Abstract
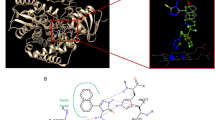
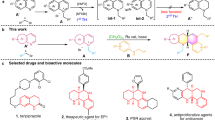
The acylhydrazone unit is well represented in screening databases used to find ligands for biological targets, and numerous bioactive acylhydrazones have been reported. However, potential E/Z isomerization of the C=N bond in these compounds is rarely examined when bioactivity is assayed. Here we analysed two ortho-hydroxylated acylhydrazones discovered in a virtual drug screen for modulators of N-methyl-d-aspartate receptors and other bioactive hydroxylated acylhydrazones with structurally defined targets reported in the Protein Data Bank. We found that ionized forms of these compounds, which are populated under laboratory conditions, photoisomerize readily and the isomeric forms have markedly different bioactivity. Furthermore, we show that glutathione, a tripeptide involved with cellular redox balance, catalyses dynamic E⇄Z isomerization of acylhydrazones. The ratio of E to Z isomers in cells is determined by the relative stabilities of the isomers regardless of which isomer was applied. We conclude that E/Z isomerization may be a common feature of the bioactivity observed with acylhydrazones and should be routinely analysed.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Source data include Excel files containing data for Figs. 2c,d,g,h, 3b,e–g, 4a–d, 5b–g and 6b,c,f–h and Pymol files for models (Figs. 1d,e, 2b,i and 3d). Computed structures included in the tables in Supplementary Information are attached as .xyz files. The X-ray crystallographic coordinates for structures reported in this study have been deposited at the Cambridge Crystallographic Data Centre (CCDC): E-43 (deposition number 2143493) and E-217 (deposition number 2143494). These data can be obtained free of charge from The Cambridge Crystallographic Data Centre via www.ccdc.cam.ac.uk/data_request/cif. Source data are provided with this paper.
References
He, L. Y. et al. Discovering potent inhibitors against the β-hydroxyacyl-acyl carrier protein dehydratase (FabZ) of Helicobacter pylori: structure-based design, synthesis, bioassay, and crystal structure determination. J. Med. Chem. 52, 2465–2481 (2009).
Ahmad, M. F. et al. Potent competitive inhibition of human ribonucleotide reductase by a nonnucleoside small molecule. Proc. Natl Acad. Sci. USA 117, 30858–30858 (2020).
Huff, S. E. et al. Structure-guided synthesis and mechanistic studies reveal sweetspots on naphthyl salicyl hydrazone scaffold as non-nucleosidic competitive, reversible inhibitors of human ribonucleotide reductase. J. Med. Chem. 61, 666–680 (2018).
Mota Fernandes, C. et al. The future of antifungal drug therapy: novel compounds and targets. Antimicrob. Agents Chemother. https://doi.org/10.1128/AAC.01719-20 (2021).
Herschhorn, A. et al. A broad HIV-1 inhibitor blocks envelope glycoprotein transitions critical for entry. Nat. Chem. Biol. 10, 845–852 (2014).
Sorna, V. et al. High-throughput virtual screening identifies novel N′-(1-phenylethylidene)-benzohydrazides as potent, specific, and reversible LSD1 inhibitors. J. Med. Chem. 56, 9496–9508 (2013).
Bhat, V. T. et al. Nucleophilic catalysis of acylhydrazone equilibration for protein-directed dynamic covalent chemistry. Nat. Chem. 2, 490–497 (2010).
Jaegle, M. et al. Protein-templated fragment ligations—from molecular recognition to drug discovery. Angew. Chem. Int. Ed. Engl. 56, 7358–7378 (2017).
Canal-Martin, A. et al. Insights into real-time chemical processes in a calcium sensor protein-directed dynamic library. Nat. Commun. 10, 2798 (2019).
Senger, M. R., Fraga, C. A., Dantas, R. F. & Silva, F. P. Jr. Filtering promiscuous compounds in early drug discovery: is it a good idea? Drug Discov. Today 21, 868–872 (2016).
Baell, J. B. & Nissink, J. W. M. Seven year itch: pan-assay interference compounds (PAINS) in 2017—utility and limitations. ACS Chem. Biol. 13, 36–44 (2018).
