Abstract
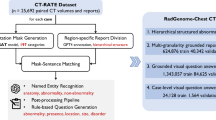
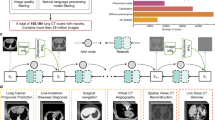
Advancements in medical imaging AI, particularly in 3D imaging, have been limited due to the scarcity of comprehensive datasets. We introduce CT-RATE, a public dataset that pairs 3D medical images with corresponding textual reports. CT-RATE comprises 25,692 non-contrast 3D chest CT scans from 21,304 unique patients. Each scan is accompanied by its corresponding radiology report. Leveraging CT-RATE, we develop CT-CLIP, a CT-focused contrastive language–image pretraining framework designed for broad applications without the need for task-specific training. We demonstrate how CT-CLIP can be used in multi-abnormality detection and case retrieval, and outperforms state-of-the-art fully supervised models across all key metrics. By combining CT-CLIP’s vision encoder with a pretrained large language model, we create CT-CHAT, a vision–language foundational chat model for 3D chest CT volumes. Fine-tuned on over 2.7 million question–answer pairs derived from the CT-RATE dataset, CT-CHAT underscores the necessity for specialized methods in 3D medical imaging. Collectively, the open-source release of CT-RATE, CT-CLIP and CT-CHAT not only addresses critical challenges in 3D medical imaging but also lays the groundwork for future innovations in medical AI and improved patient care.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Our CT-RATE dataset, comprising paired chest CT volumes and radiology text reports, is publicly accessible via Hugging Face at https://huggingface.co/datasets/ibrahimhamamci/CT-RATE (ref. 66). The external validation set, RAD-ChestCT, is also openly accessible through Zenodo at https://doi.org/10.5281/zenodo.6406114 (ref. 67). Source data are provided with this paper.
Code availability
Our trained models and codebase are publicly available in GitHub at https://github.com/ibrahimethemhamamci/CT-CLIP (ref. 68) and https://github.com/ibrahimethemhamamci/CT-CHAT (ref. 69) for further research.
References
Salvolini, L., Secchi, E. B., Costarelli, L. & Nicola, M. D. Clinical applications of 2D and 3D CT imaging of the airways—a review. Eur. J. Radiol. 34, 9–25 (2000).
McCollough, C. H. & Rajiah, P. S. Milestones in CT: past, present, and future. Radiology 309, e230803 (2023).
Claessens, Y. E. et al. Early chest computed tomography scan to assist diagnosis and guide treatment decision for suspected community-acquired pneumonia. Am. J. Respir. Crit. Care Med. 192, 974–982 (2015).
Esteva, A. et al. Deep learning-enabled medical computer vision. npj Digit. Med. 4, 5 (2021).
Qin, C., Yao, D., Shi, Y. & Song, Z. Computer-aided detection in chest radiography based on artificial intelligence: a survey. Biomed. Eng Online 17, 113 (2018).
Lipková, J. et al. Deep learning-enabled assessment of cardiac allograft rejection from endomyocardial biopsies. Nat. Med. 28, 575–582 (2022).
Hamamci, I. E. et al. Diffusion-based hierarchical multi-label object detection to analyze panoramic dental X-rays. In International Conference on Medical Image Computing and Computer-Assisted Intervention (eds Greenspan, H. et al.) 389–399 (Springer, 2023).
Pati, S. et al. GaNDLF: the generally nuanced deep learning framework for scalable end-to-end clinical workflows. Commun. Eng 2, 23 (2021).
Johnson, A. et al. MIMIC-CXR, a de-identified publicly available database of chest radiographs with free-text reports. Sci. Data 6, 317 (2019).
Wang, X. et al. Chestx-ray8: hospital-scale chest X-ray database and benchmarks on weakly-supervised classification and localization of common thorax diseases. In 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 3462–3471 (IEEE, 2017).
Nguyen, H. Q. et al. VinDr-CXR: an open dataset of chest X-rays with radiologist’s annotations. Sci. Data 9, 429 (2020).
