Abstract
Lymph nodes are the primary sites where adaptive immunity is initiated, yet most messenger RNA cancer vaccines reach them inefficiently and instead accumulate in organs such as the liver, limiting therapeutic potency and increasing systemic toxicity. Here we developed a transferrin receptor-associating polyplex formed by electrostatic complexation of mRNA with low-molecular-weight polyethylenimine that had been chemically modified with cyclic disulfide monomers to enhance nucleic acid binding stability, enable thiol-based transferrin receptor engagement and reduce off-target liver uptake. After subcutaneous administration, these polyplexes activated innate immunity, rapidly recruited monocytes with high transferrin receptor expression and bound these cells through cyclic disulfide-mediated interactions. Monocytes then trafficked the vaccine to draining lymph nodes, where mRNA translation and antigen presentation occurred. Delivery of ovalbumin and interleukin 12 mRNA elicited strong antigen-specific cytotoxic T cell responses and inhibited melanoma progression and metastatic disease. Studies using Survivin and human papillomavirus antigens in distinct tumour models demonstrated broad applicability. This monocyte-driven lymph node-targeting strategy enables potent and selective delivery of mRNA cancer vaccines.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All data supporting the findings of this study are available within the Article, Supplementary Information and Source Data. Source data are provided with this paper. Additional data are available from the corresponding authors on reasonable request.
References
Chaudhary, N., Weissman, D. & Whitehead, K. A. mRNA vaccines for infectious diseases: principles, delivery and clinical translation. Nat. Rev. Drug Discov. 20, 817–838 (2021).
Edwards, D. K. & Carfi, A. Messenger ribonucleic acid vaccines against infectious diseases: current concepts and future prospects. Curr. Opin. Immunol. 77, 102214 (2022).
Pardi, N., Hogan, M. J., Porter, F. W. & Weissman, D. mRNA vaccines—a new era in vaccinology. Nat. Rev. Drug Discov. 17, 261–279 (2018).
Zhang, G., Tang, T., Chen, Y., Huang, X. & Liang, T. mRNA vaccines in disease prevention and treatment. Signal Transduct. Target. Ther. 8, 365 (2023).
Bitounis, D., Jacquinet, E., Rogers, M. A. & Amiji, M. M. Strategies to reduce the risks of mRNA drug and vaccine toxicity. Nat. Rev. Drug Discov. 23, 281–300 (2024).
Kon, E., Ad-El, N., Hazan-Halevy, I., Stotsky-Oterin, L. & Peer, D. Targeting cancer with mRNA–lipid nanoparticles: key considerations and future prospects. Nat. Rev. Clin. Oncol. 20, 739–754 (2023).
Xia, Y., Fu, S., Ma, Q., Liu, Y. & Zhang, N. Application of nano-delivery systems in lymph nodes for tumor immunotherapy. Nanomicro Lett. 15, 145 (2023).
Aldén, M. et al. Intracellular reverse transcription of Pfizer BioNTech COVID-19 mRNA vaccine BNT162b2 in vitro in human liver cell line. Curr. Issues Mol. Biol. 44, 1115–1126 (2022).
Boettler, T. et al. SARS-CoV-2 vaccination can elicit a CD8 T-cell dominant hepatitis. J. Hepatol. 77, 653–659 (2022).
Loughrey, D. & Dahlman, J. E. Non-liver mRNA delivery. Acc. Chem. Res. 55, 13–23 (2022).
Cosentino, M. & Marino, F. The spike hypothesis in vaccine-induced adverse effects: questions and answers. Trends Mol. Med. 28, 797–799 (2022).
Trougakos, I. P. et al. Adverse effects of COVID-19 mRNA vaccines: the spike hypothesis. Trends Mol. Med. 28, 542–554 (2022).
Chen, J. et al. Lipid nanoparticle-mediated lymph node-targeting delivery of mRNA cancer vaccine elicits robust CD8+ T cell response. Proc. Natl Acad. Sci. USA 119, e2207841119 (2022).
