Abstract
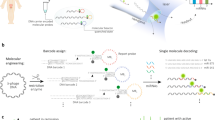
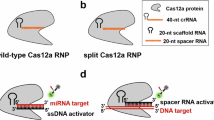
The use of microRNAs as clinical cancer biomarkers is hindered by the absence of accurate, fast and inexpensive assays for their detection in biofluids. Here we report a one-step and one-pot isothermal assay that leverages rolling-circle amplification and the endonuclease Cas12a for the accurate detection of specific miRNAs. The assay exploits the cis-cleavage activity of Cas12a to enable exponential rolling-circle amplification of target sequences and its trans-cleavage activity for their detection and for signal amplification. In plasma from patients with pancreatic ductal adenocarcinoma, the assay detected the miRNAs miR-21, miR-196a, miR-451a and miR-1246 in extracellular vesicles at single-digit femtomolar concentrations with single-nucleotide specificity. The assay is rapid (sample-to-answer times ranged from 20 min to 3 h), does not require specialized instrumentation and is compatible with a smartphone-based fluorescence detection and with the lateral-flow format for visual readouts. Simple assays for the detection of miRNAs in blood may aid the development of miRNAs as biomarkers for the diagnosis and prognosis of cancers.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The main data supporting the findings of this study are available within the paper and its Supplementary Information. The raw and analysed datasets generated during the study are available for research purposes from the corresponding author on reasonable request. Source data are provided with this paper.
References
Ha, M. & Kim, V. N. Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 15, 509–524 (2014).
Di Leva, G., Garofalo, M. & Croce, C. M. MicroRNAs in cancer. Annu. Rev. Pathol. 9, 287–314 (2014).
Inui, M., Martello, G. & Piccolo, S. MicroRNA control of signal transduction. Nat. Rev. Mol. Cell Biol. 11, 252–263 (2010).
Lin, S. & Gregory, R. I. MicroRNA biogenesis pathways in cancer. Nat. Rev. Cancer 15, 321–333 (2015).
Mori, M. A., Ludwig, R. G., Garcia-Martin, R., Brandao, B. B. & Kahn, C. R. Extracellular miRNAs: from biomarkers to mediators of physiology and disease. Cell Metab. 30, 656–673 (2019).
Jet, T., Gines, G., Rondelez, Y. & Taly, V. Advances in multiplexed techniques for the detection and quantification of microRNAs. Chem. Soc. Rev. 50, 4141–4161 (2021).
Dave, V. P. et al. MicroRNA amplification and detection technologies: opportunities and challenges for point of care diagnostics. Lab. Invest. 99, 452–469 (2019).
Kilic, T., Erdem, A., Ozsoz, M. & Carrara, S. microRNA biosensors: opportunities and challenges among conventional and commercially available techniques. Biosens. Bioelectron. 99, 525–546 (2018).
Ouyang, T., Liu, Z., Han, Z. & Ge, Q. MicroRNA detection specificity: recent advances and future perspective. Anal. Chem. 91, 3179–3186 (2019).
Forero, D. A., Gonzalez-Giraldo, Y., Castro-Vega, L. J. & Barreto, G. E. qPCR-based methods for expression analysis of miRNAs. Biotechniques 67, 192–199 (2019).
Hunt, E. A., Broyles, D., Head, T. & Deo, S. K. MicroRNA detection: current technology and research strategies. Annu. Rev. Anal. Chem. 8, 217–237 (2015).
Deng, R., Zhang, K. & Li, J. Isothermal amplification for microRNA detection: from the test tube to the cell. Acc. Chem. Res. 50, 1059–1068 (2017).
Cao, H., Zhou, X. & Zeng, Y. Microfluidic exponential rolling circle amplification for sensitive microRNA detection directly from biological samples. Sens. Actuators B 279, 447–457 (2019).
Yan, H., Xu, Y., Lu, Y. & Xing, W. Reduced graphene oxide-based solid-phase extraction for the enrichment and detection of microRNA. Anal. Chem. 89, 10137–10140 (2017).
Deng, R. et al. Toehold-initiated rolling circle amplification for visualizing individual microRNAs in situ in single cells. Angew. Chem. Int. Ed. Engl. 53, 2389–2393 (2014).
Qian, J., Zhang, Q., Liu, M., Wang, Y. & Lu, M. A portable system for isothermal amplification and detection of exosomal microRNAs. Biosens. Bioelectron. 196, 113707 (2021).
