Abstract
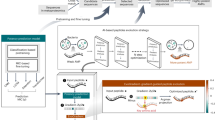
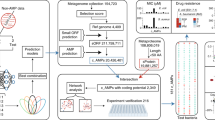
Identifying evolutionarily remote antimicrobial peptides (AMPs) is crucial for discovering underexplored clinical candidates to combat antibiotic resistance. Existing experimental and computational methods are limited by their reliance on sequence identity to known AMPs, missing distant homologues. Here we introduce HMD-AMP, a protein language model-based approach for AMP discovery. HMD-AMP outperforms previous methods in identifying evolutionarily distant AMPs and enables the discovery of unknown and highly potent AMPs from metagenomic data. Applied to host and gut microorganism genomes of nine mammals, HMD-AMP revealed over 37 million predicted AMPs. Of 91 high-confidence sequences experimentally validated, 74 showed strong antibacterial activity and 48 were evolutionarily remote from known AMPs. Four of these AMPs exhibited broad-spectrum antibacterial activity at low effective concentrations and showed low toxicity, with the most potent peptide demonstrating therapeutic efficacy in a mouse model of peritoneal Escherichia coli infection. This study introduces an effective strategy to uncover AMPs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All the datasets we used are provided in the Methods and are publicly available. The training dataset for HMD-AMP and predicted sORFs of all nine mammals is available via Zenodo at https://doi.org/10.5281/zenodo.15622525 (ref. 69). All other relevant data supporting the key findings of this study, such as the results in the AMP discovery, are available within this Article and its Supplementary Information. Source data are provided with this paper.
Code availability
The open source codes of HMD-AMP, including trained weights, inference scripts, and training scripts, are available via GitHub at https://github.com/ml4bio/HMD-AMP. Logo plots were created in WebLogo (https://weblogo.threeplusone.com/). Visualization of multiple sequence alignment of peptides used MEGA11. Structure visualizations used PyMOL (https://pymol.org/). Helical wheel plots used HeliQuest (https://heliquest.ipmc.cnrs.fr).
References
Holmes, A. H. et al. Understanding the mechanisms and drivers of antimicrobial resistance. Lancet 387, 176–187 (2016).
De Breij, A. et al. The antimicrobial peptide saap-148 combats drug-resistant bacteria and biofilms. Sci. Transl. Med. 10, eaan4044 (2018).
Mwangi, J. et al. The antimicrobial peptide zy4 combats multidrug-resistant Pseudomonas aeruginosa and Acinetobacter baumannii infection. Proc. Natl Acad. Sci. USA 116, 26516–26522 (2019).
Thapa, R. K., Diep, D. B. & Tønnesen, H. H. Topical antimicrobial peptide formulations for wound healing: current developments and future prospects. Acta Biomater. 103, 52–67 (2020).
Zhang, R. et al. Antimicrobial peptide dp7 with potential activity against sars coronavirus infections. Signal Transduct. Target. Ther. 6, 140 (2021).
Pierre, J. F. et al. Peptide yy: a paneth cell antimicrobial peptide that maintains candida gut commensalism. Science 381, 502–508 (2023).
Fjell, C. D., Hiss, J. A., Hancock, R. E. & Schneider, G. Designing antimicrobial peptides: form follows function. Nat. Rev. Drug Discov. 11, 37–51 (2012).
Porto, W. F. et al. In silico optimization of a guava antimicrobial peptide enables combinatorial exploration for peptide design. Nat. Commun. 9, 1490 (2018).
Ma, Y. et al. Identification of antimicrobial peptides from the human gut microbiome using deep learning. Nat. Biotechnol. 40, 921–931 (2022).
Huang, J. et al. Identification of potent antimicrobial peptides via a machine-learning pipeline that mines the entire space of peptide sequences. Nat. Biomed. Eng. 7, 797–810 (2023).