Han, X. et al. Adjuvant lipidoid-substituted lipid nanoparticles augment the immunogenicity of SARS-CoV-2 mRNA vaccines. Nat. Nanotechnol. 18, 1105–1114 (2023).
Liu, M. et al. Lymph-targeted high-density lipoprotein-mimetic nanovaccine for multi-antigenic personalized cancer immunotherapy. Sci. Adv. 10, eadk2444 (2024).
Wang, S. et al. Macrophage-tumor chimeric exosomes accumulate in lymph node and tumor to activate the immune response and the tumor microenvironment. Sci. Transl. Med. 13, eabb6981 (2021).
Li, Y. et al. Rapid surface display of mRNA antigens by bacteria-derived outer membrane vesicles for a personalized tumor vaccine. Adv. Mater. 34, e2109984 (2022).
Zhang, D. et al. Targeted delivery of mRNA with one-component ionizable amphiphilic Janus dendrimers. J. Am. Chem. Soc. 143, 17975–17982 (2021).
Zhang, D. et al. The unexpected importance of the primary structure of the hydrophobic part of one-component ionizable amphiphilic Janus dendrimers in targeted mRNA delivery activity. J. Am. Chem. Soc. 144, 4746–4753 (2022).
Jiang, H., Wang, Q. & Sun, X. Lymph node targeting strategies to improve vaccination efficacy. J. Control. Release 267, 47–56 (2017).
Lei, J. et al. Development of mannosylated lipid nanoparticles for mRNA cancer vaccine with high antigen presentation efficiency and immunomodulatory capability. Angew. Chem. Int. Ed. 63, e202318515 (2024).
Zhou, L. et al. Tumor cell-released kynurenine biases MEP differentiation into megakaryocytes in individuals with cancer by activating AhR–RUNX1. Nat. Immunol. 24, 2042–2052 (2023).
Zhu, L. et al. Microbiota-assisted iron uptake promotes immune tolerance in the intestine. Nat. Commun. 14, 2790 (2023).
Hansen, F. J. et al. CD71 expressing circulating neutrophils serve as a novel prognostic biomarker for metastatic spread and reduced outcome in pancreatic ductal adenocarcinoma patients. Sci. Rep. 14, 21164 (2024).
Candelaria, P. V., Leoh, L. S., Penichet, M. L. & Daniels-Wells, T. R. Antibodies targeting the transferrin receptor 1 (TfR1) as direct anti-cancer agents. Front. Immunol. 12, 607692 (2021).
Daniels, T. R., Delgado, T., Rodriguez, J. A., Helguera, G. & Penichet, M. L. The transferrin receptor part I: biology and targeting with cytotoxic antibodies for the treatment of cancer. Clin. Immunol. 121, 144–158 (2006).
Zhang, D. et al. Transferrin receptor targeting chimeras for membrane protein degradation. Nature 638, 787–795 (2025).
Baeyens, A. et al. Monocyte-derived S1P in the lymph node regulates immune responses. Nature 592, 290–295 (2021).
Chen, K. Y. et al. Inflammation switches the chemoattractant requirements for naive lymphocyte entry into lymph nodes. Cell 188, 1019–1035 (2024).
Elewaut, A. et al. Cancer cells impair monocyte-mediated T cell stimulation to evade immunity. Nature 637, 716–725 (2024).
Langlet, C. et al. CD64 expression distinguishes monocyte-derived and conventional dendritic cells and reveals their distinct role during intramuscular immunization. J. Immunol. 188, 1751–1760 (2012).
León, B., López-Bravo, M. & Ardavín, C. Monocyte-derived dendritic cells formed at the infection site control the induction of protective T helper 1 responses against Leishmania. Immunity 26, 519–531 (2007).
Zhang, F. et al. Genetic programming of macrophages to perform anti-tumor functions using targeted mRNA nanocarriers. Nat. Commun. 10, 3974 (2019).
Kang, M. et al. Nanocomplex-mediated in vivo programming to chimeric antigen receptor–M1 macrophages for cancer therapy. Adv. Mater. 33, e2103258 (2021).
Tang, Y. et al. Precise delivery of nanomedicines to M2 macrophages by combining ‘eat me/don’t eat me’ signals and its anticancer application. ACS Nano 15, 18100–18112 (2021).
