Abstract
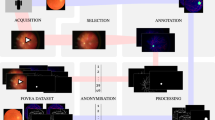
Foundation models in artificial intelligence are revolutionizing healthcare by utilizing large-scale unlabelled data for pretraining. However, their intraoperative applications remain underexplored owing to limited surgical data and the challenges of real-time deployment. Here we show the development of the ophthalmic video foundation model (OVFM), designed for microscopic ophthalmic surgical recognition and navigation. Leveraging a self-supervised video transformer structure and trained on an ophthalmic video dataset comprising 1.1 million clips across 144 surgical types, OVFM learns the spatiotemporal motion features of ophthalmic procedures. We demonstrate OVFM’s superior performance across seven downstream tasks. To enable real-time use, we applied knowledge distillation, reducing the model’s size while retaining its accuracy, which allows for deployment on surgical microscope units. In cataract surgeries performed by ten surgeons on wet-lab porcine eyes, the OVFM-powered system enhanced surgical performance and reduced skill gaps, demonstrating notable potential for real-time, intraoperative applications across various surgical fields.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The Cataract-1K dataset for pretraining is available at https://www.synapse.org/Synapse:syn53404507. The Cataract-1K surgical step recognition dataset is available at https://www.synapse.org/Synapse:syn53395146. The CATARACTS dataset, used for tool presence recognition, is available at https://ieee-dataport.org/open-access/cataracts. The Cataract-1K complications detection dataset is available at https://www.synapse.org/Synapse:syn53395402, and the Cataract-1K surgical scene segmentation dataset is available at https://www.synapse.org/Synapse:syn53395479. All in-house datasets can be available upon request from the corresponding authors, subject to a signed Data Use Agreement and Non-Disclosure Agreement with the respective hospital. Source data are provided with this paper.
Code availability
The code is publicly available at https://github.com/puxuntu/OVFM.
References
He, Y. et al. Foundation model for advancing healthcare: challenges, opportunities and future directions. IEEE Rev. Biomed. Eng. 18, 172–191 (2025).
Zhou, C. et al. A comprehensive survey on pretrained foundation models: a history from BERT to ChatGPT. Int. J. Mach. Learn. Cybern. 16, 9851–9915 (2025).
Moor, M. et al. Foundation models for generalist medical artificial intelligence. Nature 616, 259–265 (2023).
Christensen, M. et al. Vision–language foundation model for echocardiogram interpretation. Nat. Med. 30, 1481–1488 (2024).
Zhou, Y. et al. A foundation model for generalizable disease detection from retinal images. Nature 622, 156–163 (2023).
Xu, H. et al. A whole-slide foundation model for digital pathology from real-world data. Nature 630, 181–188 (2024).
Chen, R. J. et al. Towards a general-purpose foundation model for computational pathology. Nat. Med. 30, 850–862 (2024).
Kim, C. et al. Transparent medical image AI via an image–text foundation model grounded in medical literature. Nat. Med. 30, 1154–1165 (2024).
Li, J. et al. Integrated image-based deep learning and language models for primary diabetes care. Nat. Med. 30, 2886–2896 (2024).
Varghese, C. et al. Artificial intelligence in surgery. Nat. Med. 30, 1257–1268 (2024).
Wang, Z. et al. Foundation model for endoscopy video analysis via large-scale self-supervised pre-train. In International Conference on Medical Image Computing and Computer-Assisted Intervention 101–111 (Springer, 2023).
Hirsch, R. et al. Self-supervised learning for endoscopic video analysis. In International Conference on Medical Image Computing and Computer-Assisted Intervention 569–578 (Springer, 2023).
Bahri, Y. et al. Explaining neural scaling laws. Proc. Natl Acad. Sci. USA 121, e2311878121 (2024).
Hu, M. et al. OphNet: a large-scale video benchmark for ophthalmic surgical workflow understanding. In European Conference on Computer Vision 481–500 (Springer, 2024).
Ghamsarian, N. et al. Cataract-1k dataset for deep-learning-assisted analysis of cataract surgery videos. Sci. Data 11, 373 (2024).
Al Hajj, H. et al. Cataracts: challenge on automatic tool annotation for cataract surgery. Med. Image Anal. 52, 24–41 (2019).
Schoeffmann, K. et al. Cataract-101: video dataset of 101 cataract surgeries. In Proc. 9th ACM Multimedia Systems Conference 421–425 (ACM, 2018).
Caron, M. et al. Emerging properties in self-supervised vision transformers. In Proc. IEEE/CVF International Conference on Computer Vision 9650–9660 (IEEE, 2021).