Jia, H. X., Li, Z. P., Liu, C. H. & Cheng, Y. Q. Ultrasensitive detection of microRNAs by exponential isothermal amplification. Angew. Chem. Int. Ed. Engl. 49, 5498–5501 (2010).
Li, C., Li, Z., Jia, H. & Yan, J. One-step ultrasensitive detection of microRNAs with loop-mediated isothermal amplification (LAMP). Chem. Commun. 47, 2595–2597 (2011).
Park, C. R., Park, S. J., Lee, W. G. & Hwang, B. H. Biosensors using hybridization chain reaction—design and signal amplification strategies of hybridization chain reaction. Biotechnol. Bioprocess Eng. 23, 355–370 (2018).
Choi, H. M. et al. Programmable in situ amplification for multiplexed imaging of mRNA expression. Nat. Biotechnol. 28, 1208–1212 (2010).
Wu, H. et al. Label-free and enzyme-free colorimetric detection of microRNA by catalyzed hairpin assembly coupled with hybridization chain reaction. Biosens. Bioelectron. 81, 303–308 (2016).
Cheglakov, Z., Cronin, T. M., He, C. & Weizmann, Y. Live cell microRNA imaging using cascade hybridization reaction. J. Am. Chem. Soc. 137, 6116–6119 (2015).
Aita, A. et al. Serum miRNA profiling for early PDAC diagnosis and prognosis: a retrospective study. Biomedicines 9, 845 (2021).
Shukla, N., Yan, I. K. & Patel, T. Multiplexed detection and quantitation of extracellular vesicle RNA expression using NanoString. Methods Mol. Biol. 1740, 177–185 (2018).
Srinivasan, S., Duval, M. X., Kaimal, V., Cuff, C. & Clarke, S. H. Assessment of methods for serum extracellular vesicle small RNA sequencing to support biomarker development. J. Extracell. Vesicles 8, 1684425 (2019).
Guerau-de-Arellano, M., Alder, H., Ozer, H. G., Lovett-Racke, A. & Racke, M. K. miRNA profiling for biomarker discovery in multiple sclerosis: from microarray to deep sequencing. J. Neuroimmunol. 248, 32–39 (2012).
Hong, L. Z. et al. Systematic evaluation of multiple qPCR platforms, NanoString and miRNA-Seq for microRNA biomarker discovery in human biofluids. Sci. Rep. 11, 4435 (2021).
Kaminski, M. M., Abudayyeh, O. O., Gootenberg, J. S., Zhang, F. & Collins, J. J. CRISPR-based diagnostics. Nat. Biomed. Eng. 5, 643–656 (2021).
Li, Y., Li, S., Wang, J. & Liu, G. CRISPR/Cas systems towards next-generation biosensing. Trends Biotechnol. 37, 730–743 (2019).
Gootenberg, J. S. et al. Nucleic acid detection with CRISPR–Cas13a/C2c2. Science 356, 438–442 (2017).
Chen, J. S. et al. CRISPR–Cas12a target binding unleashes indiscriminate single-stranded DNase activity. Science 360, 436–439 (2018).
Joung, J. et al. Detection of SARS-CoV-2 with SHERLOCK one-pot testing. N. Engl. J. Med. 383, 1492–1494 (2020).
Li, L. et al. HOLMESv2: a CRISPR–Cas12b-assisted platform for nucleic acid detection and DNA methylation quantitation. ACS Synth. Biol. 8, 2228–2237 (2019).
Wang, G., Tian, W., Liu, X., Ren, W. & Liu, C. New CRISPR-derived microRNA sensing mechanism based on Cas12a self-powered and rolling circle transcription-unleashed real-time crRNA recruiting. Anal. Chem. 92, 6702–6708 (2020).
Xu, L. Q. et al. Accurate MRSA identification through dual-functional aptamer and CRISPR–Cas12a assisted rolling circle amplification. J. Microbiol. Methods 173, 105917 (2020).
Zhang, G., Zhang, L., Tong, J. T., Zhao, X. X. & Ren, J. L. CRISPR–Cas12a enhanced rolling circle amplification method for ultrasensitive miRNA detection. Microchem. J. 158, 105239 (2020).
Zhang, M., Wang, H. H., Wang, H., Wang, F. F. & Li, Z. P. CRISPR/Cas12a-assisted ligation-initiated loop-mediated isothermal amplification (CAL-LAMP) for highly specific detection of microRNAs. Anal. Chem. 93, 7942–7948 (2021).
