Abstract
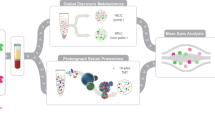
Profiling molecular panorama from massive omics data identifies regulatory networks in cells but requires mechanistic interpretation and experimental follow up. Here we combine deep learning and large language model reasoning to develop a hybrid workflow for omics interpretation, called LyMOI. LyMOI incorporates GPT-3.5 for biological knowledge reasoning and a large graph model with graph convolutional networks (GCNs). The large graph model integrates evolutionarily conserved protein interactions and uses hierarchical fine-tuning to predict context-specific molecular regulators from multi-omics data. GPT-3.5 then generates machine chain-of-thought (CoT) to mechanistically interpret their roles in biological systems. Focusing on autophagy, LyMOI mechanistically interprets 1.3 TB transcriptomic, proteomic and phosphoproteomic data and expands the knowledge of autophagy regulators. We also show that LyMOI highlights two human oncoproteins, CTSL and FAM98A, for enhancing autophagy upon treatment with disulfiram (DSF), an antitumour agent. Silencing these genes in vitro attenuates DSF-mediated autophagy and suppresses cancer cell proliferation. Strikingly, DSF treatment with Z-FY-CHO, a CTSL-specific inhibitor previously used for preventing SARS-CoV-2 infection, potently inhibits tumour growth in vivo.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The data supporting the results in this study are available within the paper and its Supplementary Information. The RNA-seq data of yeast were deposited into the NCBI Sequence Read Archive (SRA, https://www.ncbi.nlm.nih.gov/sra) with the dataset identifier PRJNA912308 (ref. 90). The raw MS datasets of the proteome and phosphoproteome of yeast were submitted to integrated proteome resources (iProX, http://www.iprox.org/) with the dataset identifier PXD038804 (ref. 91). Source data are provided with this paper.
Code availability
The source code for LyMOI in this study is available on GitHub (https://github.com/BioCUCKOO/LyMOI)92.
References
Stefely, J. A. et al. Mitochondrial protein functions elucidated by multi-omic mass spectrometry profiling. Nat. Biotechnol. 34, 1191–1197 (2016).
Karczewski, K. J. & Snyder, M. P. Integrative omics for health and disease. Nat. Rev. Genet. 19, 299–310 (2018).
Rhodes, D. R. & Chinnaiyan, A. M. Integrative analysis of the cancer transcriptome. Nat. Genet. 37, S31–S37 (2005).
Chung, M. et al. Best practices on the differential expression analysis of multi-species RNA-seq. Genome Biol. 22, 121 (2021).
Yamada, R. et al. Interpretation of omics data analyses. J. Hum. Genet. 66, 93–102 (2021).
Subramanian, I. et al. Multi-omics data integration, interpretation, and its application. Bioinform. Biol. Insights 14, 1177932219899051 (2020).
Shu, T. et al. Plasma proteomics identify biomarkers and pathogenesis of COVID-19. Immunity 53, 1108–1122.e5 (2020).
Shui, K. et al. Small-sample learning reveals propionylation in determining global protein homeostasis. Nat. Commun. 14, 2813 (2023).
Yuan, Y. et al. PIM1 promotes hepatic conversion by suppressing reprogramming-induced ferroptosis and cell cycle arrest. Nat. Commun. 13, 5237 (2022).
Hirschberg, J. & Manning, C. D. Advances in natural language processing. Science 349, 261–266 (2015).
ChatGPT: Optimizing Language Models for Dialogue (OpenAI, 2022).
Christiano, P. F. et al. Deep reinforcement learning from human preferences. In Proc. 31st International Conference on Neural Information Processing Systems (eds von Luxburg, U. et al.) 4302–4310 (Curran, 2017).
Brown, T. B. et al. Language models are few-shot learners. In Proc. 34th Conference on Neural Information Processing Systems (eds Larochelle, H. et al.) 1–25 (2020).
