Abstract
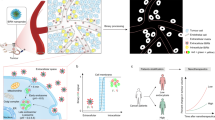
Nanoparticles are promising for drug delivery applications, with several clinically approved products. However, attaining high nanoparticle accumulation in solid tumours remains challenging. Here we show that tumour cell-derived small extracellular vesicles (sEVs) block nanoparticle delivery to tumours, unveiling another barrier to nanoparticle-based tumour therapy. Tumour cells secrete large amounts of sEVs in the tumour microenvironment, which then bind to nanoparticles entering tumour tissue and traffic them to liver Kupffer cells for degradation. Knockdown of Rab27a, a gene that controls sEV secretion, decreases sEV levels and improves nanoparticle accumulation in tumour tissue. The therapeutic efficacy of messenger RNAs encoding tumour suppressing and proinflammatory proteins is greatly improved when co-encapsulated with Rab27a small interfering RNA in lipid nanoparticles. Together, our results demonstrate that tumour cell-derived sEVs act as a defence system against nanoparticle tumour delivery and that this system may be a potential target for improving nanoparticle-based tumour therapies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All relevant data of this article are available within the paper and its Supplementary Information files. A dataset is provided with this paper. Transcriptomics sequencing data is available from the Sequence Read Archive under accession code PRJNA1086632. Mouse single-cell RNA sequencing datasets were downloaded from Gene Expression Omnibus (GSE109774). The images in Supplementary Figs. 19, 20, 23 and 24b were downloaded from The Human Protein Atlas (primary publication: Uhlén M., Fagerberg L., Hallström B.M., et al. Science, 2015, 347(6220): 1260419.). Source data are provided with this paper.
References
D’Mello, S. R. et al. The evolving landscape of drug products containing nanomaterials in the United States. Nat. Nanotech. 12, 523–529 (2017).
van der Meel, R. et al. Smart cancer nanomedicine. Nat. Nanotech. 14, 1007–1017 (2019).
Sindhwani, S. et al. The entry of nanoparticles into solid tumours. Nat. Mater. 19, 566–575 (2020).
Wilhelm, S. et al. Analysis of nanoparticle delivery to tumours. Nat. Rev. Mater. 1, 1–12 (2016).
Sykes, E. A., Chen, J., Zheng, G. & Chan, W. C. Investigating the impact of nanoparticle size on active and passive tumor targeting efficiency. ACS Nano 8, 5696–5706 (2014).
Zhang, Y. R. et al. Strategies to improve tumor penetration of nanomedicines through nanoparticle design. Wires Nanomed. Nanobi. 11, e1519 (2019).
Choi, K. Y. et al. Self-assembled hyaluronic acid nanoparticles for active tumor targeting. Biomaterials 31, 106–114 (2010).
Kong, S. M., Costa, D. F., Jagielska, A., Van Vliet, K. J. & Hammond, P. T. Stiffness of targeted layer-by-layer nanoparticles impacts elimination half-life, tumor accumulation, and tumor penetration. Proc. Natl Acad. Sci. USA 118, e2104826118 (2021).
Acharya, S. & Sahoo, S. K. PLGA nanoparticles containing various anticancer agents and tumour delivery by EPR effect. Adv. Drug Del. Rev. 63, 170–183 (2011).
Bazak, R., Houri, M., El Achy, S., Kamel, S. & Refaat, T. Cancer active targeting by nanoparticles: a comprehensive review of literature. J. Cancer Res. Clin. Oncol. 141, 769–784 (2015).
Cedervall, T. et al. Understanding the nanoparticle–protein corona using methods to quantify exchange rates and affinities of proteins for nanoparticles. Proc. Natl Acad. Sci. USA 104, 2050–2055 (2007).
Zhang, T. et al. Magnetothermal regulation of in vivo protein corona formation on magnetic nanoparticles for improved cancer nanotherapy. Biomaterials 276, 121021 (2021).
Zhang, Y.-N., Poon, W., Tavares, A. J., McGilvray, I. D. & Chan, W. C. Nanoparticle–liver interactions: cellular uptake and hepatobiliary elimination. J. Control. Release 240, 332–348 (2016).
Nia, H. T., Munn, L. L. & Jain, R. K. Physical traits of cancer. Science 370, eaaz0868 (2020).
Stylianopoulos, T. et al. Causes, consequences, and remedies for growth-induced solid stress in murine and human tumors. Proc. Natl Acad. Sci. USA 109, 15101–15108 (2012).
Chen, Y. et al. Vasodilator hydralazine promotes nanoparticle penetration in advanced desmoplastic tumors. ACS Nano 13, 1751–1763 (2019).