Sun, H. H., He, F., Wang, T., Yin, B. C. & Ye, B. C. A Cas12a-mediated cascade amplification method for microRNA detection. Analyst 145, 5547–5552 (2020).
Ding, X. et al. Ultrasensitive and visual detection of SARS-CoV-2 using all-in-one dual CRISPR–Cas12a assay. Nat. Commun. 11, 4711 (2020).
Li, S. et al. A one-step, one-pot CRISPR nucleic acid detection platform (CRISPR-top): application for the diagnosis of COVID-19. Talanta 233, 122591 (2021).
Lu, S. H. et al. Fast and sensitive detection of SARS-CoV-2 RNA using suboptimal protospacer adjacent motifs for Cas12a. Nat. Biomed. Eng. 6, 286–297 (2022).
Bruch, R. et al. CRISPR/Cas13a‐powered electrochemical microfluidic biosensor for nucleic acid amplification‐free miRNA diagnostics. Adv. Mater. 31, 1970365 (2019).
Zheng, F. et al. A highly sensitive CRISPR-empowered surface plasmon resonance sensor for diagnosis of inherited diseases with femtomolar-level real-time quantification. Adv. Sci. 9, e2105231 (2022).
Hajian, R. et al. Detection of unamplified target genes via CRISPR–Cas9 immobilized on a graphene field-effect transistor. Nat. Biomed. Eng. 3, 427–437 (2019).
Yan, H., Li, Y., Cheng, S. & Zeng, Y. Advances in analytical technologies for extracellular vesicles. Anal. Chem. 93, 4739–4774 (2021).
Kalluri, R. & LeBleu, V. S. The biology, function, and biomedical applications of exosomes. Science 367, eaau6977 (2020).
Thery, C., Zitvogel, L. & Amigorena, S. Exosomes: composition, biogenesis and function. Nat. Rev. Immunol. 2, 569–579 (2002).
Moller, A. & Lobb, R. J. The evolving translational potential of small extracellular vesicles in cancer. Nat. Rev. Cancer 20, 697–709 (2020).
van Niel, G., D’Angelo, G. & Raposo, G. Shedding light on the cell biology of extracellular vesicles. Nat. Rev. Mol. Cell Biol. 19, 213–228 (2018).
Garcia-Martin, R. et al. MicroRNA sequence codes for small extracellular vesicle release and cellular retention. Nature 601, 446–451 (2022).
Anfossi, S., Babayan, A., Pantel, K. & Calin, G. A. Clinical utility of circulating non-coding RNAs—an update. Nat. Rev. Clin. Oncol. 15, 541–563 (2018).
Ko, J. et al. miRNA profiling of magnetic nanopore-isolated extracellular vesicles for the diagnosis of pancreatic cancer. Cancer Res. 78, 3688–3697 (2018).
Pfeffer, S. R., Yang, C. H. & Pfeffer, L. M. The role of miR-21 in cancer. Drug Dev. Res. 76, 270–277 (2015).
Singh, D. et al. Real-time observation of DNA target interrogation and product release by the RNA-guided endonuclease CRISPR Cpf1 (Cas12a). Proc. Natl Acad. Sci. USA 115, 5444–5449 (2018).
Nallur, G. et al. Signal amplification by rolling circle amplification on DNA microarrays. Nucleic Acids Res. 29, E118 (2001).
Baner, J., Nilsson, M., Mendel-Hartvig, M. & Landegren, U. Signal amplification of padlock probes by rolling circle replication. Nucleic Acids Res. 26, 5073–5078 (1998).
Dahl, F. et al. Circle-to-circle amplification for precise and sensitive DNA analysis. Proc. Natl Acad. Sci. USA 101, 4548–4553 (2004).
Ali, M. M. et al. Rolling circle amplification: a versatile tool for chemical biology, materials science and medicine. Chem. Soc. Rev. 43, 3324–3341 (2014).
Lohman, G. J., Zhang, Y., Zhelkovsky, A. M., Cantor, E. J. & Evans, T. C. Jr. Efficient DNA ligation in DNA-RNA hybrid helices by Chlorella virus DNA ligase. Nucleic Acids Res. 42, 1831–1844 (2014).
Jin, J. M., Vaud, S., Zhelkovsky, A. M., Posfai, J. & McReynolds, L. A. Sensitive and specific miRNA detection method using SplintR Ligase. Nucleic Acids Res. 44, e116 (2016).
Wee, E. J. H. & Trau, M. Simple isothermal strategy for multiplexed, rapid, sensitive, and accurate miRNA detection. ACS Sens. 1, 670–675 (2016).