Wei, J. S. et al. Chain-of-thought prompting elicits reasoning in large language models. In Proc. 36th International Conference on Neural Information Processing Systems (eds Koyejo, S. et al.) 24824–24837 (Curran, 2022).
Klionsky, D. J. et al. Guidelines for the use and interpretation of assays for monitoring autophagy (4th edition). Autophagy 17, 1–382 (2021).
Ulgherait, M. et al. Circadian autophagy drives iTRF-mediated longevity. Nature 598, 353–358 (2021).
Skrott, Z. et al. Alcohol-abuse drug disulfiram targets cancer via p97 segregase adaptor NPL4. Nature 552, 194–199 (2017).
Zhao, M.-M. et al. Novel cleavage sites identified in SARS-CoV-2 spike protein reveal mechanism for cathepsin L-facilitated viral infection and treatment strategies. Cancer Discov. 8, 53 (2022).
Podgorski, J. & Berg, M. Global threat of arsenic in groundwater. Science 368, 845–850 (2020).
Diamantopoulou, Z. et al. The metastatic spread of breast cancer accelerates during sleep. Nature 607, 156–162 (2022).
Obradović, M. M. S. et al. Glucocorticoids promote breast cancer metastasis. Nature 567, 540–544 (2019).
Hirota, T. & King, B. H. Autism spectrum disorder: a review. JAMA 329, 157–168 (2023).
Chen, A. et al. Spatiotemporal transcriptomic atlas of mouse organogenesis using DNA nanoball-patterned arrays. Cell 185, 1777–1792.e21 (2022).
Tang, F. et al. A pan-cancer single-cell panorama of human natural killer cells. Cell 186, 4235–4251.e20 (2023).
Velmeshev, D. et al. Single-cell analysis of prenatal and postnatal human cortical development. Science 382, eadf0834 (2023).
Deng, W. et al. THANATOS: an integrative data resource of proteins and post-translational modifications in the regulation of autophagy. Autophagy 14, 296–310 (2018).
Han, Z. et al. Model-based analysis uncovers mutations altering autophagy selectivity in human cancer. Nat. Commun. 12, 3258 (2021).
Santos, A. et al. A knowledge graph to interpret clinical proteomics data. Nat. Biotechnol. 40, 692–702 (2022).
Zhang, Z. et al. Large graph models: a perspective. Preprint at https://doi.org/10.48550/arXiv.2308.14522 (2023).
Yu, H. et al. Annotation transfer between genomes: protein–protein interologs and protein–DNA regulogs. Genome Res. 14, 1107–1118 (2004).
Lyu, Y., Huang, X. & Zhang, Z. Revisiting 2D convolutional neural networks for graph-based applications. IEEE Trans. Pattern Anal. Mach. Intell. 45, 6909–6922 (2023).
Díez, J., Walter, D., Muñoz-Pinedo, C. & Gabaldón, T. DeathBase: a database on structure, evolution and function of proteins involved in apoptosis and other forms of cell death. Cell Death Differ. 17, 735–736 (2010).
Homma, K., Suzuki, K. & Sugawara, H. The Autophagy Database: an all-inclusive information resource on autophagy that provides nourishment for research. Nucleic Acids Res. 39, D986–D990 (2011).
Moussay, E. et al. The acquisition of resistance to TNFα in breast cancer cells is associated with constitutive activation of autophagy as revealed by a transcriptome analysis using a custom microarray. Autophagy 7, 760–770 (2011).
Xu, J. & Li, Y. H. miRDeathDB: a database bridging microRNAs and the programmed cell death. Cell Death Differ. 19, 1571 (2012).
Arntzen, M., Bull, V. H. & Thiede, B. Cell death proteomics database: consolidating proteomics data on cell death. J. Proteome Res. 12, 2206–2213 (2013).
Wanichthanarak, K., Cvijovic, M., Molt, A. & Petranovic, D. yApoptosis: yeast apoptosis database. Database 2013, bat068 (2013).
Türei, D. et al. Autophagy Regulatory Network—a systems-level bioinformatics resource for studying the mechanism and regulation of autophagy. Autophagy 11, 155–165 (2015).
