Abstract
Bacterial biofilms, prevalent in human infections, present a major barrier to effective antibacterial therapy due to limited drug permeability and resistance. Here we introduce a ‘trick-bacteria-with-bacteria’ strategy that employs bacteria modified via calcium chloride treatment and antibiotic loading, followed by ultraviolet inactivation. These modified bacteria integrate selectively into biofilms of the same species, enabling targeted intra-biofilm drug release triggered by local pH and hydrogen peroxide. Species-specific integration is essential, as mismatched strains exhibit spatial segregation due to differences in surface adhesins and protein profiles. The strategy is effective against polymicrobial biofilms and demonstrated efficacy in treating biofilms formed by Staphylococcus aureus, Escherichia coli and Candida albicans. It also reinvigorates biofilm-associated macrophages by inducing the release of biofilm-derived l-arginine, enhancing immune responses. In vivo studies using subcutaneous and bone implant infection models showed stronger biofilm eradication and longer-term immunity in animals treated with modified bacteria compared with those treated with antibiotics, including resistance to re-infection. This approach could be adapted to modify infection-related bacteria from patients for personalized intra-biofilm drug delivery.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All data are available in the main text or Supplementary Information. Source data are provided with this paper.
References
Choi, V., Rohn, J. L., Stoodley, P., Carugo, D. & Stride, E. Drug delivery strategies for antibiofilm therapy. Nat. Rev. Microbiol. 21, 555–572 (2023).
Mah, T. F. & O’Toole, G. A. Mechanisms of biofilm resistance to antimicrobial agents. Trends Microbiol. 9, 34–39 (2001).
Ciofu, O., Moser, C., Jensen, P. O. & Hoiby, N. Tolerance and resistance of microbial biofilms. Nat. Rev. Microbiol. 20, 621–635 (2022).
Masters, E. A. et al. Skeletal infections: microbial pathogenesis, immunity and clinical management. Nat. Rev. Microbiol. 20, 385–400 (2022).
Wouthuyzen-Bakker, M. et al. How to handle concomitant asymptomatic prosthetic joints during an episode of hematogenous periprosthetic joint infection, a multicenter analysis. Clin. Infect. Dis. 73, e3820–e3824 (2021).
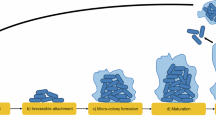
Sauer, K. et al. The biofilm life cycle: expanding the conceptual model of biofilm formation. Nat. Rev. Microbiol. 20, 608–620 (2022).
Bayles, K. W. The biological role of death and lysis in biofilm development. Nat. Rev. Microbiol. 5, 721–726 (2007).
Winstel, V., Kühner, P., Rohde, H. & Peschel, A. Genetic engineering of untransformable coagulase-negative staphylococcal pathogens. Nat. Protoc. 11, 949–959 (2016).
Gurbatri, C. R., Arpaia, N. & Danino, T. Engineering bacteria as interactive cancer therapies. Science 378, 858–864 (2022).
Chen, Y. E. et al. Engineered skin bacteria induce antitumor T cell responses against melanoma. Science 380, 203–210 (2023).
Stary, G. et al. VACCINES. A mucosal vaccine against Chlamydia trachomatis generates two waves of protective memory T cells. Science 348, aaa8205 (2015).
Jaiaue, P. et al. Mathematical modeling for evaluating inherent parameters affecting UVC decontamination of indicator bacteria. Appl. Environ. Microbiol. 88, e0214821 (2022).
Arciola, C. R., Campoccia, D. & Montanaro, L. Implant infections: adhesion, biofilm formation and immune evasion. Nat. Rev. Microbiol. 16, 397–409 (2018).
Guo, G. et al. Space-selective chemodynamic therapy of CuFe5O8 nanocubes for implant-related infections. ACS Nano 14, 13391–13405 (2020).
Schilcher, K. & Horswill, A. R. Staphylococcal biofilm development: structure, regulation, and treatment strategies. Microbiol. Mol. Biol. Rev. 84, e00026–00019 (2020).
Geiger, R. et al. L-arginine modulates T cell metabolism and enhances survival and anti-tumor activity. Cell 167, 829–842.e813 (2016).
Canale, F. P. et al. Metabolic modulation of tumours with engineered bacteria for immunotherapy. Nature 598, 662–666 (2021).
Gao, L. et al. Orchestrating nitric oxide and carbon monoxide signaling molecules for synergistic treatment of MRSA infections. Angew. Chem. Int. Ed. 61, e202112782 (2022).
