Abstract
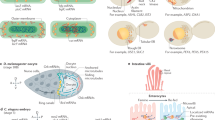
The importance of subcellular mRNA localization is well established, but the underlying mechanisms mostly remain an enigma. Early studies suggested that specific mRNA sequences recruit RNA-binding proteins (RBPs) to regulate mRNA localization. However, despite the observation of thousands of localized mRNAs, only a handful of these sequences and RBPs have been identified. This suggests the existence of alternative, and possibly predominant, mechanisms for mRNA localization. Here I re-examine currently described mRNA localization mechanisms and explore alternative models that could account for its widespread occurrence.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Jeffery, W. R., Tomlinson, C. R. & Brodeur, R. D. Localization of actin messenger RNA during early ascidian development. Dev. Biol. 99, 408–417 (1983).
Briese, M. et al. Whole transcriptome profiling reveals the RNA content of motor axons. Nucleic Acids Res. 44, e33 (2016).
Cajigas, I. J. et al. The local transcriptome in the synaptic neuropil revealed by deep sequencing and high-resolution imaging. Neuron 74, 453–466 (2012).
Ciolli Mattioli, C. et al. Alternative 3′ UTRs direct localization of functionally diverse protein isoforms in neuronal compartments. Nucleic Acids Res. 47, 2560–2573 (2019).
Farris, S. et al. Hippocampal subregions express distinct dendritic transcriptomes that reveal differences in mitochondrial function in CA2. Cell Rep. 29, 522–539 e526 (2019).
Maciel, R. et al. The human motor neuron axonal transcriptome is enriched for transcripts related to mitochondrial function and microtubule-based axonal transport. Exp. Neurol. 307, 155–163 (2018).
Middleton, S. A., Eberwine, J. & Kim, J. Comprehensive catalog of dendritically localized mRNA isoforms from sub-cellular sequencing of single mouse neurons. BMC Biol. 17, 5 (2019).
Minis, A. et al. Subcellular transcriptomics-dissection of the mRNA composition in the axonal compartment of sensory neurons. Dev. Neurobiol. 74, 365–381 (2014).
Rotem, N. et al. ALS along the axons—expression of coding and noncoding RNA differs in axons of ALS models. Sci. Rep. 7, 44500 (2017).
Taliaferro, J. M. et al. Distal alternative last exons localize mRNAs to neural projections. Mol. Cell 61, 821–833 (2016).
Tushev, G. et al. Alternative 3′ UTRs modify the localization, regulatory potential, stability and plasticity of mRNAs in neuronal compartments. Neuron 98, 495–511 (2018).
Zappulo, A. et al. RNA localization is a key determinant of neurite-enriched proteome. Nat. Commun. 8, 583 (2017).
Mili, S., Moissoglu, K. & Macara, I. G. Genome-wide screen reveals APC-associated RNAs enriched in cell protrusions. Nature 453, 115–119 (2008).
Poon, M. M., Choi, S. H., Jamieson, C. A., Geschwind, D. H. & Martin, K. C. Identification of process-localized mRNAs from cultured rodent hippocampal neurons. J. Neurosci. 26, 13390–13399 (2006).
Zhong, J., Zhang, T. & Bloch, L. M. Dendritic mRNAs encode diversified functionalities in hippocampal pyramidal neurons. BMC Neurosci. 7, 17 (2006).
Lecuyer, E. et al. Global analysis of mRNA localization reveals a prominent role in organizing cellular architecture and function. Cell 131, 174–187 (2007).
Gumy, L. F. et al. Transcriptome analysis of embryonic and adult sensory axons reveals changes in mRNA repertoire localization. RNA 17, 85–98 (2011).
Zivraj, K. H. et al. Subcellular profiling reveals distinct and developmentally regulated repertoire of growth cone mRNAs. J. Neurosci. 30, 15464–15478 (2010).
Shigeoka, T. et al. Dynamic axonal translation in developing and mature visual circuits. Cell 166, 181–192 (2016).
Jambor, H. et al. Systematic imaging reveals features and changing localization of mRNAs in Drosophila development. eLife 4, e05003 (2015).
von Kugelgen, N. & Chekulaeva, M. Conservation of a core neurite transcriptome across neuronal types and species. Wiley Interdiscip. Rev. RNA 11, e1590 (2020).
