lipid and amino acid profiles) to enhance flavor remain poorly understood. A systematic understanding of these interactions is also crucial for optimizing the sensory quality of low-nitrite sausages.
Lacticaseibacillus rhamnosus is a facultatively heterofermentative LAB widely valued in the food industry for its excellent stress tolerance and probiotic properties8,9,10. Previous studies have confirmed its potential in nitrite reduction or flavor formation within meat matrices. For example, Lcb. rhamnosus YL-1 shows certain nitrite-degrading ability (83% degradation rate in 48 h) and flavor-enhancing effects11. Sionek et al. reported that inoculation of Lcb. rhamnosus LOCK900 in sausages enhanced the production of esters and acids12. While existing studies confirm the degradation efficiency, the underlying metabolic linkages—particularly how these high-degrading strains modulate lipid hydrolysis and oxidation in a low-nitrite system—have not been fully elucidated.
In our previous work, we isolated a novel strain, Lcb. rhamnosus H7, which exhibited superior nitrite degradation capabilities (94.6% degradation after 36 h of cultivation) and significantly reduced nitrosamine and biogenic amine contents in preliminary sausage trials (unpublished data). Despite these promising safety characteristics, the specific linkages between Lcb. rhamnosus H7 inoculation and the development of characteristic flavor compounds in dry fermented sausages have not been detailed. To address this gap, this study employed an integrated multi-omics approach—combining microbiomics, flavoromics, lipidomics, and amino acid analysis—to systematically investigate the effects of Lcb. rhamnosus H7 on dry fermented sausages. We aimed to elucidate how this strain modulates the microbial community structure and regulates lipid and amino acid metabolism to drive the formation of VFCs. By establishing the correlation between the microbial community and metabolic profiles, this study provides a theoretical basis for the targeted modulation of flavor in low-nitrite dry fermented sausages using functional LAB starter cultures.
Results and discussion
Electronic nose (E-nose) analysis
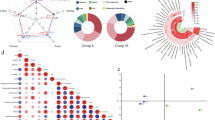
To investigate the effects of Lcb. rhamnosus H7 on the overall aroma characteristics of dry fermented sausages, an E-nose was employed to analyze the aroma differences among different treatment groups of final products. As shown in Fig. 1a, the response pattern was dominated by the W1C and W5S sensors, suggesting the possible presence of aromatic compounds and nitrogen oxides13. The response of the W5S sensor was significantly higher in the inoculated groups than in the C0 and C1 groups (p < 0.05). This distinct response pattern suggests that nitrogen oxides may be a potential contributor to the observed flavor differences between the groups.
a E-nose sensor response intensity. Data are presented as mean ± SD. b Principal component analysis (PCA). C0: sausages without nitrite or inoculation; C1: sausages with nitrite (100 mg/kg) without inoculation; LR1: sausages supplemented with nitrite and Lcb. rhamonosus H7 at 106 CFU/g; LR2: sausages supplemented with nitrite and Lcb. rhamonosus H7 at 107 CFU/g.
To further elucidate the flavor distinctions among the samples, Principal component analysis (PCA) was applied to reduce the dimensionality of the E-nose data14. As shown in Fig. 1b, PC1 and PC2 accounted for 67.7% and 21.5% of the total variance, respectively, with a cumulative contribution rate of 89.2%. This value exceeds 80%, indicating that the model fits well. The LR1 and LR2 groups were clustered in the positive quadrant of PC1, clearly separated from the C0 and C1 groups, suggesting that microbial inoculation significantly influences the flavor profile of dry fermented sausages. In summary, E-nose combined with PCA analysis effectively discriminated the flavor differences in dry fermented sausages induced by inoculation. To further elucidate the specific effects of Lcb. rhamnosus H7 on sausage flavor, the concentrations of VFCs were analyzed using GC–MS.
VFCs analysis
As shown in Table S2, a total of 43 VFCs were identified in the dry fermented sausages, which were classified into 10 alcohols, 8 aldehydes, 6 acids, 14 esters, and 5 other compounds. To elucidate the overall variations in flavor profiles, PCA was conducted on the VFC dataset obtained on day 12 (Fig. 2a). The first three principal components (PC1, PC2, and PC3) cumulatively explained 99.728% of the total variance, indicating that the model robustly captured the major compositional differences among samples. A distinct separation was observed between the treatment groups at the fermentation endpoint, suggesting that inoculation with Lcb. rhamnosus H7 substantially modulated the volatile composition. Further quantitative analysis revealed a progressive accumulation of total VFCs throughout the fermentation process across all groups (Fig. 2b and Table S2). Notably, the inoculated groups (LR1 and LR2) exhibited superior levels of total VFCs upon completion of fermentation, a trend primarily driven by the enrichment of acids and esters. These findings imply a strong correlation between Lcb. rhamnosus H7 inoculation and the enhanced production of these flavor-active metabolites.
a PCA of VFCs. b Content of each kind of VFC. c Heatmap of VFCs. d VFCs with VIP > 1 at the end of fermentation. e Content of differential VFCs at the end of fermentation. 0D, 3D, and 12D represent sausages sampled on day 0, day 3, and day 12, respectively. Sample abbreviations are defined in Fig. 1. Specific VFC identities are listed in Table S2. Data are presented as mean ± SD.
Alcohols, which generally impart mild aromatic notes, are typically derived from fatty acid oxidation and amino acid catabolism15,16. In the present study, phenylethanol (A1) and pentanol (A5) emerged as the predominant alcohols post-fermentation (Fig. 2c and Table S2). The inoculated groups demonstrated significantly higher concentrations of these compounds compared to the control (p < 0.05), which may be attributed to the inoculation of Lcb. rhamnosus H7. Phenylethanol is widely recognized for its contribution of rose-like notes to fermented sausages17, while pentanol provides an oily nuance18. Moreover, inoculation significantly elevated the concentration of hexanol (A2) (p < 0.05), associated with green, nutty, and popcorn-like descriptors. It is also noteworthy that (E)-2-decenol (A9), a characteristic volatile in dry-cured meat products19, was exclusively detected in the inoculated samples. Collectively, these results suggest that inoculation not only intensified the concentration of common alcohols but also introduced unique volatile markers such as (E)-2-decenol to the flavor profile.
