Abstract
Bifidobacteria are beneficial saccharolytic microbes that are widely used as probiotics or in synbiotic formulations, yet individual responses to supplementation can vary with strain type, microbiota composition, diet and lifestyle, underscoring the need for strain-level insights into glycan metabolism. Here we reconstructed 68 pathways for the utilization of mono-, di-, oligo- and polysaccharides by analysing the distribution of 589 curated metabolic gene functions (catabolic enzymes, transporters and transcriptional regulators) across 3,083 non-redundant Bifidobacterium genomes of human origin. Thirty-eight predicted phenotypes were validated in vitro for 30 geographically diverse strains, supporting genomics-based predictions. Our analysis uncovered extensive inter- and intraspecies functional heterogeneity, including a distinct clade within Bifidobacterium longum that metabolizes α-glucans and Bangladeshi isolates carrying unique gene clusters for xyloglucan and human milk oligosaccharide utilization. This large-scale genomic compendium advances our understanding of bifidobacterial carbohydrate metabolism and can inform the rational design of probiotic and synbiotic formulations tailored to strain-specific nutrient preferences.
Similar content being viewed by others
Main
Bifidobacteria are Gram-positive, saccharolytic microorganisms that predominantly inhabit animal gastrointestinal tracts1. Multiple Bifidobacterium species colonize the human gut throughout life, with dietary carbohydrate intake playing a key role in shaping this process2. Breastfeeding fosters the dominance of specific Bifidobacterium taxa within the neonatal gut microbiota owing to their evolutionary adaptation to metabolize human milk oligosaccharides (HMOs)3,4,5. Weaning drives a gradual succession of bifidobacterial taxa from those tuned for HMO utilization to those more adapted to foraging dietary glycans (oligo- and polysaccharides) of plant origin6,7,8.
Geographic and cultural dietary differences also profoundly influence the bifidobacterial composition of neonatal microbiotas. For example, Bifidobacterium longum subsp. infantis (Bl. infantis), a specialist HMO utilizer, constitutes up to 90% of the gut microbial composition of healthy breastfed infants from non-Westernized populations9,10,11. Infants from Westernized populations often lack Bl. infantis and instead harbour less proficient HMO utilizers such as Bifidobacterium longum subsp. longum (Bl. longum), Bifidobacterium breve and Bifidobacterium pseudocatenulatum; this appears to be due to a combination of lifestyle and cultural factors9,12,13.
Despite these differences, the predominance of bifidobacteria in gut communities is associated with multiple health benefits, particularly in infancy. Bifidobacterial fermentation products, lactate and acetate, can inhibit pathogen colonization14,15 and serve as substrates for cross-feeding among microbial community members16,17. In addition, multiple Bifidobacterium species produce aromatic lactic acids modulating the immune system18,19. These beneficial traits underpin the widespread use of bifidobacterial strains as probiotics20,21,22,23, often supplemented with complementary prebiotics to facilitate engraftment23,24.
Further development is needed to rationally select probiotic strains and prebiotic glycans tailored for different populations. For instance, a probiotic Bl. infantis strain did not durably engraft in the microbiota of malnourished Bangladeshi infants whose diets were low in breast milk compared with complementary foods23. Moreover, strains isolated from children in this population may harbour distinctive genomic adaptations for metabolizing glycans common in weaning diets23. Thus, strain-level insights into bifidobacterial carbohydrate metabolism, especially in understudied populations, may be instrumental in developing locally adapted pro- and synbiotics.
Genomics-based approaches, including the analysis of carbohydrate-active enzyme (CAZyme) repertoires7,25,26,27, genotype-to-phenotype matching28,29,30 and genome-scale metabolic models31,32,33, have been used to predict the carbohydrate utilization capabilities of bifidobacteria. However, functional gene annotations, particularly for glycan transporters, remain insufficiently curated and integrated into metabolic reconstructions34. In addition, current genomic resources may not fully capture the strain-level diversity within the Bifidobacterium genus.
We used a combination of curated metabolic reconstruction and machine learning to map 68 carbohydrate utilization pathways across 263 reference and 2,820 additional Bifidobacterium genomes by analysing the representation of 589 distinct functional roles (catabolic enzymes, CAZymes, transporters and transcriptional regulators). Thirty-eight predicted glycan utilization phenotypes were validated in vitro in 30 diverse strains. Our findings reveal remarkable inter- and intraspecies variability, including a distinct Bifidobacterium longum clade that metabolizes α-glucans and Bangladeshi strains with unique gene clusters for plant hemicellulose and HMO catabolism. These insights establish a genomic framework for predicting glycan utilization networks across Bifidobacterium lineages and may guide the development of targeted probiotics and synbiotics.
Results
Curated metabolic reconstruction from reference genomes
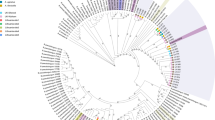
The curated reconstruction encompassed a non-redundant reference set of 263 Bifidobacterium genomes from cultured isolates, including 19 genomes from Bangladeshi strains sequenced in this study (Supplementary Tables 1 and 2). To refine taxonomic assignments, we first constructed a maximum-likelihood phylogenetic tree based on 487 core genes identified through a pangenome analysis (Supplementary Fig. 1). Additional pairwise average nucleotide identity (ANI) comparisons for 39 genomes allowed us to delineate the infraspecific structure of Bifidobacterium longum and Bifidobacterium catenulatum species (Extended Data Fig. 1 and Supplementary Note 1). Overall, the reference set spanned 19 taxa (Supplementary Table 1).
We leveraged a subsystem-based comparative genomics framework35 to reconstruct carbohydrate utilization pathways and predict associated metabolic phenotypes (Fig. 1a). We first mined 222 publications to identify 433 orthologous groups of carbohydrate utilization genes, whose encoded proteins were either experimentally characterized in bifidobacteria (210 metabolic functional roles), shared substantial sequence similarity with proteins characterized in other microbial taxa (80 roles) or were previously assigned putative functions (143 roles). Genomic context analysis, including in silico transcriptional regulon reconstruction (Supplementary Notes 2 and 3), enabled tentative functional predictions for 156 additional gene groups involved in glycan metabolism. The resulting curated set of 589 roles—comprising 235 components of glycan-specific transporters, 197 catabolic CAZymes, 72 downstream catabolic enzymes and 85 transcription factors—was used to functionally annotate 39,589 of 541,418 protein-coding genes across 263 reference genomes (Supplementary Tables 4 and 8). Manual curation improved 76.6% and 69% of annotations over Prokka and EggNOG-mapper, respectively, including more than 90% of annotations for transporters and transcriptional regulators (Fig. 1b). The metabolic reconstruction also captured 82.2% of catabolic CAZymes identified by dbCAN (Fig. 1c).
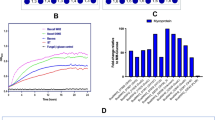
a, An overview of the computational framework. We analysed the distribution of 589 metabolic functional roles (glycan transporters, catabolic enzymes and transcription factors) across 263 reference Bifidobacterium genomes. Manual reconstruction of 68 carbohydrate utilization pathways enabled the assignment of binary pathway variants corresponding to predicted glycan utilization phenotypes based on specific genomic signatures (Extended Data Fig. 2). This curated dataset was used to train an automated pathway prediction pipeline (glycobif), which outputs (1) the distribution of functional roles across additional 2,820 Bifidobacterium genomes and (2) a BPM reflecting the inferred presence or absence of pathways in each genome. Predicted phenotypes for 20 reference and 10 additional strains were compared with in vitro growth profiles to evaluate prediction accuracy. b, A comparison of functional gene annotations obtained via manual curation versus automated tools in the 263 reference Bifidobacterium genomes. Stacked bar plots show the distribution of 39,589 manually curated annotations across four categories: ‘new’, annotations that update non-specific predictions (for example, hypothetical protein); ‘corrected’, annotations that replace specific but incorrect predictions; ‘refined’, annotations that add functional precision (for example, substrate and linkage specificity for GHs); ‘same’, annotations that are essentially identical to automated output (Supplementary Table 13). c, The percentage of catabolic CAZymes (GHs, CEs and polysaccharide lyases) captured in the reconstructed metabolic pathways across the 263 reference Bifidobacterium genomes. Each point represents a genome.
The genome-wide distribution of genes assigned to 589 functional roles was used to reconstruct 68 catabolic pathways: 18 for monosaccharides and their derivatives (sugar alcohols and acids), 39 for di- and oligosaccharides and 11 for polysaccharides (Supplementary Table 5). To link gene conservation patterns with metabolic phenotypes, we first established rules that define genomic signatures distinguishing metabolic pathway variants (Methods, Extended Data Fig. 2 and Supplementary Table 6). Each pathway variant was then converted into a binary phenotype, classifying strains as predicted utilizers (‘1’) or non-utilizers (‘0’) of specific carbohydrates. These assignments formed a binary phenotype matrix (BPM), summarizing the predicted utilization profiles of 263 reference Bifidobacterium strains spanning 68 glycans (Supplementary Table 7). As a preliminary validation of the reconstruction, we compared predicted phenotypes with in vitro growth data for 33 strains from previous studies8,28,36, yielding 94% accuracy (Supplementary Table 17a–d).
Automated pathway prediction across large genomic datasets
We utilized the metabolic reconstruction for 263 reference Bifidobacterium genomes to analyse the representation of glycan utilization pathways in an additional set of 2,820 non-redundant genomes (Supplementary Table 3). This set included 364 isolate genomes and 2,456 high-quality metagenome-assembled genomes (MAGs) with completeness ≥97% and contamination ≤3%. Twelve genomes represented four taxa absent from the reference collection, and nine were from Bangladeshi and Malawian strains isolated in previous studies37,38,39 and sequenced in this work (Supplementary Table 1). We used annotated protein sequences of reference strains to assign functions to a subset of 419,055 protein-coding genes in the 2,820 genomes. These functional annotations, combined with the reference BPM, were used to train a random forest model that predicted the presence of 68 reconstructed carbohydrate utilization pathways (Methods, Fig. 1a and Supplementary Tables 9 and 10).
Interspecies diversity in glycan utilization
The BPMs for 263 reference and 2,820 additional genomes were merged to assess the distribution of glycan utilization pathways across the Bifidobacterium genus. Non-metric multidimensional scaling (NMDS) of the Hamming distance matrix derived from the BPM of 3,083 genomes revealed grouping by species and subspecies, with taxonomy explaining 91% of the variation (permutational multivariate analysis of variance (PERMANOVA) R2 = 0.91, P = 0.001; Fig. 2a). A significant dispersion effect (F = 81.06, P = 0.001) indicated that strain-level variability in pathway representation differed across taxa, probably also influencing the separation.
a, NMDS of a Hamming distance matrix derived from the presence–absence patterns of 68 predicted carbohydrate utilization pathways across 627 isolate genomes plus 2,456 MAGs. Points represent genomes; spider lines connect genomes to their group (taxon) centroid. Colours and shapes of centroids indicate taxonomic assignments. b, Predicted phenotypic richness (the total number of carbohydrate utilization pathways) at species and strain levels. Each point represents a genome. Box plots show the median (centre line), interquartile range (IQR; box bounds) and full data range excluding outliers (whiskers, defined as 1.5× IQR). Statistical comparisons were performed using a two-sided GLM with a Poisson distribution, followed by post-hoc pairwise comparisons with Bonferroni correction. The sample size (n) per group corresponds to the number of genomes analysed. c, A heat map of the proportion of genomes within each taxon encoding the 68 predicted carbohydrate utilization pathways. The colour intensity indicates the percentage of genomes that encode each pathway. Annotation rows at the bottom indicate pathway and phenotype classifications. Full names are provided in Supplementary Table 5.
We observed significant differences in predicted phenotypic richness (total number of predicted carbohydrate utilization pathways) between taxa, including phylogenetically close subspecies sharing over 95% ANI (Poisson generalized linear model (GLM), P < 2.2 × 10−16; Fig. 2b). For example, Bifidobacterium catenulatum subsp. catenulatum (Bc. catenulatum) had significantly lower phenotypic richness than subspecies kashiwanohense (Bc. kashiwanohense) and kashiwanohense_A (Bc. kashiwanohense_A) due to the absence of utilization pathways for fucosylated HMOs (for example, 2′-fucosyllactose (2′FL) and 3-fucosyllactose (3FL)) and certain plant oligosaccharides (for example, β-mannose oligosaccharides, bMnOS) in >95% of genomes (Extended Data Fig. 3 and Supplementary Note 4).