Fazal, F. M. et al. Atlas of subcellular RNA localization revealed by APEX-Seq. Cell 178, 473–490 (2019).
Das, S., Vera, M., Gandin, V., Singer, R. H. & Tutucci, E. Intracellular mRNA transport and localized translation. Nat. Rev. Mol. Cell Biol. 22, 483–504 (2021).
Engel, K. L., Arora, A., Goering, R., Lo, H. G. & Taliaferro, J. M. Mechanisms and consequences of subcellular RNA localization across diverse cell types. Traffic 21, 404–418 (2020).
Chouaib, R. et al. A dual protein-mRNA localization screen reveals compartmentalized translation and widespread co-translational RNA targeting. Dev. Cell 54, 773–791 (2020).
Lasko, P. Patterning the Drosophila embryo: a paradigm for RNA-based developmental genetic regulation. Wiley Interdiscip. Rev. RNA 11, e1610 (2020).
Micheva, K. D., Vallee, A., Beaulieu, C., Herman, I. M. & Leclerc, N. β-actin is confined to structures having high capacity of remodelling in developing and adult rat cerebellum. Eur. J. Neurosci. 10, 3785–3798 (1998).
Bassell, G. J. et al. Sorting of β-actin mRNA and protein to neurites and growth cones in culture. J. Neurosci. 18, 251–265 (1998).
Chowdhury, S. et al. Arc/Arg3.1 interacts with the endocytic machinery to regulate AMPA receptor trafficking. Neuron 52, 445–459 (2006).
Peebles, C. L. et al. Arc regulates spine morphology and maintains network stability in vivo. Proc. Natl Acad. Sci. USA 107, 18173–18178 (2010).
Kislauskis, E. H. & Singer, R. H. Determinants of mRNA localization. Curr. Opin. Cell Biol. 4, 975–978 (1992).
Kislauskis, E. H., Zhu, X. & Singer, R. H. Sequences responsible for intracellular localization of β-actin messenger RNA also affect cell phenotype. J. Cell Biol. 127, 441–451 (1994).
Ross, A. F., Oleynikov, Y., Kislauskis, E. H., Taneja, K. L. & Singer, R. H. Characterization of a β-actin mRNA zipcode-binding protein. Mol. Cell. Biol. 17, 2158–2165 (1997).
Oleynikov, Y. & Singer, R. H. Real-time visualization of ZBP1 association with β-actin mRNA during transcription and localization. Curr. Biol. 13, 199–207 (2003).
Tiruchinapalli, D. M. et al. Activity-dependent trafficking and dynamic localization of zipcode binding protein 1 and β-actin mRNA in dendrites and spines of hippocampal neurons. J. Neurosci. 23, 3251–3261 (2003).
Patel, V. L. et al. Spatial arrangement of an RNA zipcode identifies mRNAs under post-transcriptional control. Genes Dev. 26, 43–53 (2012).
Niessing, D., Jansen, R. P., Pohlmann, T. & Feldbrugge, M. mRNA transport in fungal top models. Wiley Interdiscip. Rev. RNA 9, e1453 (2018).
McClintock, M. A. RNA-directed activation of cytoplasmic dynein-1 in reconstituted transport RNPs. eLife 7, e36312 (2018).
Jambor, H., Mueller, S., Bullock, S. L. & Ephrussi, A. A stem–loop structure directs oskar mRNA to microtubule minus ends. RNA 20, 429–439 (2014).
Van De Bor, V., Hartswood, E., Jones, C., Finnegan, D. & Davis, I. gurken and the I factor retrotransposon RNAs share common localization signals and machinery. Dev. Cell 9, 51–62 (2005).
Dienstbier, M., Boehl, F., Li, X. & Bullock, S. L. Egalitarian is a selective RNA-binding protein linking mRNA localization signals to the dynein motor. Genes Dev. 23, 1546–1558 (2009).
Muntjes, K., Devan, S. K., Reichert, A. S. & Feldbrugge, M. Linking transport and translation of mRNAs with endosomes and mitochondria. EMBO Rep. 22, e52445 (2021).
Otis, J. P. & Mowry, K. L. Hitting the mark: localization of mRNA and biomolecular condensates in health and disease. Wiley Interdiscip. Rev. RNA 14, e1807 (2023).