He, K. et al. Momentum contrast for unsupervised visual representation learning. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 9729–9738 (IEEE, 2020).
He, K. et al. Masked autoencoders are scalable vision learners. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 16000–16009 (IEEE, 2022).
Ranasinghe, K. et al. Self-supervised video transformer. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 2874–2884 (IEEE, 2022).
Gou, J. et al. Knowledge distillation: a survey. Int. J. Comput. Vis. 129, 1789–1819 (2021).
Bai, Y. et al. Masked autoencoders enable efficient knowledge distillers. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 24256–24265 (IEEE, 2023).
Huang, W. et al. Generic-to-specific distillation of masked autoencoders. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 15996–16005 (IEEE, 2023).
Jiao, X. et al. TinyBERT: distilling BERT for natural language understanding. In Findings of the Association for Computational Linguistics: EMNLP 4163–4174 (ACL, 2020).
Kozak, I. & Rahn, U. Navigation technology/eye-tracking in ophthalmology: principles, applications and benefits—a narrative review. Ann. Eye Sci. 6, 6 (2021).
Kose, B. et al. Results of Callisto eye system in toric intraocular lens alignment. Beyoglu Eye J. 5, 1–4 (2020).
Carreira, J. & Zisserman, A. Quo vadis, action recognition? A new model and the kinetics dataset. In Proc. IEEE Conference on Computer Vision and Pattern Recognition 6299–6308 (IEEE, 2017).
Protserov, S. et al. Development, deployment and scaling of operating room-ready artificial intelligence for real-time surgical decision support. npj Digit. Med. 7, 231 (2024).
Wang, Z. et al. Autolaparo: a new dataset of integrated multi-tasks for image-guided surgical automation in laparoscopic hysterectomy. In International Conference on Medical Image Computing and Computer-Assisted Intervention 486–496 (Springer, 2022).
Tong, B. C. et al. Outcomes of video-assisted thoracoscopic decortication. Ann. Thorac. Surg. 89, 220–225 (2010).
Ma, L. & Fei, B. Comprehensive review of surgical microscopes: technology development and medical applications. J. Biomed. Opt. 26, 010901 (2021).
Cobianchi, L. et al. Artificial intelligence and surgery: ethical dilemmas and open issues. J. Am. Coll. Surg. 235, 268–275 (2022).
Dosovitskiy, A. et al. An image is worth 16 × 16 words: transformers for image recognition at scale. In International Conference on Learning Representations (ICLR, 2021).
Bertasius, G., Wang, H. & Torresani, L. Is space-time attention all you need for video understanding? In International Conference on Machine Learning 813-824 (PMLR, 2021).
Thomsen, A. S. S. et al. Update on simulation-based surgical training and assessment in ophthalmology: a systematic review. Ophthalmology 122, 1111–1130 (2015).
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (82330063 to X.C., 823B2045 to P.T. and 82571270 to C.Z.), the Foundation of Science and Technology Commission of Shanghai Municipality (24490710300 to X.C.), the Explorers Program of Shanghai (Basic Research Funding, number 24TS1413000 to X.C.), the Shanghai Leading Talent Program of Eastern Talent Plan (BJKJ2024003 to X.C.), the Hospital Funded Clinical Research, Xinhua Hospital Affiliated to Shanghai Jiao Tong University School of Medicine (21XJMR02 to C.Z.), and the China Postdoctoral Science Foundation (2025M772924 to P.T.).
Author information
Authors and Affiliations
Contributions
P.T., C.Z., P.Z., M.Z. and X.C. conceptualized the project. P.T., C.Z. and X.C. acquired the funding. P.T., Y. Wei, C.W. and X.C. developed the algorithms and the system. P.T., X.X., J.L., M.X., J.G., Y. Wang, C.W. and X.C. designed and performed the experiments. P.T., C.Z., Y. Wei, H.C. and X.C. analysed the data. S.Y., K.Q., J.C., W.M., X.Z., D.J., K.P., W.Z. and F.T. constructed the dataset. P.T., C.Z. and X.C. wrote and edited the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Geographic distribution of datasets used for pretraining.
Seven in-house datasets were collected from different medical centers across China, while one publicly available dataset was sourced from Austria.
