Abstract
Understanding cellular functions in health and disease requires dissecting spatiotemporal variations in the subcellular transcriptome. Existing methods for mitochondrial RNA profiling suffer from limitations, including low resolution, contamination and dependence on genetic manipulation. Here we present a bioorthogonal photocatalytic labelling and sequencing strategy (CAT-seq) that enables high-resolution, in situ profiling of mitochondrial RNA in living cells without genetic manipulation. We identified a quinone methide probe for efficient RNA labelling. Rigorous validation and optimization enabled CAT-seq to successfully profile mitochondrial RNA and track RNA dynamics in HeLa cells. We further applied CAT-seq to the challenging RAW 264.7 macrophages, revealing an underlying mitochondrial translational remodelling pathway. By leveraging the chemistry of quinone methide warheads, we established an orthogonal labelling system enabling synchronous RNA and protein multi-omics profiling within the same sample. Together, assisted by bioorthogonal photocatalytic chemistry, CAT-seq offers a general, non-genetic and well-compatible approach for subcellular-resolved RNA and multi-omics investigations, particularly in studies of intact primary living samples that are otherwise challenging to access.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data supporting the findings of this study are available within the Article, the Supplementary Information, and the Source data. Uncropped Western blot scans for figures in the Article and extended data are provided in the Source data; those for supplementary figures are included in the Supplementary Information. Raw microscopy image stacks underlying the imaging experiments (for example, confocal/fluorescence time-lapse files in proprietary formats such as .czi) are available from the corresponding author upon reasonable request, as the datasets are very large and require conversion from proprietary formats before sharing. The mass spectrometry proteomics data generated in this study have been deposited with the ProteomeXchange Consortium via the iProX partner repository under accession PXD065542. The transcriptomics data have been deposited in GEO under accession GSE255007. Public reference proteomes were obtained from the UniProt database. Source data are provided with this paper.
References
Buxbaum, A. R., Haimovich, G. & Singer, R. H. In the right place at the right time: visualizing and understanding mRNA localization. Nat. Rev. Mol. Cell Biol. 16, 95–109 (2015).
Mercer, T. R. et al. The human mitochondrial transcriptome. Cell 146, 645–658 (2011).
Jedynak-Slyvka, M., Jabczynska, A. & Szczesny, R. J. Human mitochondrial RNA processing and modifications: overview. Int. J. Mol. Sci. 22, 7999 (2021).
Grochowska, J., Czerwinska, J., Borowski, L. S. & Szczesny, R. J. Mitochondrial RNA, a new trigger of the innate immune system. Wiley Interdiscip. Rev. RNA 13, e1690 (2022).
Zhu, X. et al. Non-coding 7S RNA inhibits transcription via mitochondrial RNA polymerase dimerization. Cell 185, 2309–2323.e2324 (2022).
Buck, M. D. et al. Mitochondrial dynamics controls T cell fate through metabolic programming. Cell 166, 63–76 (2016).
Delaunay, S. et al. Mitochondrial RNA modifications shape metabolic plasticity in metastasis. Nature 607, 593–603 (2022).
Lisci, M. et al. Mitochondrial translation is required for sustained killing by cytotoxic T cells. Science 374, eabe9977 (2021).
Hooftman, A. et al. Macrophage fumarate hydratase restrains mtRNA-mediated interferon production. Nature 615, 490–498 (2023).
Wigger, L. et al. Multi-omics profiling of living human pancreatic islet donors reveals heterogeneous beta cell trajectories towards type 2 diabetes. Nat. Metab. 3, 1017–1031 (2021).
Walsh, L. A. & Quail, D. F. Decoding the tumor microenvironment with spatial technologies. Nat. Immunol. 24, 1982–1993 (2023).
Zhu, C. et al. Joint profiling of histone modifications and transcriptome in single cells from mouse brain. Nat. Methods 18, 283–292 (2021).
Asp, M., Bergenstrahle, J. & Lundeberg, J. Spatially resolved transcriptomes-next generation tools for tissue exploration. Bioessays 42, e1900221 (2020).
Singha, M., Spitalny, L., Nguyen, K., Vandewalle, A. & Spitale, R. C. Chemical methods for measuring RNA expression with metabolic labeling. Wiley Interdiscip. Rev. RNA 12, e1650 (2021).