Kolb, M. E., Horne, M. L. & Martz, R. Dantrolene in human malignant hyperthermia. Anesthesiology 56, 254–262 (1982).
Thota, S. et al. N-acylhydrazones as drugs. Bioorg. Med. Chem. Lett. 28, 2797–2806 (2018).
Maia, R. D., Tesch, R. & Fraga, C. A. M. Acylhydrazone derivatives: a patent review. Expert Opin. Ther. Pat. 24, 1161–1170 (2014).
Padwa, A. Photochemistry of carbon–nitrogen double-bond. Chem. Rev. 77, 37–68 (1977).
Lu, C. C., Htan, B., Ma, C. M., Liao, R. Z. & Gan, Q. Acylhydrazone switches: E/Z stability reversed by introduction of hydrogen bonds. Eur. J. Org. Chem. 2018, 7046–7050 (2018).
Yuan, Y. X. & Zheng, Y. S. New acylhydrazone photoswitches with quantitative conversion and high quantum yield but without hydrogen bond stabilizing (Z)-isomer. ACS Appl. Mater. Interfaces 11, 7303–7310 (2019).
Cvrtila, I., Fanlo-Virgos, H., Schaeffer, G., Santiago, G. M. & Otto, S. Redox control over acyl hydrazone photoswitches. J. Am. Chem. Soc. 139, 12459–12465 (2017).
Ratjen, L. & Lehn, J. M. Reversible photo-, metallo- and thermo-induced morphological dynamics of bis-acylhydrazones. RSC Adv. 4, 50554–50557 (2014).
Belov, D. G., Rogachev, B. G., Tkachenko, L. I., Smirnov, V. A. & Aldoshin, S. M. Photochemistry of arylhydrazides in solution. Russ. Chem. Bull. 49, 666–668 (2000).
van Dijken, D. J., Kovaricek, P., Ihrig, S. P. & Hecht, S. Acylhydrazones as widely tunable photoswitches. J. Am. Chem. Soc. 137, 14982–14991 (2015).
Moreno, M., Gelabert, R. & Lluch, J. M. Deciphering the grounds of the suitability of acylhydrazones as efficient photoswitches. Phys. Chem. Chem. Phys. 21, 16075–16082 (2019).
Borowiak, M. et al. Photoswitchable inhibitors of microtubule dynamics optically control mitosis and cell death. Cell 162, 403–411 (2015).
Donau, S. S., Bechmann, M., Müller, N., Nielsen, T. T. & Wimmer, R. (Z), not (E)—an end to a century of confusion about the double-bond stereoisomers of 3-amino-2-cyanoacrylates. Eur. J. Org. Chem. 2017, 6408–6412 (2017).
Katzenellenbogen, B. S., Norman, M. J., Eckert, R. L., Peltz, S. W. & Mangel, W. F. Bioactivities, estrogen receptor interactions, and plasminogen activator-inducing activities of tamoxifen and hydroxy-tamoxifen isomers in MCF-7 human breast cancer cells. Cancer Res. 44, 112–119 (1984).
Jumde, V. R. et al. Design and synthesis of bioisosteres of acylhydrazones as stable inhibitors of the aspartic protease endothiapepsin. ChemMedChem 13, 2266–2270 (2018).
Toro, P. M. et al. Spectroscopic study of the E/Z photoisomerization of a new cyrhetrenyl acylhydrazone: a potential photoswitch and photosensitizer. Photochem. Photobiol. 97, 61–70 (2021).
Yin, T. T., Zhao, Z. X., Yu, L. Y. & Zhang, H. X. Theoretical study on the thermal cis–trans isomerization of novel acylhydrazone photoswitches. Org. Electron. 48, 154–164 (2017).
Liu, Z. et al. A reversible fluorescent probe for real-time quantitative monitoring of cellular glutathione. Angew. Chem. Int. Ed. Engl. 56, 5812–5816 (2017).
Broichhagen, J., Frank, J. A. & Trauner, D. A roadmap to success in photopharmacology. Acc. Chem. Res. 48, 1947–1960 (2015).
