Abstract
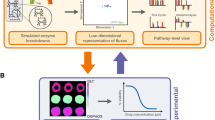
KRAS-mutant colorectal cancer (CRC) undergoes metabolic reprogramming that promotes tumor progression and drug resistance. Cancer-associated fibroblasts (CAFs), a major component of the tumor microenvironment (TME), play a pivotal role in modulating these metabolic adaptations in CRC. This study applies flux sampling combined with representation learning and hierarchical clustering to a computational model of central carbon metabolism to understand how CAFs influence KRAS-mutant CRC metabolic reprogramming following targeted enzyme knockdowns. Focusing on 12 key enzymes involved in glycolysis and the pentose phosphate pathway, knockdowns were simulated under normal CRC media and CAF-conditioned media (CCM) conditions. Analysis revealed CCM induces greater metabolic heterogeneity, with knockdown models exhibiting more variable and distinct metabolic states compared to those cultured in normal CRC media, indicating CAF-derived factors diversify the metabolic responses of CRC cells to enzyme perturbations. Pathway-level flux analysis demonstrated media-specific shifts in central carbon metabolism. Predicted biomass flux showed enzyme knockdowns reduced growth across both conditions, but CCM models indicated a protective effect against perturbation. Overall, simulations illustrated CCM enhances the metabolic adaptability of KRAS-mutant CRC cells to perturbations, emphasizing the importance of including TME components in metabolic modeling and therapeutic development and suggesting that targeting tumor-CAF metabolic interactions may improve treatment strategies.

Similar content being viewed by others
Data availability
All relevant datasets and scripts for analysis are available on GitHub at: https://github.com/FinleyLabUSC/CRC_Flux_Sampling.
References
Biller, L. H. & Schrag, D. Diagnosis and treatment of metastatic colorectal cancer: a review. JAMA 325, 669–685 (2021).
Ye, J. et al. Tissue gene mutation profiles in patients with colorectal cancer and their clinical implications. Biomed. Rep. 13, 43–48 (2020).
Meng, M. et al. The current understanding on the impact of KRAS on colorectal cancer. Biomed. Pharmacother. 140, 111717 (2021).
Tria, S. M., Burge, M. E. & Whitehall, V. L. J. The therapeutic landscape for KRAS-mutated colorectal cancers. Cancers 15, 2375 (2023).
Ma, Q., Zhang, W., Wu, K. & Shi, L. The roles of KRAS in cancer metabolism, tumor microenvironment and clinical therapy. Mol. Cancer 24, 14 (2025).
Heiden, M. G. V. & DeBerardinis, R. J. Understanding the intersections between metabolism and cancer biology. Cell 168, 657–669 (2017).
Baghban, R. et al. Tumor microenvironment complexity and therapeutic implications at a glance. Cell Commun. Signal. 18, 59 (2020).
Liang, L. et al. ‘Reverse Warburg effect’ of cancer‑associated fibroblasts (Review). Int. J. Oncol. 60, 67 (2022).
Pang, B. & Wu, H. Metabolic reprogramming in colorectal cancer: a review of aerobic glycolysis and its therapeutic implications for targeted treatment strategies. Cell Death Discov. 11, 321 (2025).
Wang, J. et al. Elucidating tumor-stromal metabolic crosstalk in colorectal cancer through integration of constraint-based models and LC-MS metabolomics. Metab. Eng. 69, 175–187 (2022).
Tavakoli, N. et al. Merging metabolic modeling and imaging for screening therapeutic targets in colorectal cancer. Npj Syst. Biol. Appl. 11, 12 (2025).
Orth, J. D., Thiele, I. & Palsson, B. Ø What is flux balance analysis? Nat. Biotechnol. 28, 245–248 (2010).
Sahu, A., Blätke, M. A., Szymański, J. J. & Töpfer, N. Advances in flux balance analysis by integrating machine learning and mechanism-based models. Comput. Struct. Biotechnol. J. 19, 4626–4640 (2021).
Sebastian, C. et al. A non-dividing cell population with high pyruvate dehydrogenase kinase activity regulates metabolic heterogeneity and tumorigenesis in the intestine. Nat. Commun. 13, 1503 (2022).
Kubicka, A., Matczak, K. & Łabieniec-Watała, M. More than meets the eye regarding cancer metabolism. Int. J. Mol. Sci. 22, 9507 (2021).
Gelbach, P. E. & Finley, S. D. Genome-scale modeling predicts metabolic differences between macrophage subtypes in colorectal cancer. iScience 26, 107569 (2023).