Graversen, J. H. et al. Targeting the hemoglobin scavenger receptor CD163 in macrophages highly increases the anti-inflammatory potency of dexamethasone. Mol. Ther. 20, 1550–1558 (2012).
Zheng, L. et al. In vivo monocyte/macrophage-hitchhiked intratumoral accumulation of nanomedicines for enhanced tumor therapy. J. Am. Chem. Soc. 142, 382–391 (2020).
Liu, C., Shang, C., Ma, L., Zhou, X. & Guo, C. Abstract 377: Evaluating delivery of TFR1-targeting therapies to the CNS in a novel humanized TFR1 mouse model. Cancer Res. 82, 377–377 (2022).
Cheng, Y., Zak, O., Aisen, P., Harrison, S. C. & Walz, T. Structure of the human transferrin receptor–transferrin complex. Cell 116, 565–576 (2004).
Lebrón, J. A. et al. Crystal structure of the hemochromatosis protein HFE and characterization of its interaction with transferrin receptor. Cell 93, 111–123 (1998).
Abraham, J., Corbett, K. D., Farzan, M., Choe, H. & Harrison, S. C. Structural basis for receptor recognition by New World hemorrhagic fever arenaviruses. Nat. Struct. Mol. Biol. 17, 438–444 (2010).
Gruszczyk, J. et al. Cryo-EM structure of an essential Plasmodium vivax invasion complex. Nature 559, 135–139 (2018).
Choi, Y. S. et al. Beyond hydrophilic polymers in amphiphilic polymer-based self-assembled NanoCarriers: small hydrophilic carboxylate-capped disulfide drug delivery system and its multifunctionality and multispatial targetability. Biomaterials 280, 121307 (2022).
Zhang, P. et al. Systemic multifunctional nanovaccines for potent personalized immunotherapy of acute myeloid leukemia. Adv. Mater. 36, e2407189 (2024).
Qu, L. et al. A biomimetic autophagosomes-based nanovaccine boosts anticancer immunity. Adv. Mater. 36, e2409590 (2024).
Kong, H. et al. An antifouling membrane-fusogenic liposome for effective intracellular delivery in vivo. Nat. Commun. 15, 4267 (2024).
Brown, D. W. et al. Safe and effective in vivo delivery of DNA and RNA using proteolipid vehicles. Cell 187, 5357–5375 (2024).
Zhao, C., Wang, C., Shan, W., Wang, W. & Deng, H. Fusogenic lipid nanovesicle for biomacromolecular delivery. Nano Lett. 24, 8609–8618 (2024).
Zhuo, Y. et al. Direct cytosolic delivery of siRNA via cell membrane fusion using cholesterol-enriched exosomes. Nat. Nanotechnol. 19, 1858–1868 (2024).
Laurent, Q. et al. Thiol-mediated uptake. JACS Au 1, 710–728 (2021).
Du, S., Liew, S. S., Li, L. & Yao, S. Q. Bypassing endocytosis: direct cytosolic delivery of proteins. J. Am. Chem. Soc. 140, 15986–15996 (2018).
Derakhshankhah, H. & Jafari, S. Cell penetrating peptides: a concise review with emphasis on biomedical applications. Biomed. Pharmacother. 108, 1090–1096 (2018).
Guo, J. et al. Rational design of poly(disulfide)s as a universal platform for delivery of CRISPR–Cas9 machineries toward therapeutic genome editing. ACS Cent. Sci. 7, 990–1000 (2021).
Shu, Z. et al. Disulfide-unit conjugation enables ultrafast cytosolic internalization of antisense DNA and siRNA. Angew. Chem. Int. Ed. 58, 6611–6615 (2019).
Martinent, R. et al. Dithiolane quartets: thiol-mediated uptake enables cytosolic delivery in deep tissue. Chem. Sci. 12, 13922–13929 (2021).
Abegg, D. et al. Strained cyclic disulfides enable cellular uptake by reacting with the transferrin receptor. J. Am. Chem. Soc. 139, 231–238 (2017).
