Abstract
Vision foundation models have demonstrated vast potential in achieving generalist medical segmentation capability, providing a versatile, task-agnostic solution through a single model. However, current generalist models involve simple pre-training on various medical data containing irrelevant information, often resulting in the negative transfer phenomenon and degenerated performance. Furthermore, the practical applicability of foundation models across diverse open-world scenarios, especially in out-of-distribution (OOD) settings, has not been extensively evaluated. Here we construct a publicly accessible database, MedSegDB, based on a tree-structured hierarchy and annotated from 129 public medical segmentation repositories and 5 in-house datasets. We further propose a Generalist Medical Segmentation model (MedSegX), a vision foundation model trained with a model-agnostic Contextual Mixture of Adapter Experts (ConMoAE) for open-world segmentation. We conduct a comprehensive evaluation of MedSegX across a range of medical segmentation tasks. Experimental results indicate that MedSegX achieves state-of-the-art performance across various modalities and organ systems in in-distribution (ID) settings. In OOD and real-world clinical settings, MedSegX consistently maintains its performance in both zero-shot and data-efficient generalization, outperforming other foundation models.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The main data supporting the results in this study are available within the paper and its Supplementary Information. The pre-training, ID validation and OOD validation datasets from our MedSegDB database are curated from open-source datasets and can be accessed via the weblinks provided in Supplementary Table 1. Among these, the data in MedSegDB that permit redistribution are available on HuggingFace60 (https://huggingface.co/datasets/medicalai/MedSegDB). Each dataset used in the study is also provided in Supplementary Table 1. The validation dataset from MedSegDB that consists of real-world data collected from hospitals cannot be fully released in a public repository due to privacy regulations. We have deposited a minimum dataset of de-identified real-world data on GitHub (https://github.com/MedSegX/MedSegX-code).
Code availability
The source code to train MedSegX and reproduce the results is available in GitHub at https://github.com/MedSegX/MedSegX-code (ref. 61).
References
Wang, S. et al. Annotation-efficient deep learning for automatic medical image segmentation. Nat. Commun. 12, 5915 (2021).
Cao, K. et al. Large-scale pancreatic cancer detection via non-contrast CT and deep learning. Nat. Med. 29, 3033–3043 (2023).
Mei, X. et al. Artificial intelligence-enabled rapid diagnosis of patients with COVID-19. Nat. Med. 26, 1224–1228 (2020).
Wang, G. et al. A deep-learning pipeline for the diagnosis and discrimination of viral, non-viral and COVID-19 pneumonia from chest X-ray images. Nat. Biomed. Eng. 5, 509–521 (2021).
Wang, S. et al. Mining whole-lung information by artificial intelligence for predicting EGFR genotype and targeted therapy response in lung cancer: a multicohort study. Lancet Digit. Health 4, e309–e319 (2022).
Zhang, K. et al. Clinically applicable AI system for accurate diagnosis, quantitative measurements, and prognosis of COVID-19 pneumonia using computed tomography. Cell 181, 1423–1433.e11 (2020).
Xu, Y. et al. Improving artificial intelligence pipeline for liver malignancy diagnosis using ultrasound images and video frames. Brief. Bioinform. 24, bbac569 (2023).
Hatamizadeh, A. et al. Swin UNETR: Swin transformers for semantic segmentation of brain tumors in MRI images. In International MICCAI Brainlesion Workshop (eds Crimi, A. et al.) 272–284 (Springer, 2021).
Isensee, F., Jaeger, P. F., Kohl, S. A., Petersen, J. & Maier-Hein, K. H. nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nat. Methods 18, 203–211 (2021).
Lee, H. H., Bao, S., Huo, Y. & Landman, B. A. 3D UX-Net: a large kernel volumetric convnet modernizing hierarchical transformer for medical image segmentation. In International Conference on Learning Representations 21891-21905 (ICLR, 2023).
Ronneberger, O., Fischer, P. & Brox, T. U-Net: convolutional networks for biomedical image segmentation. In International Conference on Medical Image Computing and Computer-Assisted Intervention (eds Navab, N. et al.) 234–241 (Springer, 2015).
Zhou, Z., Siddiquee, M. M. R., Tajbakhsh, N. & Liang, J. UNet++: redesigning skip connections to exploit multiscale features in image segmentation. IEEE Trans. Med. Imaging 39, 1856–1867 (2019).
Zhang, J., Xie, Y., Xia, Y. & Shen, C. DoDNet: learning to segment multi-organ and tumors from multiple partially labeled datasets. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 1195–1204 (IEEE, 2021).