Pathways for glucose (Glc), galactose (Gal), fructose (Fru), lactose (Lac) and galactooligosaccharide (GOS) utilization were identified in over 98% of analysed genomes, defining the core catabolic potential of human-colonizing bifidobacteria (Fig. 2c). Pathways for sucrose (Scr), maltose (Mal), maltooligosaccharide (MOS), isomaltooligosaccharide (IMO), melibiose (Mel), raffinose-family oligosaccharide (RFO) and short-chain fructooligosaccharide (scFOS) metabolism were encoded in over 84% of genomes, indicating broad conservation across most species, except Bifidobacterium bifidum. Other pathways showed more sporadic distribution patterns, reflecting species-level adaptations for metabolizing dietary glycans of different origins and structures. For example, B. bifidum exhibited a notable genomic specialization towards the metabolism of host mucin O-glycans, HMOs and their degradation products lacto-N-biose (LNB), galacto-N-biose (GNB) and N-acetyl-d-glucosamine (GlcNAc)—consistent with prior reports40 (Fig. 2c). At the same time, most B. bifidum genomes lacked complete pathways for utilizing plant-derived mono-, di-, oligo- and polysaccharides. Other bifidobacteria, including the specialist HMO utilizer Bl. infantis, were more versatile in their predicted glycan utilization profiles, although complete pathways for plant polysaccharide degradation were less common than those for catabolizing the corresponding mono- and oligosaccharide components (Fig. 2c).
Hierarchical clustering of the BPM for 263 reference genomes showed a moderate correlation with core-gene phylogeny (cophenetic correlation 0.58, permutation test, P < 0.001; Supplementary Fig. 2), indicating incomplete concordance between predicted glycan utilization capabilities and phylogenetic relatedness. For example, the predicted phenotypic profiles of Bl. infantis and Bl. longum, two phylogenetically related subspecies within the B. longum species, were markedly different. The representation of glycan utilization pathways in Bl. infantis more closely resembled that of B. breve—a more distantly related species inhabiting the neonatal human gut. Given the importance of B. longum in infant microbiota development, we next conducted a more focused analysis of pathway variability within this heterogeneous species.
Diversity of glycan metabolism within the B. longum species
The B. longum species comprises multiple subspecies distinguished by phylogeny and specific phenotypic traits26,41,42,43. Our phylogenomic and ANI analyses clustered B. longum genomes into three clades matching previously described subspecies—infantis (Bl. infantis), longum (Bl. longum) and suis (Bl. suis)—and a distinct clade hereafter referred to as Bl. nov. (Extended Data Fig. 1a, Supplementary Fig. 1 and Supplementary Note 1). Bl. nov. exhibited significantly lower predicted phenotypic richness than other subspecies (Fig. 2b), lacking pathways for LNB, GNB, N-glycan, HMO and T-antigen (Tan) metabolism (Fig. 2c and Supplementary Note 5). Conversely, only Bl. nov. genomes encoded extracellular amylopullulanase ApuB (GH13_14_32), a bifunctional glycoside hydrolase (GH) cleaving both α-1,4 and α-1,6-glycosidic bonds in soluble starch (ST) and pullulan (PUL)44, and a pathway for difructose dianhydride (DFA) metabolism45. These findings suggest that Bl. nov. has a reduced capacity to metabolize host-derived glycans but can degrade α-glucans of plant and fungal origin.
Comparative analysis revealed stark differences in the repertoire of carbohydrate utilization pathways between Bl. infantis and Bl. longum (Fig. 2c). Consistent with previous studies, Bl. infantis genomes were distinguished by the presence of H146 and FL1/2 (ref. 4) gene clusters, which enable the utilization of lacto-N-neotetraose (LNnT), 2′FL, 3FL, lactodifucotetraose (LDFT), lacto-N-fucopentaose I (LNFP I) and sialylated HMOs (SHMOs; Supplementary Tables 7–10). Predicted HMO utilization potential of Bl. longum was more limited: 35% of genomes encoded extracellular lacto-N-biosidase LnbX (GH136) that cleaves lacto-N-tetraose (LNT) and LNFP I47, and only 2.3% carried a gene cluster driving intracellular utilization of 2′FL, 3FL, LDFT and LNFP I48,49. Beyond HMO metabolism, Bl. infantis exclusively encoded catabolic pathways for glucuronate (GlcA), inositol (Ino) and gluconate (Gco) in 63%, 48% and 23% of genomes, respectively (Extended Data Fig. 4a,b,d,e and Supplementary Note 3). Bl. longum genomes, by contrast, commonly encoded pathways for l-arabinose (Ara), α/β-arabinooligosaccharides (aAOS/bAOS), type II arabinogalactan (AGII), arabinan (AR), arabinoxylan (AX) and host- or plant-derived O-glycans (Tan and HRGP; Fig. 2c). These findings illustrate an ecological divergence between Bl. infantis and Bl. longum shaped by their respective adaptations to thrive on milk glycans during breastfeeding versus plant-derived carbohydrates after weaning.
Predicted glycan utilization profiles within the Bl. suis group were highly heterogeneous. Most genomes encoded a set of metabolic pathways similar to that of Bl. longum but more frequently included pathways for the utilization of N-acetylneuraminic acid (Neu5Ac; 44% versus 2%), l-fucose (Fuc; 61% versus 1.7%) and fucosylated HMOs (2′FL, 3FL, LDFT and LNFP I; 56% versus 2.3%), while lacking genes encoding extracellular α-l-arabinofuranosidases required for AX degradation50,51 (Fig. 2c and Supplementary Tables 8 and 10). By contrast, the Bangladeshi isolate Bg131.S11_17.F6 shared multiple genomic features with Bl. infantis, including the presence of the H1 gene cluster and the absence of araBDA genes (Supplementary Table 8). Consequently, this strain was predicted to metabolize more HMO structures (for example, LNnT and SHMOs) than other Bangladeshi Bl. suis strains, while lacking the capacity to utilize arabinose-containing glycans of plant origin (for example, Ara, aAOS and AGII; Supplementary Fig. 3). Genomic analysis of ten additional animal-derived Bl. suis strains confirmed the uniqueness of the Bg131.S11_17.F6 isolate and further underscored the phenotypic heterogeneity within this group (Extended Data Fig. 5 and Supplementary Note 5). Collectively, these findings reveal pronounced differences in carbohydrate utilization across B. longum subspecies and underscore pervasive strain-level heterogeneity within each clade.
Strain-level heterogeneity of glycan metabolism
Beyond interspecies differences, we observed extensive strain-level variability in predicted carbohydrate utilization capabilities. Among the 68 reconstructed pathways, 66 exhibited variability within at least one taxonomic group (Fig. 2c and Supplementary Tables 7 and 9). The genomic differences driving this heterogeneity ranged from individual genes encoding extracellular GHs enabling polysaccharide degradation to multigene clusters comprising up to 20 genes encoding complete metabolic pathways (Supplementary Table 8). By contrast, biosynthetic pathways for essential metabolites, such as amino acids and B vitamins, were largely conserved. A few exceptions included riboflavin (B2) biosynthesis, which varied across Bl. suis and Bc. kashiwanohense_A, and thiamine (B1) and niacin (B3) biosynthesis in Bifidobacterium adolescentis (Extended Data Fig. 6 and Supplementary Table 11).
The observed heterogeneity reflects that, while most Bifidobacterium taxa follow general ecological strategies centred on the utilization of specific core glycans, individual strains can exhibit substantial metabolic tuning. For instance, Bl. infantis Bg064.S07_13.C6 harboured pathways for xylooligosaccharide (XOS) and long-chain fructooligosaccharide (lcFOS) utilization, suggesting an enhanced capacity to metabolize dietary plant glycans compared with most other Bl. infantis strains (Supplementary Fig. 3). Conversely, several B. adolescentis isolates carried pathways for the utilization of LNB, GNB, N-glycans (strains AF96-10M2bTA and UN03-88) and fucosylated HMOs (strain M56B_1C3), highlighting the presence of traits characteristic of infant-adapted taxa in a species commonly associated with the adult gut (Supplementary Tables 9 and 10). Additional examples of notable strain-specific variability of genomic features related to carbohydrate utilization were found in Bangladeshi isolates and are detailed below.
Unique glycan utilization features of Bangladeshi strains
Our previous study described distinctive genomic features of Bangladeshi Bl. infantis strains related to N-glycan and β-glucoside catabolism23. Here, we identified a distinct gene cluster (xgl) in Bc. kashiwanohense Bg42221_1E1 and Bc. kashiwanohense_A Bg42221_1D3, two isolates from a Bangladeshi infant (Fig. 3a). This cluster encoded multiple GHs (families 3, 5_4, 29, 31, 42 and 43_12), an unclassified carbohydrate esterase (CE), a β-glucoside kinase and an ABC transport system, which together may catabolize plant hemicelluloses such as xyloglucans (XGLs). The reconstructed XGL pathway involved the hydrolysis of the XGL backbone to oligosaccharides by extracellular endo-β-1,4-glucanase Xgl5A (GH5_4), which shares catalytic and glycan-binding residues with xyloglucanase PpXG5 from Paenibacillus pabuli XG552 (Extended Data Fig. 7a). Released oligosaccharides would be imported into the cell and degraded sequentially to individual monosaccharides and cellobiose by the coordinated action of GHs and CEs via a mechanism similar to that described for Ruminiclostridium cellulolyticum53 (Fig. 3b). Regulon reconstruction of XglT, a putative TetR-family transcription factor, suggested potential co-regulation of the xgl cluster and cbpA, which encodes a GH94-family cellobiose phosphorylase (Fig. 3a–c and Supplementary Table 19). The xgl cluster was identified in only 3 of 110 studied B. catenulatum genomes but was conserved in Bifidobacterium dentium and Bifidobacterium tsurumirense (Fig. 2c).
a, Gene clusters potentially driving XGL degradation in Bc. kashiwanohense Bg42221_1E1. b, Reconstructed XGL degradation pathway in Bc. kashiwanohense Bg42221_1E1: (1) XGL chains are potentially cleaved by extracellular endo-β-1,4-glucanase Xgl5A, and (2) released oligosaccharides are imported and metabolized inside the cell by a coordinated action of GHs, CEs and downstream catabolic enzymes. c, The predicted DNA-binding motif of the XglT transcriptional regulator potentially controlling XGL metabolism genes. d, Growth curves of Bifidobacterium strains in MRS-AC supplemented with 0.5% tamarind XGL. Data represent the mean ± s.d. of three biological replicates. e, A volcano plot showing differential gene expression (log2 fold change (FC) versus −log10-adjusted P value) in Bc. kashiwanohense Bg42221_1E1 grown in MRS-AC with tamarind XGL versus MRS-AC with Lac. Differential expression was assessed using moderated two-sided t-tests with empirical Bayes variance moderation. P values were adjusted for multiple comparisons using the Benjamini–Hochberg procedure. Genes were considered differentially expressed at Padj < 0.01 and |log2FC| > 2. Genes belonging to the reconstructed XglT, XylR and XosR regulons, as well as xyn cluster genes, are highlighted. Exact log2FC values, test statistics and adjusted P values are provided in Supplementary Table 21a.
