Abstract
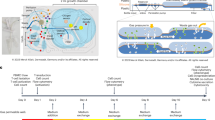
Chimaeric antigen receptor (CAR) T cells can generate durable clinical responses in B-cell haematologic malignancies. The manufacturing of these T cells typically involves their activation, followed by viral transduction and expansion ex vivo for at least 6 days. However, the activation and expansion of CAR T cells leads to their progressive differentiation and the associated loss of anti-leukaemic activity. Here we show that functional CAR T cells can be generated within 24 hours from T cells derived from peripheral blood without the need for T-cell activation or ex vivo expansion, and that the efficiency of viral transduction in this process is substantially influenced by the formulation of the medium and the surface area-to-volume ratio of the culture vessel. In mouse xenograft models of human leukaemias, the rapidly generated non-activated CAR T cells exhibited higher anti-leukaemic in vivo activity per cell than the corresponding activated CAR T cells produced using the standard protocol. The rapid manufacturing of CAR T cells may reduce production costs and broaden their applicability.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The main data supporting the findings of this study are available within the article and its Supplementary Information. All raw data generated during the study are available from the corresponding authors on reasonable request. Source data are provided with this paper.
References
Salter, A. I., Pont, M. J. & Riddell, S. R. Chimeric antigen receptor-modified T cells: CD19 and the road beyond. Blood 131, 2621–2629 (2018).
Brudno, J. N. et al. T cells genetically modified to express an anti-B-cell maturation antigen chimeric antigen receptor cause remissions of poor-prognosis relapsed multiple myeloma. J. Clin. Oncol. 36, 2267–2280 (2018).
Cohen, A. D. et al. B cell maturation antigen-specific CAR T cells are clinically active in multiple myeloma. J. Clin. Investig. 129, 2210–2221 (2019).
D’Angelo, S. P. et al. Antitumor activity associated with prolonged persistence of adoptively transferred NY-ESO-1 (c259)T cells in synovial sarcoma. Cancer Discov. 8, 944–957 (2018).
Raje, N. et al. Anti-BCMA CAR T-cell therapy bb2121 in relapsed or refractory multiple myeloma. N. Engl. J. Med. 380, 1726–1737 (2019).
Kalos, M. et al. T cells with chimeric antigen receptors have potent antitumor effects and can establish memory in patients with advanced leukemia. Sci. Transl. Med. 3, 95ra73 (2011).
Ghassemi, S. et al. Reducing ex vivo culture improves the antileukemic activity of chimeric antigen receptor (CAR) T cells. Cancer Immunol. Res. 6, 1100–1109 (2018).
Berger, C. et al. Adoptive transfer of effector CD8+ T cells derived from central memory cells establishes persistent T cell memory in primates. J. Clin. Investig. 118, 294–305 (2008).
Graef, P. et al. Serial transfer of single-cell-derived immunocompetence reveals stemness of CD8+ central memory T cells. Immunity 41, 116–126 (2014).
Hinrichs, C. S. et al. Adoptively transferred effector cells derived from naive rather than central memory CD8+ T cells mediate superior antitumor immunity. Proc. Natl Acad. Sci. USA 106, 17469–17474 (2009).
Hinrichs, C. S. et al. Human effector CD8+ T cells derived from naive rather than memory subsets possess superior traits for adoptive immunotherapy. Blood 117, 808–814 (2011).
Klebanoff, C. A. et al. Memory T cell-driven differentiation of naive cells impairs adoptive immunotherapy. J. Clin. Invest. 126, 318–334 (2016).
Crompton, J. G. et al. Akt inhibition enhances expansion of potent tumor-specific lymphocytes with memory cell characteristics. Cancer Res. 75, 296–305 (2015).
van der Waart, A. B. et al. Inhibition of Akt signaling promotes the generation of superior tumor-reactive T cells for adoptive immunotherapy. Blood 124, 3490–3500 (2014).
Gattinoni, L. et al. Wnt signaling arrests effector T cell differentiation and generates CD8+ memory stem cells. Nat. Med. 15, 808–813 (2009).
Gattinoni, L. et al. A human memory T cell subset with stem cell-like properties. Nat. Med. 17, 1290–1297 (2011).
Muralidharan, S. et al. Activation of Wnt signaling arrests effector differentiation in human peripheral and cord blood-derived T lymphocytes. J. Immunol. 187, 5221–5232 (2011).