Sang, L. et al. Mitochondrial long non-coding RNA GAS5 tunes TCA metabolism in response to nutrient stress. Nat. Metab. 3, 90–106 (2021).
Villanueva, E. et al. System-wide analysis of RNA and protein subcellular localization dynamics. Nat. Methods 21, 60–71 (2024).
Ren, J. et al. Spatiotemporally resolved transcriptomics reveals the subcellular RNA kinetic landscape. Nat. Methods 20, 695–705 (2023).
Zeng, H. et al. Spatially resolved single-cell translatomics at molecular resolution. Science 380, eadd3067 (2023).
Fazal, F. M. et al. Atlas of subcellular RNA localization revealed by APEX-Seq. Cell 178, 473–490 e426 (2019).
Wang, P. et al. Mapping spatial transcriptome with light-activated proximity-dependent RNA labeling. Nat. Chem. Biol. 15, 1110–1119 (2019).
Engel, K. L. et al. Analysis of subcellular transcriptomes by RNA proximity labeling with Halo-seq. Nucleic Acids Res. 50, e24 (2022).
Huang, Z. et al. Bioorthogonal photocatalytic decaging-enabled mitochondrial proteomics. J. Am. Chem. Soc. 143, 18714–18720 (2021).
Liu, Z. et al. Spatially resolved cell tagging and surfaceome labeling via targeted photocatalytic decaging. Chem 8, 2179–2191 (2022).
Liu, Z. et al. Bioorthogonal photocatalytic proximity labeling in primary living samples. Nat. Commun. 15, 2712 (2024).
Zhang, Y. et al. Bioorthogonal quinone methide decaging enables live-cell quantification of tumor-specific immune interactions. J. Am. Chem. Soc. 146, 15186–15197 (2024).
Guo, F., Qin, S., Liu, Z., Chen, P. R. & Fan, X. Decaging-to-labeling: development and investigation of quinone methide warhead for protein labeling. Bioorg. Chem. 143, 107088 (2024).
Rokita, S. E., Yang, J., Pande, P. & Greenberg, W. A. Quinone methide alkylation of deoxycytidine. J. Org. Chem. 62, 3010–3012 (1997).
Ali, K. et al. Reactivity vs. selectivity of quinone methides: synthesis of pharmaceutically important molecules, toxicity and biological applications. Chem. Commun. (Camb.) 58, 6160–6175 (2022).
Pande, P., Shearer, J., Yang, J. H., Greenberg, W. A. & Rokita, S. E. Alkylation of nucleic acids by a model quinone methide. J. Am. Chem. Soc. 121, 6773–6779 (1999).
Boughanem, H. et al. The emergent role of mitochondrial RNA modifications in metabolic alterations. Wiley Interdiscip. Rev. RNA 14, e1753 (2023).
Ali, A. T. et al. Nuclear genetic regulation of the human mitochondrial transcriptome. eLife 8, e41927 (2019).
Wang, N., Liang, H. & Zen, K. Molecular mechanisms that influence the macrophage m1–m2 polarization balance. Front. Immunol. 5, 614 (2014).
Aegerter, H., Lambrecht, B. N. & Jakubzick, C. V. Biology of lung macrophages in health and disease. Immunity 55, 1564–1580 (2022).
Tannahill, G. M. et al. Succinate is an inflammatory signal that induces IL-1β through HIF-1α. Nature 496, 238–242 (2013).
Mills, E. L. et al. Succinate dehydrogenase supports metabolic repurposing of mitochondria to drive inflammatory macrophages. Cell 167, 457–470 e413 (2016).
Li, J. et al. YTHDF2 mediates the mRNA degradation of the tumor suppressors to induce AKT phosphorylation in N6-methyladenosine-dependent way in prostate cancer. Mol. Cancer 19, 152 (2020).
Feng, Y. et al. Stabilization of RRBP1 mRNA via an m6A-dependent manner in prostate cancer constitutes a therapeutic vulnerability amenable to small-peptide inhibition of METTL3. Cell. Mol. Life Sci. 81, 414 (2024).
Kazuhito, T. & Wei, F. Y. Posttranscriptional modifications in mitochondrial tRNA and its implication in mitochondrial translation and disease. J. Biochem. 168, 435–444 (2020).