Szymanski, W., Beierle, J. M., Kistemaker, H. A., Velema, W. A. & Feringa, B. L. Reversible photocontrol of biological systems by the incorporation of molecular photoswitches. Chem. Rev. 113, 6114–6178 (2013).
Perszyk, R. E. et al. Biased modulators of NMDA receptors control channel opening and ion selectivity. Nat. Chem. Biol. 16, 188–196 (2020).
Ahmed, H., Haider, A. & Ametamey, S. M. N-methyl-d-aspartate (NMDA) receptor modulators: a patent review (2015–present). Expert Opin. Ther. Pat. 30, 743–767 (2020).
Karakas, E., Simorowski, N. & Furukawa, H. Subunit arrangement and phenylethanolamine binding in GluN1/GluN2B NMDA receptors. Nature 475, 249–253 (2011).
Thapaliya, E. R. et al. Photochemical control of drug efficacy: a comparison of uncaging and photoswitching ifenprodil on NMDA receptors. ChemPhotoChem https://doi.org/10.1002/cptc.202000240 (2021).
Axerio-Cilies, P. et al. Inhibitors of androgen receptor activation function-2 (AF2) site identified through virtual screening. J. Med. Chem. 54, 6197–6205 (2011).
Sterling, T. & Irwin, J. J. ZINC 15—ligand discovery for everyone. J. Chem. Inf. Model. 55, 2324–2337 (2015).
Friesner, R. A. et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J. Med. Chem. 47, 1739–1749 (2004).
Zsoldos, Z., Reid, D., Simon, A., Sadjad, S. B. & Johnson, A. P. eHiTS: a new fast, exhaustive flexible ligand docking system. J. Mol. Graph. Model. 26, 198–212 (2007).
Axerio-Cilies, P. Discovery of a selective GluN2A-containing N-Methyl-d-Aspartate Glutamate Receptor Positive Modulator as a Novel Post-Stroke Neuroprotective Therapy PhD thesis, Univ. British Columbia (2016).
Chen, X., Zhu, H., Liu, X., Li, Q. & Dong, M. Design and synthesis of novel GluN2A NMDAR positive allosteric modulators via scaffold hopping strategy as anti-stroke therapeutic agents. Bioorg. Med. Chem. 83, 117236 (2023).
Cordier, C. et al. Salicylaldehyde benzoyl hydrazone: isomerization due to water. A structural analysis using a combination of NMR, IR, and theoretical investigations. Struct. Chem. 15, 295–307 (2004).
Flegontov, S. A., Titova, Z. S., Stolyarov, A. P., Buzykin, B. I. & Kitaev, Y. P. Hydrazones 62. Structures of aroylhydrazones with intra-molecular hydrogen-bonding. B Acad. Sci. USSR Div. Chem. Sci. 28, 948–953 (1979).
Patorski, P., Wyrzykiewicz, E. & Bartkowiak, G. Synthesis and conformational assignment of N-(E)-stilbenyloxymethylenecarbonyl-substituted hydrazones of acetone and o-(m- and p-) chloro- (nitro-) benzaldehydes by means of H-1 and C-13 NMR spectroscopy. J Spectrosc. https://doi.org/10.1155/2013/197475 (2013).
Lopes, A. B. et al. Characterization of amide bond conformers for a novel heterocyclic template of N-acylhydrazone derivatives. Molecules 18, 11683–11704 (2013).
Arya, N., Mishra, S. K. & Suryaprakash, N. Intramolecular hydrogen bond directed distribution of conformational populations in the derivatives of N′-benzylidenebenzohydrazide. New J. Chem. 43, 13134–13142 (2019).
Palla, G., Predieri, G., Domiano, P., Vignali, C. & Turner, W. Conformational behavior and E/Z isomerization of N-acyl and N-aroylhydrazones. Tetrahedron 42, 3649–3654 (1986).
Li, P., Chen, X. G., Shulin, E. & Asher, S. A. UV resonance raman ground and excited state studies of amide and peptide isomerization dynamics. J. Am. Chem. Soc. 119, 1116–1120 (1997).