Maasch, J. R., Torres, M. D., Melo, M. C. & de la Fuente-Nunez, C. Molecular de-extinction of ancient antimicrobial peptides enabled by machine learning. Cell Host Microbe 31, 1260–1274 (2023).
Santos-Júnior, C. D. et al. Discovery of antimicrobial peptides in the global microbiome with machine learning. Cell 187, 3761–3778 (2024).
Torres, M. D. et al. Mining human microbiomes reveals an untapped source of peptide antibiotics. Cell 187, 5453–5467 (2024).
Wan, F., Torres, M. D., Peng, J. & de la Fuente-Nunez, C. Deep-learning-enabled antibiotic discovery through molecular de-extinction. Nat. Biomed. Eng. 8, 854–871 (2024).
Das, P. et al. Accelerated antimicrobial discovery via deep generative models and molecular dynamics simulations. Nat. Biomed. Eng 5, 613–623 (2021).
Szymczak, P. et al. Discovering highly potent antimicrobial peptides with deep generative model hydramp. Nat. Commun. 14, 1453 (2023).
Li, T. et al. A foundation model identifies broad-spectrum antimicrobial peptides against drug-resistant bacterial infection. Nat. Commun. 15, 7538 (2024).
Lazzaro, B. P., Zasloff, M. & Rolff, J. Antimicrobial peptides: application informed by evolution. Science 368, eaau5480 (2020).
Loewenstein, Y. et al. Protein function annotation by homology-based inference. Genome Biol. 10, 1–8 (2009).
Pirtskhalava, M. et al. Dbaasp v3: database of antimicrobial/cytotoxic activity and structure of peptides as a resource for development of new therapeutics. Nucleic Acids Res. 49, D288–D297 (2021).
Wang, G., Li, X. & Wang, Z. Apd3: the antimicrobial peptide database as a tool for research and education. Nucleic Acids Res. 44, D1087–D1093 (2016).
Hancock, R. E. Cationic peptides: effectors in innate immunity and novel antimicrobials. Lancet Infect. Dis. 1, 156–164 (2001).
Brogden, K. A., Ackermann, M., McCray Jr, P. B. & Tack, B. F. Antimicrobial peptides in animals and their role in host defences. Int. J. Antimicrob. Agents 22, 465–478 (2003).
Rives, A. et al. Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences. Proc. Natl Acad. Sci. USA 118, e2016239118 (2021).
Hong, L. et al. Fast, sensitive detection of protein homologs using deep dense retrieval. Nat. Biotechnol. 43, 983–995 (2025).
Lin, Z. et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science 379, 1123–1130 (2023).
Zhou, Z.-H. & Feng, J. Deep forest. Natl Sci. Rev. 6, 74–86 (2019).
Gull, S., Shamim, N. & Minhas, F. Amap: hierarchical multi-label prediction of biologically active and antimicrobial peptides. Comput. Biol. Med. 107, 172–181 (2019).
Bhadra, P., Yan, J., Li, J., Fong, S. & Siu, S. W. Ampep: sequence-based prediction of antimicrobial peptides using distribution patterns of amino acid properties and random forest. Sci. Rep. 8, 1–10 (2018).
Xiao, X., Wang, P., Lin, W.-Z., Jia, J.-H. & Chou, K.-C. iamp-2l: a two-level multi-label classifier for identifying antimicrobial peptides and their functional types. Anal. Biochem. 436, 168–177 (2013).
Veltri, D., Kamath, U. & Shehu, A. Deep learning improves antimicrobial peptide recognition. Bioinformatics 34, 2740–2747 (2018).
Consortium, U. UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res. 47, D506–D515 (2019).
Zhang, Y. & Skolnick, J. TM-align: a protein structure alignment algorithm based on the TM-score. Nucleic Acids Res. 33, 2302–2309 (2005).
Monzon, V., Haft, D. H. & Bateman, A. Folding the unfoldable: using alphafold to explore spurious proteins. Bioinform. Adv. 2, vbab043 (2022).