Wu, D. et al. ncRDeathDB: a comprehensive bioinformatics resource for deciphering network organization of the ncRNA-mediated cell death system. Autophagy 11, 1917–1926 (2015).
Wang, N. N. et al. HAMdb: a database of human autophagy modulators with specific pathway and disease information. J. Cheminform. 10, 34 (2018).
Chen, K. et al. Autophagy and Tumor Database: ATdb, a novel database connecting autophagy and tumor. Database https://doi.org/10.1093/database/baaa052 (2020).
Zhou, N. & Bao, J. FerrDb: a manually curated resource for regulators and markers of ferroptosis and ferroptosis–disease associations. Database https://doi.org/10.1093/database/baaa021 (2020).
Zhang, L. et al. MCDB: a comprehensive curated mitotic catastrophe database for retrieval, protein sequence alignment, and target prediction. Acta Pharm. Sin. B 11, 3092–3104 (2021).
Sun, Y. J., Sheng, D. F., Zhou, Z. H. & Wu, Y. F. AI hallucination: towards a comprehensive classification of distorted information in artificial intelligence-generated content. Hum. Soc. Sci. Commun. https://doi.org/10.1057/S41599-024-03811-X (2024).
Bang, Y. et al. A multitask, multilingual, multimodal evaluation of ChatGPT on reasoning, hallucination, and interactivity. Preprint at https://arxiv.org/abs/2302.04023 (2023).
Zhu, W., Swaminathan, G. & Plowey, E. D. GA binding protein augments autophagy via transcriptional activation of BECN1-PIK3C3 complex genes. Autophagy 10, 1622–1636 (2014).
Sun, W., Jia, M., Feng, Y. & Cheng, X. Lactate is a bridge linking glycolysis and autophagy through lactylation. Autophagy 19, 3240–3241 (2023).
Fujioka, Y. et al. Structural basis of starvation-induced assembly of the autophagy initiation complex. Nat. Struct. Mol. Biol. 21, 513–521 (2014).
Schreiber, A. et al. Multilayered regulation of autophagy by the Atg1 kinase orchestrates spatial and temporal control of autophagosome formation. Mol. Cell 81, 5066–5081.e10 (2021).
Feng, Y. et al. Phosphorylation of Atg9 regulates movement to the phagophore assembly site and the rate of autophagosome formation. Autophagy 12, 648–658 (2016).
Cowley, M. J. et al. PINA v2.0: mining interactome modules. Nucleic Acids Res. 40, D862–D865 (2012).
Das, J. & Yu, H. HINT: high-quality protein interactomes and their applications in understanding human disease. BMC Syst. Biol. 6, 92 (2012).
Razick, S., Magklaras, G. & Donaldson, I. M. iRefIndex: a consolidated protein interaction database with provenance. BMC Bioinformatics 9, 405 (2008).
Oughtred, R. et al. The BioGRID interaction database: 2019 update. Nucleic Acids Res. 47, D529–D541 (2019).
Calderone, A., Castagnoli, L. & Cesareni, G. mentha: a resource for browsing integrated protein-interaction networks. Nat. Methods 10, 690–691 (2013).
Kotlyar, M., Pastrello, C., Malik, Z. & Jurisica, I. IID 2018 update: context-specific physical protein–protein interactions in human, model organisms and domesticated species. Nucleic Acids Res. 47, D581–D589 (2019).
Li, T. et al. A scored human protein–protein interaction network to catalyze genomic interpretation. Nat. Methods 14, 61–64 (2017).
Galluzzi, L. et al. Molecular definitions of autophagy and related processes. EMBO J. 36, 1811–1836 (2017).
Yi, C. et al. Formation of a Snf1-Mec1-Atg1 module on mitochondria governs energy deprivation-induced autophagy by regulating mitochondrial respiration. Dev. Cell 41, 59–71.e54 (2017).
Yi, C., Tong, J. J. & Yu, L. Mitochondria: the hub of energy deprivation-induced autophagy. Autophagy 14, 1084–1085 (2018).
