Abstract
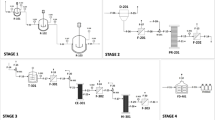
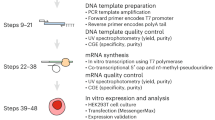
mRNA vaccines have emerged as a transformative modality for infectious disease prevention. In response to the emergence of SARS-CoV-2, large-scale in vitro transcription (IVT) of mRNA vaccines was developed. Large-scale IVT currently relies on linearized plasmid DNA (pDNA) as a template for mRNA production. Linearized pDNA production presents several challenges at manufacturing scale, including removal of residual host-cell DNA, protein, endotoxins, and antibiotics. Additionally, pDNA-derived sequences irrelevant to mRNA production must be removed from the final product. Finally, the generation of linear pDNA template is laborious, which reduces mRNA production speed, a renowned advantage of this technology. Enzymatic DNA amplification strategies such as rolling circle amplification (RCA) of a synthetic circular DNA molecule offer a rapid, isothermal reaction as an alternative to pDNA. Therefore, we have developed a fully synthetic, single-vessel mRNA manufacturing platform. Beginning with a chemically synthesized circular DNA template, we amplify via a fit-for-purpose RCA, linearize with a TypeIIS restriction enzyme (RE), and perform IVT in a single vessel. The entire process—from circular template to mRNA—can be completed in as little as two days. This method, termed Unified Sequential Template Amplification and Transcription (USTAT), eliminates bacterial components, large volume pDNA production, and enables rapid, modular mRNA production.
Similar content being viewed by others
Data availability
The data supporting the findings are contained within the manuscript or supplementary files. Genetic data is present in the supplementary data file. Additional raw data are available from the corresponding author upon request.
References
Xenopoulos, A. & Pattnaik, P. Production and purification of plasmid DNA vaccines: Is there scope for further innovation?. Expert Rev. Vaccines 13, 1537–1551 (2014).
Piao, X. et al. Supercoiled DNA percentage: a key in-process control of linear DNA template for mRNA drug substance manufacturing. Mol. Ther. Nucleic Acids 35, 102223 (2024).
Dean, F. B., Nelson, J. R., Giesler, T. L. & Lasken, R. S. Rapid amplification of plasmid and phage DNA using Phi29 DNA polymerase and multiply-primed rolling circle amplification. Genome Res. 11, 1095–1099 (2001).
Fidelity of phi 29 DNA polymerase. Comparison between protein-primed initiation and DNA polymerization. J. Biol. Chem. 268, 2719–2726 (1993).
Garmendia, C., Bernad, A., Esteban, J. A., Blanco, L. & Salas, M. The bacteriophage phi 29 DNA polymerase, a proofreading enzyme. J. Biol. Chem. 267, 2594–2599 (1992).
Kumar, G. & Chernaya, G. Cell-free protein synthesis using multiply-primed rolling circle amplification products. BioTechniques 47, 637–639 (2009).
van Emmerik, C. L. et al. Ramified rolling circle amplification for synthesis of nucleosomal DNA sequences. Anal. Biochem. 588, 113469 (2020).
Engler, C., Gruetzner, R., Kandzia, R. & Marillonnet, S. Golden gate shuffling: a one-pot DNA shuffling method based on type IIs restriction enzymes. PLoS One 4, e5553 (2009).
Bauer, R. J. et al. Comparative analysis of the end-joining activity of several DNA ligases. PLoS One 12, e0190062 (2017).
Strzelecki, P. et al. Enhanced Golden Gate Assembly: evaluating overhang strength for improved ligation efficiency. Nucleic Acids Res. 52, e95 (2024).
Shi, K. et al. T4 DNA ligase structure reveals a prototypical ATP-dependent ligase with a unique mode of sliding clamp interaction. Nucleic Acids Res. 46, 10474–10488 (2018).
Inoue, H., Nojima, H. & Okayama, H. High efficiency transformation of E. coli with plasmids. Gene 96, 23–28 (1990).
Miyoshi, D., Karimata, H. & Sugimoto, N. The effect of 1,2-propanediol on DNA structure and stability. Nucleic Acids Symposium Series 203–204 (Oxford University Press, 2006).