The most substantial fluctuation in volatile composition occurred within the acid fraction, which exhibited a 29.60- to 313.49-fold increase by the end of fermentation (Table S2). Given that most acidic compounds—including octanoic acid (AC1), acetic acid (AC5), and 2-methylhexanoic acid (AC6)—were undetectable on day 0, their presence is principally ascribed to the fermentation process. Acetic acid, a product of carbohydrate metabolism known for its cheesy aroma20, emerged as the dominant acid in the final products, with peak levels observed in the Lcb. rhamnosus H7 inoculated groups. Additionally, inoculation specifically elevated levels of hexanoic acid (AC2) and butanoic acid (AC4), both recognized as typical sausage aroma compounds21. The concentration of these acids showed a significant positive correlation with inoculum size (p < 0.05), indicating that Lcb. rhamnosus H7 may play a role in modulating this core flavor pathway. Therefore, fermentation drives acid formation, while inoculation shapes the acid profile, leading to higher levels of characteristic acids such as acetic, hexanoic, and butanoic acids.
Esters constituted a major class of volatiles, contributing significantly to the fruity and sweet notes of the sausage. Their formation is generally linked to the esterification of alcohols and fatty acids, a reaction often catalyzed by microbial esterase activity22. Following fermentation, ethyl hexanoate (E2) emerged as the major ester, with its concentration significantly elevated by inoculation (p < 0.05). Consistent with earlier reports, this compound has been identified as a key aroma contributor in dry fermented sausages, imparting fruity and sweet aromas23. Similarly, levels of ethyl octanoate (E3), known for its vegetable-like and fruity attributes24, increased significantly in a dose-dependent manner relative to the inoculum size (p < 0.05). Beyond esters, inoculation also significantly upregulated certain terpenoids, specifically D-limonene (O1) and o-cymene (O2) (p < 0.05). These terpenoids, previously documented for their antimicrobial properties, are known to contribute citrus-like aroma notes to dry fermented sausages25.
To pinpoint the specific markers distinguishing the inoculated groups from the control (C1), a Variable Importance in Projection (VIP) analysis was performed (Fig. 2d). Thirteen VFCs with VIP values > 1 were identified as discriminative markers. Among these, phenylethanol, hexanoic acid, (E)-2-decenol, octanoic acid, butanoic acid, ethyl 2,4-hexadienoate, azulene, and o-cymene exhibited higher concentrations in inoculated groups compared to the C1 group, and their concentrations did not significantly decrease with increasing inoculum amount (Fig. 2e). This suggests that the increase in these compounds is directly related to the inoculation of Lcb. rhamnosus H7.
FAAs analysis
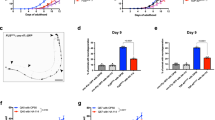
Free amino acids (FAAs) serve as pivotal precursors for flavor development in dry fermented sausages26. As illustrated in Fig. 3a, the total FAA content progressively increased during fermentation, a trend consistent with that reported by Wang et al.27, likely due to the proteolytic activity of microorganisms promoting protein breakdown and FAA release. Conversely, umami amino acids exhibited a declining trend, possibly because their degradation and conversion into flavor compounds exceeded their rate of formation28. Notably, at the end of fermentation, the total FAA content in both the nitrite-added control (C1) and the inoculated groups was higher than in the blank control (C0) (p < 0.05). This indicates that both inoculation and nitrite addition can accelerate protein degradation. Furthermore, at the end of fermentation, the bitter amino acid content of the LR1 and LR2 groups was reduced by 22.33% and 13.93%, respectively, as compared to the C1 group (p < 0.05). A similar reduction in bitter amino acids was previously noted by Wen et al., which is considered beneficial for the development of favorable taste in the sausages29. In summary, while both inoculation and nitrite addition enhance total FAA release, inoculation inhibits the accumulation of bitter amino acids, which is conducive to improving the taste of the product.
a Contents of total FAAs and each class of FAA. Data are presented as mean ± SD. The different letters represent significant differences among samples. b Heatmap of each FAA: the size of the circles represents the content of each FAA across different samples, while the color intensity of the squares reflects the relative abundance of various FAAs within the same sample.
Regarding individual amino acids (Fig. 3b), glutamic acid (Glu) and alanine (Ala) were identified as the predominant FAAs across all groups. a compositional pattern consistently observed across all experimental groups and in agreement with earlier findings30. This suggests that inoculation with Lcb. rhamnosus H7 optimizes the content of FAAs while preserving the overall amino acid composition dominated by Glu and Ala, thereby helping to maintain the typical amino acid profile and inherent flavor characteristics of dry fermented sausages. In addition, inoculation led to a marked reduction in the levels of leucine (Leu), valine (Val), and isoleucine (Ile) (p < 0.05). Studies have shown that these amino acids serve as key precursors for the characteristic flavor compounds of dry fermented sausages. They can be converted into VFCs such as aldehydes, acids, and alcohols through metabolic pathways like deamination and decarboxylation. Consequently, Lcb. rhamnosus H7 appears to enhance the flavor complexity of the sausages by facilitating the metabolic conversion of specific precursor amino acids without disrupting the product’s typical amino acid profile.
Bacterial diversity analysis
Figure 4a–c illustrates the bacterial community dynamics of dry fermented sausages. At the phylum level (Fig. 4a), Firmicutes emerged as the dominant phylum, with its relative abundance increasing throughout fermentation to exceed 95% by the endpoint. This predominance aligns with observations in other fermented meat products, such as Chinese dry-cured ham31. At the genus level (Fig. 4b), the relative abundance of Lacticaseibacillus was significantly higher in the inoculated groups (p < 0.05) and showed a positive correlation with the inoculation level. Correspondingly, at the species level (Fig. 4c), Lcb. rhamnosus showed markedly higher relative abundance in the inoculated groups. These suggests that Lcb. rhamnosus H7 successfully colonized the sausage matrix. Several studies have demonstrated the potential of Lcb. rhamnosus as a starter culture to enhance the flavor of fermented foods. For example, Sionek et al. reported that inoculation with Lcb. rhamnosus LOCK90 increased the levels of acids, aldehydes, and esters, thereby enhancing the flavor profile of dry fermented sausages12. Liang et al. observed that Lcb. rhamnosus L08 intensified the flavor of blue honeysuckle juice while reducing its bitterness32. In the present study, the successful establishment of Lcb. rhamnosus H7 as a dominant population likely serves as a foundational factor for the observed flavor enhancement.
a The level of phylum. b The level of genus. c The level of species.