Wu, L., Bao, F., Li, L., Yin, X. & Hua, Z. Bacterially mediated drug delivery and therapeutics: strategies and advancements. Adv. Drug. Deliv. Rev. 187, 114363 (2022).
Dubnau, D. & Blokesch, M. Mechanisms of DNA uptake by naturally competent bacteria. Annu. Rev. Genet. 53, 217–237 (2019).
Huang, R. & Reusch, R. N. Genetic competence in Escherichia coli requires poly-beta-hydroxybutyrate/calcium polyphosphate membrane complexes and certain divalent cations. J. Bacteriol. 177, 486–490 (1995).
Aich, P., Patra, M., Chatterjee, A. K., Roy, S. S. & Basu, T. Calcium chloride made E. coli competent for uptake of extraneous DNA through overproduction of OmpC protein. Protein J. 31, 366–373 (2012).
Turnbull, L. et al. Explosive cell lysis as a mechanism for the biogenesis of bacterial membrane vesicles and biofilms. Nat. Commun. 7, 11220 (2016).
Flemming, H. C. et al. The biofilm matrix: multitasking in a shared space. Nat. Rev. Microbiol. 21, 70–86 (2022).
Davies, M. J. & Truscott, R. J. W. in Comprehensive Series in Photosciences Vol. 3 (ed. Giacomoni, P. U.) Ch. 12 (Elsevier, 2001).
Vollmer, W., Blanot, D. & De Pedro, M. A. Peptidoglycan structure and architecture. FEMS Microbiol. Rev. 32, 149–167 (2008).
Pasquina-Lemonche, L. et al. The architecture of the Gram-positive bacterial cell wall. Nature. 582, 294–297 (2020).
Patel, M., Andoy, N. M. O., Tran, S. M., Jeon, K. & Sullan, R. M. A. Different drug loading methods and antibiotic structure modulate the efficacy of polydopamine nanoparticles as drug nanocarriers. J. Mater. Chem. B 11, 11335–11343 (2023).
Chen, C. et al. Single-particle assessment of six different drug-loading strategies for incorporating doxorubicin into small extracellular vesicles. Anal. Bioanal. Chem. 415, 1287–1298 (2022).
Chang, L. et al. Micro-/nanoscale electroporation. Lab Chip 16, 4047–4062 (2016).
Vajente, M., Clerici, R., Ballerstedt, H., Blank, L. M. & Schmidt, S. Using Cupriavidus necator H16 to provide a roadmap for increasing electroporation efficiency in nonmodel bacteria. ACS Synth. Biol. 14, 4330–4341 (2024).
Makabenta, J. M. V. et al. Nanomaterial-based therapeutics for antibiotic-resistant bacterial infections. Nat. Rev. Microbiol. 19, 23–36 (2021).
Hamad, C. D. et al. Comparing the in vitro efficacy of commonly used surgical irrigants for the treatment of implant-associated infections. J. Bone Joint. Surg. Am. 107, 1818–1824 (2025).
Uberoi, A., McCready-Vangi, A. & Grice, E. A. The wound microbiota: microbial mechanisms of impaired wound healing and infection. Nat. Rev. Microbiol. 22, 507–521 (2024).
Ruiz-Sorribas, A., Poilvache, H. & Van Bambeke, F. Pharmacodynamics of moxifloxacin, meropenem, caspofungin, and their combinations against in vitro polymicrobial interkingdom biofilms. Antimicrob. Agents Chemother. 66, e0214921 (2022).
Vlaeminck, J. et al. The dynamic transcriptome during maturation of biofilms formed by methicillin-resistant Staphylococcus aureus. Front. Microb. 13, 882346 (2022).
Mishra, S. & Imlay, J. Why do bacteria use so many enzymes to scavenge hydrogen peroxide? Arch. Biochem. Biophys. 525, 145–160 (2012).
Gehrke, T. et al. Organism profile in periprosthetic joint infection: pathogens differ at two arthroplasty infection referral centers in Europe and in the United States. J. Knee Surg. 27, 399–406 (2014).
Savage, V. J., Chopra, I. & O’Neill, A. J. Staphylococcus aureus biofilms promote horizontal transfer of antibiotic resistance. Antimicrob. Agents Chemother. 57, 1968–1970 (2013).
Wang, Y. et al. CCR2 contributes to host defense against Staphylococcus aureus orthopedic implant-associated infections in mice. J. Orthop. Res. 40, 409–419 (2021).
