Abstract
Hydrogen gas is naturally produced by microorganisms from renewable feedstocks, yet industrial hydrogenation relies almost entirely on fossil fuel-derived H2. Despite advances in engineering biology and increasing demand for greener manufacturing, microbial H2 has seen limited application in chemical synthesis. Here we demonstrate that genetically unmodified microorganisms can generate H2 in situ to drive biocompatible alkene hydrogenation at the cell membrane using membrane-bound Pd catalysts. When combined with de novo alkene biosynthesis in engineered Escherichia coli, this system enables the simultaneous in vivo production of both substrate (alkene) and reagent (H2), followed by membrane-associated biohydrogenation to yield new metabolic end products. Quantitative life cycle assessment reveals that hybrid chemo-microbial systems utilizing waste feedstocks can outperform electrolytic hydrogenation and achieve carbon-negative outcomes. Together, this work demonstrates how microbial metabolites can be generated, intercepted and metabolically multiplexed to support biocompatible transition metal catalysis and sustainable chemical synthesis in living cells.

Similar content being viewed by others
Main
In the absence of oxygen many obligate and facultative anaerobic microorganisms produce hydrogen gas (H2) via heterotrophic growth on organic carbon. In all known examples, hydrogen is produced via the two-electron reduction of 2H+ within the active site of iron–sulfur cluster ([Fe–S]) dependent metalloenzymes1,2. The production of intracellular reducing equivalents as H2 can be used to support respiration3, cofactor regeneration4, carbon fixation5 or may be released from the cell to enable syntrophic metabolic interactions in multicellular environments6. In Escherichia spp., H2 biosynthesis is largely governed by genes encoded by the hyc operon and σ54/hycA regulon7. Initially, pyruvate and coenzyme A (CoA) are converted, non-oxidatively, to formate and acetyl-CoA by the pyruvate formate lyase PflAB8. Acidification of the cytosol by formate accumulation is mitigated by the FocAB exporter before active transport of formate back into the cell9 and conversion to CO2 and H2 by the formate dehydrogenase FdhF and [2Fe–2S] dependent [NiFe] hydrogenase HycE (Fig. 1a)10.
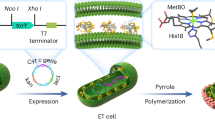
a, The native hyc-encoded H2 biosynthesis pathway in Escherichia spp. (i) and associated FHL-encoded metabolic pathway (ii). b, Proposed alkene biohydrogenation using a biocompatible Pd catalyst and microbial H2(g) from native encoded pathways (i), and multiplexed microbial metabolism for the cometabolic generation of alkene substrates and H2 in a single cell for the one-pot synthesis of hydrogenation products from D-glucose (ii).
Hydrogen gas is also ubiquitous in the field of synthetic organic chemistry and is used throughout the chemical industry for diverse functional group transformations11. However, H2 for chemical synthesis applications is currently produced by the water–gas shift reaction during steam reforming of coal, via processes that rely on diminishing natural resources and emit 15–20 kg CO2 equivalents (CO2e) per kg H2 into the atmosphere12. One industrial application of H2 is in hydrogenation reactions, where unsaturated and oxidized functional groups are reduced in the presence of H2 and a metal catalyst to produce valuable synthetic intermediates and products. Such reactions are widespread in process chemistry routes in the pharmaceutical industry (14% of all reactions13) as well as being used in the manufacture of preserved foods (C=C reduction for vegetable oils), fuels (coal liquefaction; Bergius process), hydrocracking to diesel and plastic materials (polyurethane synthesis). Many of these reactions also require organic solvents which, when combined with H2 and a transition metal catalyst, can pose safety concerns at scale.
For these reasons, combined with the need to develop more sustainable chemical manufacturing processes, greener methods for chemical hydrogenation have been a focus of research in recent years. Much of this work has focused on the generation of H2 from water by electrolysis or the use of earth-abundant metal catalysts and small organic reductants to replace H2 from fossil fuels. However, large-scale electrolysis reactors are energy-intensive and only moderately efficient (~75%)14; consequently, most organic methods continue to rely on petrochemical reagents and solvents under harsh reaction conditions15. Biocatalytic methods for many C=C, NO2 and imine reductions exist and utilize biological reducing equivalents (for example, NAD(P)H/FMNred)16,17 but often possess narrow substrate scopes and require expensive cofactors when applied in vitro. Despite the advantages of using biological catalysts for chemical hydrogenation, the use of living cells for H2 production in hydrogenation reactions has received little attention. This is despite the advantages of microbial hydrogenation being that one could: (1) interface H2-producing microorganisms with biocompatible or biogenic Pd catalysts to hydrogenate a range of substrates, (2) hydrogenate metabolites in vivo to generate new-to-nature products by fermentation and (3) generate H2 for hydrogenation from D-glucose and waste feedstocks to create circular economies.
To this end, a seminal study in this area from Sirasani et al.18 reported the use of a biocompatible Pd catalyst and H2 generated by Escherichia coli DD-2 for alkene hydrogenation. Here, H2 was generated via a heterologous hydrogen production circuit developed by Agapakis et al.19 in a modified E. coli BL21(DE3) strain. This engineered pathway used three plasmids and four chromosomal gene knockouts to generate H2 from D-glucose via an insulated pathway consisting of a heterologous pyruvate ferredoxin oxidoreductase from Desulfovibrio africanus, ferredoxin and [FeFe] hydrogenase from Clostridium acetobutylicum, and hydrogenase maturation factors HydEF and HydG from Chlamydomonas reinhardtii. However, this strain is metabolically impaired and requires prolonged growth in the presence of three antibiotics. E. coli DD-2 is therefore challenging to further manipulate to increase H2 titres, enable use of alternative feedstocks or further engineer to enable the metabolic coproduction of alkene substrates. This is despite the presence of a native metabolic pathway to H2 in Escherichia spp. and other microorganisms that could enable biohydrogenation in unmodified strains.
In this study, we report the use of unmodified E. coli strains for biocompatible hydrogenation. We demonstrate that readily available laboratory strains outperform E. coli DD-2, enable preparative-scale reactions under optimized conditions and can be applied to H2 generation from waste bread-derived carbohydrate feedstocks. Finally, by engineering E. coli to produce multiple alkene-containing metabolites, we demonstrate combined substrate (alkene) and reagent (H2) biosynthesis from D-glucose in a single cell for in situ membrane-associated biohydrogenation, yielding valuable metabolic end products (Fig. 1b). This work integrates biocompatible chemistry with multiple metabolic pathways simultaneously, offering an approach to alkane biosynthesis from sustainable feedstocks in engineered bacteria.