As fermentation progressed, the relative abundance of three genera—Lactococcus, Weissella, and Latilactobacillus—increased markedly, rising from an initial range of 7.36%–74.00% to 22.55%–94.22%, becoming predominant in both the inoculated and control groups (Fig. 4b). Species-level identification (Fig. 4c) further revealed that these genera were represented primarily by Lactococcus lactis, Weissella hellenica, and Latilactobacillus sakei, respectively. Lactococcus, Latilactobacillus, and Weissella are key functional microbial groups that contribute significantly to the flavor development in fermented sausages33,34. Specifically, L. lactis contributes to the formation of aldehydes, alcohols, and organic acids, and also exhibits probiotic properties35. Lat. sakei demonstrates a strong competitive advantage in the dry fermented sausage environment, efficiently fermenting carbohydrates to produce lactic and acetic acids, thereby influencing the fermentation process and significantly enhancing the overall aroma36. Meanwhile, W. hellenica promotes flavor generation by facilitating the release of FAAs, which serve as precursors for VFCs such as acetaldehyde and ethanol37,38. Notably, at the end of fermentation, although Lacticaseibacillus exhibited higher abundance in the inoculated groups due to the inoculation of Lcb. rhamnosus H7, the relative abundance of Weissella (predominantly W. hellenica) remained comparable across all groups. Specifically, W. hellenica accounted for 8.54%, 10.80%, and 9.08% in the C1, LR1, and LR2 groups, respectively. These results suggest that the colonization of Lcb. rhamnosus H7 was compatible with the persistence of indigenous functional bacteria, creating a symbiotic community environment that supported their co-occurrence and metabolic activity.
Furthermore, inoculation was associated with a marked decrease in the relative abundance of spoilage-associated taxa, such as Pseudocitrobacter and Klebsiella (Fig. 4b). This pattern was also evident at the species level for Pseudomonas psychrophila, Hafnia alvei, Pantoea agglomerans, and Klebsiella oxytoca. Pseudocitrobacter, H. alvei, P. agglomerans, and K. oxytoca all belong to the Enterobacteriaceae family and are common spoilage bacteria in food, readily causing meat product deterioration39,40,41. Meanwhile, P. psychrophila, as a psychrophilic bacterium, has also been reported as a key species contributing to meat spoilage42. The observed reduction in their relative abundance suggests that Lcb. rhamnosus H7 exerts a competitive exclusion effect against these undesirable microorganisms.
In summary, inoculation with Lcb. rhamnosus H7 induced a favorable shift in the microbial community, characterized by the successful establishment of the starter strain, the preservation of key indigenous functional groups, and a reduced relative abundance of several spoilage-associated taxa.
Lipid composition analysis
As shown in Fig. 5a, a total of 1218 lipids were detected in dry fermented sausages, classified into 14 categories: phosphatidylcholine (PC), sphingomyelin (SM), lyso-phospatidylcholines (LPC), triacylglycerol (TG), diglyceride (DG), phosphatidylethanolamin (PE), lyso-phosphatidylethanolamine (LPE), free fatty acids (FA), ceramides (Cer), hexosylceramide (HexCer), phosphatidylinositol (PI), phosphatidylglycerol (PG), cholesteryl ester (CE), and phosphatidylserine (PS). Following fermentation, a significant decrease in the relative content of PC and PE was observed. This decrease appeared more pronounced in the inoculated groups compared to the C1 group. The degradation of phospholipids such as PC and PE during fermentation is a recognized phenomenon, often associated with the activity of microbial or endogenous enzymes, and has been linked to flavor development as these lipids can serve as precursors for volatile compounds43,44. Thus, the observed reduction in PC and PE is consistent with typical fermentation-induced lipid metabolism, and the data suggest that inoculation with Lcb. rhamnosus H7 may have promoted this process.
a Relative contents of each lipid class. b Partial least squares-discriminant analysis (PLS-DA) of lipids. c Venn diagram of differential lipids (VIP > 1) at day 3 and day 12 in each group. d Heatmap showing the contents of common differential lipids among the inoculated groups as identified by Venn diagram analysis.
Furthermore, PCA was performed to analyze the overall lipid profile of the dry fermented sausages (Fig. 5b). The results showed that at day 0 and day 3, samples from different groups were closely clustered with substantial overlap, indicating minimal differences in lipid composition. By day 12, however, the LR1 and LR2 groups separated from each other and were clearly distinguished from the C1 group. The results indicate that the addition of Lcb. rhamnosus H7 significantly affects the lipid profile of dry fermented sausages, with its effect primarily occurring during the later stages of fermentation. This pattern aligns with findings from other studies where starter cultures induced more pronounced lipid profile shifts during the mid-to-late stages of meat fermentation45. Additionally, different addition levels also lead to variations in the lipid profile. Based on this, we employed a PLS-DA model to screen for differential lipids in each group at days 3 and 12 of fermentation. As shown in Fig. 5c, 720, 772, 747, and 700 differential lipids were identified in the C0, C1, LR1, and LR2 groups, respectively. Among these, the inoculated groups had 37 unique differential lipids, including PE, TG, PC, Cer, SM, HexCer, DG, and PI (Fig. 5d). Specifically, compared to the C1 group, inoculation promoted the accumulation of PC 40:1 and Cer 42:5. The accumulation of PC 40:1, a phospholipid containing unsaturated fatty acids, may be associated with phospholipid transformation processes, a process that can influence subsequent lipid oxidation and flavor generation pathways as suggested in prior research46. The increased abundance of Cer 42:5 may result from fat decomposition during fermentation47. As a key component of sphingolipids, Cer plays an important role in maintaining cell membrane integrity48. Therefore, its increased level may improve the textural properties and storage stability of the product. Thus, inoculation facilitates the accumulation of PC 40:1 and Cer 42:5, possibly exerting positive effects on both the quality and flavor of dry fermented sausages. Furthermore, PE 36:4, PE 32:2, PC O-27:0, PE 40:6, TG 58:8, PE 42:6, and PI 36:0 were degraded in the inoculated groups during the later stages of fermentation. The fatty acyl chains in these PE species contain multiple unsaturated double bonds, making them susceptible to further degradation into aromatic compounds, thereby contributing to the development of meat product flavor49. TG can be further converted into DG, and both can undergo hydrolysis to release free fatty acids, promoting the formation of flavor compounds50. Collectively, these shifts in specific lipid species suggest that inoculation with Lcb. rhamnosus H7 is associated with a modulation of the lipid metabolic network, potentially providing more precursor substances for the generation of flavor compounds.