Chung, C.-R., Kuo, T.-R., Wu, L.-C., Lee, T.-Y. & Horng, J.-T. Characterization and identification of antimicrobial peptides with different functional activities. Brief. Bioinform. 21, 1098–1114 (2020).
Wylensek, D. et al. A collection of bacterial isolates from the pig intestine reveals functional and taxonomic diversity. Nat. Commun. 11, 6389 (2020).
Chen, C. et al. Expanded catalog of microbial genes and metagenome-assembled genomes from the pig gut microbiome. Nat. Commun. 12, 1106 (2021).
Scicchitano, D. et al. Dispersion of antimicrobial resistant bacteria in pig farms and in the surrounding environment. Anim. Microbiome 6, 17 (2024).
Holman, D. B., Brunelle, B. W., Trachsel, J. & Allen, H. K. Meta-analysis to define a core microbiota in the swine gut. mSystems 2, 10–1128 (2017).
van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
VanderWaal, K. & Deen, J. Global trends in infectious diseases of swine. Proc. Natl Acad. Sci. USA 115, 11495–11500 (2018).
Pillai, A., Ueno, S., Zhang, H., Lee, J. M. & Kato, Y. Cecropin p1 and novel nematode cecropins: a bacteria-inducible antimicrobial peptide family in the nematode ascaris suum. Biochem. J. 390, 207–214 (2005).
Wang, J. et al. The conserved domain database in 2023. Nucleic Acids Res. 51, D384–D388 (2023).
Carter, M. M. et al. Ultra-deep sequencing of hadza hunter-gatherers recovers vanishing gut microbes. Cell 186, 3111–3124 (2023).
Levin, D. et al. Diversity and functional landscapes in the microbiota of animals in the wild. Science 372, eabb5352 (2021).
Bailey, T. L. Streme: accurate and versatile sequence motif discovery. Bioinformatics 37, 2834–2840 (2021).
Mookherjee, N., Anderson, M. A., Haagsman, H. P. & Davidson, D. J. Antimicrobial host defence peptides: functions and clinical potential. Nat. Rev. Drug Discov. 19, 311–332 (2020).
Shen, J. et al. Unbiased organism-agnostic and highly sensitive signal peptide predictor with deep protein language model. Nat. Comput. Sci. 4, 29–42 (2024).
Yu, T. et al. Enzyme function prediction using contrastive learning. Science 379, 1358–1363 (2023).
Ahn, S., Kim, S., Ko, J. & Yun, S. -Y. Fine-tuning pre-trained models for robustness under noisy labels. In Proc. Thirty-Third International Joint Conference on Artificial Intelligence 3643–3651 (International Joint Conferences on Artificial Intelligence Organization, 2024).
Chen, C. H. & Lu, T. K. Development and challenges of antimicrobial peptides for therapeutic applications. Antibiotics 9, 24 (2020).
Oliveira Júnior, N. G., Souza, C. M., Buccini, D. F., Cardoso, M. H. & Franco, O. L. Antimicrobial peptides: structure, functions and translational applications. Nat. Rev. Microbiol. 23, 687–700 (2025).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152 (2012).
Burdukiewicz, M. et al. Proteomic screening for prediction and design of antimicrobial peptides with ampgram. Int. J. Mol. Sci. 21, 4310 (2020).
Sidorczuk, K. et al. Benchmarks in antimicrobial peptide prediction are biased due to the selection of negative data. Brief. Bioinform. 23, bbac343 (2022).
Su, X., Xu, J., Yin, Y., Quan, X. & Zhang, H. Antimicrobial peptide identification using multi-scale convolutional network. BMC Bioinformatics 20, 1–10 (2019).
Chaumeil, P.-A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. GTBD-Tk: a toolkit to classify genomes with the Genome Taxonomy Database. Bioinformatics 36, 1925–1927 (2019).
Cao, K., Wei, C., Gaidon, A., Arechiga, N. & Ma, T. Learning imbalanced datasets with label-distribution-aware margin loss. In Proc. 33rd Advances in Neural Information Processing Systems 1567–1578 (Curran Associates, Inc., 2019).