Forrest, K. M. & Gavis, E. R. Live imaging of endogenous RNA reveals a diffusion and entrapment mechanism for nanos mRNA localization in Drosophila. Curr. Biol. 13, 1159–1168 (2003).
Gutzeit, H. The role of microtubules in the differentiation of ovarian follicles during vitellogenesis in Drosophila. Rouxs Arch. Dev. Biol. 195, 173–181 (1986).
Chang, P. et al. Localization of RNAs to the mitochondrial cloud in Xenopus oocytes through entrapment and association with endoplasmic reticulum. Mol. Biol. Cell 15, 4669–4681 (2004).
Aviram, N. & Schuldiner, M. Targeting and translocation of proteins to the endoplasmic reticulum at a glance. J. Cell Sci. 130, 4079–4085 (2017).
Safieddine, A. et al. A choreography of centrosomal mRNAs reveals a conserved localization mechanism involving active polysome transport. Nat. Commun. 12, 1352 (2021).
Anadolu, M. N. et al. Ribosomes in RNA granules are stalled on mRNA sequences that are consensus sites for FMRP association. J. Neurosci. 43, 2440–2459 (2023).
Ding, D., Parkhurst, S. M., Halsell, S. R. & Lipshitz, H. D. Dynamic Hsp83 RNA localization during Drosophila oogenesis and embryogenesis. Mol. Cell. Biol. 13, 3773–3781 (1993).
Semotok, J. L. et al. Smaug recruits the CCR4/POP2/NOT deadenylase complex to trigger maternal transcript localization in the early Drosophila embryo. Curr. Biol. 15, 284–294 (2005).
Zaessinger, S., Busseau, I. & Simonelig, M. Oskar allows nanos mRNA translation in Drosophila embryos by preventing its deadenylation by Smaug/CCR4. Development 133, 4573–4583 (2006).
Mendonsa, S. et al. Massively parallel identification of mRNA localization elements in primary cortical neurons. Nat. Neurosci. 26, 394–405 (2023).
Schwamborn, J. C., Berezikov, E. & Knoblich, J. A. The TRIM-NHL protein TRIM32 activates microRNAs and prevents self-renewal in mouse neural progenitors. Cell 136, 913–925 (2009).
Zou, Y. et al. Developmental decline in neuronal regeneration by the progressive change of two intrinsic timers. Science 340, 372–376 (2013).
Li, S. et al. Let-7 microRNAs regenerate peripheral nerve regeneration by targeting nerve growth factor. Mol. Ther. 23, 423–433 (2015).
Caygill, E. E. & Johnston, L. A. Temporal regulation of metamorphic processes in Drosophila by the let-7 and miR-125 heterochronic microRNAs. Curr. Biol. 18, 943–950 (2008).
Edbauer, D. et al. Regulation of synaptic structure and function by FMRP-associated microRNAs miR-125b and miR-132. Neuron 65, 373–384 (2010).
Feltrin, D. & Pertz, O. Assessment of Rho GTPase signaling during neurite outgrowth. Methods Mol. Biol. 827, 181–194 (2012).
Lein, E. S. et al. Genome-wide atlas of gene expression in the adult mouse brain. Nature 445, 168–176 (2007).
Poulopoulos, A. et al. Subcellular transcriptomes and proteomes of developing axon projections in the cerebral cortex. Nature 565, 356–360 (2019).
Saal, L., Briese, M., Kneitz, S., Glinka, M. & Sendtner, M. Subcellular transcriptome alterations in a cell culture model of spinal muscular atrophy point to widespread defects in axonal growth and presynaptic differentiation. RNA 20, 1789–1802 (2014).
Taylor, A. M. et al. Axonal mRNA in uninjured and regenerating cortical mammalian axons. J. Neurosci. 29, 4697–4707 (2009).
Toth, E. N. et al. Single-cell nanobiopsy reveals compartmentalization of mRNAs within neuronal cells. J. Biol. Chem. 293, 4940–4951 (2018).
Arora, A. et al. High-throughput identification of RNA localization elements in neuronal cells. Nucleic Acids Res. 50, 10626–10642 (2022).
Mikl, M. et al. A massively parallel reporter assay reveals focused and broadly encoded RNA localization signals in neurons. Nucleic Acids Res. 50, 10643–10664 (2022).
