Abstract
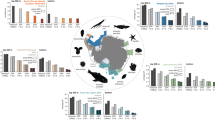
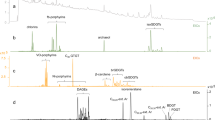
Chemical pollution has profound impacts on marine ecosystem health and services. Most investigations of the distributions of anthropogenic organic chemicals (xenobiotics) have been regionally focused, which limits our understanding of the extent of chemical pollution in the world’s largest biome. To address this gap, we mapped the presence of xenobiotics across marine ecosystems. Here we present a meta-analysis of 21 public non-targeted tandem mass spectrometry datasets, which are not restricted to a predefined set of compounds but rather capture thousands of chemicals. These datasets comprise 2,315 seawater samples, spanning coastal to open ocean environments across three ocean basins. Our analysis revealed that common pollutants such as pesticides and pharmaceuticals were predominantly detected in estuaries and coastal areas but declined with distance from shore, whereas industrial chemicals and additives, including polyalkylene glycols, phthalates and organophosphates, were widely distributed across marine ecosystems. A total of 248 annotated xenobiotic features contributed a median of 2% to the total detected peak area per sample. We observed highest median levels of xenobiotic contribution in coastal datasets (up to 20%) and lowest levels (0.5%) in open ocean datasets, which indicates that anthropogenic organic substances contribute substantially to the dissolved organic matter pool in the surface ocean.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All datasets are publicly available through the MassIVE repository (massive.ucsd.edu) under the following identifiers: Aruba 2019 (UCSD, Q Exactive): MSV000085891; Baltic Sea 2021 (Tuebingen, Q Exactive HF): MSV000089481; California Current 2017 (UCSD, Q Exactive): MSV000082082; California Current 2019 (UCSD, Q Exactive): MSV000085852; California Current 2020 (UCSD, Q Exactive): MSV000086236; California Current 2021 (UCSD, Q Exactive): MSV000090123; Caribbean Netherlands 2022 (NIOZ, Q Exactive): MSV000094015; Curaçao 2018 (UCSD, Q Exactive): MSV000083522; Curaçao 2021/22 (NIOZ, Q Exactive): MSV000094018; Gulf of Maine 2022 (Tuebingen, Q Exactive HF): MSV000090696; Hawaii 2019 (UCSD, Q Exactive): MSV000082085; Moorea 2017 (UCSD, Q Exactive): MSV000082083; Moorea 2022 (UCSD, Q Exactive): MSV000090798; Northern Pacific 2018 (UCSD, Q Exactive): MSV000083365; Northern San Diego Coast 2017/18 (UCSD, Q Exactive): MSV000082312; Northern San Diego Coast 2020 (UCSD, Q Exactive): MSV000085555 and MSV000085779; Pacific OMZ 2018 (UCSD, Q Exactive): MSV000085480; Puerto Rico 2022 (Tuebingen, Q Exactive HF): MSV000092677; South Africa 2018/19 (UCSD, Q Exactive): MSV000085843; Southern San Diego Coast 2019 (UCSD, Q Exactive): MSV000083889; Patagonia 2018 (UCSD, Q Exactive): MSV000083521. All processed data and mzmine batch files used in this study are available via Zenodo at https://doi.org/10.5281/zenodo.16687158 (ref. 51). Source data are provided with this paper.
Code availability
All code, including python scripts, is available via Github at https://github.com/Functional-Metabolomics-Lab/Marine-Xenobiotics and via Zenodo at https://doi.org/10.5281/zenodo.16687158 (ref. 51).
References
Halpern, B. S. et al. A global map of human impact on marine ecosystems. Science 319, 948–952 (2008).
Zhou, W., Li, M. & Achal, V. A comprehensive review on environmental and human health impacts of chemical pesticide usage. Emerg. Contam. 11, 100410 (2025).
Sigmund, G. et al. Addressing chemical pollution in biodiversity research. Glob. Change Biol. 29, 3240–3255 (2023).
Nguyen, M.-K. et al. A comprehensive review on ecological effects of microplastic pollution: an interaction with pollutants in the ecosystems and future perspectives. TrAC, Trends Anal. Chem. 168, 117294 (2023).
Yang, Y. et al. Which micropollutants in water environments deserve more attention globally?. Environ. Sci. Technol. 56, 13–29 (2022).
Stubbins, A., Law, K. L., Munoz, S. E., Bianchi, T. S. & Zhu, L. Plastics in the Earth system. Science 373, 51–55 (2021).
Tian, Z. et al. Suspect and nontarget screening for contaminants of emerging concern in an urban estuary. Environ. Sci. Technol. 54, 889–901 (2020).