Ye, Y., Xie, Y., Zhang, J., Chen, Z. & Xia, Y. UniSeg: a prompt-driven universal segmentation model as well as a strong representation learner. In International Conference on Medical Image Computing and Computer-Assisted Intervention (eds Greenspan, H. et al.) 508–518 (Springer, 2023).
Kirillov, A. et al. Segment anything. In Proc. IEEE/CVF International Conference on Computer Vision 4015–4026 (IEEE, 2023).
Oquab, M. et al. DINOv2: learning robust visual features without supervision. Trans. Mach. Learn. Res. https://openreview.net/forum?id=a68SUt6zFt (2024).
Ma, J. & Wang, B. Towards foundation models of biological image segmentation. Nat. Methods 20, 953–955 (2023).
Ma, J. et al. Segment anything in medical images. Nat. Commun. 15, 654 (2024).
Cheng, J. et al. SAM-med2D. Preprint at https://arxiv.org/abs/2308.16184 (2023).
Zhili, L. et al. Task-customized self-supervised pre-training with scalable dynamic routing. In Proc. AAAI Conference on Artificial Intelligence 1854–1862 (AAAI, 2022).
Wang, Z., Dai, Z., Póczos, B. & Carbonell, J. Characterizing and avoiding negative transfer. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 11293–11302 (IEEE, 2019).
Senushkin, D., Patakin, N., Kuznetsov, A. & Konushin, A. Independent component alignment for multi-task learning. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 20083–20093 (IEEE, 2023).
Langlotz, C. P. RadLex: a new method for indexing online educational materials. Radiographics 26, 1595–1597 (2006).
Shazeer, N. et al. Outrageously large neural networks: the sparsely-gated mixture-of-experts layer. In International Conference on Learning Representations 878–896 (ICLR, 2017).
Ma, J., Li, F. & Wang, B. U-Mamba: enhancing long-range dependency for biomedical image segmentation. Preprint at https://arxiv.org/abs/2401.04722 (2024).
Li, T. et al. SrSNet: accurate segmentation of stroke lesions by a two-stage segmentation framework with asymmetry information. Expert Syst. Appl. 254, 124329 (2024).
Leclerc, S. et al. Deep learning for segmentation using an open large-scale dataset in 2D echocardiography. IEEE Trans. Med. Imaging 38, 2198–2210 (2019).
Azizi, S. et al. Robust and data-efficient generalization of self-supervised machine learning for diagnostic imaging. Nat. Biomed. Eng. 7, 756–779 (2023).
Hupkes, D. et al. A taxonomy and review of generalization research in NLP. Nat. Mach. Intell. 5, 1161–1174 (2023).
Shiraishi, J. et al. Development of a digital image database for chest radiographs with and without a lung nodule: receiver operating characteristic analysis of radiologists’ detection of pulmonary nodules. Am. J. Roentgenol. 174, 71–74 (2000).
Stirenko, S. et al. Chest X-ray analysis of tuberculosis by deep learning with segmentation and augmentation. In International Conference on Electronics and Nanotechnology 422–428 (IEEE, 2018).
Siegel, R. L., Giaquinto, A. N. & Jemal, A. Cancer statistics, 2024. CA Cancer J. Clin. 74, 12–49 (2024).
Zhou, H.-Y. et al. nnFormer: volumetric medical image segmentation via a 3D transformer. IEEE Trans. Image Process. 32, 4036–4045 (2023).
Wang, N. et al. MISSU: 3D medical image segmentation via self-distilling TransUNet. IEEE Trans. Med. Imaging 42, 2740–2750 (2023).
Zhu, J., Qi, Y. & Wu, J. Medical SAM 2: segment medical images as video via Segment Anything Model 2. Preprint at https://arxiv.org/abs/2408.00874 (2024).
Ma, J. et al. Segment anything in medical images and videos: benchmark and deployment. Preprint at https://arxiv.org/abs/2408.03322 (2024).
Ravi, N. et al. SAM 2: segment anything in images and videos. In International Conference on Learning Representations 41175–41218 (ICLR, 2025).
Huang, Y. et al. Segment anything model for medical images? Med. Image Anal. 92, 103061 (2024).
Ye, J. et al. SA-med2D-20M dataset: segment anything in 2D medical imaging with 20 million masks. Preprint at https://arxiv.org/abs/2311.11969 (2023).
Ma, J. et al. AbdomenCT-1K: is abdominal organ segmentation a solved problem. IEEE Trans. Pattern Anal. Mach. Intell. 44, 6695–6714 (2021).
Ji, Y. et al. AMOS: a large-scale abdominal multi-organ benchmark for versatile medical image segmentation. Adv. Neural Inf. Process. Syst. 35, 36722–36732 (2022).