Kanjilal, P., Dutta, K. & Thayumanavan, S. Thiol-disulfide exchange as a route for endosomal escape of polymeric nanoparticles. Angew. Chem. Int. Ed. 61, e202209227 (2022).
Cai, T., Liu, H., Zhang, S., Hu, J. & Zhang, L. Delivery of nanovaccine towards lymphoid organs: recent strategies in enhancing cancer immunotherapy. J. Nanobiotechnol. 19, 389 (2021).
Najibi, A. J. & Mooney, D. J. Cell and tissue engineering in lymph nodes for cancer immunotherapy. Adv. Drug Deliv. Rev. 161–162, 42–62 (2020).
Wang, Q. et al. Lymph node-targeting nanovaccines for cancer immunotherapy. J. Control. Release 351, 102–122 (2022).
Wauters, A. C. et al. Polymersomes with splenic avidity target red pulp myeloid cells for cancer immunotherapy. Nat. Nanotechnol. 19, 1735–1744 (2024).
Griffith, J. W., Sokol, C. L. & Luster, A. D. Chemokines and chemokine receptors: positioning cells for host defense and immunity. Annu. Rev. Immunol. 32, 659–702 (2014).
Murdoch, C. & Finn, A. Chemokine receptors and their role in inflammation and infectious diseases. Blood 95, 3032–3043 (2000).
Leal, J. M. et al. Innate cell microenvironments in lymph nodes shape the generation of T cell responses during type I inflammation. Sci. Immunol. 6, eabb9435 (2021).
Drakesmith, H. et al. The hemochromatosis protein HFE inhibits iron export from macrophages. Proc. Natl Acad. Sci. USA 99, 15602–15607 (2002).
Hu, Y. Z. et al. Supramolecular assembly of polycation/mRNA nanoparticles and in vivo monocyte programming. Proc. Natl Acad. Sci. USA 121, e2400194121 (2024).
Cheong, C. et al. Microbial stimulation fully differentiates monocytes to DC-SIGN/CD209+ dendritic cells for immune T cell areas. Cell 143, 416–429 (2010).
Menezes, S. et al. The heterogeneity of Ly6Chi monocytes controls their differentiation into iNOS+ macrophages or monocyte-derived dendritic cells. Immunity 45, 1205–1218 (2016).
Zigmond, E. et al. Ly6Chi monocytes in the inflamed colon give rise to proinflammatory effector cells and migratory antigen-presenting cells. Immunity 37, 1076–1090 (2012).
Yona, S. et al. Fate mapping reveals origins and dynamics of monocytes and tissue macrophages under homeostasis. Immunity 38, 79–91 (2013).
Elsner, R. A., Smita, S. & Shlomchik, M. J. IL-12 induces a B cell-intrinsic IL-12/IFNγ feed-forward loop promoting extrafollicular B cell responses. Nat. Immunol. 25, 1283–1295 (2024).
Li, C. et al. Mechanisms of innate and adaptive immunity to the Pfizer-BioNTech BNT162b2 vaccine. Nat. Immunol. 23, 543–555 (2022).
Garcia, E. & Ismail, S. Spatiotemporal regulation of signaling: focus on T cell activation and the immunological synapse. Int. J. Mol. Sci. 21, 3283 (2020).
Hashimoto, M., Im, S. J., Araki, K. & Ahmed, R. Cytokine-mediated regulation of CD8 T-cell responses during acute and chronic viral infection. Cold Spring Harb. Perspect. Biol. 11, a028464 (2019).
Brunner, P., Kiwitz, L., Li, L. & Thurley, K. Diffusion-limited cytokine signaling in T cell populations. iScience 27, 110134 (2024).
Sterner, R. M. et al. GM-CSF inhibition reduces cytokine release syndrome and neuroinflammation but enhances CAR-T cell function in xenografts. Blood 133, 697–709 (2019).
Zhang, C., Wu, Z., Li, J. W., Zhao, H. & Wang, G. Q. Cytokine release syndrome in severe COVID-19: interleukin-6 receptor antagonist tocilizumab may be the key to reduce mortality. Int. J. Antimicrob. Agents 55, 105954 (2020).