Results and discussion
Our studies began by screening various unmodified laboratory strains of E. coli for H2 production and biohydrogenation activity. We selected caffeic acid as a model substrate and the Royer Pd catalyst (3% w/w Pd on polyethylenimine). We chose to screen the common K-12-derived strains E. coli MG1655, BW25113, MC4100 and TOP10 and the B-strain E. coli BL21(DE3). The cells were initially grown in minimal media supplemented with casamino acids (M9CA media) containing 0.5% w/v glucose under aerobic conditions at 37 °C in a shaking incubator. The substrate and catalyst were added at mid-log phase of growth (optical density at 600 nm (OD600) 0.5–0.6) before the cultures were sparged with N2 to create an anaerobic environment (Fig. 2a,b). Reactions were sealed and incubated at 37 °C at 220 rpm for 21 h before organic extraction and analysis by 1H nuclear magnetic resonance (NMR) spectroscopy. To our surprise, E. coli TOP10 was comparable to the engineered H2 producer E. coli DD-2, producing dihydrocaffeic acid (DHCA) in 64% and 67% yield, respectively. However, E. coli BW25113, MG1655 and MC4100 substantially outperformed E. coli DD-2, affording DHCA in 90%, 91% and 94% yield, respectively.
a, The model biohydrogenation reaction of caffeic acid to DHCA. b, The hyc- and FHL-encoded H2 pathway in Escherichia coli. c, Screen of native E. coli strains for biohydrogenation activity and confirmation of hyc-encoded H2 synthesis using pflB gene knockouts. d, TEM images of E. coli cells under biohydrogenation reaction conditions. Imaging was performed on a single representative replicate at the conclusion of the hydrogenation reaction. e, H2 detection by headspace analysis of E. coli BW25113 and E. coli BW25113∆pflB cultures by GC-TCD. f, Plate-count assay of Royer Pd toxicity to E. coli MG1655. g, Biohydrogenation activity of other microorganisms. The product concentrations were determined by 1H NMR relative to an internal standard of 1,3,5-trimethoxybenzene. The data shown are an average of three independent experiments to one standard deviation.
Product formation was abolished in the control strain E. coli BW25113∆pflB lacking the active subunit of Pfl—and so cannot endogenously produce formate—and recapitulated upon the addition of exogenous formate (Fig. 2c,e). Biohydrogenation was also abolished in the absence of the Pd catalyst and was reduced to 41% yield in E. coli BW25113∆hycE, which lacks the hydrogen-forming subunit of the formate hydrogenlyase (FHL) complex (Supplementary Fig. 10). The latter observation is probably due to H2 production from formate by Hyd-2 and Hyd-4 in the absence of Hyd-320,21,22. Together, these experiments confirm H2 generation from native hyc-encoded pathways in E. coli and that caffeic acid reduction occurs via a Pd-catalysed hydrogenation as opposed to biocatalysis by a native alkene reductase or transfer hydrogenation from metabolic formate. Hydrogen production was confirmed in E. coli MG1655, MC4100, BW25113 and TOP10 by gas chromatography with a thermal conductivity detector (GC-TCD) headspace analysis (Supplementary Fig. 11). Interestingly, E. coli BL21(DE3) was an inefficient biohydrogenation strain, producing DHCA in 5% yield. As well as being the parental strain of E. coli DD-2, E. coli BL21(DE3) also possesses mutations in fnr, a global regulator of anaerobic metabolism impacting several hundred genes23, in addition to genes essential for the transport and maturation of transition metal cofactors required for formate dehydrogenase and hydrogenase activity24.
Analysis of cells by transmission electron microscopy (TEM) revealed a distinct interaction between the Royer Pd catalyst and the cell surface, probably due to ionic interactions between the positively charged polyethylenimine support of the Royer catalyst and the negatively charged phospholipid bilayer of E. coli (Fig. 2d and Supplementary Figs. 39–48). Intriguingly, this catalyst–microorganism interaction was not observed when E. coli was cultured in the presence of the Royer Pd catalyst under aerobic conditions (Supplementary Figs. 49–51). Although the exact reason(s) for this are currently unclear, this suggests that the protein content of the outer membrane also plays a role in facilitating catalyst–microorganism interactions and catalyst activity in vivo. Plate-count analysis of cells cultured in the presence and absence of the Royer Pd catalyst showed no difference in optical density (OD600) or colony forming unit (cfu ml−1) values, indicating that the catalyst–microorganism interaction does not impact cell growth or viability (Fig. 2f). E. coli and related strains were also found to be the optimal laboratory microorganisms for this chemistry (Fig. 2g). Saccharomyces cerevisiae is not known to produce H2 under fermentation conditions and did not yield any DHCA under optimized biohydrogenation reaction conditions. Similarly, Pseudomonas putida, an obligate aerobe, was unable to produce H2 via fermentation and did not yield DHCA under the reaction conditions. Bacillus spp. have been reported to produce H2 but Bacillus subtilis did not produce H2 by GC-TCD analysis nor yield any DHCA under the reaction conditions. The Gram-negative facultative anaerobe Shewanella oneidensis is known to exhibit robust H2 production via metal reduction and fermentative pathways. However, minimal H2 was detected under optimized reaction conditions, resulting in 1.6% yield of DHCA. Intriguingly, no H2 was detected from cultures of Vibrio natriegens yet DHCA was produced in 2% yield. Upon further investigation, V. natriegens was found to possess a H2-evolving hydrogenase (WP_065301636), which may enable low-level H2 production and biohydrogenation. The highest conversions to DHCA (>99%) were obtained by Citrobacter freundii—the closest phylogenetic relative to E. coli, which also possesses the hyc operon—and attributed to faster induction of H2 biosynthesis in this species (Supplementary Figs. 28–34).
