Abstract
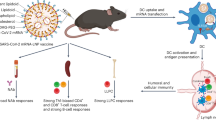
Modulating metabolism in immune cells is an effective approach to induce desired immune responses. Here we develop a lipid nanoparticle (LNP) capable of metabolic reprogramming of dendritic cells for mRNA vaccine applications. Using imidoester-based conjugation chemistry, we design a crosslinked ionizable lipid, C12-2aN, which possesses intrinsic metabolic modulatory properties. This multifunctional ionizable lipid not only promotes effective mRNA expression by facilitating endosomal escape but also stimulates glycolysis through mTORC2 pathway activation. As both an mRNA carrier and a metabolic modulator, C12-2aN LNPs lead to potent vaccine efficacy in both SARS-CoV-2 and OVA cancer vaccine models, resulting in stronger neutralization of pseudovirus infection and improved survival rates, respectively, compared with control LNPs without the crosslinker. Moreover, C12-2aN LNPs outperformed FDA-approved LNPs in terms of reduced off-target delivery and lower immunogenicity. Overall, the integration of mRNA delivery and metabolic reprogramming induced by the ionizable lipid component presents significant potential for next-generation mRNA LNP vaccines.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others

Data availability
The data supporting the findings of this study are available within the Article and its Supplementary Information. Bulk RNA-sequencing data have been deposited in the NCBI Sequence Read Archive (accession number PRJNA1183400). Single-cell RNA-sequencing data has been deposited in the NCBI Sequence Read Archive (accession number PRJNA1314823). Source data are provided with this paper.
References
Paludan, S. R., Pradeu, T., Masters, S. L. & Mogensen, T. H. Constitutive immune mechanisms: mediators of host defence and immune regulation. Nat. Rev. Immunol. 21, 137–150 (2021).
Leone, R. D. & Powell, J. D. Metabolism of immune cells in cancer. Nat. Rev. Cancer 20, 516–531 (2020).
Jung, J., Zeng, H. & Horng, T. Metabolism as a guiding force for immunity. Nat. Cell Biol. 21, 85–93 (2019).
Ganeshan, K. & Chawla, A. Metabolic regulation of immune responses. Annu. Rev. Immunol. 32, 609–634 (2014).
Menk, A. V. et al. Early TCR signaling induces rapid aerobic glycolysis enabling distinct acute T cell effector functions. Cell Rep. 22, 1509–1521 (2018).
Pearce, E. L. et al. Enhancing CD8 T-cell memory by modulating fatty acid metabolism. Nature 460, 103–107 (2009).
Kastenmüller, W., Kastenmüller, K., Kurts, C. & Seder, R. A. Dendritic cell-targeted vaccines—hope or hype?. Nat. Rev. Immunol. 14, 705–711 (2014).
Giovanelli, P., Sandoval, T. A. & Cubillos-Ruiz, J. R. Dendritic cell metabolism and function in tumors. Trends Immunol. 40, 699–718 (2019).
Pearce, E. J. & Everts, B. Dendritic cell metabolism. Nat. Rev. Immunol. 15, 18–29 (2015).
Perez, C. R. & De Palma, M. Engineering dendritic cell vaccines to improve cancer immunotherapy. Nat. Commun. 10, 5408 (2019).
Everts, B. et al. TLR-driven early glycolytic reprogramming via the kinases TBK1-IKKɛ supports the anabolic demands of dendritic cell activation. Nat. Immunol. 15, 323–332 (2014).
Cullis, P. R. & Felgner, P. L. The 60-year evolution of lipid nanoparticles for nucleic acid delivery. Nat. Rev. Drug Discov. 23, 709–722 (2024).
Kon, E., Ad-El, N., Hazan-Halevy, I., Stotsky-Oterin, L. & Peer, D. Targeting cancer with mRNA-lipid nanoparticles: key considerations and future prospects. Nat. Rev. Clinc. Oncol. 20, 739–754 (2023).