Dempsey, L. A. Immunoregulatory itaconate. Nat. Immunol. 19, 511 (2018).
Villar, J. & Segura, E. The more, the merrier: DC3s join the human dendritic cell family. Immunity 53, 233–235 (2020).
Terashima, A. et al. Sepsis-induced osteoblast ablation causes immunodeficiency. Immunity 44, 1434–1443 (2016).
Godbey, W. T., Wu, K. K. & Mikos, A. G. Poly(ethylenimine) and its role in gene delivery. J. Control. Release 60, 149–160 (1999).
Kunath, K. et al. Low-molecular-weight polyethylenimine as a non-viral vector for DNA delivery: comparison of physicochemical properties, transfection efficiency and in vivo distribution with high-molecular-weight polyethylenimine. J. Control. Release 89, 113–125 (2003).
Lee, J. et al. CHARMM-GUI input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field. J. Chem. Theory Comput. 12, 405–413 (2016).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2, 19–25 (2015).
Best, R. B. et al. Optimization of the additive CHARMM all-atom protein force field targeting improved sampling of the backbone φ, ψ and side-chain χ1 and χ2 dihedral angles. J. Chem. Theory Comput. 8, 3257–3273 (2012).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Bussi, G., Zykova-Timan, T. & Parrinello, M. Isothermal-isobaric molecular dynamics using stochastic velocity rescaling. J. Chem. Phys. 130, 074101 (2009).
Bernetti, M. & Bussi, G. Pressure control using stochastic cell rescaling. J. Chem. Phys. 153, 114107 (2020).
Hess, B., Bekker, H., Berendsen, H. J. C. & Fraaije, J. G. E. M. LINCS: a linear constraint solver for molecular simulations. J. Comput. Chem. 18, 1463–1472 (1997).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Li, L. et al. Burst release of encapsulated annexin A5 in tumours boosts cytotoxic T-cell responses by blocking the phagocytosis of apoptotic cells. Nat. Biomed. Eng. 4, 1102–1116 (2020).
Acknowledgements
We acknowledge financial support from National Natural Science Foundation of China (grant numbers U25A20260 and 52233007), Jiangsu Provincial Department of Science and Technology (grant numbers BG2025060 and BG2025053), Natural Science Foundation of Jiangsu Province (grant number SBK20250405084). We thank L. Wang and J. Yang for synthesizing mRNA, and J. Zhu for assistance in the preliminary cell experiments. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Q.R., X.Z. and L.Z. performed experiments, analysed data and wrote the manuscript. R.Y., L.C., K.R., X.P., Y.Z., Y.Q., K.C.C., L.C., L.D. and P.G. assisted with the preparation of reagents, data analysis and manuscript preparation. Z.Z., C.X., F.Z., C.D., B.Y. and F.M. conceptualized the project, designed and supervised the research, and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Z.Z., C.X., R.Y., Q.R. and F.M. have filed a provisional patent (China, CN202510375982.0) related to DTC-modified PEI for mRNA delivery, which is relevant to the mRNA delivery platform investigated in this study. K.R. is a co-founder of Catug Biotechnology Co., Ltd, and X.P. is an employee of Catug Biotechnology Co., Ltd, which is involved in the synthesis of mRNA used in this work. B.Y. is the founder of Suzhou Abogen Biosciences, and L.D. and P.G. are employees of Suzhou Abogen Biosciences. The other authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks Lin Mei, Xianzhu Yang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
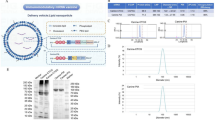
Extended Data Fig. 1 Physicochemical characterization and transfection performance of TRAP–mRNA complexes.