The biohydrogenation reaction was also found to reduce a range of alkene-containing substrates (Fig. 3, 1–13). These included ortho-, meta- and para- coumaric acids in 24-86% yield; ortho-, meta- and para-vinyl benzoic acids in 81–100% yield; di-acid O-methylcatechol and pyridine-substituted cinnamic acid derivatives in 79%, 67% and 80%, respectively; and aliphatic polyene substrates cis,cis-muconic acid and docosahexaenoic acid (DHA) generating the nylon-6,6 precursor adipic acid and the natural product behenic acid in 76% and 42% yield, respectively. Biohydrogenation of (4′-hydroxyphenyl)-but-3-en-2-one yielded the high-value aroma compound raspberry ketone in 87% yield. Of note, E. coli BW25113∆fadE—deficient in the first step of the fatty acid assimilation pathway—was required for biohydrogenation of DHA to avoid competing catabolism of the substrate and product via β-oxidation pathways. The microbial hydrogenation of caffeic and cinnamic acids were also performed in a large-scale bacterial culture (0.5 g scale) and isolated in 91% and 68% yield (Fig. 2d and Supplementary Table 7). Yields could also be improved by enhancing flux from pyruvate to H2 through the hyc pathway and overexpressing the transcriptional activator fhlA. These engineered strains are denoted as E. coli BW25113∆frdC∆ldhA∆fdnG∆ppc∆narG∆mgsA-hycAΩkanR (XH2-1), which has reduced H2 uptake and produces less 1,2 propanediol, lactate, succinate and CO2 when grown anaerobically on glycerol25, and E. coli BW25113∆hyaB∆hybC∆hycA∆fdoG_pCA24N-fhlA (XH2-2), with reduced H2 uptake and increased flux of anaerobically produced formate through the H2-producing FHL complex when grown on glucose26. Pleasingly, H2 production was increased in both XH2-1 and XH2-2 strains by total headspace volume and GC-TCD analysis relative to MG1655 (Supplementary Fig. 11). Quantification of hydrogen production showed comparable levels among wildtype strains (72–80 µmol ml−1), excluding E. coli BL21(DE3) (0 µmol ml−1), and significantly elevated production in XH2-1 (86 µmol ml−1, P = 0.02) and XH2-2 (95 µmol ml−1, P = 0.002) relative to the baseline E. coli MG1655 strain, as anticipated (Supplementary Fig. 13). Notably, hydrogen production in E. coli MG1655 was also significantly higher than in the engineered strain E. coli DD-2 (P = 0.03), underscoring the potential of native metabolic pathways for biocompatible chemistry. When cultured under optimized biohydrogenation reaction conditions the yield of DHCA increased from 90% (BW25113; parent strain) to >98% in both XH2-1 and XH2-2 after 21 h (37 °C, 220 rpm). Under these optimized conditions, the catalyst loading could also be reduced to 12 mol% Pd using E. coli MG1655 and to 8 mol% Pd for E. coli XH2-1/XH2-2 strains (Supplementary Figs. 1–9 and Supplementary Table 5).
a, E. coli XH2-1 and XH2-2 genotypes. b, GC-TCD chromatograms of H2 from native and engineered strains. c, Biocompatible hydrogenation using enhanced H2-producing strains. d, The substrate scope of biocompatible hydrogenation reaction using E. coli MG1655 and E. coli XH2-2 strains. e, Enzymatically depolymerized bread waste as a carbon source for H2 biosynthesis and biocompatible alkene hydrogenation. The product concentrations were determined by 1H NMR relative to an internal standard of 1,3,5-trimethoxybenzene. The data shown are an average of three independent experiments to one standard deviation. [a] represents the isolated yield from 0.5 g scale reaction. CA, caffeic acid. Credit: loaf of bread in e, PublicDomainPictures.net under a Creative Commons licence CC0 1.0.
Next, we moved on to examine the use of alternative carbohydrate feedstocks for H2 generation and biohydrogenation. We decided to focus on waste bread, which is an abundant source of carbohydrate waste worldwide. In the UK alone, 900,000 tonnes of waste bread are generated every year27. This waste is currently disposed in landfill or incinerated, generating 80–560 ktonne year−1 CO2 emissions. We generated a modified M9CA + bread growth media using the soluble carbohydrate fraction from an enzymatically depolymerized naan bread using an amylogluocosidase from Aspergillus niger. The resulting D-glucose concentration and purity was determined by a 3,5-dinitrosalicylic acid colorimetric assay and by 1H NMR of waste supernatant samples in D2O and was found to be 101 g l−1 (Supplementary Table 3). The growth of E. coli MG1655 was unchanged in M9CA + bread media compared with media using commercial glucose, and H2 concentration measured by headspace analysis was not substantially altered (Supplementary Fig. 11). When transferred to our optimized biohydrogenation reaction conditions (Fig. 2a and Supplementary Section 4), the yield of DHCA increased from 91% to 97% (4.55 and 4.85 mM, respectively) in E. coli MG1655 using the bread hydrolysate and achieved high conversions of >99% using E. coli XH2-1/XH2-2. This demonstrates a unique feature of biocompatible chemistry to enable the generation of synthetic reagents for sustainable chemical synthesis from renewable and waste feedstocks via microbial metabolism.
Having demonstrated the use of native and engineered H2 pathways in E. coli for biocompatible hydrogenation using exogenous alkenes and waste feedstocks, we next moved on to metabolically engineer these strains to generate alkene substrates in situ. We envisioned that cometabolic generation of an alkene and H2 could proceed simultaneously and then be intercepted at the cell membrane by a biocompatible Pd catalyst (Fig. 4a). This concept of multiplexed microbial metabolism for both substrate and reagent biosynthesis in biocompatible chemistry is completely unexplored but would enable both the generation of hydrogenation substrates and reagents from D-glucose in a single cell, followed by the generation of novel products in vivo. This is particularly salient for alkene-containing metabolites such as terpenes and flavonoids of interest to the fragrance and biofuel communities where alkene reductase enzymes are not known, necessitating their semi-synthetic production via fermentation, extraction and subsequent chemical manipulation in vitro.
a, Overall reaction scheme. b, Multiplexed metabolism from phosphoenolpyruvate enables simultaneous H2 biosynthesis from formate and cinnamic/coumaric acid biosynthesis from Phe/Tyr via heterologous expression of an ammonia-lyase. c, Testing the activity of each pathway and combined pathways in the absence of Pd catalyst. d, GC-FID chromatograms of metabolite products from multiplexed strains cultured in the presence and absence of Royer Pd catalyst. Stars refer to an internal standard of 1,3,5-trimethoxybenzene. e, HCA production upon delayed Pd catalyst addition in H2 and cinnamic acid coproducing strains. f, In vivo biohydrogenation of metabolically generated ccMA from D-glucose and bread waste carbon feedstocks. The data shown are an average of three independent experiments to one standard deviation.
To test this hypothesis, we chose to generate trans-cinnamic acid (tCA) and H2 in E. coli NST74, generating hydrocinnamic acid (HCA) after metabolite biohydrogenation. E. coli NST74 is a metabolically evolved L-Phe overproducer (aroH367, tyrR366, tna-2, lacY5, aroF394(fbr), malT384, pheA101(fbr), pheO352, aroG397(fbr))28, enabling increased flux from D-glucose via phosphoenolpyruvate into shikimic acid and phenylalanine biosynthesis. The conversion of L-Phe then occurs via a heterologous 4-methylideneimidazole-5-one-dependent phenylalanine ammonia-lyase PAL2 from Arabidopsis thaliana28. The PAL2 gene was assembled on a pTrc99a plasmid (ptCA), overexpressed in E. coli NST74 and confirmed to confer increased L-Phe and tCA production (pathway 1, Fig. 4b) relative to E. coli MG1655(DE3)_ptCA and E. coli MG1655(DE3)_pET22b(+) empty plasmid control under aerobic and anaerobic conditions. Hydrogen gas biosynthesis (pathway 2, Fig. 4b) was also confirmed in these engineered strains and in the absence of the required pathway genes. To test the metabolic biohydrogenation of tCA in vivo, we grew cultures of E. coli NST74_ptCA to mid-log phase (OD600 0.5–0.6) in MM1 media containing D-glucose (optimal for Phe overproduction by E. coli NST74) (Supplementary Section 2.2), induced the tCA pathway with isopropyl ß-D-1-thiogalactopyranoside (IPTG), added Pd catalysts and then sparged with N2 to remove oxygen and induce H2 biosynthesis. The cells were then incubated under anaerobic conditions for 90 h before being extracted and analysed by 1H NMR spectroscopy. Titres of tCA were decreased in these multiplexed strains, probably due to increased metabolic burden, and although 57% conversion to HCA could be detected, the overall titres were reduced to 11 mg l−1. Recapitulating the pathway into E. coli MG1655(DE3) and using L-Phe as the pathway substrate at 5 mM increased overall titres to ~2.5 mM, but conversion from tCA to HCA was reduced to 27%. Extensive optimization of the reaction conditions focusing on host strain engineering (including deregulating FeS biogenesis in E. coli BW25113∆iscR strains), media composition, reaction time, addition of pyruvate/formate, IPTG concentration and temperature resulted in varying tCA metabolite levels from 0.2 to 2.5 mM but did not increase the mean conversion of tCA to HCA beyond 28%.