Naldini, L. et al. In vivo gene delivery and stable transduction of nondividing cells by a lentiviral vector. Science 272, 263–267 (1996).
Plesa, G. et al. Addition of deoxynucleosides enhances human immunodeficiency virus type 1 integration and 2LTR formation in resting CD4+ T cells. J. Virol. 81, 13938–13942 (2007).
Swiggard, W. J. et al. Human immunodeficiency virus type 1 can establish latent infection in resting CD4+ T cells in the absence of activating stimuli. J. Virol. 79, 14179–14188 (2005).
Amirache, F. et al. Mystery solved: VSV-G-LVs do not allow efficient gene transfer into unstimulated T cells, B cells, and HSCs because they lack the LDL receptor. Blood 123, 1422–1424 (2014).
Finkelshtein, D., Werman, A., Novick, D., Barak, S. & Rubinstein, M. LDL receptor and its family members serve as the cellular receptors for vesicular stomatitis virus. Proc. Natl Acad. Sci. USA 110, 7306–7311 (2013).
Agosto, L. M. et al. The CXCR4-tropic human immunodeficiency virus envelope promotes more-efficient gene delivery to resting CD4+ T cells than the vesicular stomatitis virus glycoprotein G envelope. J. Virol. 83, 8153–8162 (2009).
Pace, M. J., Agosto, L. & O’Doherty, U. R5 HIV env and vesicular stomatitis virus G protein cooperate to mediate fusion to naive CD4+ T cells. J. Virol. 85, 644–648 (2011).
Unutmaz, D., KewalRamani, V. N., Marmon, S. & Littman, D. R. Cytokine signals are sufficient for HIV-1 infection of resting human T lymphocytes. J. Exp. Med. 189, 1735–1746 (1999).
Cavalieri, S. et al. Human T lymphocytes transduced by lentiviral vectors in the absence of TCR activation maintain an intact immune competence. Blood 102, 497–505 (2003).
Trinite, B., Chan, C. N., Lee, C. S. & Levy, D. N. HIV-1 Vpr- and reverse transcription-induced apoptosis in resting peripheral blood CD4 T cells and protection by common gamma-chain cytokines. J. Virol. 90, 904–916 (2016).
Verhoeyen, E. et al. IL-7 surface-engineered lentiviral vectors promote survival and efficient gene transfer in resting primary T lymphocytes. Blood 101, 2167–2174 (2003).
Korin, Y. D. & Zack, J. A. Nonproductive human immunodeficiency virus type 1 infection in nucleoside-treated G0 lymphocytes. J. Virol. 73, 6526–6532 (1999).
Rausell, A. et al. Innate immune defects in HIV permissive cell lines. Retrovirology 13, 43 (2016).
Descours, B. et al. SAMHD1 restricts HIV-1 reverse transcription in quiescent CD4+ T-cells. Retrovirology 9, 87 (2012).
Pierson, T. C. et al. Molecular characterization of preintegration latency in human immunodeficiency virus type 1 infection. J. Virol. 76, 8518–8531 (2002).
Milone, M. C. et al. Chimeric receptors containing CD137 signal transduction domains mediate enhanced survival of T cells and increased antileukemic efficacy in vivo. Mol. Ther. 17, 1453–1464 (2009).
Burnie, J. & Guzzo, C. The incorporation of host proteins into the external HIV-1 envelope. Viruses 11, 85 (2019).
Eyquem, J. et al. Targeting a CAR to the TRAC locus with CRISPR/Cas9 enhances tumour rejection. Nature 543, 113–117 (2017).
Wilen, C. B., Tilton, J. C. & Doms, R. W. HIV: cell binding and entry. Cold Spring Harb. Perspect. Med. 2, a006866 (2012).
Brown, M. S. & Goldstein, J. L. A receptor-mediated pathway for cholesterol homeostasis. Science 232, 34–47 (1986).
Tran, R. et al. Microfluidic transduction harnesses mass transport principles to enhance gene transfer efficiency. Mol. Ther. 25, 2372–2382 (2017).
Maude, S. L. et al. Chimeric antigen receptor T cells for sustained remissions in leukemia. N. Engl. J. Med. 371, 1507–1517 (2014).
Berry, C. C. et al. INSPIIRED: quantification and visualization tools for analyzing integration site distributions. Mol. Ther. Methods Clin. Dev. 4, 17–26 (2017).