Another gene cluster, unique to Bc. kashiwanohense Bg42221_1E1, contained orthologues of H1 cluster HMO utilization genes from Bl. infantis and the ‘outlier’ Bangladeshi Bl. suis Bg131.S11_17.F6 strain (Fig. 4a). The H1 variant identified in Bg42221_1E1 encoded two ABC transporters, one of which (HmoABC) is probably involved in the uptake of LNnT and other type II HMO structures54. In addition, this cluster encoded orthologues of characterized β-N-acetylglucosaminidase (GH20), two α-fucosidases (GH29 and GH95) and α-sialidase (GH33), which mediate intracellular HMO hydrolysis55,56,57, along with downstream catabolic pathways for GlcNAc and Fuc. Regulon reconstruction suggested the potential transcriptional control of H1 cluster genes in Bc. kashiwanohense Bg42221_1E1 by NagR, a GlcNAc-responsive ROK-family repressor58 (Fig. 4a and Supplementary Table 19).
a, A schematic representation of HMO utilization genes in selected Bangladeshi Bifidobacterium strains and their homology to H1 cluster genes from Bl. infantis ATCC 15697. The H1 cluster encodes multiple paralogous substrate-binding proteins (SBPs) of putative HMO transporters (Hmo), whereas FHMO encodes a characterized, non-orthologous transporter (FL2_ABC) specific for fucosylated HMOs. b, Growth curves of Bifidobacterium strains in MRS-AC supplemented with pooled HMOs. Data represent the mean ± s.d. of three biological replicates. Curves are colour-coded based on the presence of specific HMO utilization gene clusters in the respective strains. c, HPLC-based quantification of HMO depletion from culture supernatants after 24 h. Data represent the percentage of utilized HMOs (mean of three biological replicates) relative to the medium control. Total HMO, total HMO utilized; total FHMO, total fucosylated HMO utilized; total SHMO, total sialylated HMO utilized. Concentrations of individual HMOs (nmol ml−1) are provided in Supplementary Table 18b. d, PCA of combined growth metrics (maximum OD600, area under the curve and maximum growth rate) and per cent utilization of individual HMOs. Each point represents a strain, colour-coded by the presence of specific HMO utilization gene clusters. Axes represent principal components (PCs). In b and d, the notations H1 cluster+ and FHMO cluster+ indicate that a strain carries the complete H1 or FHMO gene cluster, respectively. e, A volcano plot showing differential gene expression (log2FC versus −log10-adjusted P value) in Bc. kashiwanohense Bg42221_1E1 grown in MRS-AC with LNT versus MRS-AC with Lac. Differential expression was assessed using moderated two-sided t-tests with empirical Bayes variance moderation. P values were adjusted for multiple comparisons using the Benjamini–Hochberg procedure. Genes were considered differentially expressed at Padj < 0.01 and |log2FC| > 2. Genes belonging to the reconstructed NagR regulon are highlighted. Exact log2FC values, test statistics and adjusted P values are provided in Supplementary Table 21b.
Other notable features of Bangladeshi Bifidobacterium isolates included a putative d-galactonate utilization pathway found exclusively in B. breve Bg41721_1C11 (Extended Data Fig. 4d,e and Supplementary Note 3). This strain, along with B. breve Bg131.S11_D6, also lacked the nan gene cluster, which encodes a well-characterized Neu5Ac catabolic pathway59 (Extended Data Fig. 4f,g). This absence of the nan cluster in these two strains was unexpected, given its previously reported broad conservation in B. breve genomes29,59 and its established role in utilizing Neu5Ac via cross-feeding on SHMOs and mucin O-glycans degraded by B. bifidum59,60.
These results demonstrate that Bangladeshi Bifidobacterium strains carry unique adaptations for metabolizing dietary plant polysaccharides and HMO, while lacking pathways conserved in well-characterized strains. These traits may reflect strain-level adaptation to the diet and lifestyle of Bangladeshi children. We next investigated whether broader patterns of bifidobacterial carbohydrate utilization are associated with host age and lifestyle across diverse human populations.
Associations between pathway profiles and lifestyle
We examined the enrichment of glycan utilization pathways in Bifidobacterium genomes from different populations based on: (1) host age and stage of gut microbiota maturation (<3 years: infant or transitional; ≥3 years: adult-like), and (2) host lifestyle (‘Westernized’ versus ‘non-Westernized’ as defined by Pasolli et al.61). Host glycan (for example, HMO) utilization pathways were enriched in genomes from the ‘age <3’ group, whereas plant glycan utilization pathways were more prevalent in adult-associated (≥3 years) genomes across both lifestyle groups (Fisher’s exact test, Benjamini–Hochberg adjusted P ≤ 0.01; Extended Data Fig. 8a,b). Within the ‘age <3’ group, 14 pathways (11 for plant glycans) were enriched in the Westernized group and 24, including 11 for HMOs and their constituent blocks LNB, Neu5Ac and Fuc, in the non-Westernized group (Extended Data Fig. 8c). These differences probably stem from the uneven distribution of taxa across Westernized and non-Westernized microbiotas. For example, pathways enriched in the non-Westernized group were associated with Bl. infantis, a taxon more prevalent in that group (odds ratio 4.98, 95% confidence interval 3.24–7.51, Fisher’s exact test, P = 1.82 × 10−11).
Within taxa, pathways for sorbitol (Stl), mannitol (Mtl), lcFOS and pectic galactan (PG) metabolism were enriched in Westernized Bl. infantis genomes (Extended Data Fig. 8d). The Stl and lcFOS catabolic pathways were also more prevalent in Westernized B. adolescentis genomes, and the PG pathway was overrepresented in Westernized Bl. longum genomes. Conversely, non-Westernized B. breve genomes were enriched for melezitose (Mlz) and 1,2-β-oligoglucan (BGL12) utilization pathways but more frequently lacked the Neu5Ac pathway. These findings underscore how lifestyle-driven ecological pressures shape glycan utilization strategies among bifidobacteria. We next tested carbohydrate utilization phenotypes in representative isolates to validate predictions from the reconstruction framework.
Growth-based validation of predicted glycan utilization
To experimentally validate in silico phenotypic predictions, we conducted in vitro growth assays on 30 geographically diverse Bifidobacterium strains (15 Bangladeshi, 8 Malawian, 5 US and two European). Growth was tested on 43 substrates corresponding to 38 predicted glycan utilization phenotypes: 13 monosaccharides and derivatives, 18 di- or oligosaccharides and 7 polysaccharides (Fig. 5a). Strains were cultured in a sugar-free De Man–Rogosa–Sharpe (MRS-AC) medium4,58 supplemented with the test substrate (5–10 mg ml−1), and growth was defined using strain-specific optical density at 600 nm (OD600) thresholds (Methods and Supplementary Fig. 4). Growth outcomes were compared with predictions derived from manual curation (20 strains) or the automated pipeline (10 strains; Fig. 5b).
a, A summary of in vitro growth profiles of 30 Bifidobacterium strains on 43 substrates. Growth, weak growth and no growth were categorized on the basis of strain-specific OD600 thresholds (Methods). The annotation column on the right shows the geographical origin of each strain. The details about substrates are given in Supplementary Table 15; all growth curves are shown in Supplementary Fig. 4; raw OD600 data are provided in Supplementary Table 16. b, A comparison of 38 predicted carbohydrate utilization phenotypes with corresponding in vitro growth profiles for the same 30 strains. Predicted phenotypes were tested using a singular substrate, except IMO (panose and isomaltotriose), BglOS (cellobiose, gentiobiose, laminaritriose and sophorose) and SHMO (3′SL and 6′SL). Prediction outcomes are colour-coded to indicate agreement between predicted and observed phenotypes. The annotation column on the right shows whether genomes belong to the reference or extended dataset. The annotation row at the bottom represents phenotype classification (glycan type). Summary data are provided in Supplementary Table 17f; full names of abbreviations are provided in Supplementary Table 5.
Prediction accuracy was similar for manual and automated approaches: 95% and 94%, respectively, with Matthews correlation coefficients of 0.9 and 0.89 (Supplementary Table 17e,f). False-negative predictions (growth despite predicted non-utilization) probably stemmed from incomplete knowledge about monosaccharide and HMO transporters. For example, Bl. suis Bg131.S11_17.F6 grew on 2′FL despite lacking genes encoding known transporters for this substrate4,5 (Supplementary Table 8). Some false-positive predictions (predicted utilization but no growth) may have resulted from gene disruptions, such as a premature stop codon in the gltA gene in B. pseudocatenulatum LFYP29, which probably impaired the LNT transport function.
As predicted, Bl. nov. LFYP82 was the only B. longum strain to grow on ST and PUL (Fig. 5a). We also validated strain-specific utilization of scFOS, lcFOS, XOS and GlcA in Bl. infantis (Fig. 5 and Extended Data Fig. 4c). Similarly, while two B. breve strains grew on mannotriose (bMnOS), none grew on konjac glucomannan (bMAN), in contrast to B. dentium LFYP24 and Bifidobacterium scardovii JCM 12489, which harboured extracellular endo-β-1,4-mannanases (Extended Data Fig. 7e–g and Supplementary Note 2).
We further validated several unusual glycan utilization traits predicted in Bangladeshi and Malawian isolates. Bc. kashiwanohense Bg42221_1E1 and Bc. kashiwanohense_A Bg42221_1D3, both carrying the xgl cluster, grew in the medium supplemented with tamarind XGL (Fig. 3d). Bangladeshi B. breve isolates lacking the Neu5Ac catabolic pathway (Bg41721_1C11 and Bg131.S11_D6) failed to grow on Neu5Ac, unlike other tested B. breve strains (Extended Data Fig. 4h). Finally, consistent with our prediction, B. adolescentis M56B_1C3 grew on 2′FL, highlighting the previously unrecognized capacity of this species (Fig. 5).
Glycoprofiling of HMO utilization
The H1 cluster in Bl. infantis is believed to enable the metabolism of multiple HMOs46, although the precise range of structures cannot be confidently predicted due to limited understanding of the functions of individual transporter genes. Given the presence of homologous H1 clusters in Bl. suis Bg131.S11_17.F6 and Bc. kashiwanohense Bg42221_1E1, we compared their HMO utilization with that of (1) two Bl. infantis strains, (2) phylogenetically related strains carrying a fucosylated HMO utilization gene cluster (FHMO) instead of H1 (Bl. suis Bg41121_2E1 and Bc. kashiwanohense_A Bg42221_1D3; Fig. 4a), and (3) distantly related strains predicted to have weak (Bl. longum and B. breve) or minimal (Bl. nov. and B. pseudocatenulatum) HMO utilization capacity.
Strains with the H1 cluster reached the highest optical densities when grown in MRS-AC supplemented with an HMO mixture isolated from pooled human milk (Fig. 4b). High-performance liquid chromatography (HPLC)-based glycoprofiling of culture supernatants revealed that these strains consumed 72–86% of total HMOs by 24 h, including multiple fucosylated and sialylated structures (Fig. 4c and Extended Data Fig. 9). However, Bl. suis Bg131.S11_17.F6 and Bc. kashiwanohense Bg42221_1E1 did not efficiently utilize 2′FL, the most abundant HMO species in the mixture, in line with the absence of characterized 2′FL transporters. By contrast, strains carrying the FHMO cluster preferentially metabolized fucosylated HMOs, completely depleting 2′FL while exhibiting limited utilization of sialylated structures. Among strains negative for both H1 and FHMO clusters, Bl. longum Bg115.S08_3A11 depleted LNT, LNnT, LNFP I, LNFP III and lacto-N-hexaose (LNH), probably via partial extracellular degradation by β-galactosidase and lacto-N-biosidase, whereas B. breve Bg155.S08_4F7 metabolized LNT and LNnT intracellularly (Supplementary Fig. 5). Bl. nov. LFYP82 failed to efficiently utilize any tested HMOs, consistent with lineage-specific loss of relevant genes and pathways. Principal component analysis (PCA) of growth and HMO utilization data separated strains with H1 and FHMO clusters from each other and all others (Fig. 4d). These findings provide the experimental evidence that Bc. kashiwanohense and Bl. suis strains carrying the H1 cluster exhibit HMO utilization patterns comparable to Bl. infantis, and highlight how variation in gene repertoires drives metabolic divergence among closely related Bifidobacterium strains.