Lie, Y. S. & Macdonald, P. M. Apontic binds the translational repressor Bruno and is implicated in regulation of oskar mRNA translation. Development 126, 1129–1138 (1999).
Filardo, P. & Ephrussi, A. Bruno regulates gurken during Drosophila oogenesis. Mech. Dev. 120, 289–297 (2003).
Wang, Y., Wang, J., Gao, L., Stamm, S. & Andreadis, A. An SRp75/hnRNPG complex interacting with hnRNPE2 regulates the 5′ splice site of tau exon 10, whose misregulation causes frontotemporal dementia. Gene 485, 130–138 (2011).
Moissoglu, K. et al. RNA localization and co-translational interactions control RAB13 GTPase function and cell migration. EMBO J. 39, e104958 (2020).
Kobayashi, H., Yamamoto, S., Maruo, T. & Murakami, F. Identification of a cis-acting element required for dendritic targeting of activity-regulated cytoskeleton-associated protein mRNA. Eur. J. Neurosci. 22, 2977–2984 (2005).
Hachet, O. & Ephrussi, A. Splicing of oskar RNA in the nucleus is coupled to its cytoplasmic localization. Nature 428, 959–963 (2004).
Davis, L., Burger, B., Banker, G. A. & Steward, O. Dendritic transport: quantitative analysis of the time course of somatodendritic transport of recently synthesized RNA. J. Neurosci. 10, 3056–3068 (1990).
Fonkeu, Y. et al. How mRNA localization and protein synthesis sites influence dendritic protein distribution and dynamics. Neuron 103, 1109–1122 (2019).
Loedige, I. et al. mRNA stability and m6A are major determinants of subcellular mRNA localization in neurons. Mol. Cell 83, 2709–2725 (2023).
Gentilella, A. et al. Autogenous control of 5′TOP mRNA stability by 40S ribosomes. Mol. Cell 67, 55–70 (2017).
Shigeoka, T. et al. On-site ribosome remodeling by locally synthesized ribosomal proteins in axons. Cell Rep. 29, 3605–3619 (2019).
Fusco, C. M. et al. Neuronal ribosomes exhibit dynamic and context-dependent exchange of ribosomal proteins. Nat. Commun. 12, 6127 (2021).
Dermit, M. et al. Subcellular mRNA localization regulates ribosome biogenesis in migrating cells. Dev. Cell 55, 298–313 (2020).
Goering, R. et al. FMRP promotes RNA localization to neuronal projections through interactions between its RGG domain and G-quadruplex RNA sequences. eLife 9, e52621 (2020).
Du, H. et al. YTHDF2 destabilizes m6A-containing RNA through direct recruitment of the CCR4-NOT deadenylase complex. Nat. Commun. 7, 12626 (2016).
Zaccara, S. & Jaffrey, S. R. A unified model for the function of YTHDF proteins in regulating m6A-modified mRNA. Cell 181, 1582–1595 (2020).
Huang, H. et al. Recognition of RNA N6-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat. Cell Biol. 20, 285–295 (2018).
Heraud-Farlow, J. E. & Kiebler, M. A. The multifunctional Staufen proteins: conserved roles from neurogenesis to synaptic plasticity. Trends Neurosci. 37, 470–479 (2014).
Heraud-Farlow, J. E. et al. Staufen2 regulates neuronal target RNAs. Cell Rep. 5, 1511–1518 (2013).
Harris, K. M. & Spacek, J. in Dendrites (eds Greg Stuart, Nelson Spruston, & Michael Häusser) Ch. 1 (Oxford Univ. Press, 2016).
Park, H. Y. et al. Visualization of dynamics of single endogenous mRNA labeled in live mouse. Science 343, 422–424 (2014).
Jankowsky, E. & Harris, M. E. Specificity and nonspecificity in RNA–protein interactions. Nat. Rev. Mol. Cell Biol. 16, 533–544 (2015).
Corley, M., Burns, M. C. & Yeo, G. W. How RNA-binding proteins interact with RNA: molecules and mechanisms. Mol. Cell 78, 9–29 (2020).
Paz, I., Argoetti, A., Cohen, N., Even, N. & Mandel-Gutfreund, Y. RBPmap: a tool for mapping and predicting the binding sites of RNA-binding proteins considering the motif environment. Methods Mol. Biol. 2404, 53–65 (2022).