Sherman, E. et al. INSPIIRED: a pipeline for quantitative analysis of sites of new DNA integration in cellular genomes. Mol. Ther. Methods Clin. Dev. 4, 39–49 (2017).
Berry, C. C. et al. Estimating abundances of retroviral insertion sites from DNA fragment length data. Bioinformatics 28, 755–762 (2012).
Nobles, C. L. et al. CD19-targeting CAR T cell immunotherapy outcomes correlate with genomic modification by vector integration. J. Clin. Invest. 130, 673–685 (2019).
Schroder, A. R. et al. HIV-1 integration in the human genome favors active genes and local hotspots. Cell 110, 521–529 (2002).
Mitchell, R. S. et al. Retroviral DNA integration: ASLV, HIV, and MLV show distinct target site preferences. PLoS Biol. 2, E234 (2004).
Wang, G. P., Ciuffi, A., Leipzig, J., Berry, C. C. & Bushman, F. D. HIV integration site selection: analysis by massively parallel pyrosequencing reveals association with epigenetic modifications. Genome Res. 17, 1186–1194 (2007).
Brady, T. et al. HIV integration site distributions in resting and activated CD4+ T cells infected in culture. AIDS 23, 1461–1471 (2009).
Haining, W. N. et al. Antigen-specific T-cell memory is preserved in children treated for acute lymphoblastic leukemia. Blood 106, 1749–1754 (2005).
Halliwell, B. Cell culture, oxidative stress, and antioxidants: avoiding pitfalls. Biomed. J. 37, 99–105 (2014).
Zhang, H. et al. 4-1BB is superior to CD28 costimulation for generating CD8+ cytotoxic lymphocytes for adoptive immunotherapy. J. Immunol. 179, 4910–4918 (2007).
Klebanoff, C. A. et al. Determinants of successful CD8+ T-cell adoptive immunotherapy for large established tumors in mice. Clin. Cancer Res. 17, 5343–5352 (2011).
Klebanoff, C. A. et al. Inhibition of AKT signaling uncouples T cell differentiation from expansion for receptor-engineered adoptive immunotherapy. JCI Insight 2, e95103 (2017).
O’Doherty, U., Swiggard, W. J. & Malim, M. H. Human immunodeficiency virus type 1 spinoculation enhances infection through virus binding. J. Virol. 74, 10074–10080 (2000).
Baldauf, H. M. et al. SAMHD1 restricts HIV-1 infection in resting CD4+ T cells. Nat. Med. 18, 1682–1687 (2012).
Mauney, C. H., Perrino, F. W. & Hollis, T. Identification of inhibitors of the dNTP triphosphohydrolase SAMHD1 using a novel and direct high-throughput assay. Biochemistry 57, 6624–6636 (2018).
Trobridge, G. D. et al. Cocal-pseudotyped lentiviral vectors resist inactivation by human serum and efficiently transduce primate hematopoietic repopulating cells. Mol. Ther. 18, 725–733 (2010).
Klebanoff, C. A. et al. Central memory self/tumor-reactive CD8+ T cells confer superior antitumor immunity compared with effector memory T cells. Proc. Natl Acad. Sci. USA 102, 9571–9576 (2005).
Pollok, K. E., Kim, S. H. & Kwon, B. S. Regulation of 4-1BB expression by cell–cell interactions and the cytokines, interleukin-2 and interleukin-4. Eur. J. Immunol. 25, 488–494 (1995).
Lelek, M. et al. Chromatin organization at the nuclear pore favours HIV replication. Nat. Commun. 6, 6483 (2015).
Wong, R. W., Mamede, J. I. & Hope, T. J. Impact of nucleoporin-mediated chromatin localization and nuclear architecture on HIV integration site selection. J. Virol. 89, 9702–9705 (2015).
Couzin-Frankel, J. Supply of promising T cell therapy is strained. Science 356, 1112–1113 (2017).
O’Connor, R. S. et al. Substrate rigidity regulates human T cell activation and proliferation. J. Immunol. 189, 1330–1339 (2012).
De Spiegelaere, W. et al. Quantification of integrated HIV DNA by repetitive-sampling Alu-HIV PCR on the basis of poisson statistics. Clin. Chem. 60, 886–895 (2014).
Liszewski, M. K., Yu, J. J. & O’Doherty, U. Detecting HIV-1 integration by repetitive-sampling Alu-gag PCR. Methods 47, 254–260 (2009).