Correlation analysis of microorganisms, lipids, flavors, and FAAs driven by inoculation
LAB metabolize proteins and lipids in meat through a variety of enzymatic systems, including phospholipases, lipoxygenases, decarboxylases, and transaminases. These enzymatic activities significantly influence fatty acid metabolism and amino acid catabolism, contributing to the formation of distinctive flavor compounds in the final product. Therefore, based on Pearson correlation analysis, we investigated the relationships among the differential flavor metabolites (Fig. 2e), differential lipids (Fig. 5d), amino acids, and core microbiota following inoculation. As shown in Fig. 6a, Lcb. rhamnosus H7 showed a positive correlation with W. hellenica and Weissella minor. Previous studies have demonstrated that inoculation of starter culture can exert a promoting effect on other microorganisms in fermented foods. For instance, Guo et al. reported that inoculation with Yarrowia lipolytica promotes the growth of LAB, thereby enhancing the production of ethyl esters and improving product flavor51. Similarly, Zou et al. confirmed that the introduction of LAB increases bacterial abundance within the fermentation system, accelerating the formation of key flavor compounds52. Given the established role of Weissella in meat fermentation, particularly in pathways related to lipid and amino acid metabolism53, we hypothesize that the inoculation of Lcb. rhamnosus H7 may be related to the proliferation of Weissella, suggesting a potential microbial association within the metabolic network underlying flavor formation.
a The network diagram shows the correlations among key microorganisms, FAAs, and lipids that are closely associated with the influence of Lcb. rhamonosus H7 on flavor development of sausage. The thicker the line, the stronger the correlation between the substances. b The heatmap shows the contents of key microorganisms, FAAs, and lipids. c The schematic reveals the speculative pathway by which Lcb. rhamonosus H7 modulates FAAs, lipids, and microbial metabolism, thereby enhancing flavor development in dry fermented sausages. PLA Phospholipase A, DC decarboxylase, DH dehydrogenase, AT transaminase, LOX Lipoxygenase, ES esterase, AAT alcohol acyltransferase.
Further analysis revealed that Lcb. rhamnosus H7, W. hellenica, and W. minor were positively correlated with tyrosine (Tyr) and cysteine (Cys), and negatively correlated with PE 40:6, PE 42:6, and TG 58:8 (Fig. 6a). During fermentation, PE 40:6, PE 42:6, and TG 58:8 gradually degraded, with significantly lower levels observed in the inoculated groups compared to the C1 group, whereas Tyr and Cys continuously accumulated, with a greater increase in the inoculated groups (Fig. 6b). This inverse correlation suggests a potential link between the presence of these microorganisms and the observed shifts in lipid and amino acid. For instance, Tan et al. discovered that during refrigerated storage of sturgeon fillets, extracellular enzymes (phospholipase A (PLA)) produced by Aeromonas sobria can mediate the interconversion of PE and PC54, while lipases catalyze the hydrolysis of TG into DG and FAs55. Consistent with this, our data showed that PE 40:6, PE 42:6, and PC 40:1 were negatively correlated, as were TG 58:8 and DG 31:1 (Fig. 6a). Given the observed increase in the levels of PC 40:1 and DG 31:1 during fermentation, along with their significantly higher accumulation in the inoculated groups (Fig. 6b), it is postulated that inoculation is correlated with the degradation of PE 40:6 and PE 42:6 into PC 40:1, as well as the hydrolysis of TG 58:8 into DG 31:1, likely mediated by enzymatic systems such as phospholipases and lipases.
Furthermore, PC 40:1, DG 31:1, Tyr, and Cys exhibited positive correlations with most differential flavor metabolites that were elevated in inoculated groups (Fig. 6a, b). Concurrently, multiple free fatty acids (FA 18:1, FA 18:0, FA 13:0, FA 17:2, FA 15:0, FA 17:0, and FA 20:0) increased after fermentation, with higher concentrations observed in the inoculated groups. Theoretically, amino acids can be converted into α-keto acids under the catalysis of transaminase (AT). Subsequently, they are transformed into alcohols and acids under the action of decarboxylase (DC) and dehydrogenase (DH), and finally converted into ester compounds via the catalysis of esterase (ES) and alcohol acyltransferase (AAT)56. Meanwhile, lipoxygenase (LOX) can oxidize FAs to generate flavor compounds such as alcohols, acids, esters, and hydrocarbons57. Therefore, the correlation network links Lcb. rhamnosus H7 inoculation with shifts in core microbiota (W. hellenica and W. minor), concurrent changes in amino acid and lipid profiles, and the enhanced accumulation of characteristic flavor compounds. This integrated view suggests that inoculation may coordinate a microbial-metabolic network that contributes to the improved flavor profile of low-nitrite dry fermented sausages.
Based on the correlation network analysis, we propose a hypothetical pathway through which Lcb. rhamnosus H7 promotes flavor formation. Inoculation likely promote the growth of W. hellenica and W. minor, microorganisms that likely possess a rich repertoire of endogenous enzymes—such as phospholipase A (PLA), DC, DH, AT, and LOX—acting synergistically to modulate metabolism. Specifically, in amino acid metabolism, microbial-secreted AT may catalyze the conversion of Tyr and Cys into α-keto acids. Subsequently, these α-keto acids are further transformed into alcohols (e.g., phenylethanol) and acids (e.g., hexanoic acid) by DC and DH. The resulting alcohols and acids then undergo esterification, mediated by ES and AAT, to yield ester compounds such as ethyl 2,4-hexadienoate. In lipid metabolism, microbial-derived PLA may catalyze the hydrolysis of PE 40:6 and PE 42:6, releasing free fatty acids and generating PC 40:1. Concurrently, lipases may facilitate the hydrolysis of TG 58:8 into DG 31:1 and FAs. These released FAs undergo oxidation catalyzed by LOX, leading to the formation of various VFCs. While this coordinated microbial-metabolic network constitutes a plausible hypothesis for flavor formation driven by Lcb. rhamnosus H7 inoculation, future studies employing metagenomics, metatranscriptomics, or targeted enzyme activity measurements are warranted to validate the functional roles of these microorganisms and the specific enzymatic mechanisms underlying these correlated metabolic changes.