El Kasmi, K. C. et al. Toll-like receptor-induced arginase 1 in macrophages thwarts effective immunity against intracellular pathogens. Nat. Immunol. 9, 1399–1406 (2008).
Hanke, M. L. & Kielian, T. Deciphering mechanisms of staphylococcal biofilm evasion of host immunity. Front. Cell. Infect. Microbiol. 2, 62 (2012).
Urbano, R. et al. Host nitric oxide disrupts microbial cell-to-cell communication to inhibit staphylococcal virulence. Cell Host Microbe 23, 594–606 (2018).
Garrett, W. S. Immune recognition of microbial metabolites. Nat. Rev. Immunol. 20, 91–92 (2020).
Roewe, J. et al. Bacterial polyphosphates interfere with the innate host defense to infection. Nat. Commun. 11, 4035 (2020).
Morita, N. et al. GPR31-dependent dendrite protrusion of intestinal CX3CR1+ cells by bacterial metabolites. Nature 566, 110–114 (2019).
Thurlow, L. R. et al. Functional modularity of the arginine catabolic mobile element contributes to the success of USA300 methicillin-resistant Staphylococcus aureus. Cell Host Microbe 13, 100–107 (2013).
Yang, C. et al. Inorganic nanosheets facilitate humoral immunity against medical implant infections by modulating immune co-stimulatory pathways. Nat. Commun. 13, 4866 (2022).
Blaskovich, M. A. T. et al. A lipoglycopeptide antibiotic for Gram-positive biofilm-related infections. Sci. Transl. Med. 14, eabj2381 (2022).
Hang, S. et al. Bile acid metabolites control TH17 and Treg cell differentiation. Nature 576, 143–148 (2019).
Cortes-Penfield, N. et al. How we approach suppressive antibiotic therapy following debridement, antibiotics, and implant retention for prosthetic joint infection. Clin. Infect. Dis. 78, 188–198 (2024).
Ren, Y. et al. Evidence of bisphosphonate-conjugated sitafloxacin eradication of established methicillin-resistant S. aureus infection with osseointegration in murine models of implant-associated osteomyelitis. Bone Res. 11, 51 (2023).
Mazzolini, R. et al. Engineered live bacteria suppress Pseudomonas aeruginosa infection in mouse lung and dissolve endotracheal-tube biofilms. Nat. Biotechnol. 41, 1089–1098 (2023).
Lei, B., van den Berg, S., de Vogel, C. P., van Belkum, A. & Bakker-Woudenberg, I. A. J. M. Mild Staphylococcus aureus skin infection improves the course of subsequent endogenous S. aureus bacteremia in mice. PLoS ONE 10, e0129150 (2015).
Vincent, R. L. et al. Probiotic-guided CAR-T cells for solid tumor targeting. Science 382, 211–218 (2023).
Zhou, S., Gravekamp, C., Bermudes, D. & Liu, K. Tumour-targeting bacteria engineered to fight cancer. Nat. Rev. Cancer 18, 727–743 (2018).
Hwang, I. Y. et al. Engineered probiotic Escherichia coli can eliminate and prevent Pseudomonas aeruginosa gut infection in animal models. Nat. Commun. 8, 15028 (2017).
Tan, L. et al. Engineered probiotics biofilm enhances osseointegration via immunoregulation and anti-infection. Sci. Adv. 6, eaba5723 (2020).
Lu, J. et al. Inactive trojan bacteria as safe drug delivery vehicles crossing the blood–brain barrier. Nano Lett. 23, 4326–4333 (2023).
Li, Z. et al. Chemically and biologically engineered bacteria-based delivery systems for emerging diagnosis and advanced therapy. Adv. Mater. 33, e2102580 (2021).
Su, L. et al. Synergy between pH- and hypoxia-responsiveness in antibiotic-loaded micelles for eradicating mature, infectious biofilms. Acta Biomater. 154, 559–571 (2022).
Damyanova, T. et al. Gram negative biofilms: structural and functional responses to destruction by antibiotic-loaded mixed polymeric micelles. Microorganisms 12, 2670 (2024).
Yildirim, T. et al. A new class of polyion complex vesicles (PIC-somes) to improve antimicrobial activity of tobramycin in Pseudomonas aeruginosa biofilms. Small 20, e2401926 (2024).
Sedighi, O., Bednarke, B., Sherriff, H. & Doiron, A. L. Nanoparticle-based strategies for managing biofilm infections in wounds: a comprehensive review. ACS Omega 9, 27853–27871 (2024).