However, delayed addition of the Royer Pd catalyst by 24 h to enable optimum cell growth, and the accumulation of both tCA and H2 in the reaction media and headspace increased product conversions to 47%. Catalyst addition after 72 h maintained production titres at 136 mg l−1 and further increased HCA conversions to 82%. Replicating these experiments using enhanced H2-producing strains E. coli XH2-1 and XH2-2 further increased conversion from L-Phe to HCA to 100% (~300 mg l−1) (Fig. 4d,e and Supplementary Fig. 15). Similarly, by applying these findings to E. coli NST74_ptCA and using formate to induce H2 production after incubation under aerobic conditions29 required for tCA biosynthesis from D-glucose, we were able to achieve 96% conversion to HCA at 137 mg l−1 scale after 68 h. Finally, production of both tCA and H2 from glucose was achieved in E.coli MG1655(DE3)_pS3_pY3-pheA-specR_ptCA under both aerobic and anaerobic conditions, with an initial 48 h tCA production period followed by addition of fresh media and 24 h incubation anaerobically with Royer Pd. Under these conditions, HCA conversion reached 47% with a total titre of 97 mg l−1 after 72 h (Supplementary Table 6).
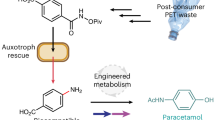
This strategy was further extended to the metabolic biohydrogenation of para-coumaric acid (pCA) to generate desaminotyrosine (DAT) and ccMA to generate adipic acid. Here, pCA was generated in E. coli MG1655(DE3) from L-Tyr by the enzyme tyrosine ammonia-lyase from Rhodotorula glutinis30. Under optimized conditions, pCA biosynthesis was induced in E. coli M6155(DE3)_ppCA under anaerobic conditions in M9CA media followed by the addition of the Royer Pd catalyst after 144 h and resulted in 154 mg l−1 of DAT (68% yield) (Supplementary Fig. 16). Further addition of two plasmids, pS3 and pY331, encoding the shikimic acid and tyrosine production pathway genes aroABCDEG*L, tyrAB, ppsA and tktA enabled elevated biosynthesis of pCA from D-glucose in E. coli (827 mg l−1) after 168 h incubation under aerobic conditions (Supplementary Figs. 17 and 18). Culturing this strain in MM1 media under anaerobic conditions in the presence of the Royer Pd catalyst enabled cometabolic pCA and H2 production and in vivo biohydrogenation yielding DAT in 226 mg l−1 and 71% yield (Supplementary Fig. 19). We next sought to expand the range of alkane products accessible by in vivo metabolite biohydrogenation to include the valuable industrial small molecule adipic acid. To achieve this, a metabolic shunt was introduced to convert 3-dehydroshikimate (DHS) to ccMA via protocatechuate and catechol32 (Fig. 4b). The plasmid pccMA, encoding the necessary DHS dehydratase, protocatechuate decarboxylase and prenyl flavin transferase (for protocatechuate decarboxylase cofactor generation), and catechol 1,2-dioxygenase was introduced into E. coli MG1655(DE3) alongside pS3 to increase DHS precursor flux. The metabolic generation of ccMA was confirmed via high-performance liquid chromatography (HPLC) analysis of aerobic cultures grown over 72 h (Supplementary Fig. 20). Extending this with an additional 168 h of anaerobic growth resulted in the cometabolic generation of H2 and conversion to adipic acid (346 mg l−1; 85% yield relative to no-catalyst controls) in the presence of the Royer Pd catalyst after 48 h. Replacing virgin glucose with waste bread-derived glucose further increased ccMA and adipic acid titres via metabolic biohydrogenation, reaching 626 mg l−1 and a >99% yield relative to mock-treated reactions (Fig. 4f). Notably, employing an H2-overproducing strain carrying pccMA led to notably lower ccMA titres (Supplementary Fig. 21), underscoring the advantage of using a non-engineered chassis for cometabolic production of both substrate and reagent for in vivo biocompatible hydrogenation.
Finally, we quantified the life cycle global warming potential (GWP) benefits of defossilizing hydrogenation product generation (Fig. 5, Supplementary Fig. 37 and Supplementary Tables 9–14). Simulating feedstocks, substrates, products and hydrogenation reactions using Aspen Plus V12.1 with the Peng-Robinson thermodynamic package and yield reactor models33 (Supplementary Table 10) revealed approximately 10% lower greenhouse gas (GHG) emissions for biohydrogenation compared to electrolytically produced hydrogen (Supplementary Fig. 37a). Greater GHG reductions were observed when evaluating the entire integrated process, including biosynthesis of substrates, reactants and hydrogenation, with GHG-negative outcomes possible for highly exothermic hydrogenation reactions (Supplementary Fig. 37c). We further assessed the GWP of DHCA synthesis from caffeic acid hydrogenation in E. coli MG1655 and E. coli XH2-1/XH2-2 using glucose and waste bread as feedstocks. Although minimal GWP differences were observed between strains, accounting for feedstock life cycle carbon footprint analysis revealed a >135% GWP improvement for both when using waste bread (Fig. 5a). For the model metabolite cinnamic acid, consolidating substrate production and hydrogenation into a one-pot process eliminated the endothermic heat requirement of the two-pot bioreduction and avoided fossil resource consumption for petrochemical and electrolytic hydrogenation routes, resulting in 3-fold and 20-fold reductions in GWP, respectively (Fig. 5b). These sustainability benefits extended beyond model substrates: the one-pot production of adipic acid via biohydrogenation using waste bread (Fig. 4f) yielded GWP savings (Fig. 5c), representing a carbon-negative route enabled by the valorization of waste feedstocks and avoidance of landfilling or incineration34,35.
a, GWP of DHCA synthesis from caffeic acid hydrogenation in E. coli MG1655 and E. coli XH2-1/XH2-2 using glucose and bread waste as feedstocks. b, GWP of one-pot HCA synthesis compared with two-pot and fossil-based synthesis from cinnamic acid. c, GWP comparisons of in vivo alkene metabolite biohydrogenation using D-glucose and waste bread feedstocks.