Lagunavicius, A., Kiveryte, Z., Zimbaite-Ruskuliene, V., Radzvilavicius, T. & Janulaitis, A. Duality of polynucleotide substrates for Phi29 DNA polymerase: 3′→5′ RNase activity of the enzyme. RNA 14, 503–513 (2008).
Bloomston, M. et al. MicroRNA expression patterns to differentiate pancreatic adenocarcinoma from normal pancreas and chronic pancreatitis. JAMA 297, 1901–1908 (2007).
Goto, T. et al. An elevated expression of serum exosomal microRNA-191,-21,-451a of pancreatic neoplasm is considered to be efficient diagnostic marker. BMC Cancer 18, 116 (2018).
Que, R., Ding, G., Chen, J. & Cao, L. Analysis of serum exosomal microRNAs and clinicopathologic features of patients with pancreatic adenocarcinoma. World J. Surg. Oncol. 11, 219 (2013).
Roldo, C. et al. MicroRNA expression abnormalities in pancreatic endocrine and acinar tumors are associated with distinctive pathologic features and clinical behavior. J. Clin. Oncol. 24, 4677–4684 (2006).
Xu, Y. F., Hannafon, B. N., Zhao, Y. D., Postier, R. G. & Ding, W. Q. Plasma exosome miR-196a and miR-1246 are potential indicators of localized pancreatic cancer. Oncotarget 8, 77028–77040 (2017).
Szafranska, A. E. et al. MicroRNA expression alterations are linked to tumorigenesis and non-neoplastic processes in pancreatic ductal adenocarcinoma. Oncogene 26, 4442–4452 (2007).
Zhang, Y. et al. Profiling of 95 microRNAs in pancreatic cancer cell lines and surgical specimens by real-time PCR analysis. World J. Surg. 33, 698–709 (2009).
Vicentini, C. et al. Exosomal miRNA signatures of pancreatic lesions. BMC Gastroenterol. 20, 137 (2020).
Takahasi, K. et al. Usefulness of exosome-encapsulated microRNA-451a as a minimally invasive biomarker for prediction of recurrence and prognosis in pancreatic ductal adenocarcinoma. J. Hepatobiliary Pancreat. Sci. 25, 155–161 (2018).
Kawamura, S. et al. Exosome-encapsulated microRNA-4525, microRNA-451a and microRNA-21 in portal vein blood is a high-sensitive liquid biomarker for the selection of high-risk pancreatic ductal adenocarcinoma patients. J. Hepatobiliary Pancreat. Sci. 26, 63–72 (2019).
Madhavan, B. et al. Combined evaluation of a panel of protein and miRNA serum-exosome biomarkers for pancreatic cancer diagnosis increases sensitivity and specificity. Int. J. Cancer 136, 2616–2627 (2015).
Zhang, P., He, M. & Zeng, Y. Ultrasensitive microfluidic analysis of circulating exosomes using a nanostructured graphene oxide/polydopamine coating. Lab Chip 16, 3033–3042 (2016).
Verel-Yilmaz, Y. et al. Extracellular vesicle-based detection of pancreatic cancer. Front. Cell. Dev. Biol. 9, 697939 (2021).
Xu, Y. F. et al. Isolation of extra-cellular vesicles in the context of pancreatic adenocarcinomas: addition of one stringent filtration step improves recovery of specific microRNAs. PLoS ONE 16, e0259563 (2021).
Mou, G., Wang, K., Xu, D. & Zhou, G. Evaluation of three RT-qPCR-based miRNA detection methods using seven rice miRNAs. Biosci. Biotechnol. Biochem. 77, 1349–1353 (2013).
Andreu, Z. et al. Comparative analysis of EV isolation procedures for miRNAs detection in serum samples. J. Extracell. Vesicles 5, 31655 (2016).
Brennan, K. et al. A comparison of methods for the isolation and separation of extracellular vesicles from protein and lipid particles in human serum. Sci. Rep. 10, 1039 (2020).
Gouin, K. et al. A comprehensive method for identification of suitable reference genes in extracellular vesicles. J. Extracell. Vesicles 6, 1347019 (2017).
Ragni, E. et al. miRNA reference genes in extracellular vesicles released from amniotic membrane-derived mesenchymal stromal cells. Pharmaceutics 12, 347 (2020).
Cai, S. et al. Single-molecule amplification-free multiplexed detection of circulating microRNA cancer biomarkers from serum. Nat. Commun. 12, 3515 (2021).