Irvin, J. A. et al. CheXpert: a large chest radiograph dataset with uncertainty labels and expert comparison. Proc. AAAI Conf. Artif. Intell. 33, 590–597 (2019).
Hamamci, I. E. et al. DENTEX: an abnormal tooth detection with dental enumeration and diagnosis benchmark for panoramic X-rays. Preprint at https://arxiv.org/abs/2305.19112 (2023).
Lu, M. Y. et al. A multimodal generative AI copilot for human pathology. Nature 634, 466–473 (2024).
Tiu, E. et al. Expert-level detection of pathologies from unannotated chest X-ray images via self-supervised learning. Nat. Biomed. Eng 6, 1399–1406 (2022).
Hu, B., Vasu, B. K. & Hoogs, A. J. X-MIR: explainable medical image retrieval. In 2022 IEEE/CVF Winter Conference on Applications of Computer Vision (WACV) 1544–1554 (IEEE, 2022).
Chen, X. et al. Recent advances and clinical applications of deep learning in medical image analysis. Med. Image Anal. 79, 102444 (2021).
Zhou, S. K. et al. A review of deep learning in medical imaging: imaging traits, technology trends, case studies with progress highlights, and future promises. Proc. IEEE Inst. Electr. Electron. Eng 109, 820–838 (2020).
Willemink, M. J. et al. Preparing medical imaging data for machine learning. Radiology 295, 4–15 (2020).
Draelos, R. L. et al. Machine-learning-based multiple abnormality prediction with large-scale chest computed tomography volumes. Med. Image Anal. 67, 101857 (2020).
Antonelli, M. et al. The medical segmentation decathlon. Nat. Commun. 13, 4128 (2022).
Bilic, P. et al. The liver tumor segmentation benchmark (LiTS). Med. Image Anal. 84, 102680 (2019).
Hamamci, I. E. et al. GenerateCT: text-conditional generation of 3D chest CT volumes. In European Conference on Computer Vision (eds Leonardis, A. et al.). https://doi.org/10.1007/978-3-031-72986-7_8 (Springer, 2024).
Gao, J. et al. GET3D: a generative model of high quality 3D textured shapes learned from images. Adv. Neural Inf. Process. Syst. 35, 31841–31854 (2022).
Xian, Y., Schiele, B. & Akata, Z. Zero-shot learning—the good, the bad and the ugly. In 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 3077–3086 (IEEE, 2017).
Liu, H., Li, C., Li, Y. & Lee, Y. J. Improved baselines with visual instruction tuning. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 26296–26306 (2024).
Li, C. et al. LLaVA-Med: training a large language-and-vision assistant for biomedicine in one day. Adv. Neural Inf. Process. Syst. 36, 28541–28564 (2023).
Lee, S., Youn, J., Kim, H., Kim, M. & Yoon, S. H. CXR-LLaVA: a multimodal large language model for interpreting chest X-ray images. Eur. Radiol. 35, 4374–4386 (2025).
Wu, C., Zhang, X., Zhang, Y., Wang, Y. & Xie, W. Towards generalist foundation model for radiology. Nat. Commun. 16, 7866 (2025).
Hamamci, I. E., Er, S. & Menze, B. H. CT2Rep: automated radiology report generation for 3D medical imaging. In International Conference on Medical Image Computing and Computer-Assisted Intervention (eds Linguraru, M.G. et al.). https://doi.org/10.1007/978-3-031-72390-2_45 (Springer, 2024).
Willemink, M. J. & Noël, P. B. The evolution of image reconstruction for CT—from filtered back projection to artificial intelligence. Eur. Radiol. 29, 2185–2195 (2018).
Xu, Y., Sun, L., Peng, W., Visweswaran, S. & Batmanghelich, K. MedSyn: text-guided anatomy-aware synthesis of high-fidelity 3D CT images. IEEE Trans. Med. Imaging 43, 3648–3660 (2023).
Yan, A. et al. RadBERT: adapting transformer-based language models to radiology. Radiol. Artif. Intell. 44, e210258 (2022).