a, Quantification of mRNA encapsulation efficiency by TRAP–mRNA and nTRAP–mRNA formulations (n = 6 independent measurements). b, Agarose gel electrophoresis showing that mRNA is effectively protected by TRAP. c, Time course of mRNA release from TRAP–mRNA complexes in the presence of GSH (0–24 h), as measured by a RibogreenTM assay (n = 3 independent measurements). d, Representative GFP protein fluorescence imaging of various cells, scale bar: 100 µm. e, Representative flow cytometry histograms of mEGFP expression in different cell lines. f, Quantification of GFP+ cells determined by flow cytometry (n = 3 biological samples). Data in a,c,f are presented as mean values ± s.d. The experiments in b and d were independently repeated at least three times with similar results. Statistical significance for panel f was determined by multiple two-sided unpaired Student’s t-tests with correction for multiple comparisons, by two-sided unpaired Student’s t-test for panel a. Exact P values are provided in the figures where applicable. Source data are provided.
Extended Data Fig. 2 mRNA translation in lymph nodes following TRAP–mRNA administration.
a, Schematic illustration of Cre mRNA-mediated tdTomato expression in cells, where TRAP-mCre induces recombination and tdTomato expression through excision of the LoxP-flanked STOP cassette. b, Representative flow cytometry plots showing tdTomato+ DCs, Macs, and MOs in inguinal lymph nodes 48 h after subcutaneous injection of TRAP-mCre. c, Quantitative summary of Cre mRNA delivery efficiency, shown as the percentage of tdTomato+ cells among each immune-cell subset (n = 3 biological samples). d, Schematic representation of Thy1.1 (membrane-anchored) reporter expression following TRAP-mediated delivery of Thy1.1 mRNA, used to trace translation events in DCs and MOs within lymph nodes. e, Representative flow cytometry plots showing the frequency of Ly6C+CD11c+cells in inguinal lymph nodes at 3 h and 24 h after subcutaneous injection of TRAP–mLuc nanoparticles. f, Representative histogram overlays depicting Thy1.1 expression in Ly6C+CD11c− monocytes at 3 h, 12 h, and 24 h post-injection of TRAP–mThy1.1, and TRAP–mLuc treated mice served as isotype controls. Quantification of the MFI of Thy1.1+ MOs (n = 5 biological samples). Data in c and f are presented as mean values ± s.d. Statistical significance in f was determined by ordinary one-way ANOVA (two-sided). Source data are provided.
Extended Data Fig. 3 TRAP enhances local immune-cell recruitment following subcutaneous administration.
a, Representative IVIS bioluminescence images showing luciferase expression after subcutaneous administration of TRAP–mLuc or non-targeted nTRAP–mLuc. b, Flow cytometry analysis of CD45+ immune-cell infiltration at the injection site in untreated (UT), nTRAP–mLuc, and TRAP–mLuc-treated mice. c, Quantification of CD45+ immune-cell proportions across groups (n = 5 biological samples). Data in c is presented as mean values ± s.d. Statistical significance in c was determined by ordinary one-way ANOVA (two-sided). Source data are provided.
Extended Data Fig. 4 TRAP–mRNA uptake and monocyte differentiation dynamics at the injection site.
a, Subcutaneous administration of TRAP–mRNA leads to increased recruitment of CD11b+ myeloid cells at the injection site, as shown by representative flow cytometry histograms and quantification (n = 6 biological samples). b, Comparative analysis of TRAP–mRNA (Cy5-labelled) uptake between immune (CD45+) and non-immune (CD45−) cell populations at the injection site, shown by representative dot plots and fluorescence intensity histograms. c, Representative flow cytometry plot showing Cy5 signal in CD45+CD11b+ myeloid cells following subcutaneous injection of TRAP-Cy5-mRNA. d, Among Cy5+ myeloid cells, monocytes (Ly6C+), dendritic cells (CD11c+), and macrophages (F4/80+) account. e, Time-course analysis of monocyte differentiation at the injection site, as evidenced by the progressive increase in CD11c expression on Ly6C+ monocytes from 0 h to 8 h post-injection and quantification of Ly6C+CD11c+ double-positive cells over time (n = 5 biological samples). f, Temporal expression of XCR1 in Ly6C+ monocytes at the injection site, as assessed by flow cytometry at 3-, and 8-h post-injection, with representative plots and quantification (n = 5 biological samples). Data in a, e and f are presented as mean values ± s.d. Statistical significance was determined by one-way ANOVA (two-sided) for panels e and f, by two-sided unpaired Student’s t-test for panel a. Source data are provided.