Dominguez, D. et al. Sequence, structure and context preferences of human RNA binding proteins. Mol. Cell 70, 854–867 (2018).
Brown, V. et al. Microarray identification of FMRP-associated brain mRNAs and altered mRNA translational profiles in fragile X syndrome. Cell 107, 477–487 (2001).
Rage, F. et al. Genome-wide identification of mRNAs associated with the protein SMN whose depletion decreases their axonal localization. RNA 19, 1755–1766 (2013).
Preitner, N. et al. APC is an RNA-binding protein, and its interactome provides a link to neural development and microtubule assembly. Cell 158, 368–382 (2014).
Batish, M., van den Bogaard, P., Kramer, F. R. & Tyagi, S. Neuronal mRNAs travel singly into dendrites. Proc. Natl Acad. Sci. USA 109, 4645–4650 (2012).
Turner-Bridger, B. et al. Single-molecule analysis of endogenous β-actin mRNA trafficking reveals a mechanism for compartmentalized mRNA localization in axons. Proc. Natl Acad. Sci. USA 115, E9697–E9706 (2018).
Mikl, M., Vendra, G. & Kiebler, M. A. Independent localization of MAP2, CaMKIIα and β-actin RNAs in low copy numbers. EMBO Rep. 12, 1077–1084 (2011).
Gao, Y., Tatavarty, V., Korza, G., Levin, M. K. & Carson, J. H. Multiplexed dendritic targeting of α calcium calmodulin-dependent protein kinase II, neurogranin and activity-regulated cytoskeleton-associated protein RNAs by the A2 pathway. Mol. Biol. Cell 19, 2311–2327 (2008).
Tauber, D., Tauber, G. & Parker, R. Mechanisms and regulation of RNA condensation in RNP granule formation. Trends Biochem. Sci. 45, 764–778 (2020).
Kato, M. & McKnight, S. L. A solid-state conceptualization of information transfer from gene to message to protein. Annu. Rev. Biochem. 87, 351–390 (2018).
Neil, C. R. et al. L-bodies are RNA-protein condensates driving RNA localization in Xenopus oocytes. Mol. Biol. Cell 32, ar37 (2021).
Bose, M., Lampe, M., Mahamid, J. & Ephrussi, A. Liquid-to-solid phase transition of oskar ribonucleoprotein granules is essential for their function in Drosophila embryonic development. Cell 185, 1308–1324 (2022).
Eichler, C. E., Li, H., Grunberg, M. E. & Gavis, E. R. Localization of oskar mRNA by agglomeration in ribonucleoprotein granules. PLoS Genet. 19, e1010877 (2023).
Knowles, R. B. et al. Translocation of RNA granules in living neurons. J. Neurosci. 16, 7812–7820 (1996).
Kohrmann, M. et al. Microtubule-dependent recruitment of Staufen-green fluorescent protein into large RNA-containing granules and subsequent dendritic transport in living hippocampal neurons. Mol. Biol. Cell 10, 2945–2953 (1999).
Bauer, K. E., de Queiroz, B. R., Kiebler, M. A. & Besse, F. RNA granules in neuronal plasticity and disease. Trends Neurosci. 46, 525–538 (2023).
Tsang, B. et al. Phosphoregulated FMRP phase separation models activity-dependent translation through bidirectional control of mRNA granule formation. Proc. Natl Acad. Sci. USA 116, 4218–4227 (2019).
Gopal, P. P., Nirschl, J. J., Klinman, E. & Holzbaur, E. L. Amyotrophic lateral sclerosis-linked mutations increase the viscosity of liquid-like TDP-43 RNP granules in neurons. Proc. Natl Acad. Sci. USA 114, E2466–E2475 (2017).
Murakami, T. et al. ALS/FTD mutation-induced phase transition of FUS liquid droplets and reversible hydrogels into irreversible hydrogels impairs RNP granule function. Neuron 88, 678–690 (2015).
Baumann, S., Pohlmann, T., Jungbluth, M., Brachmann, A. & Feldbrugge, M. Kinesin-3 and dynein mediate microtubule-dependent co-transport of mRNPs and endosomes. J. Cell Sci. 125, 2740–2752 (2012).
Schmid, M., Jaedicke, A., Du, T. G. & Jansen, R. P. Coordination of endoplasmic reticulum and mRNA localization to the yeast bud. Curr. Biol. 16, 1538–1543 (2006).