Extended Data Fig. 2 Characteristics of the pretraining dataset.
a, Distribution of video durations in the dataset. The left histogram shows the number of videos for each duration interval (in minutes). A magnified view of the long-tail distribution is provided for video durations exceeding 200 minutes. b, Age distribution of patients associated with the surgical videos. The histogram shows the number of videos across different patient age groups, excluding cases where age was not recorded. c, Sex distribution of patients featured in the dataset.
Extended Data Fig. 3 Frequency of videos per calendar year.
Panels a–g present the Xinhua-pretrain, ST-pretrain, Aier-SH-pretrain, Aier-HZ-pretrain, Aier-JF-pretrain, GZ-pretrain, and Kandze-pretrain datasets, respectively.
Extended Data Fig. 4 Schematics of the seven downstream tasks used to evaluate OVFM.
Spatiotemporal-level tasks include surgical step recognition, tool presence recognition, complication detection, and surgical skill assessment, where OVFM is connected to a linear layer for classification. Spatial-level tasks include surgical scene segmentation, limbus boundary segmentation, and nucleus localization, where OVFM is connected to a decoder.
Extended Data Fig. 5 Extended performance evaluation of OVFM.
ROC curves for surgical step recognition on the Xinhua-Cata dataset (n = 23 videos) (a) and Aier-Cata dataset (n = 26 videos) (b), comparing OVFM with four models across four steps: incision, capsulorhexis, lens implant, and others. Solid lines show empirical ROC curves from the full test set; shaded regions denote 95% bootstrap CI.
Extended Data Fig. 6 Effects of surgical data diversity and fine-tuning strategies on OVFM performance.
a, Comparison of fine-tuning strategies for OVFM across multiple tasks. Two strategies—freezing the backbone vs. fine-tuning the backbone—are evaluated on tasks: surgical step recognition (n = 28 videos), complication detection (n = 49 videos), surgical skill assessment (n = 82 videos), surgical scene segmentation (n = 10 videos), limbus segmentation (n = 28 videos), and nucleus block localization (n = 20 videos). b, Fine-tuning strategy comparison for tool existence recognition. We evaluate the same two fine-tuning strategies—freezing vs. fine-tuning the backbone—for tool presence detection (n = 25 videos), with detailed performance reported per tool type. c-f, Impact of surgical type diversity on OVFM performance in surgical step recognition (n = 28 videos). The performance of OVFM pretraining using only anterior segment video data, only posterior segment video data, and a combination of both anterior and posterior segment video data are compared. Box plots summarize the distribution of performance metrics across clustered bootstrap iterations. The center line denotes the median, the box spans the interquartile range (25th–75th percentiles), and whiskers extend to the minimum and maximum values within 1.5 × the interquartile range. Statistical comparisons were performed using a two-sided paired clustered bootstrap hypothesis test.
Extended Data Fig. 7 Performance analysis of the distilled OVFM model on retrospective clinical video cases.
This figure presents the surgical step recognition and limbus boundary segmentation performance of the distilled OVFM model on two clinical cases (a and b). For each case, the left panel displays the confusion matrix for surgical step recognition, which illustrates the relationship between the model’s predictions and the annotated labels for each surgical step. Each cell reports both the actual count (outside the parentheses) and the proportion relative to the total number of samples for that class (inside the parentheses, ranging from 0 to 1). The right panel presents qualitative results for surgical step recognition and limbus boundary segmentation. The color bar illustrates the alignment between predicted surgical steps and ground truth annotations. Limbus boundary segmentation overlays predicted boundaries (red) with ground truth (green) on the original video frames.
Extended Data Fig. 8 Intraoperative comparisons of incision and capsulorhexis steps with and without navigation.
a, Incision step comparison. Representative surgical scenes for two surgeons, comparing performance with and without navigation. The intraoperative incision targets and post-surgery incision lines are shown, with zoomed-in views highlighting discrepancies. b, Capsulorhexis step comparison. Surgical scenes from the capsulorhexis step, demonstrating the differences between with and without navigation. Target capsulorhexis ranges and actual outcomes are compared post-surgery.
Supplementary information
Supplementary Information (download PDF )
Supplementary Notes 1–16, Tables 1–11 and Figs. 1 and 2.
Supplementary Video 1 (download MP4 )
The diversity of ophthalmic surgical types in the pretraining dataset.
Supplementary Video 2 (download MP4 )
OVFM-guided navigation scenes in porcine eye surgeries.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tu, P., Zheng, C., Xie, X. et al. An ophthalmic video foundation model for surgical recognition and navigation with wet-lab porcine eye validation. Nat. Biomed. Eng (2026). https://doi.org/10.1038/s41551-026-01622-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41551-026-01622-w