Carve, M., Singh, N., Askeland, M., Allinson, G. & Shimeta, J. Salting-out assisted liquid–liquid extraction combined with LC–MS/MS for the simultaneous determination of seven organic UV filters in environmental water samples: method development and application. Environ. Sci. Pollut. Res. 30, 104870–104885 (2023).
Papadopoulos Lambidis, S. et al. Two-dimensional liquid chromatography tandem mass spectrometry untangles the deep metabolome of marine dissolved organic matter. Environ. Sci. Technol. 58, 19289–19304 (2024).
Patrone, J. et al. Enhancing molecular characterization of dissolved organic matter by integrative direct infusion and liquid chromatography nontargeted workflows. Environ. Sci. Technol. 58, 12454–12466 (2024).
Schramm, T., Kalinski, J.-C. J., Arini, G. S., Da Silva, R. R. & Petras, D. Uncovering the structural space of marine dissolved organic matter. Annu. Rev. Mar. Sci. 18, 165–194 (2026).
Petras, D. et al. Non-targeted tandem mass spectrometry enables the visualization of organic matter chemotype shifts in coastal seawater. Chemosphere 271, 129450 (2021).
Kalinski, J.-C. J. et al. Urban and agricultural influences on the coastal dissolved organic matter pool in the Algoa Bay estuaries. Chemosphere 355, 141782 (2024).
Kalinski, J.-C. et al. Comparability of liquid chromatography tandem mass spectrometry analysis of dissolved organic matter across laboratories. Environ. Sci. Technol. 60, 4814–4829 (2026).
Schmid, R. et al. Integrative analysis of multimodal mass spectrometry data in MZmine 3. Nat. Biotechnol. 41, 447–449 (2023).
Heuckeroth, S. et al. Reproducible mass spectrometry data processing and compound annotation in MZmine 3. Nat. Protoc. 19, 2597–2641 (2024).
Wang, M. et al. Sharing and community curation of mass spectrometry data with global natural products social molecular Networking. Nat. Biotechnol. 34, 828–837 (2016).
Sumner, L. W. et al. Proposed minimum reporting standards for chemical analysis: Chemical Analysis Working Group (CAWG) Metabolomics Standards Initiative (MSI). Metabolomics 3, 211–221 (2007).
Costanzo, S. D., Watkinson, A. J., Murby, E. J., Kolpin, D. W. & Sandstrom, M. W. Is there a risk associated with the insect repellent DEET (N,N-diethyl-m-toluamide) commonly found in aquatic environments?. Sci. Total Environ. 384, 214–220 (2007).
Celis, R., Gámiz, B., Adelino, M. A., Hermosín, M. C. & Cornejo, J. Environmental behavior of the enantiomers of the chiral fungicide metalaxyl in Mediterranean agricultural soils. Sci. Total Environ. 444, 288–297 (2013).
Küster, A. et al. Environmental risk assessment of human pharmaceuticals in the European Union: a case study with the β-blocker atenolol. Integr. Environ. Assess. Manage. 6, 514–523 (2010).
Han, B. et al. Prevalence of synthetic phenolic antioxidants in food contact materials from China and their implications for human dietary exposure through take-away food. J. Hazard. Mater. 473, 134599 (2024).
Kwon, B. & Choi, K. Occurrence of major organic UV filters in aquatic environments and their endocrine disruption potentials: a mini-review. Integr. Environ. Assess. Manage. 17, 940–950 (2021).
Weaver, J. A. et al. Hazards of diethyl phthalate (DEP) exposure: a systematic review of animal toxicology studies. Environ. Int. 145, 105848 (2020).
Su, Y. et al. Ecological risk of galaxolide and its transformation product galaxolidone: evidence from the literature and a case study in Guangzhou waterways. Environ. Sci. Processes Impacts 25, 1337–1346 (2023).
Cropper, M., Dunlop, S., Hinshaw, H. & Symeonides, C. The benefits of removing toxic chemicals from plastics. Proc. Natl Acad. Sci. USA 121, e2412714121 (2024).
ten Hietbrink, S., Materić, D., Holzinger, R., Groeskamp, S. & Niemann, H. Nanoplastic concentrations across the North Atlantic. Nature 643, 412–416 (2025).
Fogliano, C., Motta, C. M., Acloque, H., Avallone, B. & Carotenuto, R. Water contamination by delorazepam induces epigenetic defects in the embryos of the clawed frog Xenopus laevis. Sci. Total Environ. 896, 165300 (2023).
Hashemi, S. H., Kaykhaii, M., Jamali Keikha, A. & Naruie, N. Application of molecularly imprinted polymer pipette tip micro-solid phase extraction of nalidixic acid and acetaminophen from pills and seawater samples and their determination by spectrophotometry. Chem. Pap. 74, 4009–4023 (2020).