Tian, L. et al. Zipcode RNA-binding proteins and membrane trafficking proteins cooperate to transport Glutelin mRNAs in rice endosperm. Plant Cell 32, 2566–2581 (2020).
Liao, Y. C. et al. RNA granules hitchhike on lysosomes for long-distance transport, using annexin A11 as a molecular tether. Cell 179, 147–164 (2019).
Harbauer, A. B. et al. Neuronal mitochondria transport Pink1 mRNA via synaptojanin 2 to support local mitophagy. Neuron 110, 1516–1531 (2022).
Schuhmacher, J. S. et al. The Rab5 effector FERRY links early endosomes with mRNA localization. Mol. Cell 83, 1839–1855 (2023).
Cioni, J. M. et al. Late endosomes act as mRNA translation platforms and sustain mitochondria in axons. Cell 176, 56–72 (2019).
Baumann, S. et al. A reconstituted mammalian APC-kinesin complex selectively transports defined packages of axonal mRNAs. Sci. Adv. 6, eaaz1588 (2020).
Nunez, L. et al. Tagged actin mRNA dysregulation in IGF2BP1 mice. Proc. Natl Acad. Sci. USA 119, e2208465119 (2022).
Heym, R. G. et al. In vitro reconstitution of an mRNA-transport complex reveals mechanisms of assembly and motor activation. J. Cell Biol. 203, 971–984 (2013).
Zimyanin, V. L. et al. In vivo imaging of oskar mRNA transport reveals the mechanism of posterior localization. Cell 134, 843–853 (2008).
Ghosh, S., Marchand, V., Gaspar, I. & Ephrussi, A. Control of RNP motility and localization by a splicing-dependent structure in oskar mRNA. Nat. Struct. Mol. Biol. 19, 441–449 (2012).
Gaspar, I., Sysoev, V., Komissarov, A. & Ephrussi, A. An RNA-binding atypical tropomyosin recruits kinesin-1 dynamically to oskar mRNPs. EMBO J. 36, 319–333 (2017).
Brendza, R. P., Serbus, L. R., Duffy, J. B. & Saxton, W. M. A function for kinesin I in the posterior transport of oskar mRNA and Staufen protein. Science 289, 2120–2122 (2000).
Munro, T. P. et al. Mutational analysis of a heterogeneous nuclear ribonucleoprotein A2 response element for RNA trafficking. J. Biol. Chem. 274, 34389–34395 (1999).
Shan, J., Munro, T. P., Barbarese, E., Carson, J. H. & Smith, R. A molecular mechanism for mRNA trafficking in neuronal dendrites. J. Neurosci. 23, 8859–8866 (2003).
Lyons, D. A., Naylor, S. G., Scholze, A. & Talbot, W. S. Kif1b is essential for mRNA localization in oligodendrocytes and development of myelinated axons. Nat. Genet. 41, 854–858 (2009).
Costa, G., Bradbury, J. J., Tarannum, N. & Herbert, S. P. RAB13 mRNA compartmentalisation spatially orients tissue morphogenesis. EMBO J. 39, e106003 (2020).
Pichon, X. et al. The kinesin KIF1C transports APC-dependent mRNAs to cell protrusions. RNA 27, 1528–1544 (2021).
Norris, M. L. & Mendell, J. T. Localization of Kif1c mRNA to cell protrusions dictates binding partner specificity of the encoded protein. Genes Dev. 37, 191–203 (2023).
Wang, T., Hamilla, S., Cam, M., Aranda-Espinoza, H. & Mili, S. Extracellular matrix stiffness and cell contractility control RNA localization to promote cell migration. Nat. Commun. 8, 896 (2017).
Trovisco, V. et al. bicoid mRNA localises to the Drosophila oocyte anterior by random dynein-mediated transport and anchoring. eLife 5, e17537 (2016).
Vanzo, N. F. & Ephrussi, A. Oskar anchoring restricts pole plasm formation to the posterior of the Drosophila oocyte. Development 129, 3705–3714 (2002).
Delanoue, R., Herpers, B., Soetaert, J., Davis, I. & Rabouille, C. Drosophila Squid/hnRNP helps Dynein switch from a gurken mRNA transport motor to an ultrastructural static anchor in sponge bodies. Dev. Cell 13, 523–538 (2007).