Huang, X. et al. The landscape of mRNA nanomedicine. Nat. Med. 28, 2273–2287 (2022).
Hou, Z., Zaks, T., Langer, R. & Dong, Y. Lipid nanoparticles for mRNA delivery. Nat. Rev. Mater. 12, 1078–1094 (2021).
Hand, E. S. & Jencks, W. P. Mechanism of the reaction of imido esters with amines. J. Am. Chem. Soc. 84, 3505–3514 (1962).
Hunter, M. J. & Ludwig, M. L. The reaction of imidoesters with proteins and related small molecules. J. Am. Chem. Soc. 84, 3419–3504 (1962).
Liu, S. et al. Membrane destabilizing ionizable phospholipids for organ selective mRNA delivery and CRISPR/Cas gene editing. Nat. Mater. 20, 701–710 (2021).
Xu, Y. & Szoka, F. C. Mechanism of DNA release from cationic liposome/DNA complexes used in cell transfection. Biochemistry 35, 5616–5623 (1996).
Zainal Abidin, A. et al. Amidine containing compounds: antimicrobial activity and its potential in combating antimicrobial resistance. Heliyon 10, e32010 (2024).
Sondi, S. M., Rani, R., Roy, P., Agrawal, S. K. & Saxena, A. K. Synthesis, anti-inflammatory, and anticancer activity evaluation of some heterocyclic amidine and bis-amidine derivatives. J. Heterocyclic. Chem. 48, 921–926 (2011).
Samsonowicz-Górski, J., Brodzka, A., Ostaszewski, R. & Koszelewski, D. Screening for amidoxime reductases in plant roots and Saccharomyces cerevisiae—development of biocatalytic method for chemoselective amidine synthesis. Bioorg. Chem. 124, 105815 (2022).
Foretz, M., Guigas, B. & Viollet, B. Metformin: update on mechanisms of action and repurposing potential. Nat. Rev. Endocrinol. 19, 460–476 (2023).
Jang, J. Y. et al. Structural basis for the enhanced anti-diabetic efficacy of lobeglitazone on PPARγ. Sci. Rep. 8, 31 (2018).
Zhang, Z. et al. Brain-restricted mTOR inhibition with binary pharmacology. Nature 609, 822–828 (2022).
Saxton, R. A. & Sabatini, D. M. mTOR signaling in growth, metabolism, and disease. Cell 168, 960–976 (2017).
Kazyken, D. et al. AMPK directly activates mTORC2 to promote cell survival during acute energetic stress. Sci. Signal. 12, eaav3249 (2019).
Zhang, P. et al. WSSV exploits AMPK to activate mTORC2 signaling for proliferation by enhancing aerobic glycolysis. Commun. Biol. 6, 361 (2023).
Masui, K. et al. mTOR complex 2 controls glycolytic metabolism in glioblastoma through FoxO acetylation and upregulation of c-Myc. Cell Metab. 18, 726–739 (2013).
Li, M. et al. mTORC2-AKT signaling to PFKFB2 activates glycolysis that enhances stemness and tumorigenicity of intestinal epithelial cells. FASEB J. 38, e23532 (2024).
Hagiwara, A. et al. Hepatic mTORC2 activates glycolysis and lipogenesis through Akt, glucokinase, and SREBP1c. Cell Metab. 15, 725–738 (2012).
Lee, Y., Jeong, M., Park, J., Jung, H. & Lee, H. Immunogenicity of lipid nanoparticles and its impact on the efficacy of mRNA vaccines and therapeutics. Exp. Mol. Med. 55, 2085–2096 (2023).
Margraf, A., Lowell, C. A. & Zarbock, A. Neutrophils in acute inflammation: current concepts and translational implications. Blood 139, 2130–2144 (2022).
Binici, B., Rattray, Z., Zinger, A. & Perrie, Y. Exploring the impact of commonly used ionizable and pegylated lipids on mRNA-LNPs: a combined in vitro and preclinical perspective. J. Control. Release 377, 162–173 (2025).