Fulaz, S., Vitale, S., Quinn, L. & Casey, E. Nanoparticle–biofilm interactions: the role of the EPS matrix. Trends Microbiol. 27, 915–926 (2019).
Panthi, V. K., Fairfull-Smith, K. E. & Islam, N. Liposomal drug delivery strategies to eradicate bacterial biofilms: challenges, recent advances, and future perspectives. Int. J. Pharm. 655, 124046 (2024).
Koo, H., Allan, R. N., Howlin, R. P., Stoodley, P. & Hall-Stoodley, L. Targeting microbial biofilms: current and prospective therapeutic strategies. Nat. Rev. Microbiol. 15, 740–755 (2017).
Liu, Y. et al. Nanotechnology-based antimicrobials and delivery systems for biofilm-infection control. Chem. Soc. Rev. 48, 428–446 (2019).
Stacy, A., McNally, L., Darch, S. E., Brown, S. P. & Whiteley, M. The biogeography of polymicrobial infection. Nat. Rev. Microbiol. 14, 93–105 (2015).
Chew, S. C. et al. Dynamic remodeling of microbial biofilms by functionally distinct exopolysaccharides. mBio 5, e01536–01514 (2014).
Kim, W., Racimo, F., Schluter, J., Levy, S. B. & Foster, K. R. Importance of positioning for microbial evolution. Proc. Natl Acad. Sci. USA 111, E1639–E1647 (2014).
Korgaonkar, A., Trivedi, U., Rumbaugh, K. P. & Whiteley, M. Community surveillance enhances Pseudomonas aeruginosa virulence during polymicrobial infection. Proc. Natl Acad. Sci. USA 110, 1059–1064 (2012).
Yang, L. et al. Pattern differentiation in co-culture biofilms formed by Staphylococcus aureus and Pseudomonas aeruginosa. FEMS Immunol. Med. Microbiol. 62, 339–347 (2011).
Harding, J. L. & Reynolds, M. M. Combating medical device fouling. Trends Biotechnol. 32, 140–146 (2014).
Zecconi, A. & Scali, F. Staphylococcus aureus virulence factors in evasion from innate immune defenses in human and animal diseases. Immunol. Lett. 150, 12–22 (2013).
Scherr, T. D. et al. Staphylococcus aureus biofilms induce macrophage dysfunction through leukocidin AB and alpha-toxin. mBio 6, e01021–01015 (2015).
Alboslemy, T., Yu, B., Rogers, T., Kim, M.-H. & Freitag, N. E. Staphylococcus aureus biofilm-conditioned medium impairs macrophage-mediated antibiofilm immune response by upregulating KLF2 expression. Infect. Immun. 87, e00643–00618 (2019).
Marullo, R. et al. The metabolic adaptation evoked by arginine enhances the effect of radiation in brain metastases. Sci. Adv. 7, eabg1964 (2021).
Ralph, A. P., Kelly, P. M. & Anstey, N. M. L-arginine and vitamin D: novel adjunctive immunotherapies in tuberculosis. Trends Microbiol. 16, 336–344 (2008).
Rossato, A. M., Primon-Barros, M., Dias, C. A. G. & d’Azevedo, P. A. Vancomycin MIC and agr dysfunction in invasive MRSA infections in southern Brazil. Braz. J. Microbiol. 51, 1819–1823 (2020).
Britt, N. S., Hazlett, D. S., Horvat, R. T., Liesman, R. M. & Steed, M. E. Activity of pulmonary vancomycin exposures versus planktonic and biofilm isolates of methicillin-resistant Staphylococcus aureus from cystic fibrosis sputum. Int. J. Antimicrob. Agents 55, 105898 (2020).
Diaz, R., Afreixo, V., Ramalheira, E., Rodrigues, C. & Gago, B. Evaluation of vancomycin MIC creep in methicillin-resistant Staphylococcus aureus infections—a systematic review and meta-analysis. Clin. Microbiol. Infect. 24, 97–104 (2018).
Hartsell, E. M., Gillespie, M. N. & Langley, R. J. Does acute and persistent metabolic dysregulation in COVID-19 point to novel biomarkers and future therapeutic strategies? Eur. Clin. Respir. J. 59, 2102417 (2022).
Li, Y. et al. Untargeted metabolomics of saliva in caries-active and caries-free children in the mixed dentition. Front. Cell. Infect. Microbiol. 13, 1104295 (2023).
Gordon, O. et al. Dynamic PET-facilitated modeling and high-dose rifampin regimens for Staphylococcus aureus orthopedic implant-associated infections. Sci. Transl. Med. 13, eabl6851 (2021).