Asano, K. et al. Metabolic and chemical regulation of tRNA modification associated with taurine deficiency and human disease. Nucleic Acids Res. 46, 1565–1583 (2018).
Morscher, R. J. et al. Mitochondrial translation requires folate-dependent tRNA methylation. Nature 554, 128–132 (2018).
Chen, D. et al. Deletion of Gtpbp3 in zebrafish revealed the hypertrophic cardiomyopathy manifested by aberrant mitochondrial tRNA metabolism. Nucleic Acids Res. 47, 5341–5355 (2019).
Peng, G. X. et al. The human tRNA taurine modification enzyme GTPBP3 is an active GTPase linked to mitochondrial diseases. Nucleic Acids Res. 49, 2816–2834 (2021).
Shinoda, S. et al. Mammalian NSUN2 introduces 5-methylcytidines into mitochondrial tRNAs. Nucleic Acids Res. 47, 8734–8745 (2019).
Van Haute, L. et al. NSUN2 introduces 5-methylcytosines in mammalian mitochondrial tRNAs. Nucleic Acids Res. 47, 8720–8733 (2019).
Fischer, M. et al. Early effector maturation of naïve human CD8+ T cells requires mitochondrial biogenesis. Eur. J. Immunol. 48, 1632–1643 (2018).
Gaudet, P., Livstone, M. S., Lewis, S. E. & Thomas, P. D. Phylogenetic-based propagation of functional annotations within the Gene Ontology consortium. Brief. Bioinform. 12, 449–462 (2011).
Basu, U., Bostwick, A. M., Das, K., Dittenhafer-Reed, K. E. & Patel, S. S. Structure, mechanism, and regulation of mitochondrial DNA transcription initiation. J. Biol. Chem. 295, 18406–18425 (2020).
Ren, Z., Tang, W., Peng, L. & Zou, P. Profiling stress-triggered RNA condensation with photocatalytic proximity labeling. Nat. Commun. 14, 7390 (2023).
Queiroz, R. M. L. et al. Comprehensive identification of RNA-protein interactions in any organism using orthogonal organic phase separation (OOPS). Nat. Biotechnol. 37, 169–178 (2019).
Rao, A., Barkley, D., França, G. S. & Yanai, I. Exploring tissue architecture using spatial transcriptomics. Nature 596, 211–220 (2021).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Gu, Z., Gu, L., Eils, R., Schlesner, M. & Brors, B. circlize Implements and enhances circular visualization in R. Bioinformatics 30, 2811–2812 (2014).
Lopez-Delisle, L. et al. pyGenomeTracks: reproducible plots for multivariate genomic datasets. Bioinformatics 37, 422–423 (2021).
Acknowledgements
We acknowledge funding from the National Natural Science Foundation of China and the National Key Research and Development Program of China (grant numbers 22222701, 2022YFA1304700, 2023YFC3402200, 2024YFA0917503, 2023YFA1506501, 2024YFA1107000, 32170595, 92478119, 22321005, 92253301 and 22137001), Natural Science Foundation of Beijing Municipality (grant numbers Z240010 and Z221100007422046), Beijing Nova Program (grant number 20230484442), Beijing Hospitals Authority Clinical Medicine Development Funding (grant number YGLX202338) and a Boehringer Ingelheim Young Faculty Research Award (X.F.).
Author information
Authors and Affiliations
Contributions
X.F., J.L. and P.R.C. conceived and supervised the project. Y.B., L.Y. and Q.D. performed the experiments and analysed the data. Y.Z., F.G., L.K. and R.W. helped with material preparation and data analysis. Y.B., L.Y., Q.D., X.F., J.L. and P.R.C. wrote the paper with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
X.F., J.L., Y.B., L.Y. and Q.D. are inventors on a patent application (application number 202510356028.7) filed by Peking University with the China National Intellectual Property Administration, covering the CAT-seq method reported in this article. The application is currently under examination. The other authors declare no competing interests.
Peer review
Peer review information
Nature Chemistry thanks Ivan Corrêa Jr and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Investigation of in vitro RNA tagging on oligo and small molecule.