Clement, S. T., Dixit, G. & Dohlman, H. G. Regulation of yeast G protein signaling by the kinases that activate the AMPK homolog Snf1. Sci. Signal. 6, ra78 (2013).
Mok, J. et al. Deciphering protein kinase specificity through large-scale analysis of yeast phosphorylation site motifs. Sci. Signal. 3, ra12 (2010).
Asano, S. et al. Direct phosphorylation and activation of a Nim1-related kinase Gin4 by Elm1 in budding yeast. J. Biol. Chem. 281, 27090–27098 (2006).
Hu, Y. et al. The disulfiram/copper complex induces autophagic cell death in colorectal cancer by targeting ULK1. Front. Pharmacol. 12, 752825 (2021).
Jivan, R. et al. Disulfiram with or without metformin inhibits oesophageal squamous cell carcinoma in vivo. Cancer Lett. 417, 1–10 (2018).
Wu, X. et al. Suppressing autophagy enhances disulfiram/copper-induced apoptosis in non-small cell lung cancer. Eur. J. Pharmacol. 827, 1–12 (2018).
Xu, S. et al. Inhibition of cathepsin L alleviates the microglia-mediated neuroinflammatory responses through caspase-8 and NF-κB pathways. Neurobiol. Aging 62, 159–167 (2018).
Liu, H. et al. Oxidized DJ-1 activates the p-IKK/NF-κB/Beclin1 pathway by binding PTEN to induce autophagy and exacerbate myocardial ischemia-reperfusion injury. Eur. J. Pharmacol. 971, 176496 (2024).
Tate, J. G. et al. COSMIC: the Catalogue Of Somatic Mutations In Cancer. Nucleic Acids Res. 47, D941–D947 (2019).
Kenig, S., Frangež, R., Pucer, A. & Lah, T. Inhibition of cathepsin L lowers the apoptotic threshold of glioblastoma cells by up-regulating p53 and transcription of caspases 3 and 7. Apoptosis 16, 671–682 (2011).
Zhao, M. M. et al. Cathepsin L plays a key role in SARS-CoV-2 infection in humans and humanized mice and is a promising target for new drug development. Signal Transduct. Target. Ther. https://doi.org/10.1038/s41392-021-00558-8 (2021).
Sudhan, D. R., Pampo, C., Rice, L. & Siemann, D. W. Cathepsin L inactivation leads to multimodal inhibition of prostate cancer cell dissemination in a preclinical bone metastasis model. Int. J. Cancer 138, 2665–2677 (2016).
Richard, V. et al. The double agents in liquid biopsy: promoter and informant biomarkers of early metastases in breast cancer. Mol. Cancer 21, 95 (2022).
Xu, J. et al. ATP11B inhibits breast cancer metastasis in a mouse model by suppressing externalization of nonapoptotic phosphatidylserine. J. Clin. Invest. https://doi.org/10.1172/jci149473 (2022).
Jiang, C. C. et al. Signalling pathways in autism spectrum disorder: mechanisms and therapeutic implications. Signal Transduct. Target. Ther. 7, 229 (2022).
Bheda, A., Creek, K. E. & Pirisi, L. Loss of p53 induces epidermal growth factor receptor promoter activity in normal human keratinocytes. Oncogene 27, 4315–4323 (2008).
Linder, M. et al. EGFR is required for FOS-dependent bone tumor development via RSK2/CREB signaling. EMBO Mol. Med. https://doi.org/10.15252/emmm.201809408 (2018).
Rives, A. et al. Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences. Proc. Natl Acad. Sci. USA 118, e2016239118 (2021).
Madani, A. et al. Large language models generate functional protein sequences across diverse families. Nat. Biotechnol. 41, 1099–1106 (2023).
Yang, F. et al. scBERT as a large-scale pretrained deep language model for cell type annotation of single-cell RNA-seq data. Nat. Mach. Intell. 4, 852 (2022).
Theodoris, C. V. et al. Transfer learning enables predictions in network biology. Nature 618, 616–624 (2023).