Kenderian, S. S. et al. CD33-specific chimeric antigen receptor T cells exhibit potent preclinical activity against human acute myeloid leukemia. Leukemia 29, 1637–1647 (2015).
Acknowledgements
We thank the University of Pennsylvania Cell and Vaccine Production Facility for technical support as well as the University of Pennsylvania Stem Cell and Xenograft Core and University of Pennsylvania Flow Cytometry Core and Human Immunology Core, which is supported by NIH grants nos AI045008 and CA016520, for providing de-identified human T cells. We thank J. K. Everett and A. M. Roche for their technical expertise. This work was supported in part through funding provided by Novartis Pharmaceuticals through a research alliance with the University of Pennsylvania (M.C.M.) as well as a St. Baldrick’s Foundation Scholar Award (S.G.), a National Blood Foundation Scientific Research Grant Award (S.G.), the Office of the Assistant Secretary of Defense for Health Affairs through the Peer Reviewed Cancer Research Program under award nos W81XWH-20-1-0417 (S.G.) and RO1CA226983 (R.S.O’C.).
Author information
Authors and Affiliations
Contributions
S.G. and M.C.M. designed the study. S.G., M.C.M., J.S.D., R.S.O’C., U.O’D., F.D.B. and S.I.G. provided conceptual guidance. S.G., J.S.D, R.S.O’C, S.N.-C., J.P., K.D.C., F.S., M.P., G.P., V.A.C., S.R. and J.L. performed the experiments. S.G. and M.C.M. analysed the data. S.G. and M.C.M. wrote the manuscript. R.S.O’C and J.S.D. read and made comments on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
M.C.M. is an inventor on patent applications related to CAR technology (US patents 9,481,728 and 9,499,629 B2) and has received licencing royalties from the Novartis corporation. S.G. and M.C.M. are inventors on a patent application related to methods of manufacturing non-activated CAR T cells (provisional patent PCT/US2020/027734). The other authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks Bryon Johnson and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Non-activated T cells expressing a CD33-specific CAR exhibit an antileukemic effect in vivo in the Aml xenograft model.
a, Schematic of the xenograft model and CART33 cell treatment in NSG mice. b, c, Serial quantification of disease burden by bioluminescence imaging. b, Total bioluminescence flux in mice with no treatment. c, Total bioluminescence flux in mice treated with 5×106 non-activated T cells transduced as in panel (a) (d1), 5×106 CAR + T cells stimulated with anti- CD3/CD28 microbeads and expanded over 9 days (d9), and 5×106 non-transduced (NTD) control non-activated T cells (n = 10 per group). d, e, Absolute peripheral blood CD45+ T cell counts in blood collected from mice shown in panels (c) at d, 13 days (P = 0.0065), and e, 26 days (P = 0.0012) following T cell transfer measured by a TruCount assay. The mean of each group is indicated by the solid black line. Groups were compared using the two-tailed, unpaired Mann–Whitney test.
Supplementary information
Supplementary Information (download PDF )
Supplementary figures and report.
Source data
Source Data Fig. 3 (download XLSX )
Source data for tumour growth.
Source Data Fig. 5 (download XLSX )
Source data for tumour growth.
Source Data Fig. 6 (download XLSX )
Source data for tumour growth.
Source Data Extended Data Fig. 1 (download XLSX )
Source data for tumour growth.
Rights and permissions
About this article
Cite this article
Ghassemi, S., Durgin, J.S., Nunez-Cruz, S. et al. Rapid manufacturing of non-activated potent CAR T cells. Nat Biomed Eng 6, 118–128 (2022). https://doi.org/10.1038/s41551-021-00842-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41551-021-00842-6
This article is cited by
-
Chimeric antigen receptor T‑cell therapy in chronic viral infections: a review
Journal of Translational Medicine (2026)
-
Microsystem technologies for accelerating the discovery and translation of immunotherapies
Nature Reviews Drug Discovery (2026)
-
Strategies and mechanisms for the enhancement of chimeric antigen receptor T-cell functions
Science China Life Sciences (2026)
-
Optimizing viral transduction in immune cell therapy manufacturing: key process design considerations
Journal of Translational Medicine (2025)
-
In vivo CAR-T cell engineering: concept, research progress, potential challenges and enhancement strategies
Experimental Hematology & Oncology (2025)