Extended Data Fig. 5 Systemic depletion of macrophages by clodronate liposome administration.
a, Schematic illustration of the in vivo macrophage depletion protocol. C57BL/6 mice were intraperitoneally injected with clodronate liposomes (CL-Lipo; 200 µl on day 0 and 100 µl on day 4), and tissues were harvested on day 5 for analysis. b–e, Representative flow cytometry plots and quantification of macrophages (F4/80+) and monocytes (Ly6C+) in (b) peripheral blood, (c) peritoneal lavage fluid, (d) spleens and (e) lymph nodes from untreated (UT) and CL-Lipo–treated mice (n = 5 mice per group). Data are presented as mean values ± s.d. Statistical significance was determined by a two-sided unpaired Student’s t-test. Exact P values are provided in the figure where applicable. Source data are provided.
Extended Data Fig. 6 Tracking in situ recruitment and TRAP-Thy1.1 mRNA expression in myeloid cells at the injection site.
a, Flow cytometry analysis of CD11b+ cells at 3 h, 12 h, and 24 h following subcutaneous injection of TRAP–mThy1.1. Representative gating on CD11b+ cells is shown, with quantification of DCs, Macs, and MOs among the recruited CD11b+ population at the injection site over time (n = 6 biological samples). b, Representative flow cytometry plots showing Thy1.1+ cells at 3 h, 12 h, and 24 h post-injection. Quantification of DCs, Macs, and MOs within the Thy1.1+ population reveals the distribution of translation-active myeloid subsets at the injection site (n = 6 biological samples). Thy1.1 encodes a membrane-anchored reporter protein, enabling detection of mRNA translation following TRAP-mediated delivery. Data in a and b are presented as mean values ± s.d. Statistical significance was determined by one-way ANOVA (two-sided). Exact P values are provided in the figure where applicable. Source data are provided.
Extended Data Fig. 7 Comprehensive safety evaluation of TRAP–mRNA vaccination.
a, Complete blood count (CBC) parameters were analysed 24 h post-injection in mice receiving subcutaneous administration of TRAP–mRNA nanoparticles (3 µg mRNA, N/P = 10) or PBS (n = 6 mice per group), including WBC, NEU, LYM, RBC, HGB, HCT, PLT, MPV, NRBC, and ALY. No significant differences were observed between groups. b, Serum biochemical indicators of liver function (ALT and AST) measured 24 h post-injection (n = 4 mice per group) showed no evidence of hepatotoxicity in TRAP–mRNA-treated mice compared with PBS controls. c, Representative hematoxylin and eosin (H&E)-stained sections of major organs (heart, liver, spleen, lung, and kidney) collected 24 h post-injection from mice treated with TRAP–mLuc (control) or TRAP–mHPV revealed no apparent histopathological abnormalities, indicating minimal off-target tissue toxicity. Data are presented as mean ± s.d. Statistical significance was assessed using a two-tailed unpaired Student’s t-test. Exact P values are provided in the figure where applicable. Source data are provided.
Supplementary information
Supplementary Information (download PDF )
Supplementary Table. 1, Supplementary Figs. 1–38 and Uncropped blots and gels of Supplementary Figs. 6a, 15 and 26c.
Supplementary Video 1 (download GIF )
Cellular uptake of nanoparticles.
Supplementary Video 2 (download GIF )
Migration of nanoparticles within lymph nodes in the vasculature.
Supplementary Data 1 (download XLSX )
Coding sequences of mRNA used in this study.
Supplementary Data 2 (download XLSX )
Uncropped gels of Supplementary Figs. 15 and 26c.
Source data
Source Data Figs. 1–7 (download XLSX )
Source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ren, Q., Zhao, X., Zhou, L. et al. Polymer–mRNA complexes for monocyte-trafficked, lymph node-targeted cancer vaccination. Nat. Biomed. Eng (2026). https://doi.org/10.1038/s41551-026-01672-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41551-026-01672-0