Abstract
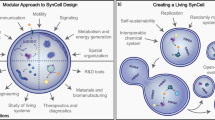
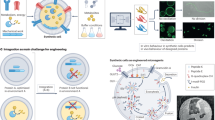
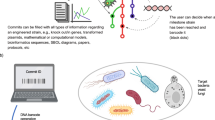
Building synthetic versions of biological cells from the bottom up offers an unprecedented opportunity to understand the rules of life and harness cellular capabilities in biotechnology. Whereas substantial progress has been made in recapitulating elementary cell functions, we argue that accelerating the engineering of synthetic cells requires a shift in research practices. The dominant approach—rationally designing and integrating functional modules—becomes restrictive when dealing with the massively complex biochemical pathways associated with life, especially when design principles remain unclear. We advocate moving away from theoretical rational design towards a data-driven model that is centred on library generation. Inspired by a systems chemistry perspective, this strategy prioritizes the systematic creation and distribution of composition–function libraries. To enable this, experimental strategies must integrate high-throughput synthetic cell generation, automation and closed-feedback control of workflows. Broad adoption will also require greater emphasis on quantitative benchmarking, and the de-skilling of techniques, supporting effective laboratory-to-laboratory collaboration.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Elani, Y. & Seddon, J. M. What it means to be alive: a synthetic cell perspective. Interface Focus https://doi.org/10.1098/rsfs.2023.0036 (2023).
Sampson, K., Sorenson, C. & Adamala, K. P. Preparing for the future of precision medicine: synthetic cell drug regulation. Synth. Biol. 9, ysae004 (2024).
Rothschild, L. J. et al. Building synthetic cells—from the technology infrastructure to cellular entities. ACS Synth. Biol. 13, 974–997 (2024).
Boyd, M. A., Thavarajah, W., Lucks, J. B. & Kamat, N. P. Robust and tunable performance of a cell-free biosensor encapsulated in lipid vesicles. Sci. Adv. 9, eadd6605 (2023).
Osawa, M., Anderson, D. E. & Erickson, H. P. Reconstitution of contractile FtsZ rings in liposomes. Science 320, 792–794 (2008).
Zhan, P., Jahnke, K., Liu, N. & Göpfrich, K. Functional DNA-based cytoskeletons for synthetic cells. Nat. Chem. 14, 958–963 (2022).
Berhanu, S., Ueda, T. & Kuruma, Y. Artificial photosynthetic cell producing energy for protein synthesis. Nat. Commun. 10, 1325 (2019).
Deng, N.-N., Yelleswarapu, M., Zheng, L. & Huck, W. T. S. Microfluidic assembly of monodisperse vesosomes as artificial cell models. JACS 139, 587–590 (2017).
Steinkühler, J. et al. Controlled division of cell-sized vesicles by low densities of membrane-bound proteins. Nat. Commun. 11, 905 (2020).
Peruzzi, J. A., Galvez, N. R. & Kamat, N. P. Engineering transmembrane signal transduction in synthetic membranes using two-component systems. Proc. Natl Acad. Sci. USA 120, 2218610120 (2023).
Rampioni, G. et al. Synthetic cells produce a quorum sensing chemical signal perceived by Pseudomonas aeruginosa. Chem. Commun. 54, 2090–2093 (2018).
Göpfrich, K., Platzman, I. & Spatz, J. P. Mastering complexity: towards bottom-up construction of multifunctional eukaryotic synthetic cells. Trends Biotechnol. 36, 938–951 (2018).
Guindani, C., da Silva, L. C., Cao, S., Ivanov, T. & Landfester, K. Synthetic cells: from simple bio-inspired modules to sophisticated integrated systems. Angew. Chem. 134, e202110855 (2022).
Elani, Y. Interfacing living and synthetic cells as an emerging frontier in synthetic biology. Angew. Chem. 133, 5662–5671 (2021).
Hürtgen, D., Härtel, T., Murray, S. M., Sourjik, V. & Schwille, P. Functional modules of minimal cell division for synthetic biology. Adv. Biosyst. 3, 1800315 (2019).