Transcriptional profiles of glycan utilization in Bc. kashiwanohense
We used RNA sequencing (RNA-seq) to test regulon predictions and pathway assignments in Bc. kashiwanohense Bg42221_1E1, which harbours both xgl and H1 gene clusters. Transcriptomic comparison of cultures grown in MRS-AC-XGL versus MRS-AC-Lac revealed strong induction (100–550 fold) of most xgl genes and cbpA, supporting their co-regulation by XglT and a shared role in XGL metabolism (Fig. 3a,e and Supplementary Table 21a). Genes involved in xylose, (arabino)xylooligosaccharide and (arabino)xylan metabolism showed moderate upregulation (4–30 fold), probably in response to intracellular xylose release, which may serve as a transcriptional effector of predicted transcriptional repressors XylR (ROK family) and XosR62 (LacI family; Extended Data Fig. 7b,c, Supplementary Table 19 and Supplementary Note 2). Next, we compared the transcriptomes of Bc. kashiwanohense Bg42221_1E1 grown in MRS-AC-LNT and MRS-AC-Lac. All H1 cluster genes except nanA2 and nanH2 were significantly upregulated in the presence of LNT (Fig. 4e and Supplementary Table 21b), consistent with the proposed regulation by NagR, a GlcNAc-responsive repressor implicated in the control of HMO utilization in bifidobacteria58. Overall, the observed transcriptomes suggest that the regulatory networks of this strain are adapted for foraging on mixtures of HMOs and plant oligo- and polysaccharides.
Our large-scale genomic and experimental analyses pinpoint ecological differences between glycan foraging strategies within Bifidobacterium that reflect species-level evolutionary adaptation to different habitats (for example, infant gut versus adult gut) and dietary carbohydrate composition. The results also underscore considerable strain-level variability, probably shaped by host lifestyle and local dietary exposures.
Discussion
The ability to metabolize dietary glycans is central to bifidobacterial fitness in the human gut, with 8–12% of their gene content dedicated to carbohydrate metabolism63. Understanding this process has fundamental and translational relevance, including for the development of probiotics and synbiotics. While interspecies differences in glycan preferences within bifidobacteria are well documented25,64,65, within-species variability, particularly in populations from low- and middle-income countries, remains understudied. The growing availability of Bifidobacterium genomes, driven by culturomics66,67,68,69 and metagenomics70,71,72,73,74,75, provides an opportunity to address this gap. However, accurately predicting carbohydrate utilization phenotypes from genomic data remains challenging due to imprecise functional annotations (particularly of glycan transporters) generated by widely used pipelines, as well as incomplete representation of metabolic pathways in public databases34,76,77. To address this gap, we reconstructed 68 carbohydrate utilization pathways encoded in 3,083 Bifidobacterium genomes and MAGs and validated 38 predicted phenotypes in vitro, with an overall accuracy exceeding 94%. Several false-negative predictions for specific mono- and oligosaccharides, including HMOs, suggest the existence of yet uncharacterized glycan transport mechanisms in bifidobacteria.
Our analysis revealed taxon-specific distribution of glycan utilization pathways, reflecting the adaptation of Bifidobacterium species and subspecies to distinct ecological niches, such as the infant and adult gut, and expanding on earlier observations25,30,64,65. For instance, we identified a distinct clade within the B. longum species (Bl. nov.) specialized in metabolizing plant and fungal α-glucans but incapable of utilizing host glycans such as LNB, GNB and HMOs. As most Bl. nov. genomes originated from infants in Westernized populations, the unique glycan preferences of this clade might reflect an adaptation to differences in early-life diet and feeding practices. However, certain pathways, such as RFO utilization, were widely conserved. A recent study demonstrated that RFO metabolism supports B. breve colonization and persistence in vivo19, suggesting that conservation of this pathway across bifidobacterial lineages may promote their persistence and transmission beyond infancy.
Another important observation was extensive strain-level variability driven by accessory genome differences ranging from single genes to gene cassettes encoding entire carbohydrate utilization pathways. Particularly notable was the heterogeneity of Bl. suis isolates of Bangladeshi origin. Similar strains were recently predicted to utilize HMOs and plant fibres based on the analysis of CAZyme repertoires and reclassified as B. longum subsp. iuvenis7,43. Our analysis revealed nuanced metabolic differences within this clade, suggesting the presence of two distinct ‘ecotypes’. The first, exemplified by strain Bg131.S11_17.F6, metabolizes a wide array of HMOs—including LNT, LNnT, long-chain fucosylated and sialylated species—but not arabinose and plant polysaccharides, mimicking the glycan preferences of Bl. infantis. The second ecotype, represented by strain Bg41121_2E1, preferentially utilizes fucosylated HMO structures, including the abundant 2′FL, but can metabolize arabinose-containing oligo- and polysaccharides, phenotypically resembling previously characterized Bl. longum strains APC1478, SC596 and MC1000728,48,49. These distinct and complementary strategies suggest ecological specialization within the Bl. suis group, potentially linked to diet and weaning status.
Further within-species variability is illustrated by Bangladeshi isolates carrying unique gene clusters. One notable example is the xgl cluster identified in Bc. kashiwanohense Bg42221_1E1 and Bc. kashiwanohense_A Bg42221_1D3 isolated from the same infant. This cluster enables XGL degradation, a capability previously suggested only for Bifidobacterium species inhabiting captive marmosets78. Another unexpected discovery was the presence of an H1 cluster variant in Bc. kashiwanohense Bg42221_1E1, formerly considered to be exclusive for Bl. infantis46 and rare Bangladeshi Bl. suis group strains7,23. Although the precise substrate specificities of transporters encoded by H1 cluster remain to be characterized, our data confirm that its presence enables the utilization of major HMOs found in human milk. Bc. kashiwanohense strains have been shown to metabolize fucosylated HMOs (2′FL and LNFP I) and plant hemicelluloses (xylan and AX)8. Our findings expand the catabolic potential of Bc. kashiwanohense to XGL and a broader range of HMOs (LNT, LNnT and SHMOs). These results indicate an evolutionary adaptation of Bangladeshi strains to thrive in the microbiota of weaning children, whose diets may combine breast milk with complementary foods rich in plant polysaccharides.
Although we did not explicitly reconstruct cross-feeding interactions, the lower prevalence of complete catabolic pathways for polysaccharides, relative to those targeting their mono- and oligomeric breakdown products, suggests that many Bifidobacterium strains may depend on syntrophic relationships, both within the genus and with keystone degraders such as Bacteroides and Segatella (formerly Prevotella)79,80,81. Recent evidence for widespread HMO-degrading capacity across gut microbes82 further supports the potential for cross-feeding involving bifidobacteria and other community members.
The genomic dataset analysed in this study is skewed towards samples from high-income countries, highlighting the need for initiatives to expand collections of Bifidobacterium isolates and genomes from underrepresented populations in an ethical and culturally sensitive manner to capture global metabolic diversity83. Some observed within-species variability may reflect artefacts introduced by incomplete or contaminated MAGs and different assembly methods, underscoring the importance of expanding collections of high-quality MAGs and isolate genomes.
In summary, our study reveals how glycan foraging strategies vary across and within Bifidobacterium species, shaped by ecological factors, including host age, diet and lifestyle. The comprehensive metabolic reconstruction and an automated pathway prediction pipeline provide a scalable framework for accurate functional annotation of bifidobacterial genomes and MAGs. This genomic compendium may inform the development of pro- and synbiotic formulations, including multistrain consortia, whose members might efficiently colonize the gut across diverse lifestyle and dietary contexts.
Methods
Collection of Bifidobacterium genomes
Reference genomes for the manual in silico metabolic reconstruction were retrieved from Bacterial and Viral Bioinformatics Resource Center (BV-BRC)84 and Integrated Microbial Genomes (IMG)85 databases as of October 2020. We selected 335 genomes on the basis of the following criteria: (1) human or probiotic product-derived Bifidobacterium spp. isolates, (2) number of contigs ≤200 and (3) CheckM86 completeness ≥97% and contamination ≤3%. In addition, we sequenced 31 genomes of cultured isolates from faecal samples obtained from Bangladeshi children37,38. The resulting reference set of 366 genomes was clustered using dRep (v3.4.2)87 at a 99.95% ANI threshold (‘dRep dereplicate -pa 0.9 -sa 0.9995 -nc 0.35 --S_algorithm ANImf’), yielding 263 non-redundant genomes (Supplementary Table 2).
Additional human gut-derived Bifidobacterium genomes were collected from prior studies. These included isolate genomes from Human Gastrointestinal Bacteria Culture Collection (HBC)66, Broad Institute-OpenBiome Microbiome Library (BIO-ML)67, Culturable Genome Reference (CGR)68 and Human intestinal Bacteria Collection (HiBC)69, and from Unified Human Gastrointestinal Genome (UHGG)70, Human Reference Gut Microbiome (HRGM)71, Early-Life Gut Genomes (ELGG)72, Singapore Platinum Metagenomes Project (SPMP)73 and Inner Mongolian Gut Genome (IMGG)74 collections and other datasets75. We used the same selection criteria for both isolate genomes and MAGs: (1) Bifidobacterium spp. based on Genome Taxonomy Database (GTDB) taxonomy88 excluding Bifidobacterium leopoldii and Bifidobacterium vaginale, (2) number of contigs ≤200 and (3) CheckM completeness ≥97% and contamination ≤3%. The resulting 4,944 genomes were clustered via dRep using the command ‘dRep dereplicate -pa 0.9 -sa 0.999 -nc 0.35 --S_algorithm fastANI’, yielding 2,811 non-redundant genomes (Supplementary Table 3). We also sequenced nine genomes of strains isolated from Bangladeshi and Malawian infants previously37,38,39, resulting in 2,820 additional genomes.
Isolation and sequencing of Bifidobacterium strains from Bangladeshi and Malawian donors
Faecal samples used for culturing Bangladeshi bifidobacterial strains were collected during studies conducted by the International Centre for Diarrhoeal Disease Research (icddr,b). These were (1) the MAL-ED birth cohort study of children aged 0–24 months (Interactions of Enteric Infections and Malnutrition and the Consequences for Child Health and Development; ClinicalTrials.gov identifier NCT02441426) and (2) a cohort of healthy 12–24-month-old Bangladeshi children enrolled in parallel with children with acute malnutrition in a study of microbiota-directed complementary food prototypes (ClinicalTrials.gov identifier NCT03084731)37,38. Both studies were approved by the Ethical Review Committee of the icddr,b. Bifidobacterial strains were also isolated from faecal samples collected in a previously reported study of Malawian twins discordant for acute malnutrition39. The protocol was approved by the College of Medicine Research Ethics Committee of the University of Malawi and by the Human Research Protection Office of Washington University in St. Louis. Written informed consent, including provisions for future use of materials, was provided by the parents or guardians of participating children before enrolment. The details of genome sequencing and assembly are provided in Supplementary Methods.
Taxonomy inference
Initial taxonomic assignments for the 263 reference Bifidobacterium genomes were retrieved from the National Center for Biotechnology Information and BV-BRC for public genomes or inferred from 16S rRNA gene sequencing for Bangladeshi isolates. To refine these assignments, we built a pangenome via Panaroo (v1.3.2)89. Concatenated nucleotide sequences of 487 identified core genes (Supplementary Table 12) were aligned using MAFFT (v7.515)90. A maximum-likelihood phylogenetic tree was built in IQ-TREE (v2.2.0.3)91 and visualized via iTOL (v5)92. Pairwise ANI values (ANIb) were calculated using pyani (v0.2.12)93. The exact commands and parameters are provided in Supplementary Code File 1. The resulting tree topology was manually inspected to verify and correct taxonomic assignments of genomes based on their co-clustering with branches corresponding to the type or well-characterized strains of Bifidobacterium species and subspecies. ANI matrices were used to delineate the within-species structure for Bifidobacterium longum and Bifidobacterium catenulatum species. GTDB-based taxonomies of 2,820 additional genomes were retrieved from original publications and refined via ANI comparisons with 263 reference genomes. Species assignments were made if ANI to the closest reference genome was >95%, and subspecies assignments if >97%.
Gene prediction and functional annotation
Protein-coding sequences were predicted and annotated with Prokka (v1.14.6)94 using default settings. The 263 reference genomes were additionally annotated via RASTtk (v1.073)95 and EggNOG-mapper (v2.1.12)96 with default settings (Supplementary Table 13). The representation of CAZymes in 263 genomes was analysed using dbCAN (v4.0.0)97. HMMER, dbCAN-sub and DIAMOND-based searches against the CAZy database were used for CAZyme identification. Only GHs, CEs and polysaccharide lyases predicted by two or more methods were retained.