Minaee, S., Cambria, E. & Gao, J. Deep learning-based text classification: a comprehensive review. ACM Comput. Surv. 54, 1–40 (2020).
Radford, A. et al. Learning transferable visual models from natural language supervision. Proc. Mach. Learn. Res. 139, 8748–8763 (2021).
Zhang, H., Koh, J. Y., Baldridge, J., Lee, H. & Yang, Y. Cross-modal contrastive learning for text-to-image generation. In 2021 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 833–842 (IEEE, 2021).
Yong, G., Jeon, K., Gil, D. & Lee, G. Prompt engineering for zero-shot and few-shot defect detection and classification using a visual-language pretrained model. Comput. Aided Civ. Infrastruct. Eng 38, 1536–1554 (2022).
Bailly, A. et al. Effects of dataset size and interactions on the prediction performance of logistic regression and deep learning models. Comp. Methods Programs Biomed. 213, 106504 (2021).
Zhai, X., Kolesnikov, A., Houlsby, N. & Beyer, L. Scaling vision transformers. In 2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 1204–1213 (IEEE, 2021).
Wortsman, M. et al. Robust fine-tuning of zero-shot models. In 2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 7949–7961 (IEEE, 2021).
Chen, C. et al. Fast and scalable search of whole-slide images via self-supervised deep learning. Nat. Biomed. Eng 6, 1420–1434 (2022).
Zhang, S., Yang, M., Cour, T., Yu, K. & Metaxas, D. N. Query specific fusion for image retrieval. In European Conference on Computer Vision 660–673 (Springer, 2012).
Dubey, A. et al. The Llama 3 herd of models. Preprint at https://arxiv.org/abs/2407.21783 (2024).
Hu, E. J. et al. LoRA: low-rank adaptation of large language models. In Proc. 10th International Conference on Learning Representations (2022).
Hou, B. et al. Shadow and light: digitally reconstructed radiographs for disease classification. Preprint at https://arxiv.org/abs/2406.03688 (2024).
Hartung, M. P., Bickle, I. C., Gaillard, F. & Kanne, J. P. How to create a great radiology report. Radiographics 40, 1658–1670 (2020).
Kelly, C. J., Karthikesalingam, A., Suleyman, M., Corrado, G. C. & King, D. Key challenges for delivering clinical impact with artificial intelligence. BMC Med. 17, 195 (2019).
Boecking, B. et al. Making the most of text semantics to improve biomedical vision–language processing. In European Conference on Computer Vision (eds Avidan, S. et al.) 1–21 (Springer, 2022).
Kazerooni, E. A. & Gross, B. H. Cardiopulmonary Imaging Vol. 4 (Lippincott Williams & Wilkins, 2004).
Xu, M., Amiranashvili, T., Navarro, F. & Menze, B. H. CADS: Comprehensive Anatomical Dataset and Segmentation for Whole-body CT Technical Report (Department of Quantitative Biomedicine, Univ. Zurich, 2025).
Huang, Z., Bianchi, F., Yuksekgonul, M., Montine, T. J. & Zou, J. Y. A visual–language foundation model for pathology image analysis using medical Twitter. Nat. Med. 29, 2307–2316 (2023).
Zhang, Z. & Sabuncu, M. Generalized cross entropy loss for training deep neural networks with noisy labels. In Advances in Neural Information Processing Systems 31 (2018).
Alain, G. & Bengio, Y. Understanding intermediate layers using linear classifier probes. In Proc. 5th International Conference on Learning Representations (2017).
van der Maaten, L. & Hinton, G. E. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Chen, R. J. et al. Towards a general-purpose foundation model for computational pathology. Nat. Med. 30, 850–862 (2024).
Kherfi, M. L., Ziou, D. & Bernardi, A. Image retrieval from the world wide web: issues, techniques, and systems. ACM Comput. Surv. 36, 35–67 (2004).
Hegde, N. et al. Similar image search for histopathology: SMILY. npj Digit. Med. 2, 56 (2019).