In this study, by integrating amino acid analysis, lipidomics, and microbial community profiling, we investigated the potential role of the nitrite-reducing strain Lcb. rhamnosus H7 in flavor formation in dry fermented sausages. GC-MS and E-nose analyses revealed that inoculation with Lcb. rhamnosus H7 significantly altered the volatile flavor profile of the final product, as evidenced by a significant increase in the levels of characteristic alcohols (e.g., phenylethanol), acids (e.g., acetic acid), esters (e.g., ethyl hexanoate), and terpenes (e.g., D-limonene). High-throughput sequencing demonstrated that Lcb. rhamnosus H7 became the dominant microbial population throughout fermentation and supported the maintenance of the ecological niche for Weissella in the sausage matrix. Amino acid analysis revealed that inoculation promoted the release of FAAs and enhanced the metabolism of branched-chain amino acids including Leu, Val, and Ile. Lipidomics analysis indicated that inoculation was linked to the accumulation of PC 40:1 and Cer 42:5, along with accelerated degradation of lipids such as PE 36:4, PE 32:2, and TG 58:8. Integrated correlation analysis further suggested that Lcb. rhamnosus H7 may influence flavor formation by modulating the microbial community structure and coordinately regulating amino acid and lipid metabolic pathways. Collectively, these multi-omics correlations provide a plausible hypothesis for how nitrite-reducing LAB could regulate flavor development and offer a theoretical basis for the targeted modulation of flavor profiles in fermented meat products. It should be noted that the insights presented here are primarily derived from correlation analyses, which do not establish causality. Future work should therefore validate the proposed metabolic pathways through enzyme activity assays, genetic manipulation, or metagenomics to clarify the mechanistic role of Lcb. rhamnosus H7.
Methods
Materials and chemicals
Fresh lean pork and pork backfat were obtained from the local market (Dalian, Liaoning, China). Lcb. rhamnosus H7 was isolated from chouguiyu, it has been preserved in the Guangdong Microbial Culture Collection Center (Guangdong, China) with the accession number GDMCC No. 64655. The 16S rRNA gene sequence of the strain is provided in the Supplementary Information (Table S1).
Chloroform was purchased from Macklin (Shanghai, China). Acetonitrile, methanol, and isopropyl alcohol were obtained from Sigma-ALdrich (Madison, WI, USA). Ammonium acetate was purchased from Aladdin Biochemical Technology Co., Ltd. (Shanghai, China). All reagents mentioned above were of HPLC grade. 1,2-dichlorobenzene (GC-grade) was purchased from Aladdin Biochemical Technology Co., Ltd. (Shanghai, China).
Lipid standards, including phosphatidylcholine (PC 34:0), phosphatidylglycerol (PG 34:0), phosphatidylethanolamine (PE 34:0), phosphatidylinositol (PI 36:0), and triacylglycerol (TG 51:0) were obtained from Sigma ALdrich (Msdison, WI, USA). Gas chromatography (GC) standards consisting of an alkane standard mixture (C7–C30) in hexane were obtained from Aladdin Biochemical Technology Co., Ltd. (Shanghai, China). Amino acid standards containing 17 kinds of amino acids were purchased from Wako Pure Chemical Industries Ltd. (Osaka, Japan).
Sampling
The preparation of dry fermented sausages was carried out according to the method described by Yue et al.58. Lcb. rhamnosus H7 was cultured at 37 °C for 24 h, after which the cells were harvested by centrifugation at 8000 rpm for 5 min. The bacterial pellet was washed twice to remove residual growth medium and then re-suspended in sterile saline to a final concentration exceeding 10⁸ CFU/mL. The experiment was conducted in three independent production batches (biological replicates). For each batch, fresh lean pork and pork backfat were mixed with seasonings and homogenized thoroughly. The homogenized mixture from each batch was subsequently divided into four equal portions. Each portion was assigned to one of the following four experimental treatments: (1) no bacterial inoculum or nitrite added (C0 group); (2) nitrite (100 mg/kg) only (C1 group); (3) nitrite (100 mg/kg) and Lcb. rhamnosus H7 (10⁶ CFU/g) added (LR1 group);. (4) nitrite (100 mg/kg) and Lcb. rhamnosus H7 (107 CFU/g) added (LR2 group). Each portion was then stuffed into pig casings to produce sausages. The sausages were fermented for 12 days at 15 °C under 60% relative humidity. From each treatment group within each batch, samples were collected on days 0, 3, and 12 for analyses of VFCs, FAAs, bacterial diversity analysis, and lipidomics. Additionally, the final products from each batch were subjected to E-nose analysis.
E-nose analysis
The E-nose (PEN 3, Win Muster Airsense Analytics Inc., Schwerin, Germany) was used to analyze the flavor profile of the sausages according to the method of Chen et al.59. Briefly, 2 g of minced sample was weighed into a 20 mL vial and equilibrated for 20 min. The measurement parameters were set as follows: a flush time of 60 s, a chamber flow of 300 mL/min, and a measurement time of 100 s.
