Abstract
The advent of mRNA vaccines represents a breakthrough in the realm of cancer therapy and the prevention of infectious disease. Nevertheless, traditional lipid nanoparticle (LNP)-based mRNA vaccines can accumulate in the liver post-intramuscular injection, posing a risk of hepatotoxicity and reducing efficacy. Here we develop an albumin-recruiting LNP system with high lymphatic drainage and no accumulation in hepatic tissue to potentiate the efficacy and safety of mRNA vaccines. We construct a library of ionizable lipids with albumin-binding capacity as alternatives to traditional polyethylene-glycol-conjugated lipid. We identify an Evans blue-modified lipid-based LNP (EB-LNP) formulation that shows high in vivo expression, albumin-facilitated transport through intramuscular lymphatic vessels to the lymph nodes, high internalization by dendritic cells and low penetration into intramuscular blood vessels, thereby avoiding liver accumulation. EB-LNP-based mRNA vaccines demonstrate excellent antitumour and antiviral efficacy, resulting in strong cellular and humoral immune responses, including the robust activation of cytotoxic T lymphocytes and production of neutralizing antibodies post-vaccination. Overall, this system shows promise as an effective and minimally toxic platform for the development of mRNA vaccines with high efficacy and safety.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All relevant data of this study are available within the Article and its Supplementary Information. All proteomics raw data are deposited to the ProteomeXchange (https://proteomecentral.proteomexchange.org/cgi/GetDataset) with the identifier PXD061738. Source data are provided with this paper.
Change history
12 August 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41563-025-02343-2
References
Pardi, N. et al. mRNA vaccines—a new era in vaccinology. Nat. Rev. Drug Discov. 17, 261–279 (2018).
Hou, X. et al. Lipid nanoparticles for mRNA delivery. Nat. Rev. Mater. 6, 1078–1094 (2021).
Pardi, N. & Krammer, F. mRNA vaccines for infectious diseases—advances, challenges and opportunities. Nat. Rev. Drug. Discov. 23, 838–861 (2024).
Ju, Y. et al. Impact of anti-PEG antibodies induced by SARS-CoV-2 mRNA vaccines. Nat. Rev. Immunol. 23, 135–136 (2023).
Chen, J. et al. Current developments and challenges of mRNA vaccines. Annu. Rev. Biomed. Eng. 24, 85–109 (2022).
Weber, J. S. et al. Individualised neoantigen therapy mRNA-4157 (V940) plus pembrolizumab versus pembrolizumab monotherapy in resected melanoma (KEYNOTE-942): a randomised, phase 2b study. Lancet 403, 632–644 (2024).
Ndeupen, S. et al. The mRNA-LNP platform’s lipid nanoparticle component used in preclinical vaccine studies is highly inflammatory. iScience 24, 103479 (2021).
Pateev, I. et al. Biodistribution of RNA vaccines and of their products: evidence from human and animal studies. Biomedicines 12, 59 (2023).
Bahl, K. et al. Preclinical and clinical demonstration of immunogenicity by mRNA vaccines against H10N8 and H7N9 influenza viruses. Mol. Ther. 25, 1316–1327 (2017).
Ye, Z. et al. Monovalent SARS-COV-2 mRNA vaccine using optimal UTRs and LNPs is highly immunogenic and broadly protective against Omicron variants. Proc. Natl Acad. Sci. USA 120, e2311752120 (2023).
Shi, Y. et al. Structural and biochemical characteristics of mRNA nanoparticles determine anti-SARS-CoV-2 humoral and cellular immune responses. Sci. Adv. 8, eabo1827 (2022).
Chen, J. et al. Lipid nanoparticle-mediated lymph node-targeting delivery of mRNA cancer vaccine elicits robust CD8+ T cell response. Proc. Natl Acad. Sci. USA 119, e2207841119 (2022).
Kitagawa, H. et al. Adverse reactions to the BNT162b2 and mRNA-1273 mRNA COVID-19 vaccines in Japan. J. Infect. Chemother. 28, 576–581 (2022).
Schinas, G. et al. Immune-mediated liver injury following COVID-19 vaccination. World J. Virol. 12, 100–108 (2023).
Efe, C. et al. Liver injury after SARS-CoV-2 vaccination: features of immune-mediated hepatitis, role of corticosteroid therapy and outcome. Hepatology 76, 1576–1586 (2022).