Kline, M., Pierce, D. & Cheatham, S. Assignment of oxime and hydrazone configuration using (1) H-(15) N and (13) C-(15) N coupling measurements at natural abundance. Magn. Reson. Chem. 55, 154–156 (2017).
Salussolia, C. L., Prodromou, M. L., Borker, P. & Wollmuth, L. P. Arrangement of subunits in functional NMDA receptors. J. Neurosci. 31, 11295–11304 (2011).
Bernhardt, P. V., Chin, P., Sharpe, P. C. & Richardson, D. R. Hydrazone chelators for the treatment of iron overload disorders: iron coordination chemistry and biological activity. Dalton Trans. https://doi.org/10.1039/b704102k (2007).
Jalali-Yazdi, F., Chowdhury, S., Yoshioka, C. & Gouaux, E. Mechanisms for zinc and proton inhibition of the GluN1/GluN2A NMDA receptor. Cell 175, 1520–1532 e1515 (2018).
Franks, A. T., Peng, D., Yang, W. & Franz, K. J. Characterization of a photoswitching chelator with light-modulated geometric, electronic, and metal-binding properties. Inorg. Chem. 53, 1397–1405 (2014).
Hall, K. C. et al. Metal-binding hydrazone photoswitches for visible light reactivity and variable relaxation kinetics. Photochem. Photobiol. Sci. 16, 1604–1612 (2017).
Sugantino, M. & Roderick, S. L. Crystal structure of Vat(D): an acetyltransferase that inactivates streptogramin group A antibiotics. Biochemistry 41, 2209–2216 (2002).
Schehr, M. et al. 2-Azo-, 2-diazocine-thiazols and 2-azo-imidazoles as photoswitchable kinase inhibitors: limitations and pitfalls of the photoswitchable inhibitor approach. Photochem. Photobiol. Sci. 18, 1398–1407 (2019).
Gao, L. et al. A robust, GFP-orthogonal photoswitchable inhibitor scaffold extends optical control over the microtubule cytoskeleton. Cell Chem. Biol. 28, 228–241 e226 (2021).
Ifa, D. R., Rodrigues, C. R., de Alencastro, R. B., Fraga, C. A. M. & Barreiro, E. J. A possible molecular mechanism for the inhibition of cysteine proteases by salicylaldehyde N-acylhydrazones and related compounds. J Mol. Struct. THEOCHEM 505, 11–17 (2000).
Bennett, B. D. et al. Absolute metabolite concentrations and implied enzyme active site occupancy in Escherichia coli. Nat. Chem. Biol. 5, 593–599 (2009).
Casey, J. R., Grinstein, S. & Orlowski, J. Sensors and regulators of intracellular pH. Nat. Rev. Mol. Cell Biol. 11, 50–61 (2010).
Pirie, N. W. & Pinhey, K. G. The titration curve of glutathione. J. Biol. Chem. 84, 321–333 (1929).
Reynders, M. et al. PHOTACs enable optical control of protein degradation. Sci. Adv. 6, eaay5064 (2020).
Paoletti, P., Ellis-Davies, G. C. R. & Mourot, A. Optical control of neuronal ion channels and receptors. Nat. Rev. Neurosci. 20, 514–532 (2019).
Berlin, S. et al. A family of photoswitchable NMDA receptors. eLife https://doi.org/10.7554/eLife.12040 (2016).
Pittolo, S. et al. An allosteric modulator to control endogenous G protein-coupled receptors with light. Nat. Chem. Biol. 10, 813–815 (2014).
Rovira, X. et al. OptoGluNAM4.1, a photoswitchable allosteric antagonist for real-time control of mGlu4 receptor activity. Cell Chem. Biol. 23, 929–934 (2016).
Klippenstein, V., Hoppmann, C., Ye, S., Wang, L. & Paoletti, P. Optocontrol of glutamate receptor activity by single side-chain photoisomerization. eLife https://doi.org/10.7554/eLife.25808 (2017).
Dong, M. et al. Near-infrared photoswitching of azobenzenes under physiological conditions. J. Am. Chem. Soc. 139, 13483–13486 (2017).
Dong, M., Babalhavaeji, A., Hansen, M. J., Kalman, L. & Woolley, G. A. Red, far-red, and near infrared photoswitches based on azonium ions. Chem. Commun. 51, 12981–12984 (2015).