Galuzzi, B. G. et al. Coupling constrained-based flux sampling and clustering to tackle cancer metabolic heterogeneity. In 2023 31st Euromicro International Conference on Parallel, Distributed and Network-Based Processing (PDP) 185–192 (IEEE, 2023).
Raman, K. & Chandra, N. Flux balance analysis of biological systems: applications and challenges. Brief. Bioinform. 10, 435–449 (2009).
Markert, E. K. & Vazquez, A. Mathematical models of cancer metabolism. Cancer Metab. 3, 14 (2015).
Kim, J. & DeBerardinis, R. J. Mechanisms and implications of metabolic heterogeneity in cancer. Cell Metab. 30, 434–446 (2019).
Hirose, Y. & Taniguchi, K. Intratumoral metabolic heterogeneity of colorectal cancer. Am. J. Physiol. Cell Physiol. 325, C1073–C1084 (2023).
Chhabra, Y. & Weeraratna, A. T. Fibroblasts in cancer: unity in heterogeneity. Cell 186, 1580–1609 (2023).
Cheng, P. S. W., Zaccaria, M. & Biffi, G. Functional heterogeneity of fibroblasts in primary tumors and metastases. Trends Cancer 11, 135–153 (2025).
Cess, C. G. & Finley, S. D. Calibrating agent-based models to tumor images using representation learning. PLoS Comput. Biol. 19, e1011070 (2023).
Hendlisz, A. et al. The prognostic significance of metabolic response heterogeneity in metastatic colorectal cancer. PLoS ONE 10, e0138341 (2015).
Li, Z., Sun, C. & Qin, Z. Metabolic reprogramming of cancer-associated fibroblasts and its effect on cancer cell reprogramming. Theranostics 11, 8322–8336 (2021).
TeSlaa, T., Ralser, M., Fan, J. & Rabinowitz, J. D. The pentose phosphate pathway in health and disease. Nat. Metab. 5, 1275–1289 (2023).
Tu, C.-E. et al. D-ribose-5-phosphate inactivates YAP and functions as a metabolic checkpoint. J. Hematol. Oncol, 18, 2 (2025).
Moscardó García, M., Pacheco, M., Bintener, T., Presta, L. & Sauter, T. Importance of the biomass formulation for cancer metabolic modeling and drug prediction. iScience 24, 103110 (2021).
Bordel, S. Constraint based modeling of metabolism allows finding metabolic cancer hallmarks and identifying personalized therapeutic windows. Oncotarget 9, 19716–19729 (2018).
Gatto, F., Miess, H., Schulze, A. & Nielsen, J. Flux balance analysis predicts essential genes in clear cell renal cell carcinoma metabolism. Sci. Rep. 5, 10738 (2015).
Bhome, R. et al. Exosomal microRNAs derived from colorectal cancer-associated fibroblasts: role in driving cancer progression. Aging 9, 2666–2694 (2017).
Horváthová, J. et al. Inhibition of glycolysis suppresses cell proliferation and tumor progression in vivo: perspectives for chronotherapy. Int. J. Mol. Sci. 22, 4390 (2021).
Hyroššová, P. et al. Effects of metabolic cancer therapy on tumor microenvironment. Front. Oncol. 12, 1046630 (2022).
Cordani, M. et al. The role of glycolysis in tumorigenesis: from biological aspects to therapeutic opportunities. Neoplasia 58, 101076 (2024).
Koh, B., Jeon, H., Kim, D., Kang, D. & Kim, K. R. Effect of fibroblast co-culture on the proliferation, viability and drug response of colon cancer cells. Oncol. Lett. 17, 2409–2417 (2019).
AlMusawi, S., Ahmed, M. & Nateri, A. S. Understanding cell-cell communication and signaling in the colorectal cancer microenvironment. Clin. Transl. Med. 11, e308 (2021).
Rinaldi, G., Rossi, M. & Fendt, S.-M. Metabolic interactions in cancer: cellular metabolism at the interface between the microenvironment, the cancer cell phenotype and the epigenetic landscape. WIREs Syst. Biol. Med. 10, e1397 (2018).
Wang, X. et al. Overcoming cancer treatment resistance: unraveling the role of cancer-associated fibroblasts. J. Natl. Cancer Cent. 5, 237–251 (2025).
Hu, J. L. et al. CAFs secreted exosomes promote metastasis and chemotherapy resistance by enhancing cell stemness and epithelial-mesenchymal transition in colorectal cancer. Mol. Cancer 18, 91 (2019).