Chen, T. & Guestrin, C. XGBoost: a scalable tree boosting system. In Proc. 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, KDD ’16 785–794 (ACM, 2016).
Loshchilov, I. & Hutter, F. Decoupled weight decay regularization. In Proc. International Conference on Learning Representations (2019).
Rost, B. Twilight zone of protein sequence alignments. Protein Eng. 12, 85–94 (1999).
Xu, J. & Zhang, Y. How significant is a protein structure similarity with TM-score = 0.5?. Bioinformatics 26, 889–895 (2010).
Crooks, G. E., Hon, G., Chandonia, J.-M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Tamura, K., Stecher, G. & Kumar, S. MEGA11: molecular evolutionary genetics analysis version 11. Mol. Biol. Evol. 38, 3022–3027 (2021).
Mirdita, M. et al. ColabFold: making protein folding accessible to all. Nat. Methods 19, 679–682 (2022).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
The PyMOL Molecular Graphics System, Version 3.0 Schrödinger, LLC.
Deutsch, E. W. et al. The ProteomeXchange consortium at 10 years: 2023 update. Nucleic Acids Res. 51, D1539–D1548 (2023).
Yu, Q. Supporting data for: uncovering evolutionarily remote and highly potent antimicrobial peptides with protein language models. Zenodo https://doi.org/10.5281/zenodo.15622525 (2025).
Acknowledgements
This study was supported by the Shenzhen Medical Research Fund (award no. A2503002 to Y.L. and B2302036 to L.D.), the National Key R&D Program of China (grant no.2025YFA0923500 to Y.L.), the Chinese University of Hong Kong (CUHK; award nos. 4937025, 4937026, 5501517, 5501329 and SHIAE BME-p1-24 to Y.L.), the IdeaBooster Fund (award nos. IDBF23ENG05 and IDBF24ENG06 to Y.L.) and the Shenzhen Key Laboratory for the Intelligent Microbial Manufacturing of Medicines (award no. ZDSYS20210623091810032 to L.D.) and partially supported by grants from the Research Grants Council of the Hong Kong Special Administrative Region (Hong Kong SAR), China (project nos. CUHK 24204023 and 14208525 to Y.L.), and grants from the Innovation and Technology Commission of the Hong Kong SAR, China (project nos. GHP/065/21SZ, ITS/247/23FP and PRP/033/24FX to Y.L.). This research was also supported by the Research Matching Grant Scheme at CUHK (award nos. 8601603 and 8601663 to Y.L.) and the National Natural Science Foundation of China (award no. 32201313 to H.L.). We thank P. Wang and Q. Shen from the Shenzhen Institutes of Advanced Technology, Chinese Academy of Sciences, China, for their help with animal experiments. We also thank S. Qiao from the China Agricultural University, China, for providing the enterotoxigenic Escherichia coli K88 and K99 strains used in this work.
Author information
Authors and Affiliations
Contributions
Q.Y., H.L., H.S., Z.D., L.D. and Y.L. conceived the research; Q.Y., H.L. and Y.A. collected the dataset; Q.Y. implemented the main algorithm; Q.Y., H.L., H.S., J.S., C.Z. and L.S. performed the experiments; Q.Y., H.L., H.S., J.S. and L.Z. conducted the analysis; Q.Y., H.L. and H.S. wrote the paper; and L.D. and Y.L. supervised the project. All authors read and approved the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks Martin Steinegger and Dong Xu for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Notes 1–6, Figs. 1–25 and Tables 1–20.
Source data
Source Data Figs. 2–6. (download ZIP )
Source data for Figs. 2–6.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yu, Q., Liu, H., Shi, H. et al. Uncovering evolutionarily remote and highly potent antimicrobial peptides with protein language models. Nat. Biomed. Eng (2026). https://doi.org/10.1038/s41551-026-01630-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41551-026-01630-w