VFCs analysis
VFCs were determined according to the method of Zhang et al., with minor modifications60. Two grams of each sample were homogenized and transferred to a 20 mL headspace vial. 1,2-dichlorobenzene was added as an internal standard. The VFCs were extracted using a SPME fiber (Supelco Inc., Bellefonte, AL, USA) coated with a 50/30 µm composite layer of divinylbenzene, carboxen, and polydimethylsiloxane (DVB/CAR/PDMS) at 60 °C for 40 min. VFCs were analyzed using a gas chromatography-mass spectrometry system (GC-MS) system (7890B–5977B, Agilent Technologies, Inc., Santa Clara, CA, USA) system equipped with a VF-WAX capillary column (30 m × 0.25 mm × 0.25 µm). A C7–C30 n-alkane standard mixture was used to calculate the retention indices (RIs) by comparing their retention times under identical GC conditions to those used for sample analysis. Data analysis was performed using MassHunter Unknowns Analysis software (Agilent Technologies, Inc., Santa Clara, CA, USA). Qualitative identification of individual volatile compounds was achieved by matching the obtained mass spectra against the NIST11 mass spectral database.
FAAs analysis
FAAs of sausages were analyzed using an LA808 automatic amino acid analyzer (Hitachi, Japan) according to the method reported by Pei et al.61. Briefly, 3 g of the sample was blended with 12 mL of deionized water using a homogenizer. After that, the protein was precipitated by adding 5 mL of acetone to 1 mL of the homogenate. The homogenate was centrifuged and the supernatant was evaporated to dryness under a stream of nitrogen gas. The residue was then reconstituted in 1 mL of HCl and filtered through a 0.22 µm membrane filter prior to analysis. A mixed amino acid standard was utilized for both qualitative and quantitative analysis of amino acids in the samples.
Bacterial diversity analysis
The bacterial diversity of dry fermented sausages was analyzed according to the method of Lv et al.62. Total genomic DNA was extracted from the sausage samples using the TGuide S96 Magnetic Soil/Stool DNA Kit (Tiangen, Beijing, China). After the quality and quantity of the DNA were assessed, the full-length 16S rRNA gene was amplified using barcoded universal bacterial primers 27F (AGRGTTTGATYNTGGCTCAG) and 1492R (TASGGHTACCTTGTTASGACTT). The resulting amplicons were purified, quantified, pooled in equal amounts, and used for SMRTbell library preparation. Sequencing was performed on the PacBio platform (Pacific Biosciences, CA, USA). Bioinformatic analysis was performed using the platform provided by Biomarker Technologies (Beijing, China). The sequencing data have been uploaded to NCBI with the accession number of PRJNA1264108.
Lipidomics analysis
Lipids were extracted from the sausage samples with a chloroform/methanol mixture (2:1, v/v). For the analysis of lipid composition, an ultra-high-performance liquid chromatography (UHPLC) system coupled with a triple quadrupole linear ion trap mass spectrometer (QTRAP 5500, AB SCIEX, Concord, Canada) was employed, according to the method of Liu et al.63. Chromatographic separation was performed on a C8 column (ACQUITY UPLC® BEH C8, 1.7 µm, 2.1 × 100 mm; Waters, Milford, MA, USA). Lipid internal standards were used for the semi-quantitative analysis of lipids in the samples.
Statistical analysis
The experiments were conducted in three independent batches (biological replicates). Data are expressed as mean values ± standard deviation (SD, n = 3 independent batches). For data involving both treatment and time, a two-way ANOVA was performed. If the interaction was significant (p < 0.05), all treatment-time combinations were compared using Duncan’s test. PCA and partial least squares discriminant analysis (PLS-DA) were performed using SIMCA version 14.1 (Umetrics, Sweden). Heatmaps were generated using TBtools. Bar charts were created using Origin 2021 (OriginLab Corporation, USA) and GraphPad Prism version 10.4.2 (GraphPad Software, La Jolla, CA, USA). Correlation analysis was performed with SPSS 23.0 (IBM Corp., Armonk, NY, USA).
Ethics and inclusion statement
This study did not involve human participants, animal subjects, or data collection from any individual or community. Therefore, ethical approval and an inclusion statement were not required for this work.
Data availability
The data that support the findings of this study are available from the corresponding authors upon reasonable request.
Code availability
This study did not generate or utilize any custom computer code or algorithms. Therefore, a code availability statement is not applicable.
References
Froehlich, D. A., Gullett, E. A. & Usborne, W. R. Effect of nitrite and salt on the color, flavor and overall acceptability of ham. J. Food Sci. 48, 152–154 (1983).
Deveci, G. & Tek, N. A. N-nitrosamines: a potential hazard in processed meat products. J. Sci. Food Agric. 104, 13102 (2024).
Wang, Y. et al. Research update on the impact of lactic acid bacteria on the substance metabolism, flavor, and quality characteristics of fermented meat products. Foods 11, 2090 (2022).
Guo, T. et al. Microbiota interactions as critical determinants of flavor development in fermented foods. Trends Food Sci. Technol. 164, 105218 (2025).
Hierro, E., De la Hoz, L. & Ordóñez, J. A. Contribution of microbial and meat endogenous enzymes to the lipolysis of dry fermented sausages. J. Agric. Food Chem. 45, 2989–2995 (1997).
Dong, Y. Y., Liu, Y., He, N., Li, X. D. & Zhu, Y. C. Comprehensive analysis of the effect of Staphylococcus xylosus YCC3 fermentation on bacterial community, flavor and lipid metabolism of fermented sausages. Food Biosci. 69, 106920 (2025).
Shao, X. F. et al. Decoding the flavor regulation mechanism of fermented sausages inoculated with indigenous strains via metagenomic and GC-MS analysis. LWT 206, 116604 (2024).
Feng, L. et al. Changes of unique flavor substances and metabolic pathway brought by Lacticaseibacillus rhamnosus WH. FH-19 fermented milk during fermentation and storage stage: HS-SPME-GC-MS and HPLC-MS-based analysis. Food Biosci. 65, 105974 (2025).
Dzandu, B., Chotiko, A. & Sathivel, S. Antioxidant activity and viability of Lacticaseibacillus rhamnosus, Lacticaseibacillus casei, and co-culture in fermented tomato juice during refrigerated storage. Food Biosci. 50, 102085 (2022).
Alemneh, S. T., Emire, S. A. & Hitzmann, B. Teff-based probiotic functional beverage fermented with Lactobacillus rhamnosus and Lactobacillus plantarum. Foods 10, 2333 (2021).
Woldemariam, K. Y. et al. Impact of lactic acid bacteria on the flavor profile of salami and role of microbial diversity and functional gene. J. Food Biochem. 2024, 6073261 (2024).