These findings establish a strategy for coupling cogenerated metabolites via biocompatible transition metal catalysis in living cells to form novel end products. The ubiquity of H2 as a chemical reagent and reductant now enables in vivo hydrogenation of a broad range of cellular metabolites, offering routes to bio-based products through the integration of multiplexed metabolism and biocompatible chemistry. The energetic advantages of a one-pot system, combined with the ability of biological platforms to utilize waste-derived feedstocks to substantially lower the GWP of synthesis, will further appeal to researchers pursuing sustainable chemical manufacturing. Ongoing work aims to expand this approach to a broader array of alkene-containing metabolites across diverse microbial hosts, to engineer strains that overcome thiol-dependent catalyst deactivation pathways and to develop a systems-level understanding that can optimize cometabolic flux for preparative-scale synthesis.
Overall, this study demonstrates the use of native and engineered metabolic pathways in living bacteria to generate both H2(g) and alkene substrates for biocompatible hydrogenation reactions. The transformation occurs at the cell membrane using a biocompatible Pd catalyst under mild aqueous conditions (pH 7, 30–37 °C) and is applicable to a variety of alkene-containing substrates at preparative scale. Bacteria can be engineered to enhance H2 flux, boost product yields and utilize waste bread as a feedstock. By integrating H2 biosynthesis with de novo alkene pathways in E. coli, we enabled the simultaneous in vivo production of both substrate and reagent in a single cell, followed by a non-enzymatic transformation at the cell surface to yield abiotic products. Life cycle assessment confirmed a reduced GWP for this hybrid chemo-microbial process, with carbon-negative outcomes achievable using waste-derived inputs. These findings underscore the potential of biocompatible chemistry to enable the cometabolic and sustainable synthesis of new-to-nature compounds in living cells.
Methods
Culture conditions
Overnight culturing of E. coli strains was performed routinely under aerobic conditions at 37 °C, 220 rpm. E. coli strains were inoculated directly from glycerol stocks stored at −70 °C into 10 ml Luria–Bertani broth (LB) + antibiotics (where appropriate). For day culturing of E. coli strains, overnight cultures were diluted 1:100 into fresh media containing the appropriate antibiotics and routinely cultured at 37 °C, 220 rpm under aerobic conditions. Overnight cultures of standard laboratory species were inoculated with a single colony grown on solid agar, stored at 4 °C, into 10 ml broth and incubated aerobically at optimum temperature, 220 rpm for 16–20 h as required (Supplementary Table 4).
For day culturing of standard laboratory species, overnight cultures were diluted 1:100 into fresh media and cultured at optimum growth temperature (Supplementary Table 4), 220 rpm under aerobic conditions.
Hydrogenation with exogenous substrate
Day cultures in M9CA media were grown to OD600 of 0.40–0.60 then transferred to Hungate tubes (5 ml). V. natriegens day cultures were grown in M9CA with NaCl (2% (w/v)). S. cerevisiae day cultures were grown in YSM. As appropriate, Hungate tubes contained preweighed catalyst (2–20 mol%) and substrate (2–5 mM). For DD-2 reactions, IPTG (1 M, 5 µl; final concentration 1 mM) and Fe(NH4)2(SO4)2 (50 mM, 5 µl; final concentration 50 µM; Alfa Aesar) were added. For hydrogenation of DHA, 0.5% (w/v) TPSG-1000 was supplemented. The tubes were sealed with butyl rubber septa and screw caps and sparged with oxygen-free N2 (BOC) (room temperature, 15 min), using a 21-gauge, 120-mm needle as the inlet and a 25-gauge, 16-mm needle as the outlet. Reactions were then incubated horizontally (37 °C, 220 rpm, 21–48 h). After the stated reaction time, E. coli samples were uncapped and acidified with HCl (aqueous concentration, 50 µl) and extracted with EtOAc (2× 10 ml). The organic phases were dried on a rotary evaporator (bath temperature 34 °C, 250 rpm, 105 mbar) until the sample was dry. The resulting solids were resolubilized in DMSO-d6 containing 1,3,5-trimethoxybenzene (TMB) internal standard (1.8 mM) and analysed by 1H NMR spectroscopy. Industrially relevant species were analysed by HPLC. The samples (100 ml) were diluted 1:10 with 55 mM HCl, vortexed for 1 min and centrifuged to pellet cells before HPLC analysis. Analyte concentrations for caffeic acid (4.66 min) and DHCA (4.13 min) were calculated by comparison to a standard curve of known concentrations. For catalyst reuse experiments, catalyst from spent reactions was collected, washed with distilled H2O, and dried at 110 °C in a vacuum furnace for 48 h before reuse.
For hydrogenations with DHA, a FAME method was used to determine DHA and behenic acid concentration. The samples (500 µl) were mixed with tridecanoic acid internal standard (2 mM) and boiled to evaporate overnight at 95 °C. The dried samples were suspended in MeOH containing 8% HCl (500 µl) and incubated at 50 °C overnight. The derivatized fatty acids were extracted with 250 µl hexane, dried on sodium sulfate and analysed by gas chromatography coupled with a flame ionization detector (GC-FID).
Cometabolic reactions, tCA
Day cultures of E. coli MG1655(DE3)_ptCA or E. coli NST74_ptCA were grown to OD600 of ~0.5. IPTG (1 M; 1 mM final concentration) and aqueous L-Phe (125 mM; 5 mM final concentration) were added and swirled for 20 s, and the cells aliquoted to Hungate tubes (5 ml) containing Royer 3% Pd catalyst (10.7 mg) as appropriate. For production of tCA from glucose in E. coli NST74_ptCA, no L-Phe was added and the total weight/volume of glucose was adjusted to 2%. The tubes were sealed with butyl rubber septa and screw caps and sparged for 15 min with oxygen-free N2, then incubated horizontally (32 °C, 300 rpm, 90 h). After 18–20 h, the pH of samples was adjusted using NaOH (1 M aqueous) injected through the septa with a 27-gauge, 16-mm needle, returning the media to a green colour (pH ~7). pH adjustments were thereafter made as needed when the colour of the media became yellow (pH ~5).
For time course experiments, reactions were prepared as above without Royer Pd catalyst. The catalyst was added at 24 h intervals as follows: a suspension of 0.33 ml of Royer Pd catalyst in glycerol (35 mg ml−1) was injected using an 18-gauge, 1.5-inch needle (to not let gas enter the syringe). The samples were then allowed to react with the catalyst for an additional 24 h or until the end of the time course.