Subsystems-based annotation and in silico metabolic reconstruction of glycan utilization pathways
We used a subsystem-based approach implemented in the SEED platform35,95 to reconstruct carbohydrate utilization pathways in reference 263 Bifidobacterium strains. RASTtk-annotated genomes were uploaded to microbial community SEED (mcSEED), a clone of the SEED annotation environment. We created 25 subsystems that captured the representation of genes that implement functional roles (glycan transporters, CAZymes, downstream catabolic enzymes and transcriptional regulators) involved in carbohydrate utilization (Supplementary Table 4). The list of functional roles was compiled via extensive literature search and using the information from Transporter Classification (TCDB)98, Carbohydrate Active Enzyme (CAZy)99, Kyoto Encyclopedia of Genes and Genomes (KEGG)100 and RegPrecise101 databases. This knowledge was used to manually curate the automated functional annotations of protein-coding genes in a subset of reference Bifidobacterium genomes (for example, type strains) where the respective functional roles had been experimentally characterized. Additional functional gene annotation to fill gaps in metabolic pathways was based on three genome context techniques: (1) clustering of genes on the chromosome (operons), (2) co-regulation of genes by a common transcription factor and (3) co-occurrence of genes across related genomes. The approach used to reconstruct the regulons of transcription factors potentially regulating carbohydrate metabolism is outlined in Supplementary Methods.
The propagation of curated annotations (corresponding to 589 distinct functional roles) across all 263 reference genomes was performed using homology-based methods implemented in mcSEED. Orthologues were detected automatically using predefined protein family classifications such as PGFams (cross-genus protein families) and PLFams (genus-specific protein families)84. These assignments were manually refined by examining the gene neighbourhood for each functional role. Genes with conserved gene neighbourhoods were classified as orthologous, while paralogues were assigned distinct functional annotations. Overall, 39,589 out of 541,418 protein-coding gene annotations in 263 reference Bifidobacterium genomes were curated.
Next, we reconstructed 72 catabolic pathways spanning 25 subsystems (Supplementary Table 5). Many pathways included alternative biochemical modules (routes) driven by different sets of catabolic enzymes and diverse glycan transporters. For each pathway, we defined genomic signatures—sets of genes encoding functional roles that together represent the minimal gene complement required to form a complete pathway for the utilization or degradation of a specific carbohydrate (Supplementary Table 6). ‘Utilization’ was defined as a process in which a glycan molecule (mono- or oligosaccharide) is transported into the cell and then catabolized using a combination of CAZymes and downstream catabolic enzymes. ‘Degradation’ referred to the partial hydrolysis of a glycan (for example, a polysaccharide) by extracellular CAZymes, with the resulting mono-, di- and oligosaccharides subsequently transported into the cell and metabolized.
We used the presence or absence of genes matching each genomic signature to assign a detailed pathway variant to each genome: (1) transporter + catabolic pathway (‘U’), (2) catabolic pathway without transporter (‘P’) and (3) no catabolic pathway (‘N’) (Supplementary Tables 6 and 8). For the purpose of automated phenotype profiling, these assignments were simplified into binary phenotypes: a complete pathway (‘U’) indicated predicted utilization or degradation and was assigned phenotype ‘1’, while incomplete (‘P’) or missing (‘N’) pathway variants indicated no utilization and were assigned phenotype ‘0’. The resulting set of 72 predicted carbohydrate utilization phenotypes across 263 strains comprised the reference BPM (Supplementary Table 7). Four pathways (GalNAc, ManAc, Man and GalA), for which all strains in the reference set were assigned variants ‘P’ or ‘N’ and phenotype ‘0’, respectively, were retained in the BPM but excluded from the pathway prediction pipeline.
Pathway prediction pipeline (glycobif)
The workflow used to predict the carbohydrate utilization pathways encoded in additional 2,820 Bifidobacterium genomes is schematically illustrated in Fig. 1a. We first constructed a reference database containing (1) 39,589 functionally annotated protein sequences from 25 curated metabolic subsystems across 263 reference genomes and (2) an additional set of 52,990 outgroup proteins (not captured in the subsystems) clustered at 95% amino acid identity and 95% coverage using MMSeqs2 (v14.7e284)102. Proteins encoded in the 2,820 query genomes were annotated by mapping their sequences to the reference database using DIAMOND (v2.1.4)103. To handle multidomain proteins, we first selected the top 50 hits for each query based on bitscore and clustered the alignment coordinates using DBSCAN (scikit-learn v1.2.1)104. Cluster centres were treated as potential domain boundaries, which were used to split query proteins into discrete domains, with database hits attributed to each individual domain. For each resulting domain with ≥35 amino acids, we applied the Gaussian kernel density modelling (KernelDensity function from sklearn.neighbors) to the distribution of sequence identity values and used the highest local minimum (argrelextrema function from sklearn.signal) to filter out low-confidence hits. Annotations were assigned by majority vote from high-scoring, domain-specific reference hits. High-identity hits to outgroup proteins from the reference database were used as criteria to vote against applying annotation to a given query. This pipeline yielded 419,055 annotated protein sequences.
These annotations, together with the BPM for 263 reference genomes, were integrated into a machine learning-based pipeline to predict the presence (‘1’) or absence (‘0’) of carbohydrate utilization pathways in additional 2,820 genomes. We evaluated over 30 machine learning methods implemented in the Caret package (v6.0.86)105 using a leave-one-out cross-validation approach: for each reference genome, a model was trained on the remaining 262 genomes and then used to predict the binary variant for the held-out genome. This process was repeated for every pathway. Random forest was identified as the best-performing model based on prediction accuracy. Pathway-specific random forest models were then trained using the full reference set, excluding four pathways (GalNAc, ManAc, Man and GalA) for which all genomes had predicted binary phenotype ‘0’. The list of functional roles used as model predictors for each remaining pathway was manually curated to match the genomic signatures delineating pathway variants (Supplementary Table 6). Model parameters were optimized using grid search, and mock genomes with custom genomic signatures were added to the training set to ensure that rare combinations from the reference collection were adequately learned and to improve model performance on incomplete MAGs. The resulting models were used to predict the presence or absence of 68 carbohydrate utilization pathways in 2,820 genomes (Supplementary Table 9).
The representation of 29 additional metabolic pathways (for example, the biosynthesis of B vitamins and amino acids) in both 263 reference and 2,820 additional genomes was inferred via a similar annotation pipeline as recently published106 (Supplementary Table 11).
Visualization and statistical analysis
NMDS was used for the ordination of the Hamming distance matrix calculated from the merged BPM of 3,083 genomes. PERMANOVA and a test for the homogeneity of multivariate dispersions were performed using the adonis2 and betadisper functions in the vegan package (v2.6-8)107. Differences in predicted phenotypic richness were assessed via a GLM assuming Poisson distribution; post-hoc comparisons with Bonferroni correction were conducted using the emmeans package (v1.10.6)108. Fisher’s exact test was used for the enrichment analysis of pathway representation. For each of the 68 carbohydrate utilization pathways, 2 × 2 contingency tables were constructed. Rows denoted the predicted absence or presence of pathways (‘0’ or ‘1’), whereas columns represented metadata categories: age group (age <3 years versus age ≥3 years) or host lifestyle (Westernized versus non-Westernized)61. P values were corrected for multiple testing using the Benjamini–Hochberg procedure, with adjusted P ≤ 0.01 considered significant (Supplementary Table 14). Odds ratios were calculated using the fisher.test function. PCA was used for the ordination of growth and HMO consumption data. Data visualization was performed using ggplot2 (v3.5.1)109, ComplexHeatmap (v2.18.0)110 and clinker (v0.0.27)111. Code for all analyses is provided in Supplementary Code File 1.
In vitro growth of Bifidobacterium strains on selected carbohydrates
All strains except B. scardovii JCM 12489 were routinely grown in dextrose-free lactobacilli De Man–Rogosa–Sharpe broth (Alpha Biosciences) supplemented with 0.34% (wt/vol) sodium ascorbate, 0.029% (wt/vol) l-cysteine–HCl monohydrate and 10 mg ml−1 Lac (MRS-AC-Lac) in a chamber maintained with a gas mix of 10% H2, 10% CO2 and 80% N2 (Coy Laboratory Products). The MRS-C-Lac medium without sodium ascorbate was used for B. scardovii JCM 12489. The growth of 30 Bifidobacterium strains on 47 substrates (Supplementary Table 15) was measured using a custom carbohydrate array constructed in flat-bottom half-area 96-well plates (Costar). Wells were filled with 55 µl of a sterilized carbohydrate stock at 2× concentration (10–20 mg ml−1) and transferred to the chamber 48 h before the experiment. Each substrate was tested in triplicate (or duplicate in a few cases). Sterile water and Lac served as negative and positive controls.
Cultures for assay inoculations were grown overnight at 37 °C in MRS-AC-Lac, diluted 1:100 into fresh medium, and incubated for 8 h. Cultures were then adjusted to OD600 of 0.6, and 100-μl aliquots were collected at 5,000g for 5 min and resuspended in 6 ml of 2× MRS-AC (2× MRS-C for B. scardovii) without added carbohydrate. Each well in the carbohydrate array was loaded with 55 µl of inoculated 2× medium to make individual 110-µl cultures. Growth was monitored by measuring OD600 every 15 or 30 min for 40 h under constant linear shaking using a Synergy H1 Plate Reader (BioTek Instruments). Raw OD600 data were exported using Gen5 software (v2.05.5) and analysed in R (v4.3.2). For all strains except B. pseudocatenulatum M26A_2F1 growth on each glycan was classified as follows: (1) no growth (‘−’) if the mean OD600 never exceeded 10% of the strain’s highest OD600 within a panel of carbon sources, (2) weak growth (‘w’) if the mean OD600 did not exceed 25% of maximum OD600, and (3) growth (‘+’) if the mean OD600 was above 25% of maximum OD600 (Supplementary Fig. 4 and Supplementary Table 16). For B. pseudocatenulatum M26A_2F1, thresholds were increased to 22% and 55%, respectively, owing to some outgrowth in basal MRS-AC lacking an added substrate.
Comparison of predicted phenotypes with growth data
Predicted carbohydrate utilization phenotypes were compared with growth data from this work or prior literature8,28,36. A predicted binary phenotype ‘1’ was classified as a true positive (TP) if it matched growth phenotypes ‘+/w’ and as a false positive (FP) if it matched ‘−’. A predicted binary phenotype ‘0’ was considered a true negative (TN) if it matched growth phenotype ‘−’ and a false negative (FN) if it matched ‘+/w’. Most predicted utilization phenotypes were tested using individual substrates. Exceptions included phenotypes IMO, for which the growth was tested on panose and isomaltotriose, BglOS (cellobiose, gentiobiose, laminaritriose and sophorose) and SHMO (3′-sialyllactose (3′SL) and 6′-sialyllactose (6′SL)). For these cases, growth (‘+/w’) on at least one substrate was considered as a TP for a predicted binary phenotype ‘1’, and the absence of growth on all tested substrates was considered a TN for ‘0’. Standard binary classification metrics were calculated from TP, TN, FP and FN counts (Supplementary Table 17).