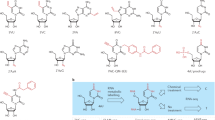
(a) Measurement of the QM-cytidine adduct formation yield in vitro. Oligo is a 28-nt DNA oligonucleotide, with a molecular weight 8,614. Biotin-oligo is a biotin-conjugated DNA oligo with the same sequence as the 28-nt oligo. 10 μg (1.16 nmol) oligo, 10 μM Ir catalyst, 200 μM QMPAB-1 were incubated in 100 μL reaction and illuminated with 4 mW/cm2 blue LED for 15 min at room temperature. Then the labeled oligo was purified by Oligo Clean & Concentrator kit. 116 pmol (~1μg) purified oligo was loaded to the membrane with biotin oligo (10, 5, 1, pmol) as reference for dot blot analysis. The intensity of 1μg labeled oligo falls between those of 1 and 5 pmol biotin-oligo, indicating that biotinylation yield was approximately once per 23 ~ 116 oligo, or approximately one biotin per 640 ~ 3,200 nucleobases. Detailed calculation was described in the Methods section. (b) Tandem MS analysis of QM-addition reaction. Representative LC-MS/MS chromatograms are shown for isolated QM-addition of QMMNP-5 probe to cytidine (right). Negative control experiments without addition product are shown on the left. cps, counts per second. (c) Infrared spectrum of isolated QM-addition of QMMNP-5 probe to cytidine. (d) Chromatography characterization of reaction between QMMNP-5 and 1-methylcytosine. The reaction mixture was compared with isolated standard N3 or N4-conjugated 1-methylcytosine. (e) Reaction mechanism of QM probe and different biomolecules. For RNA, the hydrogen bond-assisted conjugation pathway is shown during labeling and for protein, it is more possible to favor a nucleophilic addition pathway during the reaction.
Extended Data Fig. 2 Rational design of CAT-seq in HeLa cells.
(a) Workflow of in vivo RNA labeling and enrichment. (b) Fluorescence imaging of HeLa cells treated with Ir catalyst (blue) and MitoTracker Red (red). Live cells were performed for 30 min for the catalyst self-localization and then 5 min for MitoTracker Red staining for cellular mitochondria indication. Scale bar, 10 μm. Three biological replicates were repeated with similar results. (c) Immunofluorescent images of HeLa after QMPAB-1 labeling. Green, biotin labeling signal. Red, staining of mitochondrial marker TOMM20. Scale bar: 10 μm. Three biological replicates were repeated with similar results. (d) RT-qPCR analysis of enriched RNA after QMPAB-1 labeling (50 nM photocatalyst and 50 μM QMPAB-1): MTCO1 and MTND1 (mitochondrial mRNAs), GAPDH and ACTNB (cytosolic mRNA markers). Relative abundance is calculated relative to the control without QMPAB-1 probe. Error bars indicate the s.d. calculated from three biological independent biological replicates. n = 3; mean ± s.d. (e) High-resolution images of HeLa after CAT-seq labeling (50 nM photocatalyst and 50 μM QMPAB-2). Green, biotin labeling signal. Red, staining of mitochondrial marker TOMM20. Scale bar: 5 μm. Three biological replicates were repeated with similar results. (f). Chemical structure of catalyst and probe, and immunofluorescent images of HeLa after labeling (50 nM photocatalyst and 50 μM probe). Green, biotin labeling signal. Red, staining of mitochondrial marker TOMM20 or ER marker Calnexin. Blue, Hoechst staining. Scale bar: 10 μm. Three biological replicates were repeated with similar results. (g & h) Immunofluorescent images of HeLa after CAT-seq labeling with different QMPAB-2 treatment time (f) or incubation temperature (g). Green, biotin labeling signal. Red, staining of mitochondrial marker TOMM20. Scale bar: 10 μm. Error bars indicate the s.d. calculated from three independent biological replicates. n = 3; mean ± s.d. P values were calculated by one-sample t-test. Panel a created with BioRender.com.
Extended Data Fig. 3 High throughput sequencing based CAT-seq comparison with enzymatic proximity labeling techniques.