Boettler, T. et al. SARS-CoV-2 vaccination can elicit a CD8 T-cell dominant hepatitis. J. Hepatol. 77, 653–659 (2022).
Zin Tun, G. S. et al. Immune-mediated hepatitis with the Moderna vaccine, no longer a coincidence but confirmed. J. Hepatol. 76, 738–753 (2022).
Jeong, M. et al. Lipid nanoparticles (LNPs) for in vivo RNA delivery and their breakthrough technology for future applications. Adv. Drug Deliv. Rev. 200, 114990 (2023).
Liu, M. et al. Lymph-targeted high-density lipoprotein-mimetic nanovaccine for multi-antigenic personalized cancer immunotherapy. Sci. Adv. 10, eadk2444 (2024).
Sasaki, K. et al. mRNA-loaded lipid nanoparticles targeting dendritic cells for cancer immunotherapy. Pharmaceutics 14, 1572 (2022).
Cheng, Q. et al. Selective organ targeting (SORT) nanoparticles for tissue-specific mRNA delivery and CRISPR–Cas gene editing. Nat. Nanotechnol. 15, 313–320 (2020).
Liu, S. et al. Membrane-destabilizing ionizable phospholipids for organ-selective mRNA delivery and CRISPR–Cas gene editing. Nat. Mater. 20, 701–710 (2021).
Yu, X. et al. Poly(ethyl ethylene phosphate): overcoming the ‘polyethylene glycol dilemma’ for cancer immunotherapy and mRNA vaccination. ACS Nano 17, 23814–23828 (2023).
Hald Albertsen, C. et al. The role of lipid components in lipid nanoparticles for vaccines and gene therapy. Adv. Drug Deliv. Rev. 188, 114416 (2022).
Tenchov, R. et al. PEGylated lipid nanoparticle formulations: immunological safety and efficiency perspective. Bioconjug. Chem. 34, 941–960 (2023).
Dilliard, S. A. et al. On the mechanism of tissue-specific mRNA delivery by selective organ targeting nanoparticles. Proc. Natl Acad. Sci. USA 118, e210925611 (2021).
Elzoghby, A. O. et al. Albumin-based nanoparticles as potential controlled release drug delivery systems. J. Control. Release 157, 168–182 (2012).
Abdallah, M. et al. Lymphatic targeting by albumin-hitchhiking: applications and optimisation. J. Control. Release 327, 117–128 (2020).
Famta, P. et al. Albumin-hitchhiking: fostering the pharmacokinetics and anticancer therapeutics. J. Control. Release 353, 166–185 (2022).
Qi, S. et al. The bright future of nanotechnology in lymphatic system imaging and imaging-guided surgery. J. Nanobiotechnol. 20, 24 (2022).
Zhu, G. et al. Albumin/vaccine nanocomplexes that assemble in vivo for combination cancer immunotherapy. Nat. Commun. 8, 1954 (2017).
Um, W. et al. A comparative study on albumin-binding molecules for targeted tumor delivery through covalent and noncovalent approach. Bioconjug. Chem. 30, 3107–3118 (2019).
Liu, H. et al. Structure-based programming of lymph-node targeting in molecular vaccines. Nature 507, 519–522 (2014).
Song, Y. et al. Albumin nanoparticle containing a PI3Kγ inhibitor and paclitaxel in combination with α-PD1 induces tumor remission of breast cancer in mice. Sci. Transl. Med. 14, eabl3649 (2022).
Lin, T. et al. Blood–brain-barrier-penetrating albumin nanoparticles for biomimetic drug delivery via albumin-binding protein pathways for antiglioma therapy. ACS Nano 10, 9999–10012 (2016).
Liu, Z. & Chen, X. Simple bioconjugate chemistry serves great clinical advances: albumin as a versatile platform for diagnosis and precision therapy. Chem. Soc. Rev. 45, 1432–1456 (2016).
Sahin, U. et al. BNT162b2 vaccine induces neutralizing antibodies and poly-specific T cells in humans. Nature 595, 572–577 (2021).
Liu, Y. et al. Intrapleural nano-immunotherapy promotes innate and adaptive immune responses to enhance anti-PD-L1 therapy for malignant pleural effusion. Nat. Nanotechnol. 17, 206–216 (2022).