Glycoprofiling of HMO consumption
Overnight cultures of Bifidobacterium strains grown in MRS-AC-Lac were inoculated (0.5–1% (vol/vol)) into fresh medium and cultured to OD600 of 0.6. Cells were collected at 5,000g for 5 min, washed with sugar-free MRS-AC, and used to inoculate 200 µl of MRS-AC supplemented with an HMO mixture (10 mg ml−1, isolated from pooled human milk from ≥25 different donors) in 96-well plates (Costar) at OD600 of 0.005. Triplicate 100-µl samples of culture supernatant were collected at 8-h or 24-h timepoints in separate experiments, filtered through Spin-X Centrifuge Tube Filter (0.22 µm, Costar), snap-frozen in liquid nitrogen and stored at −80 °C. HMO concentrations in supernatants were analysed by HPLC with fluorescence detection112. Media samples (10 µl) were spiked with maltose as an internal standard, lyophilized and labelled with the fluorophore 2-aminobenzamide (2AB). 2AB-labelled HMOs were separated on a TSKgel Amide-80 column (15 cm length, 2 mm inner diameter, 3 μm particle size; Tosoh Bioscience) and detected at 360-nm excitation and 425-nm emission on Dionex Ultimate 3000 (Thermo Fisher Scientific). Nineteen HMO structures were annotated on the basis of standard retention times and quantified relative to the internal standard: 2′FL, 3FL, LDFT, 3′SL, 6′SL, LNT, LNnT, LNFP I, II and III, sialyllacto-N-tetraose (LST) b and c, difucosyllacto-N-tetraose (DFLNT), LNH, disialyllacto-N-tetraose (DSLNT), fucosyllacto-N-hexaose (FLNH), difucosyllacto-N-hexaose (DFLNH), fucodisialyllacto-N-hexaose (FDSLNH) and disialyllacto-N-hexaose (DSLNH). Total HMO concentration was calculated as the sum of individual HMO concentrations (Supplementary Table 18). HMO utilization at 8 or 24 h was calculated relative to HMO concentrations in the medium controls.
Transcriptional profiling (RNA-seq)
An overnight culture of Bc. kashiwanohense Bg42221_1E1 grown in MRS-AC-Glc was collected at 5,000g for 5 min, washed in sugar-free MRS-AC and used to inoculate MRS-AC supplemented with either 10 mg ml−1 of Glc, Lac, LNT or 5 mg ml−1 of tamarind XGL at OD600 of 0.01. Samples (2 ml, biological triplicates) were collected at the early exponential phase (OD600 of 0.35) and pelleted in a prechilled centrifuge at 4,800g for 5 min. Cell pellets were snap-frozen in liquid nitrogen and stored at −80 °C until further use. The detailed RNA extraction protocol is described in Supplementary Methods. Ribosomal RNA was depleted with the NEBNext rRNA depletion kit for bacteria (New England Biolabs) and a set of 20 pooled sequence-specific probes for Bc. kashiwanohense Bg42221_1E1 designed using the NEBNext Custom RNA Depletion Design Tool v1.0 (Supplementary Table 20). Barcoded libraries were made with the NEBNext Ultra II directional RNA library prep kit for Illumina (New England Biolabs). Libraries were pooled and sequenced (single-end 75-bp reads) on Illumina NextSeq 500 using the High Output V2 kit (Illumina). The details of sequencing data analysis are provided in Supplementary Methods.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Genomes of Bifidobacterium isolates sequenced in this study are available via GenBank at https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1126848. Nucleotide FASTA and annotated protein FASTA files of 263 reference genomes are available via figshare at https://doi.org/10.6084/m9.figshare.26053936 (ref.113). Additional Bifidobacterium genomes and MAGs are available via the following publicly available databases and datasets: BV-BRC (https://www.bv-brc.org), IMG (https://img.jgi.doe.gov), BIO-ML (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA544527), CGR (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA482748), CGR2 (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA903559), UHGG (http://ftp.ebi.ac.uk/pub/databases/metagenomics/mgnify_genomes), HRGM (https://www.decodebiome.org/HRGM1), ELGG (https://doi.org/10.5281/zenodo.6969519), SPMP (https://doi.org/10.6084/m9.figshare.c.5993596.v4) and IMGG (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA763692). The RNA-seq dataset is available via Gene Expression Omnibus at https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE239955. Source data are available via GitHub at https://github.com/Arzamasov/compendium_manuscript.
Code availability
Code detailing the data analysis steps is available via GitHub at https://github.com/Arzamasov/compendium_manuscript and Supplementary Code File 1. The pipeline for analysing the representation of carbohydrate utilization pathways encoded in bifidobacterial genomes (glycobif) is available via GitHub at https://github.com/Arzamasov/glycobif.
References
Alessandri, G., van Sinderen, D. & Ventura, M. The genus Bifidobacterium: from genomics to functionality of an important component of the mammalian gut microbiota. Comput. Struct. Biotechnol. J. 19, 1472–1487 (2021).
Arboleya, S., Watkins, C., Stanton, C. & Ross, R. P. Gut bifidobacteria populations in human health and aging. Front. Microbiol. 7, 1204 (2016).
Stewart, C. J. et al. Temporal development of the gut microbiome in early childhood from the TEDDY study. Nature 562, 583–588 (2018).
Sakanaka, M. et al. Evolutionary adaptation in fucosyllactose uptake systems supports bifidobacteria-infant symbiosis. Sci. Adv. 5, eaaw7696 (2019).
Arzamasov, A. A. & Osterman, A. L. Milk glycan metabolism by intestinal bifidobacteria: insights from comparative genomics. Crit. Rev. Biochem. Mol. Biol. 57, 562–584 (2023).
Kujawska, M. et al. Succession of Bifidobacterium longum strains in response to a changing early life nutritional environment reveals dietary substrate adaptations. iScience 23, 101368 (2020).
Vatanen, T. et al. A distinct clade of Bifidobacterium longum in the gut of Bangladeshi children thrives during weaning. Cell 185, 4280–4297.e12 (2022).
Orihara, K. et al. Characterization of Bifidobacterium kashiwanohense that utilizes both milk- and plant-derived oligosaccharides. Gut Microbes 15, 2207455 (2023).
Taft, D. H. et al. Bifidobacterium species colonization in infancy: a global cross-sectional comparison by population history of breastfeeding. Nutrients 14, 1423 (2022).
Olm, M. R. et al. Robust variation in infant gut microbiome assembly across a spectrum of lifestyles. Science 376, 1220–1223 (2022).
Derrien, M. et al. Gut microbiome function and composition in infants from rural Kenya and association with human milk oligosaccharides. Gut Microbes 15, 2178793 (2023).
Tannock, G. W. et al. Comparison of the compositions of the stool microbiotas of infants fed goat milk formula, cow milk-based formula, or breast milk. Appl. Environ. Microbiol. 79, 3040–3048 (2013).
Casaburi, G. et al. Metagenomic insights of the infant microbiome community structure and function across multiple sites in the United States. Sci. Rep. 11, 1472 (2021).
Fukuda, S. et al. Bifidobacteria can protect from enteropathogenic infection through production of acetate. Nature 469, 543–547 (2011).
Hirano, R. et al. Next-generation prebiotic promotes selective growth of bifidobacteria, suppressing Clostridioides difficile. Gut Microbes 13, 1973835 (2021).
Belenguer, A. et al. Two routes of metabolic cross-feeding between Bifidobacterium adolescentis and butyrate-producing anaerobes from the human gut. Appl. Environ. Microbiol. 72, 3593–3599 (2006).
Rios-Covian, D., Gueimonde, M., Duncan, S. H., Flint, H. J. & de los Reyes-Gavilan, C. G. Enhanced butyrate formation by cross-feeding between Faecalibacterium prausnitzii and Bifidobacterium adolescentis. FEMS Microbiol. Lett. 362, fnv176 (2015).
Laursen, M. F. et al. Bifidobacterium species associated with breastfeeding produce aromatic lactic acids in the infant gut. Nat. Microbiol. 6, 1367–1382 (2021).
Shiver, A. L. et al. Genome-scale resources in the infant gut symbiont Bifidobacterium breve reveal genetic determinants of colonization and host-microbe interactions. Cell 188, 2003–2021.e19 (2025).
Maldonado-Gómez, M. X. et al. Stable engraftment of Bifidobacterium longum AH1206 in the human gut depends on individualized features of the resident microbiome. Cell Host Microbe 20, 515–526 (2016).
Frese, S. A. et al. Persistence of supplemented Bifidobacterium longum subsp. infantis EVC001 in breastfed infants. mSphere 2, e00501–e00517 (2017).
Beck, L. C. et al. Strain-specific impacts of probiotics are a significant driver of gut microbiome development in very preterm infants. Nat. Microbiol. 7, 1525–1535 (2022).
Barratt, M. J. et al. Bifidobacterium infantis treatment promotes weight gain in Bangladeshi infants with severe acute malnutrition. Sci. Transl. Med. 14, eabk1107 (2022).
Button, J. E. et al. Dosing a synbiotic of human milk oligosaccharides and B. infantis leads to reversible engraftment in healthy adult microbiomes without antibiotics. Cell Host Microbe 30, 712–725 (2022).
Milani, C. et al. Bifidobacteria exhibit social behavior through carbohydrate resource sharing in the gut. Sci. Rep. 5, 15782 (2015).
Albert, K., Rani, A. & Sela, D. A. Comparative pangenomics of the mammalian gut commensal Bifidobacterium longum. Microorganisms 8, 7 (2019).
Liu, J., Li, W., Yao, C., Yu, J. & Zhang, H. Comparative genomic analysis revealed genetic divergence between Bifidobacterium catenulatum subspecies present in infant versus adult guts. BMC Microbiol. 22, 158 (2022).
Arboleya, S. et al. Gene-trait matching across the Bifidobacterium longum pan-genome reveals considerable diversity in carbohydrate catabolism among human infant strains. BMC Genomics 19, 33 (2018).
Bottacini, F. et al. Comparative genomics and genotype-phenotype associations in Bifidobacterium breve. Sci. Rep. 8, 10633 (2018).
Liu, S. et al. Gene–phenotype associations involving human-residential bifidobacteria (HRB) reveal significant species- and strain-specificity in carbohydrate catabolism. Microorganisms 9, 883 (2021).
Magnúsdóttir, S. et al. Generation of genome-scale metabolic reconstructions for 773 members of the human gut microbiota. Nat. Biotechnol. 35, 81–89 (2017).
Devika, N. T. & Raman, K. Deciphering the metabolic capabilities of Bifidobacteria using genome-scale metabolic models. Sci. Rep. 9, 18222 (2019).
Schöpping, M., Gaspar, P., Neves, A. R., Franzén, C. J. & Zeidan, A. A. Identifying the essential nutritional requirements of the probiotic bacteria Bifidobacterium animalis and Bifidobacterium longum through genome-scale modeling. npj Syst. Biol. Appl. 7, 47 (2021).
Casey, J. et al. Transporter annotations are holding up progress in metabolic modeling. Front. Syst. Biol. 4, 1394084 (2024).
Overbeek, R. et al. The subsystems approach to genome annotation and its use in the project to annotate 1000 Genomes. Nucleic Acids Res. 33, 5691–5702 (2005).
Bottacini, F. et al. Comparative genomics of the Bifidobacterium breve taxon. BMC Genomics 15, 170 (2014).
Raman, A. S. et al. A sparse covarying unit that describes healthy and impaired human gut microbiota development. Science 365, eaau4735 (2019).
Gehrig, J. L. et al. Effects of microbiota-directed foods in gnotobiotic animals and undernourished children. Science 365, eaau4732 (2019).
Smith, M. I. et al. Gut microbiomes of Malawian twin pairs discordant for kwashiorkor. Science 339, 548–554 (2013).
Turroni, F. et al. Genome analysis of Bifidobacterium bifidum PRL2010 reveals metabolic pathways for host-derived glycan foraging. Proc. Natl Acad. Sci. USA 107, 19514–19519 (2010).
Mattarelli, P., Bonaparte, C., Pot, B. & Biavati, B. Proposal to reclassify the three biotypes of Bifidobacterium longum as three subspecies: Bifidobacterium longum subsp. longum subsp. nov., Bifidobacterium longum subsp. infantis comb. nov. and Bifidobacterium longum subsp. suis comb. nov. Int. J. Syst. Evol. Microbiol. 58, 767–772 (2008).
Chaplin, A. V. et al. Intraspecies genomic diversity and long-term persistence of Bifidobacterium longum. PLoS ONE 10, e0135658 (2015).
Modesto, M. et al. Bifidobacterium longum subsp. iuvenis subsp. nov., a novel subspecies isolated from the faeces of weaning infants. Int. J. Syst. Evol. Microbiol. 73, 006013 (2023).