(a) The Pearson correlation coefficient and p values from a two-sided t-test for log2 expression levels of mitochondrial RNA in two independent high-throughput sequencing samples are presented. (b) The proportion of alignment rates between the mitochondrial genome and the whole-cell genome in sequencing data with commercial mitochondrial isolation kit. (c-e) Efficiency and Mitochondrial specificity compared with APEX-Seq: Proportion of reads aligning to the mitochondrial genome compared with the whole-cell genome of HEK293T in sequencing data of APEX-Seq (c) and CAT-Seq (d) with the columns showing the fraction of mitochondrial transcriptome reads and the Pearson correlation coefficient between log2-expression levels of CAT-seq vs. APEX-Seq was presented (e) (APEX-seq source data was acquired with the following GEO accession number in GSE116008: GSM3206981, GSM3206982, GSM3206983, GSM3206984). P values were calculated by one-sample t-test. (f & g) Labeling efficiency compared with CAP-Seq: The Pearson correlation coefficient between log2-expression levels of CAT-seq vs. CAP-Seq (f) and scatter plot of mitochondrial RNA (mRNAs, rRNAs) enrichment fold change (g) and were presented. Box plots show the median (center line), 25th and 75th percentiles (box bounds), and whiskers extending to the minimum and maximum values. Error bars indicate the s.d. calculated from three independent replicates. n = 16 (as the count of mitochondrial 13 mRNA, 2 rRNA and 1 pseudogene); mean ± s.d. P values were calculated by one-sample t-test.
Extended Data Fig. 4 Explore mitochondrial RNA variation and mechanism of metabolic remodeling in macrophage M1 polarization.
(a) Validation of LPS stimulation in RAW 264.7 cell line. RAW 264.7 cells were treated with LPS, and NLRP3 is reported to show upregulation during this process and the intensities were quantified. Error bars indicate the s.d. calculated from three independent replicates. n = 3; mean ± s.d. P values were calculated by one-sample t-test. (b) The Pearson correlation coefficient and p values from a two-sided t-test for log2 expression levels of mitochondrial RNA in two independent high-throughput sequencing samples are presented without or with LPS stimulation. (c) Genomic visualization: Overlap of the reads with the mitochondrial genome confirms accurate targeting of mitochondrial transcripts, both with and without LPS treatment. (d) Mitochondrial RNA contribution to PCA: Detailed breakdown of individual mitochondrial RNA components influencing sample discrimination. (e) Detection of mitochondrial RNA variation during different LPS stimulation time. P values were calculated by two-sided Student’s t-test. (f) RNA modification mapping: Identification seven different modifications in labeled mitochondrial large fragment RNA (mainly mRNAs and rRNAs) using tandem mass spectrometry. Abundances are calculated with standard curves. Error bars indicate the s.d. calculated from three independent replicates. n = 3; mean ± s.d. P values were calculated by one-sample t-test. (g & h) Immunoblotting: Protein levels of NSUN2 and GTPBP3 (g) in RAW 264.7 or NSUN2 in HEK293T (h) knockdown cell lines using shRNA transfection and the intensities were quantified. Error bars indicate the s.d. calculated from three independent replicates. n = 3; mean ± s.d. P values were calculated by one-sample t-test. P values were calculated by one-sample t-test. (i). Nascent protein translation analysis in mitochondria of HEK293T cells with NSUN2 knockdown. Three biological replicates were repeated with similar results.
Supplementary information
Supplementary Information (download PDF )
Supplementary Methods, Figs. 1–12, NMR spectra and References.
Supplementary Table 1 (download XLSX )
Supplementary Table 1. Sequence of nucleic acids used in the research.
Supplementary Table 2 (download XLSX )
Supplementary Table 2. Proteome data for synchronous labelling.
Source data
Source Data Fig. 1 (download ZIP )
Statistical source data and uncropped blots.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download ZIP )
Statistical source data and uncropped blots.
Source Data Fig. 4 (download ZIP )
Statistical source data and uncropped blots and gels.
Source Data Fig. 5 (download ZIP )
Statistical source data and uncropped blots.
Source Data Extended Data Fig. 1 (download ZIP )
Statistical source data and uncropped blots.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download ZIP )
Statistical source data and uncropped blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bi, Y., Yu, L., Deng, Q. et al. Photocatalytic labelling-enabled subcellular-resolved RNA profiling and synchronous multi-omics investigation. Nat. Chem. 17, 1871–1882 (2025). https://doi.org/10.1038/s41557-025-01946-1
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41557-025-01946-1