Zhang, W. et al. The expression kinetics and immunogenicity of lipid nanoparticles delivering plasmid DNA and mRNA in mice. Vaccines 11, 1580 (2023).
Yasmin, F. et al. Adverse events following COVID-19 mRNA vaccines: a systematic review of cardiovascular complication, thrombosis, and thrombocytopenia. Immun. Inflamm. Dis. 11, e807 (2023).
Stylianopoulos, T. et al. Diffusion of particles in the extracellular matrix: the effect of repulsive electrostatic interactions. Biophys. J. 99, 1342–1349 (2010).
Dilliard, S. A., Chen, Q. & Siegwart, D. J. On the mechanism of tissue-specific mRNA delivery by selective organ targeting nanoparticles. Proc. Natl Acad. Sci. USA 118, e2109256118 (2021).
Møller, S. H., Wang, L. & Ho, P. C. Metabolic programming in dendritic cells tailors immune responses and homeostasis. Cell. Mol. Immunol. 19, 370–383 (2022).
Brombacher, E. C. et al. AMPK activation induces RALDH+ tolerogenic dendritic cells by rewiring glucose and lipid metabolism. J. Cell Biol. 223, e202401024 (2024).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Castanza, A. S. et al. Extending support for mouse data in the Molecular Signatures Database (MSigDB). Nat. Methods 20, 1619–1620 (2023).
Ge, S. X., Jung, D. & Yao, R. ShinyGO: a graphical gene-set enrichment tool for animals and plants. Bioinformatics 36, 2628–2629 (2020).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Acknowledgements
M.J.M. acknowledges support from an American Cancer Society Research Scholar Grant (RSG-22-122-01-ET). E.L.H., A.M.M. and E.F. acknowledge support from an NSF Graduate Research Fellowship (award number 1845298). Data for this manuscript were generated in the University of Pennsylvania’s CDB Microscopy Core and Small Animal Imaging Core Facility (RRID:SCR_022385). Data were also generated in the Penn Cytomics and Cell Sorting Shared Resource Laboratory at the University of Pennsylvania (RRID: SCR_022376) and the Pancreatic Islet Cell Biology Core, which is supported by the University of Pennsylvania Diabetes Research Center (DRC).
Author information
Authors and Affiliations
Contributions
D.K., N.G. and M.J.M. conceived the project and designed the experiments. The experiments were performed by D.K., N.G., M.-G.A., E.L.H., I.-C.Y., H.W., E.F., Q.S. and S.-J.M. and interpreted by all authors. D.K. prepared the figures and wrote the manuscript. D.K., E.L.H., J.W., A.M.M., E.F., K.M., D.W. and M.J.M. edited and revised the manuscript. All authors reviewed the manuscript and figures and approved the final version for submission.
Corresponding author
Ethics declarations
Competing interests
D.K., N.G. and M.J.M. have filed a patent on the LNP technology discussed in this manuscript (application no. PCT/US25/51511). The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Materials thanks John T. Wilson and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Notes 1–5, Figs. 1–39, methods, discussion and references.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data for the lipid-screening results.
Source Data Fig. 2 (download XLSX )
Statistical source data for the lipid-screening results.
Source Data Fig. 3 (download XLSX )
Statistical source data for the cellular metabolic activity results.
Source Data Fig. 4 (download XLSX )
Statistical source data for the in vivo biodistribution and safety analysis.
Source Data Fig. 5 (download XLSX )
Statistical source data for the SARS-CoV-2 vaccine study.
Source Data Fig. 6 (download XLSX )
Statistical source data for the cancer vaccine study.
Source Data Fig. 3 (download JPG )
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kim, D., Gong, N., Alameh, MG. et al. Crosslinked ionizable lipids reprogram dendritic cell metabolism for potent mRNA vaccination. Nat. Mater. (2026). https://doi.org/10.1038/s41563-026-02512-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41563-026-02512-x