Bern, M. et al. The role of albumin receptors in regulation of albumin homeostasis: implications for drug delivery. J. Control. Release 211, 144–162 (2015).
Ishima, Y. et al. The new delivery strategy of albumin carrier utilizing the interaction with albumin receptors. Chem. Pharm. Bull. 70, 330–333 (2022).
Wan, Y. et al. Stable organic photosensitizer nanoparticles with absorption peak beyond 800 nanometers and high reactive oxygen species yield for multimodality phototheranostics. ACS Nano 14, 9917–9928 (2020).
Sindhwani, S. et al. The entry of nanoparticles into solid tumours. Nat. Mater. 19, 566–575 (2020).
Nguyen, L. N. M. et al. The exit of nanoparticles from solid tumours. Nat. Mater. 22, 1261–1272 (2023).
Embgenbroich, M. & Burgdorf, S. Current concepts of antigen cross-presentation. Front. Immunol. 9, 1643 (2018).
Oğuz, F. & Atmaca, H. mRNA as a therapeutics: understanding mRNA vaccines. Adv. Pharm. Bull. 12, 274–282 (2022).
Szymonowicz, K. A. & Chen, J. Biological and clinical aspects of HPV-related cancers. Cancer Biol. Med. 17, 864 (2020).
Mirabello, L. et al. HPV16 E7 genetic conservation is critical to carcinogenesis. Cell 170, 1164–1174 (2017).
Feshchenko, E. et al. Pandemic influenza vaccine: characterization of A/California/07/2009 (H1N1) recombinant hemagglutinin protein and insights into H1N1 antigen stability. BMC Biotechnol. 12, 77 (2012).
Cyster, J. G. & Allen, C. D. C. B cell responses: cell interaction dynamics and decisions. Cell 177, 524–540 (2019).
De Silva, N. S. & Klein, U. Dynamics of B cells in germinal centres. Nat. Rev. Immunol. 15, 137–148 (2015).
Mittenbühler, M. J. et al. Isolation of extracellular fluids reveals novel secreted bioactive proteins from muscle and fat tissues. Cell Metab. 35, 535–549 (2023).
Renier, N. et al. iDISCO: a simple, rapid method to immunolabel large tissue samples for volume imaging. Cell 159, 896–910 (2014).
Acknowledgements
We thank S. Zhu from the State Key Laboratory of Supramolecular Structure and Materials, Center for Supramolecular Chemical Biology, Jilin University, for his valuable suggestions and technical assistance with NIR-II imaging. G.Y. acknowledges financial support from the Beijing Municipal Science & Technology Commission (Z231100007223007), the National Key R&D Program of China (2023YFE0204900 and 2023YFC3403100), the National Natural Science Foundation of China (22175107 and 22305140) and the Starry Night Science Fund of Zhejiang University Shanghai Institute for Advanced Study (SN-ZJU-SIAS-006). X.C. acknowledges financial support from the National University of Singapore (NUHSRO/2020/133/Startup/08, NUHSRO/2023/008/NUSMed/TCE/LOA, NUHSRO/2021/034/TRP/09/Nanomedicine, NUHSRO/2021/044/Kickstart/09/LOA and 23-0173-A0001), National Medical Research Council (MOH-001388-00, CG21APR1005, MOH-001500-00 and MOH-001609-00), Singapore Ministry of Education (MOE-000387-00 and MOET32023-0005) and National Research Foundation (NRF-000352-00). G.C. acknowledges financial support from the National Key R&D Program of China (2023YFC2305900, 2021YFC2300200, 2022YFC2303200, 2021YFC2302405, 2022YFC2303400 and 2023YFC2305902), the Shenzhen Medical Research Fund (B2404002), the Yunnan Major Scientific and Technological Projects (202502AU100001), Shenzhen San-Ming Project for Prevention and Research on Vector-borne Diseases (SZSM202211023), Yunnan Provincial Science and Technology Project at Southwest United Graduate School (202302AO370010) and the New Cornerstone Science Foundation.