Sionek, B. et al. Effects of Lacticaseibacillus rhamnosus LOCK900 on development of volatile compounds and sensory quality of dry fermented sausages. Molecules 26, 6454 (2021).
Yin, X. Y. et al. Characterization of selected Harbin red sausages on the basis of their flavour profiles using HS-SPME-GC/MS combined with electronic nose and electronic tongue. Meat Sci. 172, 108345 (2021).
Yin, X. Y. et al. Collaborative analysis on differences in volatile compounds of Harbin red sausages smoked with different types of woodchips based on gas chromatography–mass spectrometry combined with electronic nose. LWT 143, 111144 (2021).
Xing, B. F. et al. Flavor evolution of normal- and low-fat Chinese sausage during natural fermentation. Food Res. Int. 169, 112937 (2023).
Sun, X. H., Hong, H., Jia, S. L., Liu, Y. M. & Luo, Y. K. Effects of phytic acid and lysozyme on microbial composition and quality of grass carp (Ctenopharyngodon idellus) fillets stored at 4 °C. Food Microbiol. 86, 103313 (2020).
Mi, R. F. et al. Predominant yeasts in Chinese Dong fermented pork (Nanx Wudl) and their aroma-producing properties in fermented sausage condition. Food Sci. Hum. Wellness 10, 231–240 (2021).
Wang, J. et al. Dynamic change of bacterial diversity, metabolic pathways, and flavor during ripening of the Chinese fermented sausage. Front. Microbiol. 13, 990606 (2022).
Hao, S. Q. et al. Characteristic flavor analysis of inner mongolia air-dried meat and the impact of vacuum tumbling curing on flavor. J. Food Biochem. 2024, 4077505 (2024).
Marco, A., Navarro, J. L. & Flores, M. Quantitation of selected odor-active constituents in dry fermented sausages prepared with different curing salts. J. Agric. Food Chem. 55, 3058–3065 (2007).
Montanari, C. et al. Effects of the diameter on physico-chemical, microbiological and volatile profile in dry fermented sausages produced with two different starter cultures. Food Biosci. 22, 9–18 (2018).
Fan, W. & Qian, M. C. Headspace solid phase microextraction and gas chromatography-olfactometry dilution analysis of young and aged Chinese “Yanghe Daqu” liquors. J. Agric. Food Chem. 53, 7931–7938 (2005).
Hu, Y. Y., Wang, H., Kong, B. H., Wang, Y. & Chen, Q. The succession and correlation of the bacterial community and flavour characteristics of Harbin dry sausages during fermentation. LWT 138, 110689 (2021).
Perea-Sanz, L., Montero, R., Belloch, C. & Flores, M. Microbial changes and aroma profile of nitrate reduced dry sausages during vacuum storage. Meat Sci. 147, 100–107 (2019).
Ozkara, K. T., Amanpour, A., Guclu, G., Kelebek, H. & Selli, S. GC-MS-olfactometric differentiation of aroma-active compounds in turkish heat-treated sausages by application of aroma extract dilution analysis. Food Anal. Methods 12, 729–741 (2019).
Zhang, Y. L., Lu, Y. Y. & Chen, F. S. Relationship between physicochemical properties and microbial structural distribution of Chinese-style and Salami fermented sausages. Food Biosci. 53, 102583 (2023).
Wang, H. P. et al. Analysis of biogenic amines in salt-reduced Harbin dry sausages: from the perspective of free amino acid profile and bacterial community composition. LWT 216, 117327 (2025).
Aro Aro, J. M. et al. The effect of starter cultures on proteolytic changes and amino acid content in fermented sausages. Food Chem. 119, 279–285 (2010).
Wen, R. X. et al. Co-inoculation of Debaryomyces hansenii and lactic acid bacteria: a strategy to improve the taste and odour profiles of dry sausages. Food Sci. Hum. Wellness 13, 3273–3283 (2024).
Xiao, Y. Q., Liu, Y. N., Chen, C. G., Xie, T. T. & Li, P. J. Effect of Lactobacillus plantarum and Staphylococcus xylosus on flavour development and bacterial communities in Chinese dry fermented sausages. Food Res. Int. 135, 109247 (2020).
Zhang, J. et al. Physicochemical property, volatile flavor quality, and microbial community composition of Jinhua fatty ham and lean ham: a comparative study. Front. Microbiol. 14, 1124770 (2023).
Liang, S. N. et al. Blue honeysuckle fermentation with Lacticaseibacillus rhamnosus L08 improves its biological activity, sensory and flavor characteristics, and storage stability. Food Chem. X 23, 101659 (2024).
Hu, Y. Y. et al. The potential correlation between bacterial diversity and the characteristic volatile flavour of traditional dry sausages from Northeast China. Food Microbiol. 91, 103505 (2020).
Wang, X. H., Zhang, Y. L., Ren, H. Y. & Zhan, Y. Comparison of bacterial diversity profiles and microbial safety assessment of salami, Chinese dry-cured sausage and Chinese smoked-cured sausage by high-throughput sequencing. LWT 90, 108–115 (2018).
Chen, L. et al. Enhancement of the flavor compound 3-Methylbutanal and sensory quality in sausages fermented by Lactococcus lactis CGMCC 31087: Leucine and α-Ketoglutaric acid as supplements. Food Chem. 473, 143091 (2025).
Zheng, S. S. et al. Innovative insights into dry fermented sausages flavor: unraveling the impact of varied lactobacillus genera-driven fermentation. Food Biosci. 60, 104331 (2024).
Huang, L. Y. et al. Thyme essential oil and sausage diameter effects on biogenic amine formation and microbiological load in smoked horse meat sausage. Food Biosci. 40, 100885 (2021).
Hu, Y. Y. et al. Improving the taste profile of reduced-salt dry sausage by inoculating different lactic acid bacteria. Food Res. Int. 145, 110391 (2021).
Guo, L. N. et al. Dynamic fermentation quality and bacterial community structure of paper mulberry silage from three regions of China. Chem. Bio. Technol. Agric. 10, 51 (2023).