Ali, M. M. et al. Rolling circle amplification: a versatile tool for chemical biology, materials science and medicine. Chem. Soc. Rev. 43, 3324–3341 (2014).
Lizardi, P. M. et al. Mutation detection and single-molecule counting using isothermal rolling-circle amplification. Nat. Genet. 19, 225–232 (1998).
Baner, J., Nilsson, M., Mendel-Hartvig, M. & Landegren, U. Signal amplification of padlock probes by rolling circle replication. Nucleic Acids Res. 26, 5073–5078 (1998).
Garafutdinov, R. R., Sakhabutdinova, A. R., Gilvanov, A. R. & Chemeris, A. V. Rolling circle amplification as a universal method for the analysis of a wide range of biological targets. Russian J. Bioorg. Chem. 47, 1172–1189 (2021).
Lenk, R. et al. Understanding the impact of in vitro transcription byproducts and contaminants. Front. Mol. Biosc. 11, 1426129 (2024).
Levy, M. S. et al. Effect of shear on plasmid DNA in solution. Bioprocess Eng. 20, 7–13 (1999).
Levy, M. S., O’Kennedy, R. D., Ayazi-Shamlou, P. & Dunnill, P. Biochemical engineering approaches to the challenges of producing pure plasmid DNA. Trends Biotechnol. 18, 296–305 (2000).
Nair, V. K., Sharma, C. & Ghosh, S. Mitigating the non-specific rolling circle amplification: a comprehensive assessment of the role of ligation and digestion methods. ChemRxiv https://doi.org/10.26434/chemrxiv-2022-mxtb3-v3 (2022).
Inoue, J., Shigemori, Y. & Mikawa, T. Improvements of rolling circle amplification (RCA) efficiency and accuracy using Thermus thermophilus SSB mutant protein. Nucleic Acids Res. 34, e69 (2006).
Baker, Y. R. et al. Expanding the chemical functionality of DNA nanomaterials generated by rolling circle amplification. Nucleic Acids Res. 49, 9042–9052 (2021).
Kieser, R. E. & Budowle, B. Rolling circle amplification: a (random) primer on the enrichment of an infinite linear DNA template. WIREs Forensic Sci. 2, e1359 (2020).
Skerra, A. Phosphorothioate primers improve the amplification of DNA sequences by DNA polymerases with proofreading activity. Nucleic Acids Res. 20, 3551–3554 (1992).
Martín, D. S. et al. Reduced amplification by phi29 DNA polymerase in the presence of unbound oligos during reaction in RCA. Biosens. Bioelectron. X 17, 100456 (2024).
Lagunavicius, A., Kiveryte, Z., Zimbaite-Ruskuliene, V., Radzvilavicius, T. & Janulaitis, A. Duality of polynucleotide substrates for Phi29 DNA polymerase: 3′→5′ RNase activity of the enzyme. RNA 14, 503–513 (2008).
Blanco, L. & Salas, M. Characterization and purification of a proofreading DNA polymerase from bacteriophage ϕ29. J. Biol. Chem. 259, XX (1984).
Krzywkowski, T., Kühnemund, M., Wu, D. & Nilsson, M. Limited reverse transcriptase activity of phi29 DNA polymerase. Nucleic Acids Res. 46, 3625–3632 (2018).
Gao, Y., Wolf, L. K. & Georgiadis, R. M. Secondary structure effects on DNA hybridization kinetics: a solution versus surface comparison. Nucleic Acids Res. 34, 3370–3377 (2006).
Klenow, H. & Henningsen, I. Effect of monovalent cations on the activity of the DNA polymerase of Escherichia coli B. Eur. J. Biochem. 9, 133–141 (1969).
Gohara, D. W. & Di Cera, E. Molecular mechanisms of enzyme activation by monovalent cations. J. Biol. Chem. 291, 20840–20848 (2016).
Sun, Y. et al. Unraveling the salt tolerance of Phi29 DNA polymerase using compartmentalized self-replication and microfluidics platform. Front. Microbiol. 14, 1267196 (2023).