Reactions were stopped by addition of HCl (aqueous concentration, 50 µl) and extracted with hexane containing 1,3,5-trimethoxybenzene as an internal standard (2 mM; 2× half-reaction volume) and analysed by GC-FID.
Cometabolic reactions, pCA
Day cultures of E. coli MG1655(DE3)ppCA containing L-Tyr (5 mM) were grown to OD600 of ~0.5. IPTG (0.5 mM) was added, mixed well, and culture aliquoted (5 ml) to Hungate tubes. The samples were sealed and sparged for 15 min with oxygen-free N2 before incubation horizontally (30 °C, 300 rpm, 24–168 h). After 18–20 h, the pH of samples was adjusted using 1 M NaOH injected through the septa with a 27-gauge, 16-mm needle, returning the media to a green colour (pH ~7). pH adjustments were thereafter made as needed when the colour of the media became yellow (pH ~5). Royer Pd catalyst was added as above 24 h before the end of the reaction.
pCA and DAT concentrations were analysed by HPLC. The samples (500 ml) were diluted 1:1 with HCl (100 mM), vortexed and centrifuged to pellet cells before HPLC analysis. Analyte concentrations for pCA (8.39 min) and DAT (7.40 min) were calculated by comparison with a standard curve of known concentrations.
Cometabolic reactions, pccMA
Day cultures of E. coli MG1655(DE3)_pS3_pccMA were grown in M9 a 37 °C to OD600 of ~0.5 and induced with 0.4 mM IPTG. After 24 h, the cultures were pH adjusted using NaOH and supplemented with a further 0.5% glucose with aerobic growth continuing for a further 24 h. A total of 2.5 ml of culture was then transferred to Hungate tubes in triplicate, 2.5 ml of M9CA media added and then sparged with oxygen-free nitrogen for 15 min before incubation horizontally (37 °C, 220 rpm, 168 h). After anaerobic growth, Royer catalyst (20 mol%) was added as a suspension in glycerol using an 18-gauge, 1.5-inch needle and incubated horizontally for a further 24 h. The reactions with bread waste glucose were conducted as above; however, 3 ml of aerobic culture and 0.75 ml of a 5× M9CA stock were used and substituting the appropriate amount of bread waste hydrolysate instead of commercial glucose in both M9 and M9CA media.
ccMA and adipic acid concentrations were determined by HPLC. The 150 ml samples were diluted 1:3 in acidified caffeine (0.1% trifluoroacetic acid (TFA), 55 mM HCl, 0.01 g l−1 caffeine), mixed thoroughly and incubated at room temperature for 20–30 min. Cell debris was removed by centrifugation (21,000 relative centrifugal force, 10 min) and the supernatant boiled for 1 h. Concentrations of ccMA and adipic acid were calculated using caffeine as an internal standard, against a standard curve of known concentrations.
Scale up, 50 and 100 ml volume
A larger scale reaction with tCA as a substrate was performed using 50 ml of M9CA media in a 125 ml Erlenmeyer flask. An overnight culture of E. coli MG1655 was diluted 1:100 into the media and grown to an OD600 of 0.5 aerobically. Once the culture reached this point, tCA (5 mM; 1 ml of 250 mM stock) and Royer Pd catalyst (107 mg, 12 mol%) were added, and the flask sealed with a rubber septum. The reaction was sparged for 20 min with N2 gas using a 21-gauge, 120-mm needle as the inlet and a 25-gauge, 16-mm needle as the outlet. The culture was then incubated horizontally (37 °C, 21 h). An aliquot (5 ml) of the final reaction was processed as in 0 to determine the conversion to HCA.
A total of 100 ml volume scale up was performed as above but with 100 ml of culture in a baffled 500 ml Erlenmeyer flask sealed with a rubber septum and screw lid, with 2 ml of 0.25 M tCA as a substrate and 214 mg of Royer Pd catalyst.
Scale up and purification, 500 ml volume
The scale up and purification using both caffeic and cinnamic acid was performed at 500 ml reactions. In a typical experiment a 500 ml day culture of E. coli MG1655 in M9CA media in a 1 litre Erlenmeyer flask was grown to OD600 of ~0.5. Then, tCA (5 mM) or CAF (5 mM) and Royer Pd catalyst (11.5 mol% and 12 mol%, respectively) were added to the culture, and the reaction mixture was sealed with a rubber septum. The reactions were then sparked for 25 min using oxygen-free N2 using a 12-inch needle as the inlet and a 25-gauge, 5/8-inch needle as the outlet. Reactions were then incubated at 37 °C, 220 rpm for 21 h.
Extraction protocol for the reactions consisted of acidifying each tube with conc. HCl (5 mL) and the product extracted in EtOAc (2–4 volumes). The organic layers were collected and dried with sodium surface (Sigma-Aldrich) before being dried on a rotary evaporator (bath temperature 34 °C, 120 rpm, 130 mbar). The sodium sulfate layer was subsequently washed with an additional volume EtOAc and dried in the same flask. A separate 20 ml aliquot of each reaction was also dried and used for NMR quantification of the crude product. The dry layers were resuspended in a minimal (10–30 ml) amount of EtOAc and dried on to 5× the expected product weight of Celite before being dry-loaded on to a silica gel column and purified by flash column chromatography. A gradient of hexane: EtOAc: 1% AcOH was used, beginning at 97% hexane and ending in 99% EtOAc: 1% AcOH to flush the column. Fractions were collected and analysed by thin-layer chromatography compared with commercial standards of the products. Product-containing fractions were combined, and the solvent removed on a rotary evaporator to give dry products. The purity of the final products was confirmed by 1H NMR quantification of a sample using TMB as an internal standard.
Headspace gas quantification
For quantification of H2 production from E. coli strains, presparged cultures at mid-log phase in M9CA media were transferred in 5 ml aliquots to N2-flushed 20 ml headspace vials (Thermo Fisher Scientific) and sealed with crimp caps fitted with PTFE septa (Thermo Fisher Scientific). The cultures were then incubated horizontally at 37 °C, 220 rpm for 24–48 h to allow H2 to accumulate before analysis.
For the preparation of H2 standards, known quantities of H2 (0.25, 0.5, 0.75 and 1 mmol) were generated in sealed headspace vials by reaction of 2 M HCl with an excess of zinc powder (Sigma-Aldrich) and copper (II) sulfate (Thermo Fisher Scientific) at room temperature until completion. 5 ml H2O was added to each vial with a 27-guage needle immediately before analysis to mimic the conditions of a culture headspace sample.
Standards and headspace gas from E. coli cultures was analysed by GC-TCD as in Supplementary Section 1.6 above.