O’Connell Motherway, M. et al. Characterization of ApuB, an extracellular type II amylopullulanase from Bifidobacterium breve UCC2003. Appl. Environ. Microbiol. 74, 6271–6279 (2008).
Kashima, T. et al. Identification of difructose dianhydride I synthase/hydrolase from an oral bacterium establishes a novel glycoside hydrolase family. J. Biol. Chem. 297, 101324 (2021).
LoCascio, R. G., Desai, P., Sela, D. A., Weimer, B. & Mills, D. A. Broad conservation of milk utilization genes in Bifidobacterium longum subsp. infantis as revealed by comparative genomic hybridization. Appl. Environ. Microbiol. 76, 7373–7381 (2010).
Sakurama, H. et al. Lacto-N-biosidase encoded by a novel gene of Bifidobacterium longum subspecies longum shows unique substrate specificity and requires a designated chaperone for its active expression. J. Biol. Chem. 288, 25194–25206 (2013).
Garrido, D. et al. A novel gene cluster allows preferential utilization of fucosylated milk oligosaccharides in Bifidobacterium longum subsp. longum SC596. Sci. Rep. 6, 35045 (2016).
Ojima, M. N. et al. Priority effects shape the structure of infant-type Bifidobacterium communities on human milk oligosaccharides. ISME J. 16, 2265–2279 (2022).
Komeno, M. et al. Two α-l-arabinofuranosidases from Bifidobacterium longum subsp. longum are involved in arabinoxylan utilization. Appl. Microbiol. Biotechnol. 106, 1957–1965 (2022).
Friess, L. et al. Two extracellular α-arabinofuranosidases are required for cereal-derived arabinoxylan metabolism by Bifidobacterium longum subsp. longum. Gut Microbes 16, 2353229 (2024).
Tm, G. et al. Characterization and three-dimensional structures of two distinct bacterial xyloglucanases from families GH5 and GH12. J. Biol. Chem. 282, 19177–19189 (2007).
Ravachol, J. et al. Mechanisms involved in xyloglucan catabolism by the cellulosome-producing bacterium Ruminiclostridium cellulolyticum. Sci. Rep. 6, 22770 (2016).
Garrido, D., Kim, J. H., German, J. B., Raybould, H. E. & Mills, D. A. Oligosaccharide binding proteins from Bifidobacterium longum subsp. infantis reveal a preference for host glycans. PLoS ONE 6, e17315 (2011).
Sela, D. A. et al. An infant-associated bacterial commensal utilizes breast milk sialyloligosaccharides. J. Biol. Chem. 286, 11909–11918 (2011).
Sela, D. A. et al. Bifidobacterium longum subsp. infantis ATCC 15697 α-fucosidases are active on fucosylated human milk oligosaccharides. Appl. Environ. Microbiol. 78, 795–803 (2012).
Garrido, D., Ruiz-Moyano, S. & Mills, D. A. Release and utilization of N-acetyl-d-glucosamine from human milk oligosaccharides by Bifidobacterium longum subsp. infantis. Anaerobe 18, 430–435 (2012).
Arzamasov, A. A. et al. Human milk oligosaccharide utilization in intestinal bifidobacteria is governed by global transcriptional regulator NagR. mSystems 7, e00343–22 (2022).
Egan, M., O’Connell Motherway, M., Ventura, M. & van Sinderen, D. Metabolism of sialic acid by Bifidobacterium breve UCC2003. Appl. Environ. Microbiol. 80, 4414–4426 (2014).
Egan, M. et al. Cross-feeding by Bifidobacterium breve UCC2003 during co-cultivation with Bifidobacterium bifidum PRL2010 in a mucin-based medium. BMC Microbiol. 14, 282 (2014).
Pasolli, E. et al. Extensive unexplored human microbiome diversity revealed by over 150,000 genomes from metagenomes spanning age, geography, and lifestyle. Cell 176, 649–662 (2019).
Khoroshkin, M. S., Leyn, S. A., Van Sinderen, D. & Rodionov, D. A. Transcriptional regulation of carbohydrate utilization pathways in the Bifidobacterium genus. Front. Microbiol. 7, 120 (2016).
Milani, C. et al. Genomic encyclopedia of type strains of the genus Bifidobacterium. Appl. Environ. Microbiol. 80, 6290–6302 (2014).
Odamaki, T. et al. Comparative genomics revealed genetic diversity and species/strain-level differences in carbohydrate metabolism of three probiotic bifidobacterial species. Int J. Genomics 2015, 567809 (2015).
Crociani, F., Alessandrini, A., Mucci, M. M. & Biavati, B. Degradation of complex carbohydrates by Bifidobacterium spp. Int. J. Food Microbiol. 24, 199–210 (1994).
Forster, S. C. et al. A human gut bacterial genome and culture collection for improved metagenomic analyses. Nat. Biotechnol. 37, 186–192 (2019).
Poyet, M. et al. A library of human gut bacterial isolates paired with longitudinal multiomics data enables mechanistic microbiome research. Nat. Med. 25, 1442–1452 (2019).
Lin, X. et al. The genomic landscape of reference genomes of cultivated human gut bacteria. Nat. Commun. 14, 1663 (2023).
Hitch, T. C. A. et al. HiBC: a publicly available collection of bacterial strains isolated from the human gut. Nat. Commun. 16, 4203 (2025).
Almeida, A. et al. A unified catalog of 204,938 reference genomes from the human gut microbiome. Nat. Biotechnol. 39, 105–114 (2021).
Kim, C. Y. et al. Human reference gut microbiome catalog including newly assembled genomes from under-represented Asian metagenomes. Genome Med. 13, 134 (2021).
Zeng, S. et al. A compendium of 32,277 metagenome-assembled genomes and over 80 million genes from the early-life human gut microbiome. Nat. Commun. 13, 5139 (2022).
Gounot, J.-S. et al. Genome-centric analysis of short and long read metagenomes reveals uncharacterized microbiome diversity in Southeast Asians. Nat. Commun. 13, 6044 (2022).
Jin, H. et al. A high-quality genome compendium of the human gut microbiome of Inner Mongolians. Nat. Microbiol. 8, 150–161 (2023).
Kim, M. et al. Higher pathogen load in children from Mozambique vs. USA revealed by comparative fecal microbiome profiling. ISME Commun. 2, 74 (2022).
de Crécy-Lagard, V. et al. A roadmap for the functional annotation of protein families: a community perspective. Database 2022, baac062 (2022).
Price, M. N. & Arkin, A. P. Interactive tools for functional annotation of bacterial genomes. Database 2024, baae089 (2024).
Zhu, L. et al. Captive common marmosets (Callithrix jacchus) are colonized throughout their lives by a community of Bifidobacterium species with species-specific genomic content that can support adaptation to distinct metabolic niches. mBio https://doi.org/10.1128/mBio.01153-21 (2021).
Chang, H.-W. et al. Prevotella copri and microbiota members mediate the beneficial effects of a therapeutic food for malnutrition. Nat. Microbiol. 9, 922–937 (2024).
Munoz, J., James, K., Bottacini, F. & Van Sinderen, D. Biochemical analysis of cross-feeding behaviour between two common gut commensals when cultivated on plant-derived arabinogalactan. Microb. Biotechnol. 13, 1733–1747 (2020).
Fernandez-Julia, P., Black, G. W., Cheung, W., Van Sinderen, D. & Munoz-Munoz, J. Fungal β-glucan-facilitated cross-feeding activities between Bacteroides and Bifidobacterium species. Commun. Biol. 6, 576 (2023).
Renwick, S. et al. Modulating the developing gut microbiota with 2′-fucosyllactose and pooled human milk oligosaccharides. Microbiome 13, 44 (2025).
Browne, H. P. et al. Boosting microbiome science worldwide could save millions of children’s lives. Nature 625, 237–240 (2024).
Davis, J. J. et al. The PATRIC Bioinformatics Resource Center: expanding data and analysis capabilities. Nucleic Acids Res. 48, D606–D612 (2020).
Markowitz, V. M. et al. IMG: the Integrated Microbial Genomes database and comparative analysis system. Nucleic Acids Res. 40, D115–D122 (2012).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Olm, M. R., Brown, C. T., Brooks, B. & Banfield, J. F. dRep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication. ISME J. 11, 2864–2868 (2017).
Parks, D. H. et al. GTDB: an ongoing census of bacterial and archaeal diversity through a phylogenetically consistent, rank normalized and complete genome-based taxonomy. Nucleic Acids Res. 50, D785–D794 (2022).
Tonkin-Hill, G. et al. Producing polished prokaryotic pangenomes with the Panaroo pipeline. Genome Biol. 21, 180 (2020).
Katoh, K. & Standley, D. M. MAFFT Multiple Sequence Alignment Software Version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
Pritchard, L., Glover, R. H., Humphris, S., Elphinstone, J. G. & Toth, I. K. Genomics and taxonomy in diagnostics for food security: soft-rotting enterobacterial plant pathogens. Anal. Methods 8, 12–24 (2015).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Overbeek, R. et al. The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res. 42, D206–D214 (2014).
Cantalapiedra, C. P., Hernández-Plaza, A., Letunic, I., Bork, P. & Huerta-Cepas, J. eggNOG-mapper v2: functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Mol. Biol. Evol. 38, 5825–5829 (2021).
Zheng, J. et al. dbCAN3: automated carbohydrate-active enzyme and substrate annotation. Nucleic Acids Res. 51, W115–W121 (2023).
Saier, M. H. et al. The Transporter Classification Database (TCDB): 2021 update. Nucleic Acids Res. 49, D461–D467 (2021).
Drula, E. et al. The carbohydrate-active enzyme database: functions and literature. Nucleic Acids Res. 50, D571–D577 (2022).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 44, D457–D462 (2016).
Novichkov, P. S. et al. RegPrecise: a database of curated genomic inferences of transcriptional regulatory interactions in prokaryotes. Nucleic Acids Res. 38, D111–D118 (2010).
Steinegger, M. & Söding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 35, 1026–1028 (2017).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60 (2015).
Abraham, A. et al. Machine learning for neuroimaging with scikit-learn. Front. Neuroinform. 8, 14 (2014).
Kuhn, M. Building predictive models in R using the caret package. J. Stat. Softw. 28, 1–26 (2008).
Hibberd, M. C. et al. Bioactive glycans in a microbiome-directed food for children with malnutrition. Nature 625, 157–165 (2024).
Oksanen, J. et al. vegan: Community Ecology Package. R package version 2.6-8 https://CRAN.R-project.org/package=vegan (2024).
Lenth, R. V. et al. emmeans: Estimated Marginal Means, aka Least-Squares Means. R package version 1.10.6 https://github.com/rvlenth/emmeans (2024).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2016).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Gilchrist, C. L. M. & Chooi, Y.-H. clinker & clustermap.js: automatic generation of gene cluster comparison figures. Bioinformatics 37, 2473–2475 (2021).
Berger, P. K. et al. Stability of human-milk oligosaccharide concentrations over 1 week of lactation and over 6 hours following a standard meal. J. Nutr. 152, 2727–2733 (2023).
Arzamasov, A. Supplementary data for the manuscript titled “Integrative genomic reconstruction reveals heterogeneity in carbohydrate utilization across human gut bifidobacteria”. figshare https://doi.org/10.6084/m9.figshare.26053936 (2025).
Acknowledgements
We thank R. Olson (Hack Biology LLC) for helping with mcSEED maintenance, K. Liu (SBP Medical Discovery Institute Genomics Core) for library preparation and sequencing, S. Nakshatri (Washington University) and K. Spann (UC San Diego) for technical assistance, Glycom A/S and dsm-firmenich for generously providing individual HMOs and T. Katayama (Kyoto University) for generously sharing Bifidobacterium scardovii JCM 12489 and insightful discussions. We are grateful to M. Manary (Washington University) and study personnel at the Malawi College of Medicine for the provision of faecal samples used to isolate the Malawian bifidobacterial strains described in this Resource. Bangladeshi isolates were provided through a long-standing collaborative research programme between Washington University and members of the International Center for Diarrheal Disease Research, Bangladesh (icddr,b) led by T. Ahmed. A material transfer agreement was established to transfer these strains to the SBP Medical Discovery Institute. This work was supported by grants from the National Institutes of Health (NIH) (grant number DK30292 to A.L.O. and J.I.G.) and the Bill & Melinda Gates Foundation (grant number INV-016367). L.B. is the UC San Diego Chair of Collaborative Human Milk Research, endowed by the Family Larsson-Rosenquist Foundation, Switzerland.