Li, L. et al. Burst release of encapsulated annexin A5 in tumours boosts cytotoxic T-cell responses by blocking the phagocytosis of apoptotic cells. Nat. Biomed. Eng. 4, 1102–1116 (2020).
Acknowledgements
We acknowledge financial support from the National Key R&D Program of China (2022YFB3804500; to H.L.), Shanghai Pilot Program for Basic Research-Chinese Academy of Science, Shanghai Branch (JCYJ-SHFY-2022-003; to H.L.), National Natural Science Foundation of China (52372276, 22335006 and 82302717; to H.L. and C.Y.), Youth Innovation Promotion Association CAS (2023262; to H.L.), Young Elite Scientists Sponsorship Program by CAST (YESS20210149; to H.L.), Shanghai Science and Technology Committee Rising-Star Program (22QA1410200; to H.L.), China Postdoctoral Science Foundation (2023M732310; to C.Y.), Shanghai Sailing Program (23YF1432200; to C.Y.), Natural Science Foundation of Shanghai (23ZR1472300; to H.L.), Harvard/Brigham Health and Technology Innovation Fund (2023A004452; to W.T.), Gillian Reny Stepping Strong Center for Trauma Innovation Breakthrough Innovator Award (113548; to W.T.), Department Basic Scientist Grant (2420 BPA075; to W.T.), Nanotechnology Foundation (2022A002721; to W.T.), and Distinguished Chair Professorship Foundation (018129; to W.T.).
Author information
Authors and Affiliations
Contributions
C.Y., M.G., H.L. and W.T. conceived of the study. C.Y., M.G., H.L. and W.T. designed the experiments. C.Y., M.G., H.L., Q.S., W.C., S.A., M.M.K., N.K., S.Z. and J.S. performed the experiments, discussed and analysed the data or provided essential experimental resources. C.Y., M.G., H.L. and W.T. wrote the paper, and the paper was revised by all the authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks Xinge Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The penetration of modified S. aureus into S. aureus biofilm.
S. aureus-eGFP biofilm (green) was treated with modified S. aureus-mcherry (red). The fluorescent images showed the penetration process of the tricker S. aureus. The experiments were repeated independently at least three times with similar results.
Extended Data Fig. 2 The penetration of modified E.coli into E.coli biofilm.
E. coli-sfgfp biofilm (green) was treated with modified E. coli-mcherry (red). The fluorescent images showed the penetration process of the tricker E. coli. The experiments were repeated independently at least three times with similar results.
Extended Data Fig. 3 The penetration of modified E.coli into S. aureus biofilm.
S. aureus-eGFP biofilm (green) was treated with modified E. coli-mcherry (red). The experiments were repeated independently at least three times with similar results.
Extended Data Fig. 4 The penetration of modified S. aureus into E.coli biofilm.
E. coli-sfgfp biofilm (green) was treated with modified S. aureus-mcherry (red). The experiments were repeated independently at least three times with similar results.
Extended Data Fig. 5 Protein bands of live and tricker bacteria.
The protein expression profile of live and tricker bacteria was determined by Coomassie Brilliant Blue staining. The experiments were repeated independently at least three times with similar results.
Extended Data Fig. 6 Penetration effect of tricker bacteria into polymicrobial biofilms.
a, Schematic of polymicrobial biofilms consists of S.aureus, E.coli and C.albicans. b, Low and high magnification of SEM images of the polymicrobial biofilms. S. aureus, E.coli and C.albicans are pseudo-colored with red, purple and brown, respectively. c, Flow cytometry plots of dislodged single-species biofilm or polymicrobial biofilms. d, Representative three-dimensional fluorescent images of biofilms (green) before and after incubation with PI-stained Van@Tr-S.a., Cip@Tr-E.c. or Flu@Tr-C.a. (red). For b, d, the experiments were repeated independently at least three times with similar results.
Supplementary information
Supplementary Information (download PDF )
Supplementary Methods, Figs. 1–34, Table 1 and References.
Supplementary Data 1 (download XLSX )
Source data for Supplementary figures.
Source data
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Unprocessed Coomassie Brilliant Blue staining gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yang, C., Saiding, Q., Chen, W. et al. Chemically modified and inactivated bacteria enable intra-biofilm drug delivery and long-term immunity against implant infections. Nat. Biomed. Eng (2026). https://doi.org/10.1038/s41551-025-01600-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41551-025-01600-8