Pogodaev, A. A., Lap, T. T. & Huck, W. T. S. The dynamics of an oscillating enzymatic reaction network is crucially determined by side reactions. ChemSystemsChem https://doi.org/10.1002/syst.202000033 (2021).
van Roekel, H. W. H. et al. Programmable chemical reaction networks: emulating regulatory functions in living cells using a bottom-up approach. Chem. Soc. Rev. 44, 7465–7483 (2015).
Jakštaitė, M., Zhou, T., Nelissen, F., Huck, W. T. S. & van Sluijs, B. Active learning maps the emergent dynamics of enzymatic reaction networks. Preprint at https://doi.org/10.26434/chemrxiv-2024-vxfkz (2024).
Cardinale, S. & Arkin, A. P. Contextualizing context for synthetic biology – identifying causes of failure of synthetic biological systems. Biotechnol. J. 7, 856–866 (2012).
Peng, Z. et al. Lipid vesicle-based molecular robots. Lab Chip 24, 996–1029 (2024).
Ghosh, S. et al. Exploring emergent properties in enzymatic reaction networks: design and control of dynamic functional systems. Chem. Rev. 124, 2553–2582 (2024).
Baltussen, M. G., van de Wiel, J., Lia Fernandez Regueiro, C., Jakstaite, M. & Huck, W. T. S. A Bayesian approach to extracting kinetic information from artificial enzymatic networks. Anal. Chem. 94, 7311–7318 (2022).
Borsley, S. Membrane transport, molecular machines, and Maxwell’s demon. ChemSystemsChem https://doi.org/10.1002/syst.202400004 (2024).
Mattia, E. & Otto, S. Supramolecular systems chemistry. Nat. Nanotechnol. 10, 111–119 (2015).
Ragazzon, G. & Prins, L. J. Energy consumption in chemical fuel-driven self-assembly. Nat. Nanotechnol. 13, 882–889 (2018).
Semenov, S. N. et al. Rational design of functional and tunable oscillating enzymatic networks. Nat. Chem. 7, 160–165 (2015).
Maguire, O. R., Wong, A. S. Y., Westerdiep, J. H. & Huck, W. T. S. Early warning signals in chemical reaction networks. Chem. Commun. 56, 3725–3728 (2020).
Sun, M., Deng, J. & Walther, A. Communication and cross-regulation between chemically fueled sender and receiver reaction networks. Angew. Chem. Int. Ed. 62, e202214499 (2023).
Jahnke, K. & Göpfrich, K. Engineering DNA-based cytoskeletons for synthetic cells. Interface Focus 13, 20230028 (2023).
Samanta, A., Baranda Pellejero, L., Masukawa, M. & Walther, A. DNA-empowered synthetic cells as minimalistic life forms. Nat. Rev. Chem. 8, 454–470 (2024).
Burns, J. R. et al. Lipid-bilayer-spanning DNA nanopores with a bifunctional porphyrin anchor. Angew. Chem. Int. Ed. 52, 12069–12072 (2013).
Langecker, M., Arnaut, V., List, J. & Simmel, F. C. DNA nanostructures interacting with lipid bilayer membranes. Acc. Chem. Res. 47, 1807–1815 (2014).
Zadeh, J. N. et al. NUPACK: analysis and design of nucleic acid systems. J. Comput. Chem. 32, 170–173 (2011).
Poppleton, E., Romero, R., Mallya, A., Rovigatti, L. & Šulc, P. OxDNA.org: a public webserver for coarse-grained simulations of DNA and RNA nanostructures. Nucleic Acids Res. 49, W491–W498 (2021).
Pandi, A. et al. A versatile active learning workflow for optimization of genetic and metabolic networks. Nat. Commun. 13, 3876 (2022).
van Sluijs, B. et al. Iterative design of training data to control intricate enzymatic reaction networks. Nat. Commun. 15, 1602 (2024).
Gan, R. et al. High-throughput regulatory part prototyping and analysis by cell-free protein synthesis and droplet microfluidics. ACS Synth. Biol. https://doi.org/10.1021/acssynbio.2c00050 (2022).