Author information
Authors and Affiliations
Contributions
A.A.A., D.A.R., M.J.B., J.I.G. and A.L.O. designed the study. A.A.A. performed data curation and metabolic reconstruction. A.A.A., D.A.R., M.D.K., S.A.L. and A.L.O. developed the pathway prediction pipeline. J.L.G. isolated bacterial strains. M.C.H., J.E.K. and M.L.E. sequenced and assembled bacterial genomes. A.A.A. performed growth and RNA-seq experiments. K.S., A.F. and L.B. isolated HMOs and conducted glycoprofiling analysis. A.A.A. wrote the initial draft with input from J.E.K. A.A.A., D.A.R., M.C.H., L.B., M.J.B., J.I.G. and A.L.O. edited the paper with invaluable assistance from co-authors.
Corresponding authors
Ethics declarations
Competing interests
D.A.R. and A.L.O. are co-founders of Phenobiome Inc., a company pursuing the development of personalized nutritional solutions to balance the gut microbiome. L.B. is a co-inventor on patent applications related to the use of HMOs in preventing necrotizing enterocolitis and other inflammatory diseases. The other authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Christian Diener, Matthew Olm and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Pairwise average nucleotide identity (ANI) analysis of reference B. longum and B. catenulatum genomes.
a, Pairwise ANI values for 28 B. longum genomes. For comparative purposes, this set contained 15 genomes from the reference collection (names are in bold) and 13 additional genomes, including the type strains of Bl. suis and Bl. suillum of non-human origin. Black lines indicate taxonomic groupings used in this study; blue lines denote the additional delineation proposed by Modesto et al.43. b, Pairwise ANI values for 11 reference B. catenulatum, B. hominis69, and B. pseudocatenulatum genomes. Black lines indicate taxonomic groupings used in this study; NCBI and GTDB taxonomy assignments are shown in blue.
Extended Data Fig. 2 Reconstruction of carbohydrate utilization pathways and associated phenotypes.
(1) A reference set of 263 Bifidobacterium genomes (Supplementary Table 2) and a list of 589 curated functional roles (Supplementary Table 4) were used to populate 25 metabolic subsystems capturing catabolic pathways for 68 glycans. (2) Functional gene annotation was performed using homology-based methods and three genome context techniques: (i) clustering of genes on the chromosome (operons), (ii) co-regulation of genes by a common transcription factor, and (iii) co-occurrence of genes across related genomes. Carbohydrate utilization pathways were reconstructed based on the distribution of functional roles. (3) Each genome was assigned a detailed pathway variant based on the presence of signature genes (Supplementary Table 6). These variants were then converted to predicted binary phenotypes (1 = utilizer, 0 = non-utilizer) to create a Binary Phenotype Matrix (BPM).
Extended Data Fig. 3 Representation of predicted carbohydrate utilization pathways in B. catenulatum and B. hominis genomes.
The heatmap shows the hierarchical clustering of the BPM for 68 carbohydrate utilization pathways (columns) predicted in 110 B. catenulatum and 26 B. hominis genomes (rows). Pathway (phenotype) classifications are shown in the bottom annotation tracks; taxonomic assignments are indicated on the right. Names of type strains are in bold. Full pathway (phenotype) names are provided in Supplementary Table 5.
Extended Data Fig. 4 Genomics-based reconstruction of pathways involved in the utilization of monosaccharide derivatives.
a, Gene clusters potentially driving D-glucuronate (GlcA) utilization by Bl. infantis JG_Bg463 and glucuronide (GluA) utilization by B. breve Bg41721_1C11. Orthologous genes are linked, and the link color represents the sequence identity between corresponding protein products. b, Reconstructed D-glucuronate and glucuronide utilization pathways. c, Growth curves of selected Bifidobacterium strains in MRS-AC supplemented with 1% D-glucuronic acid. Data represent the mean ± s.d. of three biological replicates. d, Gene clusters potentially driving the utilization of (i) sugar alcohols D-mannitol (Mtl), D-sorbitol (Stl), and myo-inositol (Ino), (ii) sugar acids D-gluconate (Gco) and D-galactonate (Glt), (iii) L-ascorbate (Asc). e, Reconstructed utilization pathways for monosaccharide derivatives. f, Gene cluster (nan) driving sialic acid (Neu5Ac) utilization by B. breve strains. g, Neu5Ac utilization pathway in B. breve. h, Growth curves of selected B. breve strains in MRS-AC supplemented with 1% N-acetylneuraminic acid. Data represent the mean ± s.d. of three biological replicates.
Extended Data Fig. 5 Representation of metabolic pathways in 28 selected B. longum isolate genomes.
For comparative purposes, this dataset includes 15 genomes from the reference collection (names in bold) and 13 additional genomes, primarily of non-human origin, including the type strains of Bl. suis and Bl. suillum. Annotation tracks at the bottom represent pathway (phenotype) classifications. Annotation columns on the right denote taxonomic groupings: the left column reflects assignments used in this study; the right column showing subspecies delineation proposed by Modesto et al.43. a, Representation of 68 carbohydrate utilization pathways and associated metabolic phenotypes. Full names are provided in Supplementary Table 5. b, Representation of 29 additional pathways, including the biosynthesis of amino acids and B vitamins, and urea utilization.
Extended Data Fig. 6 Representation of core biosynthetic pathways across 3,083 Bifidobacterium genomes.
The heatmap depicts the prevalence of metabolic pathways corresponding to: (i) B vitamin and amino acid biosynthesis, (ii) urea utilization (Urea_d) within each taxon. Color intensity indicates the proportion of genomes that encode the respective metabolic pathway.
Extended Data Fig. 7 Genomics-based reconstruction of polysaccharide degradation and oligosaccharide utilization pathways.
a, Alignment of the amino acid sequences of xyloglucan endo-β-1,4-glucanases (xyloglucanases): PpXG5 from Paenibacillus pabuli XG5 (characterized), Xgl5A and Xgl5B from Bc. kashiwanohense Bg42221_1E1 (putative). Conserved catalytic amino acid residues are highlighted in bold red, and conserved glycan-binding residues are in bold. b, Gene clusters potentially driving the degradation of (arabino)xylan and the utilization of released oligosaccharides in Bc. kashiwanohense Bg42221_1E1. c, Predicted DNA-binding motifs of transcription factors potentially controlling the expression of the gene clusters in b. Motifs were built based on operator sequences in the RegPrecise database and identified in this work (Supplementary Table 19). d, Gene clusters potentially driving β-mannose oligosaccharide utilization (phenotype bMnOS) in B. breve Bg41721_1C11 and β-mannan degradation (phenotype bMAN) in B. dentium LFYP24. Orthologous genes are linked, and the link color represents the sequence identity between corresponding protein products. e, Reconstructed β-mannose oligosaccharide utilization and β-mannan degradation pathways. Names of enzymes and transporters whose orthologues have been biochemically characterized in bifidobacteria are in bold. f, Growth curves of selected Bifidobacterium strains in the medium supplemented with 0.5% mannotriose or 0.5% konjac glucomannan. Data represent the mean ± s.d. of three biological replicates. g, Predicted MnbR DNA-binding motif based on operator sequences identified in this work (Supplementary Table 19).
Extended Data Fig. 8 Pathway enrichment analysis across 3,083 Bifidobacterium genomes.
Pathways significantly enriched in specific groups (Benjamini–Hochberg adjusted P ≤ 0.01; two-sided Fisher’s exact test) are shown. Points represent odds ratios, and horizontal lines indicate 95% confidence intervals. Pathways with infinite odds ratios are present exclusively in one group. Exact adjusted P-values are provided in Supplementary Table 14. a, Comparison between genomes from ‘Westernized (age < 3)’ vs. ‘Westernized (age ≥ 3)’ groups. b, Comparison between genomes from ‘non-Westernized (age < 3)’ vs. ‘non-Westernized (age ≥ 3)’ groups. c, Comparison between genomes from ‘Westernized (age < 3)’ vs. ‘non-Westernized (age < 3)’ groups. d, Comparison within individual taxa between genomes from ‘Westernized’ vs. ‘non-Westernized’ groups.
Extended Data Fig. 9 HPLC-based quantification of human milk oligosaccharide (HMO) utilization after 8h.
Data represent the percentage of utilized HMOs (mean of three biological replicates) relative to the medium control. Total HMO, total HMO utilized; total FHMO, total fucosylated HMO utilized; total SHMO, total sialylated HMO utilized. Concentrations of individual HMOs (nmol/mL) are provided in Supplementary Table 18a.
Supplementary information
Supplementary Information (download PDF )
Supplementary Notes 1–5, methods and Figs. 1–5.
Supplementary Tables 1–7, 9, 11–15, 17–21 (download XLSX )
Supplementary Table 1 Distribution of 3,083 Bifidobacterium genomes by taxonomic group and geographic origin. Supplementary Table 2 Metadata for 263 reference Bifidobacterium genomes used in curated metabolic reconstruction. Supplementary Table 3 Metadata for 2,820 additional Bifidobacterium genomes used in the automated pathway prediction pipeline. Supplementary Table 4 Curated list of functional roles comprising carbohydrate utilization pathways. Supplementary Table 5 Glossary of reconstructed carbohydrate utilization pathways, associated substrates, and predicted metabolic phenotypes. Supplementary Table 6 Genomic signatures used to assign pathway variants and predicted binary utilization phenotypes. Supplementary Table 7 Binary Phenotype Matrix (BPM) depicting 68 carbohydrate utilization pathways reconstructed in 263 reference Bifidobacterium genomes. Supplementary Table 9 Binary Phenotype Matrix (BPM) depicting 68 carbohydrate utilization pathways predicted in 2,820 additional Bifidobacterium genomes. Supplementary Table 11 Binary Phenotype Matrix (BPM) depicting 30 additional metabolic pathways reconstructed in 3,083 Bifidobacterium genomes. Supplementary Table 12 List of genes used to construct the phylogenetic tree of 263 reference Bifidobacterium genomes. Supplementary Table 13 Comparison of manually curated and automated (Prokka, EggNOG-mapper) annotations for 263 reference Bifidobacterium genomes. Supplementary Table 14 Pathway enrichment analysis. Supplementary Table 15 List of substrates used in in vitro growth experiments. Supplementary Table 17 Comparison between predicted carbohydrate utilization phenotypes and in vitro growth data. Supplementary Table 18 Human milk oligosaccharide (HMO) utilization by Bifidobacterium strains. Supplementary Table 19 Reconstructed regulons of transcription factors and their predicted operator sequences. Supplementary Table 20 Custom probes used to deplete Bc. kashiwanohense Bg42221_1E1 rRNA. Supplementary Table 21 Differential gene expression in Bc. kashiwanohense Bg42221_1E1 grown on various carbon sources.
Supplementary Table 8 (download XLSX )
Representation of functional roles comprising carbohydrate utilization pathways in 263 reference Bifidobacterium genomes.
Supplementary Table 10 (download XLSX )
Representation of functional roles comprising carbohydrate utilization pathways in 2,820 additional Bifidobacterium genomes.
Supplementary Table 16 (download XLSX )
Raw in vitro growth data for 30 Bifidobacterium strains.
Supplementary Code File 1 (download PDF )
Custom code used in the study.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Arzamasov, A.A., Rodionov, D.A., Hibberd, M.C. et al. Integrative genomic reconstruction reveals heterogeneity in carbohydrate utilization across human gut bifidobacteria. Nat Microbiol 10, 2031–2047 (2025). https://doi.org/10.1038/s41564-025-02056-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41564-025-02056-x
This article is cited by
-
Development of Fermented Beverages from Whole Lentils and Lentil Flour: A Comparative Study of Microbiological, Functional, and Sensory Profiles
Plant Foods for Human Nutrition (2025)