Bread waste media
Bread waste was depolymerized enzymatically according to published protocols33. Briefly, stale naan (100 g; Gousto) was torn into small pieces and suspended in MQ water (400 ml) before the pH was adjusted to 4.3 with HCl. The suspension was autoclaved (121 °C, 15 min) and allowed to cool on a room temperature orbital shaker until the temperature of the suspension reached 55 °C. Lyophilized amyloglucosidase from Aspergillus niger (1 mg g−1 bread waste; Sigma-Aldrich) was added before further incubation (room temperature, 48 h). The resulting solution was pelleted (5 min, 4,676g) and the supernatant filtered through 0.45 mM and 0.22 mM sterile syringe filters before being stored at room temperature.
Glucose concentrations were determined by assay with 3,5-dinitrosalicylic acid34. In brief, diluted hydrolysate (500 ml, 1/500) was added to 3,5-dinitrosalicylic acid (DNS) reagent (500 µl; 1% 3,5-dinitrosalicylic acid, 1% NaOH, 0.05% Na2SO3) and incubated at 90 °C for 15 min. Reactions were quenched with KNaC4H4O6·4H2O (40%; 167 µl) and cooled in a room temperature water bath. The absorbance of the sample was measured at 575 nm with a DeNovix DS11 spectrophotometer. Concentrations in grams per litre were determined by comparison to an external standard curve (R2 = 0.9971).
NMR
1H NMR spectra were recorded on a Bruker ava400, ava500 or ava600 NMR spectrometer at 298 K. Proton chemical shifts are expressed in parts per million (ppm, δ scale) and referenced to residual protium in the NMR solvent (DMSO-d6, 2.49 ppm). Coupling constants, J, are measured to the nearest 0.1 Hz and are presented as observed. The data are represented as: chemical shift, integration, multiplicity (s is singlet, d is doublet, t is triplet, q is quartet, dd is doublet of doublets, m is multiplet and/or multiple resonances), coupling constant (J) in Hertz. NMR solvents were used as purchased from commercial suppliers. For all quantitative NMR measurements, TMB was used as an internal standard. Spectra were analysed using the Mestrenova software package version 14.3.0-30573.
Gas chromatography
Gas chromatography analysis of tCA and HCA was carried out on a Shimadzu 2010-Pro system equipped with an Rxi-5HT or Rxi-5MS capillary column (30 m, ID 0.32 mm, FT 0.1–5 μM) with a flame ionization detector using hydrogen as a carrier gas. The samples were injected in split mode using 1 ml of sample at a split ratio of 1:15. Analyte concentrations were determined by comparison to a seven-point external calibration curve, calculated by LabSolutions Lite software version 5.93 (R2 = minimum 0.997). The gas chromatography programme was as follows: flow control mode: linear velocity, column flow: 3.9 mL min−1, oven programme: 50 °C, hold for 0.5 min, ramp 26 °C min−1 to 200 °C.
Headspace gas measurement was carried out on a Shimazdu GC 2030 equipped with a ShinCarbon packed column (column length 2 m; ID 0.53 mm) (Restek) and thermal conductivity detector using nitrogen as a carrier gas.
HPLC
HPLC analysis was carried out on a Thermo Fisher Vanquish HPLC system equipped with a Hypersil X Gold C18 column. The samples were held in a sample oven at 30 °C before injection of 15 μl of sample onto the column. For ccMA and adipic acid analysis, separation was performed with a gradient from 5% to 10.5% MeCN (0.1% v/v TFA) in H2O (0.1% v/v TFA), with peak areas collected at 260 nm (ccMA) and 206 nm (adipic acid, caffeine). For all other analytes the mobile phase consisted of an isocratic 85:15 H2O (0.1% trifluoroacetic acid): MeCN (0.1% trifluoroacetic acid). Analytes were detected at 275 nm with a diode array detector, and analyte concentrations were calculated compared with a seven-point external standard curve.
TOC summary
Native microbial H2 pathways can be coupled with engineered alkene biosynthesis and membrane-bound Pd catalysis to enable biocompatible hydrogenation of metabolic intermediates in living bacteria. This hybrid chemo-microbial platform supports the carbon-negative synthesis of industrial chemicals from waste-derived feedstocks, advancing sustainable manufacturing via transition metal catalysis in whole-cell systems.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data supporting the findings of this study are available from the Article and its Supplementary Information. Source data are provided with this paper.
References
Akhlaghi, N. & Najafpour-Darzi, G. A comprehensive review on biological hydrogen production. Int. J. Hydrogen Energy 45, 22492–22512 (2020).
Zheng, L. & Dean, D. R. Catalytic formation of a nitrogenase iron–sulfur cluster. J. Biol. Chem. 269, 18723–18726 (1994).
Ohmura, N., Sasaki, K., Matsumoto, N. & Saiki, H. Anaerobic respiration using Fe3+, S0, and H2 in the chemolithoautotrophic bacterium Acidithiobacillus ferrooxidans. J. Bacteriol. 184, 2081–2087 (2002).
Burgdorf, T. et al. The soluble NAD+-reducing [NiFe]-hydrogenase from Ralstonia eutropha H16 consists of six subunits and can be specifically activated by NADPH. J. Bacteriol. 187, 3122–3132 (2005).
Köpke, M. et al. Clostridium ljungdahlii represents a microbial production platform based on syngas. Proc. Natl Acad. Sci. USA 107, 13087–13092 (2010).
McInerney, M. J., Bryant, M. P., Hespell, R. B. & Costerton, J. W. Syntrophomonas wolfei gen. nov. sp. nov., an anaerobic, syntrophic, fatty acid-oxidizing bacterium. Appl. Environ. Microbiol. 41, 1029–1039 (1981).
Sawers, R. G., Ballantine, S. P. & Boxer, D. H. Differential expression of hydrogenase isoenzymes in Escherichia coli K-12: evidence for a third isoenzyme. J. Bacteriol. 164, 1324–1331 (1985).
Knappe, J. & Blaschkowski, H. P. Pyruvate formate-lyase from Escherischia coli and its activation system. Methods Enzymol. 41, 508–518 (1975).
Rossmann, R., Sawers, G. & Böck, A. Mechanism of regulation of the formate-hydrogenlyase pathway by oxygen, nitrate, and pH: definition of the formate regulon. Mol. Microbiol. 5, 2807–2814 (1991).
McDowall, J. S. et al. Bacterial formate hydrogenlyase complex. Proc. Natl Acad. Sci. USA 111, E3948–E3956 (2014).
Rambhujun, N. et al. Renewable hydrogen for the chemical industry. MRS Energy Sustain. 7, 33 (2020).
International Energy Agency. Global Hydrogen Review 2023 (IEA, 2023).
Carey, J. S., Laffan, D., Thomson, C. & Williams, M. T. Analysis of the reactions used for the preparation of drug candidate molecules. Org. Biomol. Chem. 4, 2337–2347 (2006).
Agyekum, E. B., Nutakor, C., Agwa, A. M. & Kamel, S. A critical review of renewable hydrogen production methods: factors affecting their scale-up and its role in future energy generation. Membranes 12, 173 (2022).