Gupta, P. & Rani, V. A comprehensive assessment of environmental implications, ecotoxicity and bioaccumulation potential of repurposed drug hydroxychloroquine: from challenges to sustainability. Rev. Environ. Contam. Toxicol. 262, 10 (2024).
Leite, M. et al. Pharmaceuticals in coastal sediments: validation of an ultra-high-performance liquid chromatography coupled with time-of-flight mass spectrometry (UHPLC-TOF-MS) methodology. Environ. Pollut. 380, 126571 (2025).
Goodyer, L. & Schofield, S. Mosquito repellents for the traveller: does picaridin provide longer protection than DEET?. J. Travel Med. 25, S10–S15 (2018).
Gigault, J. et al. Current opinion: what is a nanoplastic?. Environ. Pollut. 235, 1030–1034 (2018).
Rani-Borges, B. & Ando, R. A. How small a nanoplastic can be? a discussion on the size of this ubiquitous pollutant. Cambridge Prisms: Plast. 2, e23 (2024).
Li, C. & Tang, K. H. D. Effects of pH and Temperature on the Leaching of Di(2-Ethylhexyl) Phthalate and Di-n-butyl Phthalate from Microplastics in Simulated Marine Environment. Biointerface Res. Appl. Chem. 13, 269 (2022).
Henkel, C. et al. Photoaging enhances the leaching of di(2-ethylhexyl) phthalate and transformation products from polyvinyl chloride microplastics into aquatic environments. Commun. Chem. 7, 218 (2024).
Pinheiro, H. T. et al. Plastic pollution on the world’s coral reefs. Nature 619, 311–316 (2023).
Wang, W. et al. Bioaccumulation and potential risk of organophosphate flame retardants in coral reef fish from the Nansha Islands, South China Sea. Chemosphere 287, 132125 (2022).
Schupp, T. et al. in Reviews of Environmental Contamination and Toxicology vol. 244 (ed. De Voogt, P.) 53–111 (Springer International Publishing, 2017).
Massicotte, P., Asmala, E., Stedmon, C. & Markager, S. Global distribution of dissolved organic matter along the aquatic continuum: across rivers, lakes and oceans. Sci. Total Environ. 609, 180–191 (2017).
Butman, D. E., Wilson, H. F., Barnes, R. T., Xenopoulos, M. A. & Raymond, P. A. Increased mobilization of aged carbon to rivers by human disturbance. Nat. Geosci. 8, 112–116 (2015).
Cancelada, L. et al. Assessment of styrene-divinylbenzene polymer (PPL) solid-phase extraction and non-targeted tandem mass spectrometry for the analysis of xenobiotics in seawater. Limnol. Oceanogr. Methods 20, 89–101 (2022).
Mohammed Taha, H. et al. The NORMAN Suspect List Exchange (NORMAN-SLE): facilitating European and worldwide collaboration on suspect screening in high resolution mass spectrometry. Environ. Sci. Eur. 34, 104 (2022).
Sorokina, M., Merseburger, P., Rajan, K., Yirik, M. A. & Steinbeck, C. COCONUT online: collection of open natural products database. J. Cheminformatics 13, 2 (2021).
Scheubert, K. et al. Significance estimation for large scale metabolomics annotations by spectral matching. Nat. Commun. 8, 1494 (2017).
Dührkop, K. et al. SIRIUS 4: a rapid tool for turning tandem mass spectra into metabolite structure information. Nat. Methods 16, 299–302 (2019).
Lønborg, C. et al. A global database of dissolved organic matter (DOM) concentration measurements in coastal waters (CoastDOM v1). Earth Syst. Sci. Data 16, 1107–1119 (2024).
Khan, A. & Mathelier, A. Intervene: a tool for intersection and visualization of multiple gene or genomic region sets. BMC Bioinf. 18, 287 (2017).
Nothias, L.-F. et al. Feature-based molecular networking in the GNPS analysis environment. Nat. Methods 17, 905–908 (2020).
Pakkir Shah, A. K. et al. Statistical analysis of feature-based molecular networking results from non-targeted metabolomics data. Nat. Protoc. 20, 92–162 (2025).
Kalinski, J.-C. Widespread detection of xenobiotics in marine dissolved organic matter (manuscript data upload). Zenodo https://doi.org/10.5281/zenodo.16687158 (2025).