Semotok, J. L. et al. Drosophila maternal Hsp83 mRNA destabilization is directed by multiple SMAUG recognition elements in the open reading frame. Mol. Cell. Biol. 28, 6757–6772 (2008).
Merkurjev, D. et al. Synaptic N6-methyladenosine (m6A) epitranscriptome reveals functional partitioning of localized transcripts. Nat. Neurosci. 21, 1004–1014 (2018).
Goering, R., Arora, A., Pockalny, M. C. & Taliaferro, J. M. RNA localization mechanisms transcend cell morphology. eLife 12, e80040 (2023).
Muslimov, I. A., Iacoangeli, A., Brosius, J. & Tiedge, H. Spatial codes in dendritic BC1 RNA. J. Cell Biol. 175, 427–439 (2006).
Huang, Y. S., Carson, J. H., Barbarese, E. & Richter, J. D. Facilitation of dendritic mRNA transport by CPEB. Genes Dev. 17, 638–653 (2003).
Oe, S. & Yoneda, Y. Cytoplasmic polyadenylation element-like sequences are involved in dendritic targeting of BDNF mRNA in hippocampal neurons. FEBS Lett. 584, 3424–3430 (2010).
Groisman, I. et al. CPEB, maskin and cyclin B1 mRNA at the mitotic apparatus: implications for local translational control of cell division. Cell 103, 435–447 (2000).
Mori, Y., Imaizumi, K., Katayama, T., Yoneda, T. & Tohyama, M. Two cis-acting elements in the 3′ untranslated region of α-CaMKII regulate its dendritic targeting. Nat. Neurosci. 3, 1079–1084 (2000).
Atlas, R., Behar, L., Elliott, E. & Ginzburg, I. The insulin-like growth factor mRNA binding-protein IMP-1 and the Ras-regulatory protein G3BP associate with tau mRNA and HuD protein in differentiated P19 neuronal cells. J. Neurochem. 89, 613–626 (2004).
Racca, C. et al. The neuronal splicing factor Nova co-localizes with target RNAs in the dendrite. Front. Neural Circuits 4, 5 (2010).
Burute, M. & Kapitein, L. C. Cellular logistics: unraveling the interplay between microtubule organization and intracellular transport. Annu. Rev. Cell Dev. Biol. 35, 29–54 (2019).
Tas, R. P. et al. Differentiation between oppositely oriented microtubules controls polarized neuronal transport. Neuron 96, 1264–1271 e1265 (2017).
Doyle, M. & Kiebler, M. A. Mechanisms of dendritic mRNA transport and its role in synaptic tagging. EMBO J. 30, 3540–3552 (2011).
Song, T. et al. Specific interaction of KIF11 with ZBP1 regulates the transport of β-actin mRNA and cell motility. J. Cell Sci. 128, 1001–1010 (2015).
Lipka, J., Kapitein, L. C., Jaworski, J. & Hoogenraad, C. C. Microtubule-binding protein doublecortin-like kinase 1 (DCLK1) guides kinesin-3-mediated cargo transport to dendrites. EMBO J. 35, 302–318 (2016).
Huttelmaier, S. et al. Spatial regulation of β-actin translation by Src-dependent phosphorylation of ZBP1. Nature 438, 512–515 (2005).
Acknowledgements
M.C. is supported by a DFG Heisenberg grant. I thank I. Loedige and D. Niessing for their critical reading of the manuscript. BioRender was used in figure generation. I apologize to all the authors whose work could not be cited due to space limitations.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chekulaeva, M. Mechanistic insights into the basis of widespread RNA localization. Nat Cell Biol 26, 1037–1046 (2024). https://doi.org/10.1038/s41556-024-01444-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41556-024-01444-5
This article is cited by
-
Regulation of gene expression by alternative polyadenylation in health and disease
Nature Reviews Genetics (2026)
-
RNA-binding proteins and ribonucleoproteins as determinants of immunity
Nature Reviews Immunology (2026)
-
Subcellular localization as a driver of protein function
Nature Reviews Molecular Cell Biology (2026)
-
Common cell lysis procedures distort ribosome profiling analyses of gene expression
Genome Biology (2025)
-
Decoding subcellular RNA localization one molecule at a time
Genome Biology (2025)