Author information
Authors and Affiliations
Contributions
Y.F., G.C., X.C. and G.Y. conceived and designed the research. Y.F., W.T., P.H., S.Q., X.Y., Mengyao Li, X.G. and K.Y. conducted the experiments. Y.F., Mengfei Li, M.Z., F.C., B.B., J.L., M.C. and Y.L. interpreted the data and developed the discussion. Y.F., X.C. and G.Y. composed the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Materials thanks Craig Duvall, Natalie Trevaskis and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Biodistribution pattern study.
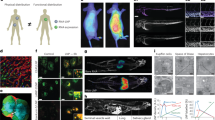
a, In vivo fluorescence imaging of mice after i.m. injection of DiR-labeled EB-LNP@mRNALuci or DiR-labeled PEG-LNP@mRNALuci in the right thigh. b, Quantification of the fluorescence signal from abdomen in a. c, Quantification of the fluorescence signal from muscle in a. d, Percentages of DiR+ cells in mice inguinal LNs by FCM analysis. e, Percentages of DiR+ cells in mice liver tissues by FCM analysis. f, Ex vivo fluorescence imaging of rabbit main organs 4 h post-injection. Abbreviations: M, muscle; LNs, lymph nodes; H, heart; Li, liver; S, spleen; Lu, lung; K, kidney. g, Immunofluorescence staining of mice LNs. Scale bar = 20 μm. Staining was repeated in n = 3 mice. Data in b–e were presented as mean ± s.d. from n = 3 biologically independent samples. Statistical significance was determined by one-way ANOVA with Dunnett’s multiple comparisons test: n.s., not significant, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Extended Data Fig. 2 Time-lapse imaging of main organs.
a, Ex vivo bioluminescence imaging of main organs at different time point post i.m. injection of DiR-labeled EB-LNP@mRNALuci or DiR-labeled PEG-LNP@mRNALuci. b, Ex vivo fluorescence imaging of main organs at different time point post i.m. injection of DiR-labeled EB-LNP@mRNALuci or DiR-labeled PEG-LNP@mRNALuci. Abbreviations: M, muscle; LN, lymph node; H, heart; Li, liver; S, spleen; Lu, lung; K, kidney. c, CLSM images of the inguinal LNs at different times post i.m. injection of DiR-labeled EB-LNP@mRNAGFP or DiR-labeled PEG-LNP@mRNAGFP. Scale bar = 300 μm. Staining was repeated in n = 3 mice. d, The content of GFP in the LNs at different time points by ELISA. e, The content of DiR in the LNs at different time points by LC-MS. Data in d and e were presented as mean ± s.d. from n = 3 biologically independent samples. Statistical significance was determined by one-way ANOVA with Dunnett’s multiple comparisons test: n.s., not significant, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Extended Data Fig. 3 Albumin receptor mediated-endocytosis.
a–c, FCM analysis of endocytosis behavior in DC2.4 cells under various cultivation conditions following incubation with DiR-labeled EB-LNP@mRNAGFP or DiR-labeled PEG-LNP@mRNAGFP: in complete RPMI culture medium (a); in serum-free RPMI culture medium (b); in serum-free RPMI culture medium supplemented with BSA (c). d–f, Statistical diagrams of mean fluorescence intensity in a–c. g–i, Quantitative assessment of GFP+ DC2.4 cells under different cultivation conditions: in complete RPMI culture medium (g); in serum-free RPMI culture medium (h); in serum-free RPMI culture medium supplemented with BSA (i). j–l, Quantitative assessment of OVA-antigen peptide presentation in DC2.4 cells after incubation with EB-LNP@mRNAOVA or PEG-LNP@mRNAOVA: in complete RPMI culture medium (j); in serum-free RPMI culture medium (k); in serum-free RPMI culture medium supplemented with BSA (l). Data in d–l were presented from n = 3 biologically independent samples. Statistical significance was determined by one-way ANOVA with Dunnett’s multiple comparisons test: n.s., not significant, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. For d–l, the center line represented the median, and the whiskers represented the maximum and minimum values.