Zhang, X. et al. Development of a large-scale grounded vision language dataset for chest CT analysis. Sci. Data 12, 1636 (2025).
Liu, H., Li, C., Wu, Q. & Lee, Y. J. Visual instruction tuning. Adv. Neural Inf. Process. Syst. 36, 34892–34916 (2023).
Wasserthal, J. et al. Totalsegmentator: robust segmentation of 104 anatomic structures in CT images. Radiol. Artif. Intell. 5, e230024 (2022).
Papineni, K., Roukos, S., Ward, T. & Zhu, W.-J. Bleu: a method for automatic evaluation of machine translation. In Proc. 40th Annual Meeting of the Association for Computational Linguistics 311–318 (Association for Computational Linguistics, 2002).
Banerjee, S. & Lavie, A. Meteor: an automatic metric for MT evaluation with improved correlation with human judgments. In Proc. ACL Workshop on Intrinsic and Extrinsic Evaluation Measures for Machine Translation and/or Summarization (eds Goldstein, J. et al.) 65–72 (Association for Computational Linguistics, 2005).
Lin, C.-Y. ROUGE: a package for automatic evaluation of summaries. In Annual Meeting of the Association for Computational Linguistics 74–81 (Association for Computational Linguistics, 2004).
Vedantam, R., Zitnick, C. L. & Parikh, D. Cider: consensus-based image description evaluation. In 2015 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 4566–4575 (IEEE, 2014).
Hamamci, I. E. et al. CRG score: a distribution-aware clinical metric for radiology report generation. In International Conference on Medical Imaging with Deep Learning 1–4 (MIDL, 2025).
Hamamci, I. E. CT-RATE: paired chest CT volumes and radiology reports. Hugging Face https://huggingface.co/datasets/ibrahimhamamci/CT-RATE (2025).
Draelos, R. L. RAD-ChestCT: radiology reports and CT imaging dataset. Zenodo https://doi.org/10.5281/zenodo.6406114 (2022).
Hamamci, I. E. CT-CLIP: contrastive language–image pretraining for CT. GitHub https://github.com/ibrahimethemhamamci/CT-CLIP (2025).
Hamamci, I. E. CT-CHAT: a vision–language foundational chat model for 3D chest CT. GitHub https://github.com/ibrahimethemhamamci/CT-CHAT (2025).
Acknowledgements
We thank the Helmut Horten Foundation for generous support, which made this work possible, and the Istanbul Medipol University for invaluable support and provision of data. Some of the HPC resources were provided by the Erlangen National High Performance Computing Center (NHR@FAU) at Friedrich-Alexander-Universität Erlangen-Nürnberg under NHR projects b143dc and b180dc. NHR is funded by federal and Bavarian state authorities, and the NHR@FAU hardware is partly funded by the German Research Foundation (DFG; grant 440719683). This research was supported in part by the Intramural Research Program of the National Library of Medicine (NLM), National Institutes of Health (NIH), and used the computational resources of the NIH high-performance computing Biowulf cluster. B.K. received research support from the ERC project MIA-NORMAL 101083647, DFG grants 513220538 and 512819079, and the state of Bavaria (HTA). C.B. received research support from the Promedica Foundation, Chur, Switzerland. C.W., W.D. and K.B. were supported by NIH grant R01 HL141813, the Hariri Institute for Computing’s Junior Faculty Fellows Program, and its Focused Research Program. Figures 1, 3, 5 and 6 were created with BioRender.com.
Author information
Authors and Affiliations
Contributions
I.E.H., S.E. and B.M. designed the study. I.E.H. and S.E. handled data collection, data analysis, model construction and validation, figure preparation and writing of the paper. F.A., A.G.S., S.N.E. and I.D. managed data anonymization and annotation. M.F.D. contributed to model construction. C.W., W.D. and K.B. carried out external evaluation. O.F.D. developed the graphical user interface and assisted with writing of the paper. M.X. and T.A. generated anatomical-segmentation labels. S.S. maintained the model repository. B.H. assisted in preparing NIfTI files with metadata and integrating nodule-segmentation masks into the report-generation pipeline. M.P. assessed report-generation performance. B.K. provided technical expertise and contributed to writing of the paper. H.R., B.W., E.S., M.S., E.B.E., A.A., A.S., A.K., Z.L., B.L., C.B. and M.K.O. provided domain-knowledge support. B.M. supervised the entire study. All authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks Imon Banerjee, Chang Min Park and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Abnormality-based t-SNE projections for chest CT volumes and radiology text reports.