Acknowledgements
We thank D. Rasher and D. Yiu for facilitating sample collection in the Gulf of Maine. D.P. was supported by the Simons Foundation International through a Simons Early Career Investigator in Aquatic Microbial Ecology and Evolution Award (SFI-LS-ECIAMEE-00013858) and the German Research Foundation through the Cluster of Excellence Controlling Microbes to Fight Infection (EXC 2124). S.P.-G. and C.C.-I. were supported by the Center for Oceanographic Research (COPAS) Coastal ANID FB210021 (Chile). R.R.T. and Z.A.Q. were supported by the National Science Foundation (NSF) through Graduate Research Fellowship and a Postdoctoral Fellowship (DGE-2038238, OCE-2308400 and award #2019252845). J.M.B. was supported through an NSF CAREER grant (OCE-1555375). We would like to thank the California Current Ecosystem Long Term Ecological Research (CCE-LTER) Program and the Captains and Crew of the RV Atlantis, RV Oceanus, RV Revelle and RV Sally Ride. We would also like to thank the team at University of California Gump Marine Station on Moorea for logistical support. Datasets presented here were supported in part by CCE-LTER funding (NSF OCE-1637632) and Moorea funding (OCE-2023509 to L.I.A. and P.C.D., OCE-2118618 to L.W.K., OCE-2023298 to C.E.N.), and NASA award 80NSSC18K0437 to C.A.C. A.H. was supported by the United States Department of Defense Environmental Security Technology Certification Program (no. CR20-5175). B.R.B.d.C. was supported by São Paulo Research Foundation (FAPESP, grant no. 2023/15215-4) through a Research Internship Abroad Fellowship. J.-C.J.K. was supported by a Rhodes University postdoctoral fellowship and by the South African Medical Research Council as well as the UK Medical Research Council, through the UK Government's Newton Fund (grant no. 96185). We thank the Shallow Marine and Coastal Research Infrastructure platform for research infrastructure support as well as the crew of the RV Ukwabelana and Koos Smith of the African Coelacanth Ecosystem Programme.
Author information
Authors and Affiliations
Contributions
J.-C.J.K., A.K.P.M.S. and D.P. conceptualized the study. L.W.K., C.E.N., C.A.C., J.M.B., R.A.D., D.T., P.C.D., L.I.A. and D.P. supervised the study. S.P.F., R.R.T., L.C., B.M.S., C.U.-T., I.K., Z.A.Q., L.W.K., C.E.N., C.A.C., C.C.-I., S.P.-G., J.M.B., A.A., A.H., X.S.N., R.A.D., D.T., A.F.H., P.C.D., L.I.A. and D.P. performed sampling. S.P.F., P.S., R.R.T., L.C., B.M.S., C.U.-T., I.K., Z.A.Q., L.W.K., C.E.N., S.P.-G., J.M.B., A.H., X.S.N., D.T., A.F.H. and D.P. acquired LC-MS/MS data. J.-C.J.K., B.R.B.d.C., L.G.G., C.E.N., M.W. and D.P. analysed the data. J.-C.J.K., A.K.P.M.S., B.R.B.d.C., S.P.F., L.G.G., T.S., Z.A.Q., C.E.N., M.W. and D.P. wrote the manuscript. All authors edited and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
P.C.D. is an advisor and holds equity in Cybele, BileOmix and Sirenas and is a scientific co-founder, advisor, holds equity in and/or received income from Ometa, Enveda and Arome with prior approval by UC-San Diego. P.C.D. also consulted for DSM animal health in 2023. M.W. is a co-founder of Ometa Labs LLC. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Geoscience thanks Eric Carmona, Mengyang Liu and Aron Stubbins for their contribution to the peer review of this work. Primary Handling Editor: James Super, in collaboration with the Nature Geoscience team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–13, Tables 1–3 and description of mass spectrometry datasets and metadata.
Supplementary Table 1 (download CSV )
Metadata of LC-MS/MS datasets.
Supplementary Table 2 (download CSV )
List of detected xenobiotics.
Source data
Source Data Fig. 2 (download XLSX )
PCoA with Bray–Curtis distance, labelled by ecosystem. Xenobiotics grouped.
Source Data Fig. 3 (download XLSX )
Top xenobiotic frequency percentages. Industrial pollutants scaled violin plot. Pesticides scaled violin plots. Xenobiotics scaled violin plots. Drugs scaled violin plots.
Source Data Fig. 4 (download XLSX )
Poly-glycols first neighbours scaled violin plot. Organophosphates first neighbours scaled violin plot. Phthalates first neighbours scaled violin plot.
Source Data Fig. 5 (download XLSX )
Results xenobiotics percentage. Results xenobiotics first neighbours percentage. Distance to coast xenobiotics percentage. Depth bucket xenobiotics percentage.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kalinski, JC.J., Pakkir Mohamed Shah, A.K., Ruiz Brandão da Costa, B. et al. Widespread presence of anthropogenic compounds in marine dissolved organic matter. Nat. Geosci. (2026). https://doi.org/10.1038/s41561-026-01928-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41561-026-01928-z