Fukumura, D. & Jain, R. K. Tumor microvasculature and microenvironment: targets for anti-angiogenesis and normalization. Microvasc. Res. 74, 72–84 (2007).
Binnewies, M. et al. Understanding the tumor immune microenvironment (TIME) for effective therapy. Nat. Med. 24, 541–550 (2018).
Dai, Y., Xu, C., Sun, X. & Chen, X. Nanoparticle design strategies for enhanced anticancer therapy by exploiting the tumour microenvironment. Chem. Soc. Rev. 46, 3830–3852 (2017).
Gong, F. et al. Tumor microenvironment-responsive intelligent nanoplatforms for cancer theranostics. Nano Today 32, 100851 (2020).
Zieren, R. C. et al. Extracellular vesicle isolation from human renal cancer tissue. Med. Oncol. 37, 1–11 (2020).
Silva, J. et al. Analysis of exosome release and its prognostic value in human colorectal cancer. Genes Chromosomes Cancer 51, 409–418 (2012).
Miller, I. V. & Grunewald, T. G. Tumour‐derived exosomes: tiny envelopes for big stories. Biol. Cell 107, 287–305 (2015).
Ostrowski, M. et al. Rab27a and Rab27b control different steps of the exosome secretion pathway. Nat. Cell Biol. 12, 19–30 (2010).
Poggio, M. et al. Suppression of exosomal PD-L1 induces systemic anti-tumor immunity and memory. Cell 177, 414–427. e413 (2019).
Chen, G. et al. Exosomal PD-L1 contributes to immunosuppression and is associated with anti-PD-1 response. Nature 560, 382–386 (2018).
Fan, Y. et al. Exosomal PD-L1 retains immunosuppressive activity and is associated with gastric cancer prognosis. Ann. Surg. Oncol. 26, 3745–3755 (2019).
Kim, D. H. et al. Exosomal PD-L1 promotes tumor growth through immune escape in non-small cell lung cancer. Exp. Mol. Med. 51, 1–13 (2019).
Monypenny, J. et al. ALIX regulates tumor-mediated immunosuppression by controlling EGFR activity and PD-L1 presentation. Cell Rep. 24, 630–641 (2018).
Ricklefs, F. L. et al. Immune evasion mediated by PD-L1 on glioblastoma-derived extracellular vesicles. Sci. Adv. 4, eaar2766 (2018).
Yang, Y. et al. Exosomal PD-L1 harbors active defense function to suppress T cell killing of breast cancer cells and promote tumor growth. Cell Res. 28, 862–864 (2018).
Gargiulo, E. et al. Extracellular vesicle secretion by leukemia cells in vivo promotes CLL progression by hampering antitumor T-cell responses. Blood Cancer Discov. 4, 54–77 (2022).
Billingsley, M. M. et al. Ionizable lipid nanoparticle-mediated mRNA delivery for human CAR T cell engineering. Nano Lett. 20, 1578–1589 (2020).
Ruiz-Alcaraz, A. J. et al. Characterization of human peritoneal monocyte/macrophage subsets in homeostasis: phenotype, GATA6, phagocytic/oxidative activities and cytokines expression. Sci. Rep. 8, 1–14 (2018).
Jing, H., Sinha, S., Sachar, H. S. & Das, S. Interactions of gold and silica nanoparticles with plasma membranes get distinguished by the van der Waals forces: implications for drug delivery, imaging, and theranostics. Colloids Surf., B 177, 433–439 (2019).
Tian, T. et al. Exosome uptake through clathrin-mediated endocytosis and macropinocytosis and mediating miR-21 delivery. J. Biol. Chem. 289, 22258–22267 (2014).
Wang, G. et al. Tumour extracellular vesicles and particles induce liver metabolic dysfunction. Nature 618, 374–382 (2023).
Springer, T., Galfré, G., Secher, D. S. & Milstein, C. Mac‐1: a macrophage differentiation antigen identified by monoclonal antibody. Eur. J. Immunol. 9, 301–306 (1979).
Kamerkar, S. et al. Exosomes facilitate therapeutic targeting of oncogenic KRAS in pancreatic cancer. Nature 546, 498–503 (2017).
Simpson, L. & Parsons, R. PTEN: life as a tumor suppressor. Exp. Cell. Res. 264, 29–41 (2001).
Huang, K. et al. Size-dependent localization and penetration of ultrasmall gold nanoparticles in cancer cells, multicellular spheroids, and tumors in vivo. ACS Nano 6, 4483–4493 (2012).
Barber, G. N. STING: infection, inflammation and cancer. Nat. Rev. Immunol. 15, 760–770 (2015).