Kuruma, Y. & Ueda, T. The PURE system for the cell-free synthesis of membrane proteins. Nat. Protoc. 10, 1328–1344 (2015).
Moga, A., Yandrapalli, N., Dimova, R. & Robinson, T. Optimization of the inverted emulsion method for high-yield production of biomimetic giant unilamellar vesicles. ChemBioChem 20, 2674–2682 (2019).
Deshpande, S., Caspi, Y., Meijering, A. E. C. & Dekker, C. Octanol-assisted liposome assembly on chip. Nat. Commun. 7, 10447 (2016).
Fletcher, M. & Elani, Y. On-the-fly microfluidic control of giant vesicle compositions validated by DNA surface charge sensors. ACS Nano 19, 13768–13778 (2025).
Kohyama, S., Frohn, B. P., Babl, L. & Schwille, P. Machine learning-aided design and screening of an emergent protein function in synthetic cells. Nat. Commun. 15, 2010 (2024).
Weakly, H. M. J. et al. Several common methods of making vesicles (except an emulsion method) capture intended lipid ratios. Biophys. J. 124, 1548 (2025).
Supramaniam, P. et al. Measuring encapsulation efficiency in cell-mimicking giant unilamellar vesicles. ACS Synth. Biol. 12, 1227–1238 (2023).
Arter, W. E. et al. Biomolecular condensate phase diagrams with a combinatorial microdroplet platform. Nat. Commun. 13, 7845 (2022).
Kumar, A. et al. Multispectral live-cell imaging with uncompromised spatiotemporal resolution. Nat. Photon. 19, 1146–1156 (2025).
Fletcher, M. et al. DNA-based optical quantification of ion transport across giant vesicles. ACS Nano 16, 17128–17138 (2022).
Al Nahas, K. et al. Measuring thousands of single-vesicle leakage events reveals the mode of action of antimicrobial peptides. Anal. Chem. 94, 9530–9539 (2022).
Genot, A. J. et al. High-resolution mapping of bifurcations in nonlinear biochemical circuits. Nat. Chem. 8, 760–767 (2016).
Sugiura, H. et al. Pulse-density modulation control of chemical oscillation far from equilibrium in a droplet open-reactor system. Nat. Commun. 7, 10212 (2016).
Rapp, J. T., Bremer, B. J. & Romero, P. A. Self-driving laboratories to autonomously navigate the protein fitness landscape. Nat. Chem. Eng. 1, 97–107 (2024).
Staufer, O. et al. Science Forum: Building a community to engineer synthetic cells and organelles from the bottom-up. eLife 10, e73556 (2021).
Frischmon, C., Sorenson, C., Winikoff, M. & Adamala, K. P. Build-a-cell: engineering a synthetic cell community. Life 11, 1176 (2021).
Zhu, K. K. et al. Magnetic modulation of biochemical synthesis in synthetic cells. J. Am. Chem. Soc. 146, 13176–13182 (2024).
Rifaie-Graham, O. et al. Photoswitchable gating of non-equilibrium enzymatic feedback in chemically communicating polymersome nanoreactors. Nat. Chem. 15, 110–118 (2023).
Ashkenasy, G., Hermans, T. M., Otto, S. & Taylor, A. F. Systems chemistry. Chem. Soc. Rev. 46, 2543–2554 (2017).
Lin, T.-Y. et al. Microsoft COCO: common objects in context. Preprint at https://arxiv.org/abs/1405.0312 (2014).
Acknowledgements
M.F. and Y.E. were supported in this project by a BBSRC award BB/Z514895/1 and BB/W00125X/1. B.D is supported by a PhD studentship awarded by the EPSRC Centre for Doctoral Training in Chemical Biology - Innovation for the Life Sciences, grant ref EP/S023518/1. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
M.F. conceived the Perspective. B.D. and Y.E. discussed and refined the ideas. M.F. wrote the manuscript. B.D. contributed Fig. 3. All authors commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemistry thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fletcher, M., Diggines, B. & Elani, Y. Molecular systems engineering of synthetic cells. Nat. Chem. 18, 14–22 (2026). https://doi.org/10.1038/s41557-025-02019-z
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41557-025-02019-z