Castaño, A., Garcı́a Fontán, M. C., Fresno, J. M., Tornadijo, M. E. & Carballo, J. Survival of Enterobacteriaceae during processing of Chorizo de cebolla, a Spanish fermented sausage. Food Control 13, 107–115 (2002).
Vasilopoulos, C. et al. Technology-induced selection towards the spoilage microbiota of artisan-type cooked ham packed under modified atmosphere. Food Microbiol. 27, 77–84 (2010).
Jia, S. L. et al. Insights into the fish protein degradation induced by the fish-borne spoiler Pseudomonas psychrophila and Shewanella putrefaciens: From whole genome sequencing to quality changes. Int. J. Food Microbiol. 416, 110675 (2024).
Gao, F. et al. Lipidomics assessment of lipid changes during the processing of fermented sausages. Available at SSRN (2023).
Wang, J. et al. Decoding the mechanism of flavor improvement mediated by inoculated starter cultures in low-salt HuangYuXiang: from lipids to volatile flavor compounds. Food Biosci. 71, 107251 (2025).
Gao, F. et al. Characterization of the effect of Lactobacillus helveticus IMAUJBH1 on lipid profiles and volatile flavors of mutton fermented sausages in Inner Mongolia based on lipomics and GC-MS. LWT 202, 116290 (2024).
Yin, H. W. et al. Characterization of the differences in lipid profiles and volatile compounds of adipose stem cells adipogenic differentiation and adipocytic transdifferentiation of muscle stem cells from large yellow croakers based on UPLC-MS/MS and GC-IMS. Food Chem. 470, 142658 (2025).
Qu, L. Y., Zhao, Y., Xu, X. D., Li, Y. F. & Lv, H. X. Untargeted lipidomics reveal quality changes in high-moisture japonica brown rice at different storage temperatures. Foods 12, 4218 (2023).
Dong, X., Shao, J., Wu, X. Y., Dong, J. L. & Tang, P. A. Lipidomic profiling reveals the protective mechanism of nitrogen-controlled atmosphere on brown rice quality during storage. Food Chem. 473, 143081 (2025).
Hou, X. H. et al. Metabolomics and lipidomics profiles related to intramuscular fat content and flavor precursors between Laiwu and Yorkshire pigs. Food Chem. 404, 134699 (2023).
Zhang, M., Fu, J. J., Mao, J. L., Dong, X. P. & Chen, Y. W. Lipidomics reveals the relationship between lipid oxidation and flavor formation of basic amnio acids participated low-sodium cured large yellow croaker. Food Chem. 429, 136888 (2023).
Guo, B. R. et al. Inoculation of Yarrowia lipolytica promotes the growth of lactic acid bacteria, Debaryomyces udenii and the formation of ethyl esters in sour meat. Food Microbiol. 119, 104447 (2024).
Zou, J. H. et al. The effect of lactic acid bacteria as a starter on the microbial community and flavors of the fermented beef-soybean paste. Food Chem. 484, 144328 (2025).
Yin, H. M. et al. Dynamic changes in volatile profiles and bacterial communities during natural fermentation of Mei yu, traditional Chinese fermented fish pieces. Food Res. Int. 194, 114882 (2024).
Tan, C. M. et al. Insights into the molecular mechanisms of lipid transformation in sturgeon fillets: interplay between specific spoilage and dominant bacteria. Food Chem. X 23, 101714 (2024).
Liu, Y. et al. Evaluation of changes in egg yolk lipids during storage based on lipidomics through UPLC-MS/MS. Food Chem. 398, 133931 (2023).
Zhao, S. S. et al. The regulation of volatile flavor compounds in fermented meat products mediated by microorganisms: a review. Food Biosci. 62, 105180 (2024).
Wang, J. et al. Unraveling the metabolic network of volatile flavor dynamics development during the golden pomfret fermentation driven by core bacteria: a combined multi-omics analysis. Food Microbiol. 133, 104864 (2026).
Yue, Y. et al. Flavour enhancement of dry fermented sausages by nitrite-degrading Levilactobacillus brevis CHOL1: combining flavouromics and lipidomics to elucidate the mechanism of aroma formation. Food Chem. 473, 143119 (2025).
Chen, X. H. et al. Aroma characterization of Sichuan and Cantonese sausages using electronic nose, gas chromatography–mass spectrometry, gas chromatography-olfactometry, odor activity values and metagenomic. Food Chem. X 24, 101924 (2024).
Zhang, X. H. et al. Insights into the balance of safety, flavor, and texture in fermented fish: a study based on full-length sequencing of fermented largemouth bass (Micropterus salmoides). Int. J. Food Microbiol. 439, 111240 (2025).
Pei, Q. X. et al. Exploring the synergistic effect of Lactiplantibacillus plantarum 1-24-LJ and lipase on improving quality, flavor, and safety of Suanzharou. Food Res. Int. 200, 115432 (2025).
Lv, J. et al. Effect of Saccharomyces cerevisiae LXPSC1 on microorganisms and metabolites of sour meat during the fermentation. Food Chem. 402, 134213 (2023).
Liu, M. Y. et al. Enhanced torularhodin production in Rhodosporidium toruloides A1-15 under salt stress: insights from multi-omics analysis. Food Biosci. 63, 105590 (2025).
Acknowledgements
This work was supported by the Xing Liao Talent Program - Outstanding Young Talents (XLYC2403129).
Author information
Authors and Affiliations
Contributions
Ying Yue: formal analysis, investigation, data curation, writing—original draft, writing—review & editing and conceptualization. Sufen Guo: data curation, visualization, investigation, writing—original draft. Hao Liu: investigation and methodology. Ning Zhao: formal analysis and data curation. Xiaohan Jia: visualization and investigation. Ning Wang: investigation and resources. Chaofan Ji: methodology and resources. Yiwei Dai: writing—review and methodology. Beiwei Zhu: writing—review & editing. Xinping Lin: writing—review & editing, supervision and funding acquisition.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yue, Y., Guo, S., Liu, H. et al. Lacticaseibacillus rhamnosus H7 shapes flavor-associated microbial-metabolic networks in low-nitrite sausages: insights from a multi-omics correlation study. npj Sci Food 10, 110 (2026). https://doi.org/10.1038/s41538-026-00757-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41538-026-00757-z