Zhang, X., Schwarze, M., Schomäcker, R., van de Krol, R. & Abdi, F. F. Life cycle net energy assessment of sustainable H2 production and hydrogenation of chemicals in a coupled photoelectrochemical device. Nat. Commun. 14, 991 (2023).
Toogood, H. S., Gardiner, J. M. & Scrutton, N. S. Biocatalytic reductions and chemical versatility of the old yellow enzyme family of flavoprotein oxidoreductases. ChemCatChem 2, 892–914 (2010).
Joo, J. C. et al. Alkene hydrogenation activity of enoate reductases for an environmentally benign biosynthesis of adipic acid. Chem. Sci. 8, 1406–1413 (2017).
Sirasani, G., Tong, L. & Balskus, E. P. A biocompatible alkene hydrogenation merges organic synthesis with microbial metabolism. Angew. Chem. Int. Ed. 53, 7785–7788 (2014).
Agapakis, C. M. et al. Insulation of a synthetic hydrogen metabolism circuit in bacteria. J. Biol. Eng. 4, 3–3 (2010).
Trchounian, K. & Trchounian, A. Escherichia coli hydrogen gas production from glycerol: effects of external formate. Renew. Energy 83, 345–351 (2015).
Sanchez-Torres, V. et al. Influence of Escherichia coli hydrogenases on hydrogen fermentation from glycerol. Int. J. Hydrogen Energy 38, 3905–3912 (2013).
Trchounian, K. Transcriptional control of hydrogen production during mixed carbon fermentation by hydrogenases 4 (hyf) and 3 (hyc) in Escherichia coli. Gene 506, 156–160 (2012).
Salmon, K. et al. Global gene expression profiling in Escherichia coli K12. The effects of oxygen availability and FNR. J. Biol. Chem. 278, 29837–29855 (2003).
Pinske, C., Bönn, M., Krüger, S., Lindenstrauß, U. & Sawers, R. G. Metabolic deficiences revealed in the biotechnologically important model bacterium Escherichia coli BL21(DE3). PLoS ONE 6, e22830–e22830 (2011).
Tran, K. T., Maeda, T. & Wood, T. K. Metabolic engineering of Escherichia coli to enhance hydrogen production from glycerol. Appl. Microbiol. Biotechnol. 98, 4757–4770 (2014).
Maeda, T., Sanchez-Torres, V. & Wood, T. K. Metabolic engineering to enhance bacterial hydrogen production. Microb. Biotechnol. 1, 30–39 (2008).
Kumar, V. et al. Bread waste—a potential feedstock for sustainable circular biorefineries. Bioresour. Technol. 369, 128449 (2023).
McKenna, R. & Nielsen, D. R. Styrene biosynthesis from glucose by engineered E. coli. Metab. Eng. 13, 544–554 (2011).
Metcalfe, G. D., Sargent, F. & Hippler, M. Hydrogen production in the presence of oxygen by Escherichia coli K-12. Microbiology 168, 001167 (2022).
Jones, J. A. et al. Complete biosynthesis of anthocyanins using E. coli polycultures. mBio https://doi.org/10.1128/mbio.00621-00617 (2017).
Juminaga, D. et al. Modular engineering of L-tyrosine production in Escherichia coli. Appl. Environ. Microbiol. 78, 89–98 (2012).
Niu, W., Draths, K. M. & Frost, J. W. Benzene-free synthesis of adipic acid. Biotechnol. Prog. 18, 201–211 (2002).
Sadhukhan, J. & Sen, S. A novel mathematical modelling platform for evaluation of a novel biorefinery design with green hydrogen recovery to produce renewable aviation fuel. Chem. Eng. Res. Des. 175, 358–379 (2021).
Hafyan, R.H., Sadhukhan, J., Kumar, V., Maity, S.K. & Gadkari, S. A comparative techno-economic feasibility of hydrogen production from sugarcane bagasse and bread waste. Fuel 388, 134469 (2025).
Hafyan, R. H. et al. Integrated biorefinery for bioethanol and succinic acid co-production from bread waste: Techno-economic feasibility and life cycle assessment. Energy Convers. Manag. 301, 118033 (2024).
Acknowledgements
This work was supported by a UKRI Future Leaders Fellowship (grant no. MR/S033882/1 to S.W.), EPSRC Sustainable Manufacturing Grant (grant no. EP/W019000/1), European Research Council (ERC Consolidator grant no. 101170985 to S.W.) and two grants from the High-Value Biorenewables Network (grant nos. BIV-HVB-2020/19 and POC-HVB-FOF 2023/08). M.F.M.W. acknowledges a PhD studentship (EP/T517884/1) and a Feasibility Fund grant from IBioIC. C.L.T. acknowledges a PhD studentship from the EASTBIO Doctoral Training Partnership (grant no. BB/T00875X/1). R.G. acknowledges a PhD studentship from the EASICAT EPSRC Centre for Doctoral Training. We thank V. Narisetty and R. Cox from C-Source Renewables Ltd. for providing samples of bread hydrolysate and Z. Gidden from the University of Edinburgh for providing Saccharomyces cerevisiae.
Author information
Authors and Affiliations
Contributions
The manuscript was written by S.W., M.F.M.W., C.L.T., J.F.C.S., E.C.H.T.L., Y.E., J.A.D. and J.S. All authors have given approval to the final version of the manuscript. The study was designed by S.W. and M.F.M.W. with contributions from C.L.T., J.F.C.S., E.C.H.T.L. and Y.E. Experimental work was performed and analysed by M.F.M.W., C.L.T., J.F.C.S., E.C.H.T.L. and Y.E. and supervised by S.W. Life cycle analysis was performed and analysed by J.S. Technical help was provided by J.A.D., N.W.J. and R.G. S.L., J.G. and S.W. provided project support and experimental guidance.
Corresponding author
Ethics declarations
Competing interests
S.L. is an employee of NCIMB Ltd. and J.G. is an employee of MiAlgae Ltd. Both may possess competing financial interests. The other authors declare no competing interests.
Peer review
Peer review information
Nature Chemistry thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
General materials and methods, Supplementary Figs. 1–51 and Supplementary Tables 1–16.
Source data
Source Data Fig. 2 (download ZIP )
Raw data for Fig. 2.
Source Data Fig. 3 (download ZIP )
Raw data for Fig. 3.
Source Data Fig. 4 (download ZIP )
Raw data for Fig. 4.
Source Data Fig. 5 (download ZIP )
Raw data for Fig. 5.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
White, M.F.M., Trotter, C.L., Steele, J.F.C. et al. Native H2 pathways enable biocompatible hydrogenation of metabolic alkenes in bacteria. Nat. Chem. 18, 535–543 (2026). https://doi.org/10.1038/s41557-025-02052-y
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41557-025-02052-y