Le Naour, J., Zitvogel, L., Galluzzi, L., Vacchelli, E. & Kroemer, G. Trial watch: STING agonists in cancer therapy. Oncoimmunology 9, 1777624 (2020).
Dane, E. L. et al. STING agonist delivery by tumour-penetrating PEG-lipid nanodiscs primes robust anticancer immunity. Nat. Mater. 21, 710–720 (2022).
De Queiroz, N. M. G. P., Xia, T., Konno, H. & Barber, G. N. Ovarian cancer cells commonly exhibit defective STING signaling which affects sensitivity to viral OncolysisSTING signaling in ovarian cancer. Mol. Cancer Res. 17, 974–986 (2019).
Liu, W. et al. Selective reactivation of STING signaling to target Merkel cell carcinoma. Proc. Natl Acad. Sci. USA 117, 13730–13739 (2020).
Jain, R. et al. MicroRNAs enable mRNA therapeutics to selectively program cancer cells to self-destruct. Nucleic Acid Ther. 28, 285–296 (2018).
Kaufman, H. L. & Maciorowski, D. Advancing oncolytic virus therapy by understanding the biology. Nat. Rev. Clin. Oncol. 18, 197–198 (2021).
Heinemann, V., Stintzing, S., Kirchner, T., Boeck, S. & Jung, A. Clinical relevance of EGFR-and KRAS-status in colorectal cancer patients treated with monoclonal antibodies directed against the EGFR. Cancer Treat. Rev. 35, 262–271 (2009).
Kimiz-Gebologlu, I., Gulce-Iz, S. & Biray-Avci, C. Monoclonal antibodies in cancer immunotherapy. Mol. Biol. Rep. 45, 2935–2940 (2018).
Ohara, Y. et al. Effective delivery of chemotherapeutic nanoparticles by depleting host Kupffer cells. Int. J. Cancer 131, 2402–2410 (2012).
Zhang, W. et al. ICAM-1-mediated adhesion is a prerequisite for exosome-induced T cell suppression. Dev. Cell 57, 329–343. e327 (2022).
Gong, N. et al. In situ PEGylation of CAR T cells alleviates cytokine release syndrome and neurotoxicity. Nat. Mater. 22, 1571–1580 (2023).
Huo, S. et al. Superior penetration and retention behavior of 50 nm gold nanoparticles in tumorscell penetration and tumor retention of gold nanoparticles. Cancer Res. 73, 319–330 (2013).
Xing, L., Zheng, H., Cao, Y. & Che, S. Coordination polymer coated mesoporous silica nanoparticles for pH‐responsive drug release. Adv. Mater. 24, 6433–6437 (2012).
Saad, W. S. & Prud’homme, R. K. Principles of nanoparticle formation by flash nanoprecipitation. Nano Today 11, 212–227 (2016).
Huo, S. et al. Ultrasmall gold nanoparticles as carriers for nucleus-based gene therapy due to size-dependent nuclear entry. ACS Nano 8, 5852–5862 (2014).
Consortium, T. M. et al.Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature 562, 367–372 (2018).
Uhlén, M. et al. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Acknowledgements
M.J.M. acknowledges support from a US National Institutes of Health (NIH) Director’s New Innovator Award (DP2 TR002776), a Burroughs Wellcome Fund Career Award at the Scientific Interface (CASI), a US National Science Foundation CAREER Award (CBET-2145491) and an American Cancer Society Research Scholar grant (RSG-22-122-01-ET). W.G. acknowledges supports from NIH R35 GM141832 and NCI P50 CA261608.
Author information
Authors and Affiliations
Contributions
N.G., W.Z. M.-G.A., D.W., W.G. and M.J.M. conceived and designed the experiments. N.G., W.Z., M.-G.A., X.H., L.X., R.E.-M., G.Z., Z.Q., F.X., A.G.H, D.K. and J.X. performed the experiments, N.G., W.Z. L.X., X.H. and X.T. analysed the data. N.G., W.Z. M.-G.A., D.W., W.G. and M.J.M. wrote and edited the manuscript. J.L. and X.-J.L. were involved in discussion. D.W., W.G. and M.J.M. supervised the entire project. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
N.G., M.J.M., W.Z. and W.G. have filed a patent (Lipid nanoparticle (LNP) compositions and methods for delivering therapeutic agents to tumour cells) related to this paper.
Peer review
Peer review information
Nature Materials thanks Joao Conde and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 LNPs delivering siRab27a and mSTING-miR-122 do not induce systemic toxicity.