Two-dimensional t-SNE visualizations showing the joint embedding space of chest CT volumes and their associated radiology text reports, color-coded by abnormality categories.
Extended Data Fig. 2 Abnormality-based t-SNE projections for chest CT volumes and radiology text reports (continued).
Additional t-SNE projections illustrating the joint embedding space of chest CT volumes and radiology text reports across further abnormality categories.
Extended Data Fig. 3 Comparison of abnormality-based performance metrics in the internal validation set.
This figure provides a detailed analysis of performance metrics, including AUROC, accuracy, precision, and F1 scores, for detecting various abnormalities with our models in the internal validation set, compared to the fully supervised baseline model. The proposed CT-CLIP based models demonstrate improvement over the baseline in almost all comparisons.
Extended Data Fig. 4 Comparison of abnormality-based performance metrics in the Rad-ChestCT validation set.
This figure provides a detailed analysis of performance metrics, including AUROC, accuracy, precision, and F1 scores, for detecting various abnormalities with our models in the Rad-ChestCT validation set, compared to the fully supervised baseline model. It highlights the models’ remarkable adaptability and superior effectiveness with distribution shifts, setting a new standard in performance compared to a fully supervised baseline model.
Extended Data Fig. 5 Comparison of abnormality-based performance metrics in the UPMC validation set.
This figure provides a detailed analysis of performance metrics, including AUROC, accuracy, precision, and F1 scores, for detecting various abnormalities with our models in the UPMC validation set, compared to the fully supervised baseline model. It highlights the models’ remarkable adaptability and superior effectiveness with distribution shifts, setting a new standard in performance compared to a fully supervised baseline model.
Extended Data Fig. 6 Comparative metrics for CT-CHAT and baselines.
This figure illustrates the performance of CT-CHAT and baselines across various tasks, demonstrating that CT-CHAT consistently outperforms the baselines, achieving the highest accuracy in every metric for each task.
Extended Data Fig. 7 Comparative metrics for CT-CHAT and baselines in report generation.
This figure presents the report generation performance of CT-CHAT compared to baseline models across multiple abnormalities, evaluated on two independent validation sets. CT-CHAT consistently outperforms the baselines, demonstrating superior clinical accuracy.
Extended Data Fig. 8 Representative examples from VQA.
This dataset includes various question types, such as long-answer (which encompass conversational, descriptive, free-response, and text-only questions), multiple-choice, short-answer, and report-generation questions.
Extended Data Fig. 9 Example outputs of CT-CHAT across different question types.
Representative CT-CHAT responses for (a) long-answer, (b) short-answer, (c) multiple-choice, and (d) report-generation questions, illustrating the model’s ability to handle diverse clinical query formats.
Supplementary information
Supplementary Information (download PDF )
Table of Contents, and Supplementary Figs. 1–11 and Tables 1–11.
Source data
Source Data Fig. 2 (download CSV )
Statistical source data.
Source Data Fig. 4 (download ZIP )
Statistical source data.
Source Data Fig. 6 (download ZIP )
Statistical source data.
Source Data Extended Data Fig. 3 (download CSV )
Statistical source data.
Source Data Extended Data Fig. 4 (download CSV )
Statistical source data.
Source Data Extended Data Fig. 5 (download CSV )
Statistical source data.
Source Data Extended Data Fig. 7 (download ZIP )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hamamci, I.E., Er, S., Wang, C. et al. Generalist foundation models from a multimodal dataset for 3D computed tomography. Nat. Biomed. Eng (2026). https://doi.org/10.1038/s41551-025-01599-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41551-025-01599-y