Bilic, P. et al. The liver tumor segmentation benchmark (lits). Med. Image Anal. 84, 102680 (2023).
Wasserthal, J. et al. TotalSegmentator: robust segmentation of 104 anatomic structures in CT images. Radiol. Artif. Intell. 5, e230024 (2023).
Pang, S. et al. SpineParseNet: spine parsing for volumetric MR image by a two-stage segmentation framework with semantic image representation. IEEE Trans. Med. Imaging 40, 262–273 (2020).
Deng, J. et al. ImageNet: a large-scale hierarchical image database. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 248–255 (IEEE, 2009).
Krishna, R. et al. Visual genome: connecting language and vision using crowdsourced dense image annotations. Int. J. Comput. Vis. 123, 32–73 (2017).
Wu, J. et al. Chest ImaGenome dataset (version 1.0.0). PhysioNet https://doi.org/10.13026/wv01-y230 (2021).
Fifty, C. et al. Efficiently identifying task groupings for multi-task learning. Adv. Neural Inf. Process. Syst. 34, 27503–27516 (2021).
Clark, K. et al. The Cancer Imaging Archive (TCIA): maintaining and operating a public information repository. J. Digit. Imaging 26, 1045–1057 (2013).
Heller, N. et al. The state of the art in kidney and kidney tumor segmentation in contrast-enhanced CT imaging: results of the KiTS19 challenge. Med. Image Anal. 67, 101821 (2021).
Heller, N. et al. The KiTS21 Challenge: automatic segmentation of kidneys, renal tumors, and renal cysts in corticomedullary-phase CT. Preprint at https://arxiv.org/abs/2307.01984 (2023).
Agarap, A. F. Deep learning using rectified linear units (ReLU). Preprint at https://arxiv.org/abs/1803.08375 (2018).
Cheng, B., Misra, I., Schwing, A. G., Kirillov, A. & Girdhar, R. Masked-attention mask transformer for universal image segmentation. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 1290–1299 (IEEE, 2022).
Milletari, F., Navab, N. & Ahmadi, S.-A. V-Net: fully convolutional neural networks for volumetric medical image segmentation. In International Conference on 3D Vision (3DV) 565–571 (IEEE, 2016).
Chen, C. et al. MA-SAM: modality-agnostic SAM adaptation for 3D medical image segmentation. Med. Image Anal. 98, 103310 (2024).
Zhai, X. et al. A large-scale study of representation learning with the visual task adaptation benchmark. Preprint at https://arxiv.org/abs/1910.04867 (2019).
Dosovitskiy, A. et al. An image is worth 16x16 words: transformers for image recognition at scale. In International Conference on Learning Representations 611–631 (ICLR, 2021).
Zhang, H., Su, Y., Xu, X. & Jia, K. Improving the generalization of segmentation foundation model under distribution shift via weakly supervised adaptation. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 23385–23395 (IEEE, 2024).
Peng, Z. et al. Parameter efficient fine-tuning via cross block orchestration for Segment Anything Model. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 3743–3752 (IEEE, 2024).
Zhang, S. MedSegDB. HuggingFace https://huggingface.co/datasets/medicalai/MedSegDB (2025).
Zhang, S. MedSegX. GitHub https://github.com/MedSegX/MedSegX-code (2025).
Acknowledgements
G.W. acknowledges funding from the National Natural Science Foundation of China (grants T2522008 and 62272055), New Cornerstone Science Foundation through the XPLORER PRIZE, and the National Key Research and Development Program of China (2024YFC3044700). Shanghang Zhang was supported by the National Science and Technology Major Project (No. 2022ZD0117800). S.W. was supported by the National Natural Science Foundation of China (62425112, 92359202 and 12326610).
Author information
Authors and Affiliations
Contributions
G.W., X.L., Siqi Zhang, Q.Z., J. Yue, H.X., J. Yao and Y.W. collected and analysed the data. M.L., Shanghang Zhang and G.W. conceived and supervised the project. G.W., Siqi Zhang, X.L., Q.Z., Shanghang Zhang, J. Yue and M.L. contributed to data interpretation, method design, and critical review and wrote the manuscript. All authors discussed the results and reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks Namkug Kim, Ongen Liao and Dong Ni for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Detailed description of MedSegDB.
It consists of 5 anatomical subtrees, including the body parts of the head and neck, torso, skeleton, blood vessel, and skin, covering 39 major organs and tissues, and 111 segmentation tasks. MedSegDB incorporates 10 medical imaging modalities including the computed tomography (CT), magnetic resonance imaging (MRI), X-ray, colonoscopy, dermoscopy, CT angiography (CTA), cone beam CT (CBCT), ultrasound (US), fundus, and endoscopy.