Veryaskina, Y. A. et al. Selection of reference genes for quantitative analysis of microRNA expression in three different types of cancer. PLoS ONE 17, e0254304 (2022).
Xiang, M. et al. U6 is not a suitable endogenous control for the quantification of circulating microRNAs. Biochem. Biophys. Res. Commun. 454, 210–214 (2014).
Lou, G. et al. Differential distribution of U6 (RNU6-1) expression in human carcinoma tissues demonstrates the requirement for caution in the internal control gene selection for microRNA quantification. Int. J. Mol. Med. 36, 1400–1408 (2015).
Yee, N. S., Zhang, S., He, H. Z. & Zheng, S. Y. Extracellular vesicles as potential biomarkers for early detection and diagnosis of pancreatic cancer. Biomedicines 8, 581 (2020).
Gootenberg, J. S. et al. Multiplexed and portable nucleic acid detection platform with Cas13, Cas12a, and Csm6. Science 360, 439–444 (2018).
Wang, N. et al. Recent advances in the rapid detection of microRNA with lateral flow assays. Biosens. Bioelectron. 211, 114345 (2022).
Cheng, Y. et al. Highly sensitive determination of microRNA using target-primed and branched rolling-circle amplification. Angew. Chem. Int. Ed. Engl. 121, 3318–3322 (2009).
Liu, H. et al. High specific and ultrasensitive isothermal detection of microRNA by padlock probe-based exponential rolling circle amplification. Anal. Chem. 85, 7941–7947 (2013).
Weber, J. A. et al. The microRNA spectrum in 12 body fluids. Clin. Chem. 56, 1733–1741 (2010).
Bhome, R. et al. Exosomal microRNAs (exomiRs): small molecules with a big role in cancer. Cancer Lett. 420, 228–235 (2018).
Salehi, M. & Sharifi, M. Exosomal miRNAs as novel cancer biomarkers: challenges and opportunities. J. Cell. Physiol. 233, 6370–6380 (2018).
O’Brien, K., Breyne, K., Ughetto, S., Laurent, L. C. & Breakefield, X. O. RNA delivery by extracellular vesicles in mammalian cells and its applications. Nat. Rev. Mol. Cell Biol. 21, 585–606 (2020).
Chevillet, J. R. et al. Quantitative and stoichiometric analysis of the microRNA content of exosomes. Proc. Natl Acad. Sci. USA 111, 14888–14893 (2014).
Krepelkova, I. et al. Evaluation of miRNA detection methods for the analytical characteristics necessary for clinical utilization. Biotechniques 66, 277–284 (2019).
Tian, W., Liu, X., Wang, G. & Liu, C. A hyperbranched transcription-activated CRISPR–Cas12a signal amplification strategy for sensitive microRNA sensing. Chem. Commun. 56, 13445–13448 (2020).
Tian, H. et al. Precise quantitation of MicroRNA in a single cell with droplet digital PCR based on ligation reaction. Anal. Chem. 88, 11384–11389 (2016).
Roy, S., Soh, J. H. & Ying, J. Y. A microarray platform for detecting disease-specific circulating miRNA in human serum. Biosens. Bioelectron. 75, 238–246 (2016).
Chen, K., Wang, Q., Kornmann, M., Tian, X. & Yang, Y. The role of exosomes in pancreatic cancer from bench to clinical application: an updated review. Front. Oncol. 11, 644358 (2021).
Park, W., Chawla, A. & O’Reilly, E. M. Pancreatic cancer: a review. JAMA 326, 851–862 (2021).
Wong, M. C. S. et al. Global temporal patterns of pancreatic cancer and association with socioeconomic development. Sci. Rep. 7, 3165 (2017).
Brezgyte, G., Shah, V., Jach, D. & Crnogorac-Jurcevic, T. Non-invasive biomarkers for earlier detection of pancreatic cancer-a comprehensive review. Cancers 13, 2722 (2021).
Yang, J. S. et al. Early screening and diagnosis strategies of pancreatic cancer: a comprehensive review. Cancer Commun. 41, 1257–1274 (2021).
Huang, L. et al. Resection of pancreatic cancer in Europe and USA: an international large-scale study highlighting large variations. Gut 68, 130–139 (2019).
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2018. CA: Cancer J. Clin. 68, 7–30 (2018).
Ballehaninna, U. K. & Chamberlain, R. S. The clinical utility of serum CA 19-9 in the diagnosis, prognosis and management of pancreatic adenocarcinoma: an evidence based appraisal. J. Gastrointest. Oncol. 3, 105–119 (2012).