Schmidt, D. et al. Axitinib: a photoswitchable approved tyrosine kinase inhibitor. ChemMedChem 13, 2415–2426 (2018).
Kirchner, S. et al. Hemipiperazines as peptide-derived molecular photoswitches with low-nanomolar cytotoxicity. Nat. Commun. 13, 6066 (2022).
Zelenitsky, S. A. & Zhanel, G. G. Phenazopyridine in urinary tract infections. Ann. Pharmacother. 30, 866–868 (1996).
Velema, W. A. et al. Optical control of antibacterial activity. Nat. Chem. 5, 924–928 (2013).
Lachmann, D., Lahmy, R. & Konig, B. Fulgimides as light-activated tools in biological investigations. Eur. J. Org. Chem. 2019, 5018–5024 (2019).
Sailer, A. et al. Hemithioindigos for cellular photopharmacology: desymmetrised molecular switch scaffolds enabling design control over the isomer-dependency of potent antimitotic bioactivity. ChemBioChem 20, 1305–1314 (2019).
Babii, O. et al. Controlling biological activity with light: diarylethene-containing cyclic peptidomimetics. Angew. Chem. Int. Ed. Engl. 53, 3392–3395 (2014).
Lentes, P. et al. Nitrogen bridged diazocines: photochromes switching within the near-infrared region with high quantum yields in organic solvents and in water. J. Am. Chem. Soc. 141, 13592–13600 (2019).
Ozcoban, C. et al. Water-soluble Py-BIPS spiropyrans as photoswitches for biological applications. Org. Lett. 17, 1517–1520 (2015).
Chai, J. D. & Head-Gordon, M. Long-range corrected hybrid density functionals with damped atom-atom dispersion corrections. Phys. Chem. Chem. Phys. 10, 6615–6620 (2008).
Lange, A. W. & Herbert, J. M. A simple polarizable continuum solvation model for electrolyte solutions. J. Chem. Phys. 134, 204110 (2011).
Karakas, E. & Furukawa, H. Crystal structure of a heterotetrameric NMDA receptor ion channel. Science 344, 992–997 (2014).
Gordon, J. C. et al. H++: a server for estimating pKas and adding missing hydrogens to macromolecules. Nucleic Acids Res. 33, W368–371 (2005).
AMBER 2016 (Univ, California, San Francisco, 2016).
Maier, J. A. et al. ff14SB: improving the accuracy of protein side chain and backbone parameters from ff99SB. J. Chem. Theory Comput. 11, 3696–3713 (2015).
Jorgensen, W., Chandrasekhar, J., Madura, J., Impey, R. & Klein, M. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926 (1983).
Wang, J., Wolf, R. M., Caldwell, J. W., Kollman, P. A. & Case, D. A. Development and testing of a general amber force field. J. Comput. Chem. 25, 1157–1174 (2004).
Gaussian 09, Revision B.01 (Gaussian, 2009).
Liu, Z., Zhou, J., Wu, R. & Xu, J. Mechanism of assembling isoprenoid building blocks 1. Elucidation of the structural motifs for substrate binding in geranyl pyrophosphate synthase. J. Chem. Theory Comput. 10, 5057–5067 (2014).
Ryckaert, J. P. C., Ciccotti, G. & Berendsen, H. J. C. Numerical integration of the Cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J. Comp. Phys. 23, 327–341 (1977).
Genheden, S. & Ryde, U. The MM/PBSA and MM/GBSA methods to estimate ligand-binding affinities. Expert Opin. Drug Discov. 10, 449–461 (2015).
Miller, B. R. 3rd et al. MMPBSA.py: an efficient program for end-state free energy calculations. J. Chem. Theory Comput. 8, 3314–3321 (2012).
Cox, G. et al. A common platform for antibiotic dereplication and adjuvant discovery. Cell Chem. Biol. 24, 98–109 (2017).
Neidhardt, F. C., Bloch, P. L. & Smith, D. F. Culture medium for enterobacteria. J. Bacteriol. 119, 736–747 (1974).