Extended Data Fig. 2 Detailed data processing for development and evaluation of MedSegDB.
a. Flow chart describing the datasets collected for the development and evaluation of ID and OOD sets. b. Details of the real-world datasets.
Extended Data Fig. 3 Detailed performance of MedSegX and other competitors on tasks related to 5 body parts in ID test set.
Comparison of 5 models on tasks in the parts of the a. head and neck (n = 67,746), b. torso (n = 205,066), c. skeleton (n = 45,981) d. blood vessel (n = 8,121), and e. skin (n = 818). P-values are calculated with two-sided t-test. Bar graphs indicate the mean ± 95% CI. The dashed horizontal line represents the Dice score achieved by MedSegX.
Extended Data Fig. 4 Generalization performance evaluation of MedSegX and other competitors on 18 cross-site tasks with different proportions of OOD fine-tuning data (n = 5,801).
a. 15% fine-tuning data. b. 25% fine-tuning data. c. 50% fine-tuning data. d.100% fine-tuning data. Tasks consist of tooth (X-ray), gallbladder (CT), left kidney (CT), right kidney (CT), left lung (CT), right lung (CT), liver (CT), pancreas (CT), stomach (CT), optic cup (Fundus), optic disc (Fundus), left atrium (MRI), prostate (MRI), right ventricle (MRI), left ventricle (MRI), spleen (MRI), left lung (X-ray), and right lung (X-ray) segmentation. In all box plots, each box shows the quartiles of the distribution, with center as the median, minimum as the first quartile, and maximum as the third quartile. The whiskers extend to the farthest data point that lies within 2× interquartile range (IQR) from the nearest quartile.
Extended Data Fig. 5 Data-efficient performance evaluation of MedSegX and SAM-Med2D on 18 cross-site tasks.
95% CI intervals are shown by the shaded area.
Extended Data Fig. 6 Generalization performance evaluation of MedSegX and competitors on seven cross-target tasks with different proportions of OOD fine-tuning data (n = 4,095).
a. 15% fine-tuning data. b. 25% fine-tuning data. c. 50% fine-tuning data. d.100% fine-tuning data. Tasks consist of the colon tumor (CT), kidney tumor (CT), liver tumor (CT), lung tumor (CT), pancreas tumor (CT), prostate tumor (MRI), and vestibular schwannoma segmentation (MRI). In all box plots, each box shows the quartiles of the distribution, with center as the median, minimum as the first quartile, and maximum as the third quartile. The whiskers extend to the farthest data point that lies within 2× interquartile range (IQR) from the nearest quartile.
Extended Data Fig. 7 Data-efficient performance evaluation of MedSegX and MedSAM on 7 unseen categories of body tumors.
95% CI intervals are shown by the shaded area.
Extended Data Fig. 8 The models’ average performance under real-world setting (n = 950).
a. Average zero-shot performance evaluation of 6 models on 5 real-world datasets. b. Average fine-tuning performance evaluation of 6 models on 5 real-world datasets. P-values are calculated with two-sided t-test. Bar graphs indicate the mean ± 95% CI. The dashed horizontal line represents the Dice score achieved by MedSegX.
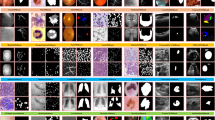
Extended Data Fig. 9 Visualization of 8 segmentation examples in OOD test sets.
T1: Left Ventricular Myocardium, T2: Teeth, T3: Colon Tumor, T4: Kidney Tumor, T5: Liver Tumor, T6: Lung Tumor, T7: Pancreas Tumor, T8: Prostate Tumor.
Extended Data Fig. 10 Visualization analysis for the ablation experiments.
8 segmentation examples from both ID and OOD test sets are incorporated (ID tasks: T1 ~ T4. OOD tasks: T5 ~ T8): T1: Left Ventricular Myocardium, T2: Optic Disc, T3: Liver, T4: Brain Core Tumor, T5: Breast Cancer, T6: Left Ventricle Epicardium, T7: Left Lung, T8: Stomach, T9: Kidney Tumor, T10: Lung Tumor.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–3 and Tables 1–41.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, S., Zhang, Q., Zhang, S. et al. A generalist foundation model and database for open-world medical image segmentation. Nat. Biomed. Eng (2025). https://doi.org/10.1038/s41551-025-01497-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41551-025-01497-3
This article is cited by
-
Bioorthogonal optimized virus immuno-nanomedicine (BOVIN)
Nature Communications (2025)