Zhang, Y. et al. Tumor markers CA19-9, CA242 and CEA in the diagnosis of pancreatic cancer: a meta-analysis. Int. J. Clin. Exp. Med. 8, 11683–11691 (2015).
Zhang, P. et al. Ultrasensitive detection of circulating exosomes with a 3D-nanopatterned microfluidic chip. Nat. Biomed. Eng. 3, 438–451 (2019).
Zhang, P. et al. Molecular and functional extracellular vesicle analysis using nanopatterned microchips monitors tumor progression and metastasis. Sci. Transl. Med. 12, eaaz2878 (2020).
Chandrasekaran, S. S. et al. Rapid detection of SARS-CoV-2 RNA in saliva via Cas13. Nat. Biomed. Eng. 6, 944–956 (2022).
Zhang, T. et al. A paper-based assay for the colorimetric detection of SARS-CoV-2 variants at single-nucleotide resolution. Nat. Biomed. Eng. 6, 957–967 (2022).
Niemeyer, C. M., Adler, M. & Wacker, R. Detecting antigens by quantitative immuno-PCR. Nat. Protoc. 2, 1918–1930 (2007).
Li, L. et al. Nucleic acid aptamers for molecular diagnostics and therapeutics: advances and perspectives. Angew. Chem. Int. Ed. Engl. 60, 2221–2231 (2021).
Han, S. et al. Primary outgrowth cultures are a reliable source of human pancreatic stellate cells. Lab. Invest. 95, 1331–1340 (2015).
Han, S. et al. The proteome of pancreatic cancer-derived exosomes reveals signatures rich in key signaling pathways. Proteomics 19, e1800394 (2019).
Tibshirani, R. Regression shrinkage and selection via the Lasso. J. R. Stat. Soc. B 58, 267–288 (1996).
Homrighausen, D. & McDonald, D. J. Leave-one-out cross-validation is risk consistent for Lasso. Mach. Learn. 97, 65–78 (2014).
Acknowledgements
We thank the University of Florida Clinical and Translational Science Institute Biorepository Facility for providing human plasma specimen. We thank Y. Li for helping with TEM analysis and G. Tushoski for helping with culturing cells and collecting conditioned medium. This study was supported in part by grants R01 CA243445, R33 CA214333, R33 CA252158A1 and R01 CA260132 from the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
Y.Z. conceived and supervised the project; H.Y. and Y.Z. designed the research; H.Y. performed technology development, mechanistic study, analytical characterization and clinical validation; H.Y. and Z.T. conducted the LFA experiments; H.Y. and N.H. constructed the POC device; H.Y. and Y.W. performed isolation and NTA analysis of EVs; S.H. conducted cell culture and provided culture media; S.H. helped with TEM analysis and was involved in experiment design and discussion; S.J.H. assisted in the clinical studies; H.Y. and Y.Z. analysed the data and wrote the paper. All authors edited the paper.
Corresponding author
Ethics declarations
Competing interests
H.Y. and Y.Z. are co-inventors on a United States provisional patent application (63/313,870) based on this work. Y.Z. holds equity interest in Clara Biotech and serves on its scientific advisory board. The other authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks Ruijie Deng and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary figures, tables, unprocessed gels and references.
Source data
Source Data Fig. 2 (download PDF )
Unprocessed gels.
Source Data Fig. 5 (download TIF )
Unprocessed blots.
Source Data Fig. 6 (download TIF )
Unprocessed blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yan, H., Wen, Y., Tian, Z. et al. A one-pot isothermal Cas12-based assay for the sensitive detection of microRNAs. Nat. Biomed. Eng 7, 1583–1601 (2023). https://doi.org/10.1038/s41551-023-01033-1
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41551-023-01033-1
This article is cited by
-
Novel serum small extracellular vesicle miRNAs with multi-target RCA-CRISPR sensor for liver cancer detection
Journal of Translational Medicine (2026)
-
PRICE: direct and robust detection of microRNAs at single-nucleotide resolution
Nature Communications (2026)
-
Direct delivery of assay reagents to extracellular vesicles in liquid biopsies for biomarker analysis
Nature Protocols (2026)
-
Versatile bioluminescence CRISPR/Cas platform for point-of-care detection of biomarkers in whole blood
Science China Chemistry (2026)
-
A multiplexed assay by self-assembled dual-target responsive DNA hydrogels for efficacy evaluation of immunotherapy
Nature Communications (2025)