Extended Data Fig. 4 Intramuscular transport pathway investigation by NIR-II imaging.
a, In vivo NIR-II fluorescence imaging of the mice post i.m. administration of ICG-labeled EB-LNP or ICG-labeled PEG-LNP. The white arrow indicated the initial injection site at right thigh. The blue arrowhead indicated illuminated blood vessels. The pink arrowhead indicated illuminated lymphatic vessels. The blue arrow pointed to the liver and the pink arrow pointed to inguinal LNs. b, In vivo NIR-II fluorescence imaging of the mice, which underwent thoracic duct ligation surgery post i.m. administration of ICG-labeled EB-LNP and ICG-labeled PEG-LNP. c,d, Quantitative analysis of the fluorescence signal intensity in lymphatic vessels (c) and blood vessels (d). e,f, Quantitative analysis of the fluorescence signal intensity in lymphatic vessel (e) and blood vessel (f) from the mice which underwent thoracic duct ligation surgery. g, Ex vivo NIR-II fluorescence imaging of the main organs 4 h post-injection. h, Quantitative analysis of the fluorescence signal intensity from main organs in g. i, Ex vivo NIR-II fluorescence imaging of the main organs from the mice underwent thoracic duct ligation surgery 4 h post-injection. j, Quantitative analysis of the fluorescence signal intensity from main organs in i. Abbreviations: M, muscle; LN, lymph node; H, heart; Li, liver; S, spleen; Lu, lung; K, kidney. Data in c–f, h, and j were presented as mean ± s.d. from n = 3 biologically independent samples. Statistical significance was determined by one-way ANOVA with Dunnett’s multiple comparisons test: n.s., not significant, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Extended Data Fig. 5 Biosafety and immune responses investigation.
a–h, Liver functions assessment of the mice with different treatments: the levels of ALT (a), ALP (b), AST (c), γ-GT (d), TBIL (e), DBIL (f), ALB (g), and TBA (h). The area between the two gray dashed lines represented the normal range (NR). i,j, Detection of PEG-specific antibodies including IgM (i) and IgG (j) in mice serum. k–m, Liver tissues staining: ORO staining for lipid accumulation (k), DHE staining for intracellular reactive oxygen species (l), and H&E staining for pathological features (m). All scale bars = 100 μm. In a–m, C57BL/6 normal mice received three i.m. injections of EB-LNP or PEG-LNP on day 0, 3, and 5. The blood samples and livers were collected on day 6 to determine liver functions. PEG-specific antibodies in serum were continuously monitored until day 28. n–p, FCM analysis of inguinal LNs dissected from C57BL/6 normal mice 7 days after a single i.m. injection of EB-Vax or PEG-Vax: the percentages of CD11c+ DCs (n), CD11c+H2Kb+ DCs (o), and CD11c+CD80+CD86+ DCs (p). q, Determination of CD8+IFN-γ+ T cells in spleen. Abbreviations: EB-Vax for EB-LNP@mRNAOVA, and PEG-Vax for PEG-LNP@mRNAOVA. Data in a–j and n–q were presented as mean ± s.d. from n = 3 biologically independent samples. Statistical significance in a–h and n–q was determined by one-way ANOVA with Tukey’s correction: n.s., not significant, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Statistical significance in i and j was determined by one-way ANOVA with Dunnett’s multiple comparisons test: n.s., not significant, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. For a–h and n–q, the center line represented the median, and the whiskers represented the maximum and minimum values.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–73, Discussion, Methods and references.
Supplementary Video 1 (download MP4 )
EB-LNP in an i.m. blood vessel.
Supplementary Video 2 (download MP4 )
EB-LNP in an i.m. lymphatic vessel.
Supplementary Video 3 (download MP4 )
PEG-LNP in an i.m. blood vessel.
Supplementary Video 4 (download MP4 )
PEG-LNP in an i.m. lymphatic vessel.
Supplementary Video 5 (download MP4 )
Tissue clearing and imaging of the LN after the i.m. injection of EB-LNP.
Supplementary Video 6 (download MP4 )
Tissue clearing and imaging of the LN after the i.m. injection of PEG-LNP.
Source data
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Feng, Y., Tai, W., Huang, P. et al. Albumin-recruiting lipid nanoparticle potentiates the safety and efficacy of mRNA vaccines by avoiding liver accumulation. Nat. Mater. 24, 1826–1839 (2025). https://doi.org/10.1038/s41563-025-02284-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41563-025-02284-w
This article is cited by
-
Albumin nanoparticles deliver mRNA on target
Nature Reviews Materials (2025)
-
Breaking the PEG barrier to boost mRNA-LNP therapeutics
Nature Reviews Materials (2025)