Wang, W., Cheng, B. & Yu, Q. Cancer-associated fibroblasts as accomplices to confer therapeutic resistance in cancer. Cancer Drug Resist. 5, 889–901 (2022).
De, P., Aske, J., Sulaiman, R. & Dey, N. Bête Noire of chemotherapy and targeted therapy: CAF-mediated resistance. Cancers 14, 1519 (2022).
Zhu, C. et al. Targeting KRAS mutant cancers: from druggable therapy to drug resistance. Mol. Cancer 21, 159 (2022).
László, L. et al. Recent updates on the significance of KRAS mutations in colorectal cancer biology. Cells 10, 667 (2021).
Kaemmerer, E., Loessner, D. & Avery, V. M. Addressing the tumour microenvironment in early drug discovery: a strategy to overcome drug resistance and identify novel targets for cancer therapy. Drug Discov. Today 26, 663–676 (2021).
McMillin, D. W., Negri, J. M. & Mitsiades, C. S. The role of tumour–stromal interactions in modifying drug response: challenges and opportunities. Nat. Rev. Drug Discov. 12, 217–228 (2013).
Brown, R. E., Short, S. P. & Williams, C. S. Colorectal cancer and metabolism. Curr. Colorectal Cancer Rep. 14, 226–241 (2018).
Hon, K. W., Zainal Abidin, S. A., Othman, I. & Naidu, R. The crosstalk between signaling pathways and cancer metabolism in colorectal cancer. Front. Pharmacol. 12, 768861 (2021).
Gong, J. et al. Reprogramming of lipid metabolism in cancer-associated fibroblasts potentiates migration of colorectal cancer cells. Cell Death Dis. 11, 267 (2020).
Zhang, D. et al. Metabolic reprogramming of cancer-associated fibroblasts by IDH3α downregulation. Cell Rep. 10, 1335–1348 (2015).
ZHONG, J. et al. The remodeling roles of lipid metabolism in colorectal cancer cells and immune microenvironment. Oncol. Res. 30, 231–242 (2023).
Herrmann, H. A., Dyson, B. C., Vass, L., Johnson, G. N. & Schwartz, J.-M. Flux sampling is a powerful tool to study metabolism under changing environmental conditions. Npj Syst. Biol. Appl. 5, 32 (2019).
Fallahi, S., Skaug, H. J. & Alendal, G. A comparison of Monte Carlo sampling methods for metabolic network models. PLoS ONE 15, e0235393 (2020).
Kaufman, D. E. & Smith, R. L. Direction choice for accelerated convergence in hit-and-run sampling. Oper. Res. 46, 84–95 (1998).
Megchelenbrink, W., Huynen, M. & Marchiori, E. optGpSampler: an improved tool for uniformly sampling the solution-space of genome-scale metabolic networks. PLoS ONE 9, e86587 (2014).
Gelbach, P. E., Cetin, H. & Finley, S. D. Flux sampling in genome-scale metabolic modeling of microbial communities. BMC Bioinformatics 25, 45 (2024).
Chicco, D. Siamese Neural Networks: An Overview. in Artificial Neural Networks (ed. Cartwright, H.) 73–94 (Springer US, New York, NY, 2021).
Zheng, L., Duffner, S., Idrissi, K., Garcia, C. & Baskurt, A. Siamese multi-layer perceptrons for dimensionality reduction and face identification. Multimed. Tools Appl. 75, 5055–5073 (2016).
Hoffer, E. & Ailon, N. Deep Metric Learning Using Triplet Network. in Similarity-Based Pattern Recognition (eds Feragen, A., Pelillo, M. & Loog, M.) 84–92 (Springer International Publishing, Cham, 2015).
Szubert, B., Cole, J. E., Monaco, C. & Drozdov, I. Structure-preserving visualisation of high dimensional single-cell datasets. Sci. Rep. 9, 8914 (2019).
Acknowledgements
The authors thank the members of the Finley Lab at USC for the deep scientific discussions and critical feedback.
Author information
Authors and Affiliations
Contributions
E.E. conducted all metabolic modeling and computational analyses. N.T. and H.C. contributed to scientific discussions of results. E.E. and S.D.F. contributed to manuscript writing and scientific discussions of results. S.D.F. was responsible for the scientific direction of this work. All authors reviewed and provided feedback on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Elton, E., Tavakoli, N., Cetin, H. et al. Cancer-associated fibroblasts drive metabolic heterogeneity in colorectal cancer cells: predictions from metabolic modeling. npj Syst Biol Appl (2026). https://doi.org/10.1038/s41540-026-00673-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41540-026-00673-8