Cui, H. T. et al. scGPT: toward building a foundation model for single-cell multi-omics using generative AI. Nat. Methods 21, 1470–1480 (2024).
Wu, A. et al. Causality for large language models. Preprint at https://arxiv.org/abs/2410.15319 (2024).
Lee, S. et al. Reasoning abilities of large language models: in-depth analysis on the abstraction and reasoning corpus. Preprint at https://arxiv.org/abs/2403.11793 (2024).
Kipf, T. N. & Welling, M. Variational graph auto-encoders. Preprint at https://arxiv.org/abs/1611.07308 (2016).
Wu, Z. H. et al. A comprehensive survey on graph neural networks. IEEE Trans. Neural Netw. Learn. Syst. 32, 4–24 (2021).
Miao, Z., Humphreys, B. D., McMahon, A. P. & Kim, J. Multi-omics integration in the age of million single-cell data. Nat. Rev. Nephrol. 17, 710–724 (2021).
Ma, A. et al. Integrative methods and practical challenges for single-cell multi-omics. Trends Biotechnol. 38, 1007–1022 (2020).
Zhang, Y. et al. DeepPhagy: a deep learning framework for quantitatively measuring autophagy activity in Saccharomyces cerevisiae. Autophagy 16, 626–640 (2020).
Tang, D., Zhang, C., Peng, D. & Xue, Y. Transcriptome of Saccharomyces cerevisiae during glucose starvation. Datasets. SRA https://www.ncbi.nlm.nih.gov/sra/?term=PRJNA912308 (2025).
Tang, D., Zhang, C., Peng, D. & Xue, Y. The proteome and phosphoproteome of Saccharomyces cerevisiae during glucose starvation. Datasets. iProX https://www.iprox.cn//page/project.html?id=IPX0005607000 (2025).
Tang, D., Zhang, C., Peng, D. & Xue, Y. LyMOI: large hybrid models for omics interpretation. Source code. GitHub https://github.com/BioCUCKOO/LyMOI (2025).
Acknowledgements
We thank L. Yang for reading the manuscript and editing the abstract, and Y. Cui for helpful suggestions on experiments. This work was supported by grants from the Natural Science Foundation of China (32341020 and 32341021 to Y.X., 32571718 to D.P.), the National Key R & D Program of China (2022YFC2704304 and 2021YFF0702000 to Y.X.), the Interdisciplinary Research Program of HUST (2023JCYJ010 and 2024JCYJ013 to Y.X.), the Hubei Province Postdoctoral Outstanding Talent Tracking Support Program (to D.P.), the Natural Science Foundation of Hubei Province of China (JCZRYB202500751 to D.P.), the start-up funding of Hubei Hongshan Laboratory (to Y.X.) and the Research Core Facilities for Life Science (HUST to Y.X.).
Author information
Authors and Affiliations
Contributions
Y.X. and D.P. initiated the project and oversaw all aspects of the project. C.Z. developed the LyMOI framework, with the help of D.T, D.P., X.H. and W.Z. D.T., C.Z. and D.P. compiled the multi-omics data and carried out data analysis. D.T. and D.P. performed the experiments with the help of D.J., H.-M.S., L.Z., L.X., D.L., S.F., F.L., C.S., J.S., M.Z., B.L. and G.C. Y.X., D.T. and C.Z. wrote the manuscript with input from all authors. All authors reviewed and approved the manuscript for publication.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Methods, Figs. 1–7 and References.
Supplementary Data 1 (download ZIP )
Supplementary Tables 1–7 and source data for Supplementary Figs. 1–3.
Supplementary Data 2 (download XLSX )
Statistical source data for Supplementary Figs. 3–6.
Source data
Source Data Figs. 1–7 (download XLSX )
Statistical source data.
Source Data Figs. 4–6 (download PDF )
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tang, D., Zhang, C., Zhang, W. et al. A deep learning and large language hybrid workflow for omics interpretation. Nat. Biomed. Eng (2026). https://doi.org/10.1038/s41551-025-01576-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41551-025-01576-5