1x106 MC38 cells were s.c. injected into the right flank of mice at day 0. At day 8, the tumour size reached 50 mm3. Mice were then treated with the following LNPs: 1) LNP co-encapsulating siRab27a and scrambled mRNA; 2) LNP co-encapsulating scrambled siRNA and mSTING-miR-122; or 3) LNP co-encapsulating siRab27a and mSTING-miR-122. These LNPs were i.v. injected (0.25mg/kg) into mice at days 7, 9, 11, 13, and 15. PBS injections into mice at different time points were used as a control group. When tumour sizes in the PBS group reached 1500 mm3 (day 20), mice were euthanized and ALT (a), AST (b), IL-6 (c), and IL-12p70 (d) in mouse blood were measured. e, H&E staining of mouse livers collected from different groups. f, H&E staining of major mouse organs collected from the PBS group and the STING-miR-122 + siRab27a group at day 50. Data in a-d was shown as mean ± s.d. (n=5 biologically independent samples). One-way ANOVA was used to determine statistical differences. e and f, Experiments were repeated independently 3 times with similar results.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–38 and Table 1.
Supplementary Data 1 (download XLSX )
Source data for Supplementary Fig. 1.
Supplementary Data 2 (download XLSX )
Source data for Supplementary Fig. 2.
Supplementary Data 3 (download XLSX )
Source data for Supplementary Fig. 3.
Supplementary Data 4 (download XLSX )
Source data for Supplementary Fig. 4.
Supplementary Data 5 (download XLSX )
Source data for Supplementary Fig. 5.
Supplementary Data 6 (download XLSX )
Source data for Supplementary Fig. 6.
Supplementary Data 7 (download XLSX )
Source data for Supplementary Fig. 7.
Supplementary Data 8 (download XLSX )
Source data for Supplementary Fig. 8.
Supplementary Data 9 (download XLSX )
Source data for Supplementary Fig. 9.
Supplementary Data 10 (download XLSX )
Source data for Supplementary Fig. 10.
Supplementary Data 11 (download XLSX )
Source data for Supplementary Fig. 11.
Supplementary Data 12 (download XLSX )
Source data for Supplementary Fig. 12.
Supplementary Data 13 (download XLSX )
Source data for Supplementary Fig. 13.
Supplementary Data 14 (download XLSX )
Source data for Supplementary Fig. 14.
Supplementary Data 16 (download XLSX )
Source data for Supplementary Fig. 16.
Supplementary Data 17 (download XLSX )
Source data for Supplementary Fig. 17.
Supplementary Data 18 (download XLSX )
Source data for Supplementary Fig. 18.
Supplementary Data 22 (download XLSX )
Source data for Supplementary Fig. 22.
Supplementary Data 25 (download XLSX )
Source data for Supplementary Fig. 25.
Supplementary Data 26 (download XLSX )
Source data for Supplementary Fig. 26.
Supplementary Data 27 (download XLSX )
Source data for Supplementary Fig. 27.
Supplementary Data 28 (download XLSX )
Source data for Supplementary Fig. 28.
Supplementary Data 29 (download XLSX )
Source data for Supplementary Fig. 29.
Supplementary Data 30 (download XLSX )
Source data for Supplementary Fig. 30.
Supplementary Data 33 (download XLSX )
Source data for Supplementary Fig. 33.
Supplementary Data 35 (download XLSX )
Source data for Supplementary Fig. 35.
Supplementary Data 36 (download XLSX )
Source data for Supplementary Fig. 36.
Supplementary Data 37 (download XLSX )
Source data for Supplementary Fig. 37.
Supplementary Data 38 (download XLSX )
Source data for Supplementary Fig. 38.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Unmodified_Gels _Fig. 2g (download PDF )
Unprocessed western blots.
Unmodified_Gels _Fig. 4h (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 1 and Table 1 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gong, N., Zhong, W., Alameh, MG. et al. Tumour-derived small extracellular vesicles act as a barrier to therapeutic nanoparticle delivery. Nat. Mater. 23, 1736–1747 (2024). https://doi.org/10.1038/s41563-024-01961-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41563-024-01961-6
This article is cited by
-
Precision nanomedicine: navigating the tumor microenvironment for enhanced cancer immunotherapy and targeted drug delivery
Molecular Cancer (2025)
-
AI-engineered multifunctional nanoplatforms: synergistically bridging precision diagnosis and intelligent therapy in next-generation oncology
Journal of Nanobiotechnology (2025)
-
Nanoparticle-mediated delivery of oncolytic viral genomes: an innovative strategy for tumor-targeted immunotherapy
Cancer Nanotechnology (2025)
-
Clinical relevance of extracellular vesicles in cancer — therapeutic and diagnostic potential
Nature Reviews Clinical Oncology (2025)
-
Dual SORT LNPs for multi-organ base editing
Nature Biotechnology (2025)