Acknowledgements
The authors gratefully acknowledge the contribution of A. Jaikaran for advice and helpful comments and S. L. Roderick for the crystallization and structure determination of VatD in complex with 3DHO. We also thank referees of the paper for helpful suggestions and, in particular, O. Thorn-Seshold for suggesting important tests. This research was funded by the Natural Science and Engineering Research Council of Canada (NSERC—RGPIN-115110) to G.A.W., a Canadian Institutes of Health Research Foundation Grant (CIHR—FDN-154286) to Y.T.W., the China Scholarship Council (CSC), the China Postdoctoral Science Foundation (2020TQ0095, and 2021M691005) and the National Natural Science Foundation of China (22101084) to Z.Z., an APRENTICE fellowship to G.L. via the Collaborative Research and Training Experience (CREATE) programme (NSERC); a National Key Research and Development Project of China (2018YFA0900804) grant to M.X.D., and a Canadian Institutes of Health Research grant (FRN-148463), and a Canada Research Chair to G.D.W. Y.T.W. is the holder of Heart and Stroke Foundation of British Columbia and Yukon Chair in Stroke Research.
Author information
Authors and Affiliations
Contributions
Z.Z., G.N.T.L., M.D. and G.A.W. developed the concepts and designed, carried out and analysed in vitro photoswitching experiments. G.N.T.L. and G.A.W. developed the concepts and designed, carried out and analysed thiol-mediated switching experiments in vitro and in cellulo. Y.G., Y.T.W. and P.A.-C. designed, carried out and analysed all electrophysiology experiments. X.T., X.C. and M.D. designed and synthesized compounds and carried out docking experiments. L.E., E.B. and G.D.W. designed, carried out and analysed enzyme activity assays for compound 3DHO. D.C.B. designed and analysed NMR experiments. S.T. provided additional measurements of ionization and photoswitching of compounds. Y.T.W., M.D. and G.A.W. supervised the project. Z.Z., G.N.T.L., Y.G., Y.T.W., M.D. and G.A.W. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
No financial interest or any direct conflict of interest exists. Y.T.W. and P.A.-C. declare an internationalized patent related to this class of NPAM compounds (N-methyl-d-aspartate receptor (NMDAR) allosteric modulators and methods for their use, WO 2018/107853 A2).
Peer review
Peer review information
Nature Chemistry thanks Nadja Simeth, Wiktor Szymanski, Oliver Thorn-Seshold and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–61, methods and Tables 1–8.
Supplementary Data 1 (download ZIP )
mol2 and .xyz files of structures in Supplementary Table 1.
Supplementary Data 2 (download ZIP )
mol2 and .xyz files of structures in Supplementary Table 2.
Supplementary Data 3 (download ZIP )
mol2 and .xyz files of structures in Supplementary Table 6.
Supplementary Data 4 (download ZIP )
mol2 and .xyz files of structures in Supplementary Table 8.
Supplementary Data 5
X-ray structure file.
Source data
Source Data Fig. 1 (download ZIP )
ChemDraw files, X-ray structure file and calculated structures files.
Source Data Fig. 2 (download ZIP )
ChemDraw files, calculated structures files; PyMol session file for Fig. 2i; Statistical Source Data Fig. 2c,d,g,h.
Source Data Fig. 3 (download ZIP )
ChemDraw files; Statistical Source Data Fig. 3b,e–g; PyMol session file for Fig. 3d.
Source Data Fig. 4 (download ZIP )
ChemDraw files; Statistical Source Data Fig. 4a–d.
Source Data Fig. 5 (download ZIP )
ChemDraw files; Statistical Source Data Fig. 5b–e,g.
Source Data Fig. 6 (download ZIP )
ChemDraw files; Statistical Source Data Fig. 6b,c,f–h.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, Z., Le, G.N.T., Ge, Y. et al. Isomerization of bioactive acylhydrazones triggered by light or thiols. Nat. Chem. 15, 1285–1295 (2023). https://doi.org/10.1038/s41557-023-01239-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41557-023-01239-5
This article is cited by
-
Hydrazone-based photocages for precise sub-organelle visualization and drug release
Communications Chemistry (2025)