Maity, A., Singh, A. & Singh, N. Differential stability of DNA based on salt concentration. Eur. Biophys. J. 46, 33–40 (2017).
Wolffs, P., Grage, H., Hagberg, O. & Rådström, P. Impact of DNA polymerases and their buffer systems on quantitative real-time PCR. J. Clin. Microbiol. 42, 408–411 (2004).
Nishimoto, N., Suzuki, M. & Izuta, S. Effect of pH on the misincorporation rate of DNA polymerase η. Biol. Pharm. Bull. 39, 953–958 (2016).
KA, E. & TA, K. DNA polymerase fidelity and the polymerase chain reaction. PCR Methods Appl. 1, 17–24 (1991).
Stephen, C. & Mishanina, T. V. Alkaline pH has an unexpected effect on transcriptional pausing during synthesis of the Escherichia coli pH-responsive riboswitch. J. Biol. Chem. 298, 102302 (2022).
Özay, B. & McCalla, S. E. A review of reaction enhancement strategies for isothermal nucleic acid amplification reactions. Sens. Actuators Rep. 3, 100033 (2021).
Mok, E., Wee, E., Wang, Y. & Trau, M. Comprehensive evaluation of molecular enhancers of the isothermal exponential amplification reaction. Sci. Rep. 6, 37837 (2016).
Karunanathie, H., Kee, P. S., Ng, S. F., Kennedy, M. A. & Chua, E. W. PCR enhancers: types, mechanisms, and applications in long-range PCR. Biochimie 197, 130–143 (2022).
Chevet, E., Lemaître, G. & Katinka, M. D. Low concentrations of tetramethylammonium chloride increase yield and specificity of PCR. Nucleic Acids Res. 23, 3343–3344 (1995).
Wang, J. et al. Exponential amplification of DNA with very low background using graphene oxide and single-stranded binding protein to suppress non-specific amplification. Microchim. Acta 182, 1095–1101 (2015).
Wartell, R. M. & Benight, A. S. Thermal denaturation of DNA molecules: a comparison of theory with experiment. Phys. Rep. 126, 67–107 (1985).
von Hippel, P. H. & Berg, O. G. On the specificity of DNA-protein interactions. Proc. Natl. Acad. Sci. USA 83, 1608–1612 (1986).
Chen, Y. G. & Hur, S. Cellular origins of dsRNA, their recognition and consequences. Nat. Rev. Mol. Cell Biol. 23, 286–301 (2021).
Baiersdörfer, M. A facile method for the removal of dsRNA contaminant from in vitro-transcribed mRNA. Mol. Ther. Nucleic Acids 15, 26–35 (2019).
Sharabrin, S. V. et al. Removal of double-stranded RNA contaminants during template-directed synthesis of mRNA. Bull. Exp. Biol. Med. 176, 751–755 (2024).
Popova, P. G. et al. Effect of in vitro transcription conditions on yield of high quality messenger and self-amplifying RNA. Eur. J. Pharm. Biopharm. 198, 114247 (2024).
Acknowledgements
We thank Erik Barton and members of Pfizer Bioprocess R&D for their thoughtful input and review of the manuscript. No funding was received for this research. The authors declare no competing financial or non-financial interests.
Author information
Authors and Affiliations
Contributions
S.G, C.L.S., S.D.R., C.H.C., J.W.S., K.N., M.J.S., and J.R. conceptualized ideas and experiments presented in this work. S.G, C.L.S., S.D.R., C.H.C., J.W.S., K.N., M.J.S., and J.R. performed experiments presented or related to this work to develop conclusions. S.G, C.L.S., S.D.R., C.H.C., J.W.S., K.N., M.J.S., and J.R. wrote the main manuscript. S.G, C.L.S., S.D.R., C.H.C., J.W.S., K.N., M.J.S., and J.R. prepared figures. S.G, C.L.S., S.D.R., C.H.C., J.W.S., K.N., M.J.S., and J.R. reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ghosh, S., Simms, C.L., Rohrer, S.D. et al. USTAT: Unified Sequential Template Amplification and Transcription—a fully synthetic mRNA manufacturing platform. npj Vaccines (2026). https://doi.org/10.1038/s41541-026-01434-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41541-026-01434-8