Abstract
Science benefits from rapid open data sharing, but current guidelines for data reuse were established two decades ago, when databases were several million times smaller than they are today. These guidelines are largely unfamiliar to the scientific community, and, owing to the rapid increase in biological data generated in the past decade, they are also outdated. As a result, there is a lack of community standards suited to the current landscape and inconsistent implementation of data sharing policies across institutions. Here we discuss current sequence data sharing policies and their benefits and drawbacks, and present a roadmap to establish guidelines for equitable sequence data reuse, developed in consultation with a data consortium of 167 microbiome scientists. We propose the use of a Data Reuse Information (DRI) tag for public sequence data, which will be associated with at least one Open Researcher and Contributor ID (ORCID) account. The machine-readable DRI tag indicates that the data creators prefer to be contacted before data reuse, and simultaneously provides data consumers with a mechanism to get in touch with the data creators. The DRI aims to facilitate and foster collaborations, and serve as a guideline that can be expanded to other data types.
Similar content being viewed by others
Main
Sequence data reuse has been an evolving topic over the past two decades. The Fort Lauderdale Agreement (FLA), a public declaration by biomedicine scientists supporting the free and unrestricted use of genome sequencing data, was coined in 2003, before the advent of metagenomics and during a time when sequencing was still too costly to be performed by individual laboratories1. The FLA concluded that large genome projects should be released before publication to allow unrestricted and immediate reuse, which would accelerate the advancement of science. The FLA strengthened the Bermuda Principles defined in 19962, which advocated for the release of sequence data 24 h after generation and before publication of research papers. In 2009, after the Human Genome Project highlighted the advantages of sharing data early and widely, the Toronto Statement (TOR)3 advocated for the prepublication release of other biological data types beyond genomics data. Finally, in 2014, 141 United Nations member states and the European Union entered into the Nagoya Protocol4 (Regulation (EU) No 511/2014), which calls on data creators and data users to develop, update and use voluntary codes of conduct, guidelines and best practices in relation to access and benefit-sharing of genetic data (see Box 1 for descriptions of the different roles of researchers working with sequence datasets).
Large-scale sequence data analysis has become mainstream with a wide array of tools available, making data mining accessible to many labs. Now, ~20 years after the FLA, GenBank holds an estimated ~5.09 terabase pairs (Tbp)5 of biological sequence data, and the Sequence Read Archive (SRA) holds 90.89 petabase pairs (Pbp) as of February 2024 (Supplementary Fig. 1 and Box 2). These databases are several million times larger than the available sequence data at the time that the FLA or TOR was formulated. With the rapid, continuous increase in public sequence data (projected to reach ~500 Pbp in 2030, Supplementary Fig. 1), data mining projects (or those requiring large public datasets for artificial intelligence training) have increased in both frequency and scope, necessitating a revisit and potential overhaul of the 20-year-old guidelines depicted in the FLA and TOR1,3.
In 2016, the FAIR principles for data management were defined, which place an emphasis on submitted data being machine actionable (that is, computational systems should be able to Find, Access, Interoperate and Reuse data with minimal human intervention)6. These principles were designed to promote good scientific practice and to serve as a guideline for those wishing to enhance the reusability of their data. The FAIR principles have since been adopted as recommendations or requirements by major funding bodies, including the US National Institute of Health and the European Commission7. The FAIR data principles prioritize data reuse and computer-driven data mining, and include a specific requirement for (meta)data to be released with a clear and accessible data reuse licence (principle R.1). To date, this aspect of the FAIR principles has not been implemented in a straightforward or machine-readable way, lacking a coordinated implementation between databases and the community.
Biological sciences, and particularly their subdisciplines associated with generating sequence data, have been at the forefront of data availability compared with the fields of earth sciences, mathematics, physics and chemistry8. For instance, astrophysics is a subdiscipline that traditionally relies on data sharing and reuse owing to the exorbitant costs of research data. A study investigating the motivational factors behind data sharing and reuse within this field identified several demotivating factors9. Among them were the lack of data standards, the lack of facilitating platforms, inconsistency between datasets, limited documentation, difficulties finding and reusing data, and last but not least, competition and fear of being ‘scooped’ (accidentally or purposefully)9. The latter point is considered in the FLA. While the FLA recommends swift prepublication of data generated by large sequencing consortia, it also states that “[…] the contributions and interests of the large-scale data producers should be recognized and respected by the users of the data, and the ability of the production centers to analyse and publish their own data should be supported by their funding agencies […]”1. This highlights one of the significant and enduring tensions between data creators and data consumers. Both data creators and data consumers are indispensable to advancing biological sciences, particularly in the realm of sequence data analysis, in which many data creators also act as data consumers. Unrestricted public use of microbiome data, on which data creators have not yet published, does not always align with the interests of data creators.
How to achieve unrestricted data reuse and, at the same time, give due credit to data creators has been discussed by the scientific community10. Data reuse in this work refers specifically to those cases in which the sequence data will be featured in a publication prepared by a data consumer, whether in figures, tables or text, or as an important aspect of the workflow that leads to new insights or conclusions (see Supplementary Table 2 for some Data Reuse Information (DRI) usage scenarios). In this spirit, a recent study thoroughly analysed the pros and cons of early data release and considered both the needs of data creators and consumers. The authors proposed immediate, unrestricted release of sequence data before publication, in parallel with the adoption of a reward system (for example, separate promotion and tenure tracks) for acknowledgement of data creators by universities and research institutions11. In addition, the authors proposed making the datasets and the protocols used for their generation citable through Digital Object Identifiers (DOIs). If implemented, these measures would create a safer environment for data sharing, benefiting all parties involved and, most importantly, supporting the advancement of science.
Mechanisms for crediting data creators beyond citing associated publications are not yet widespread in the scientific community. Creating separate tenure tracks or other incentives for data creators and data consumers requires sizable changes in evaluation criteria, and would require substantial time to propagate through institutions. DOIs for datasets, on the other hand, seem relatively easy to implement and would provide data creators with a reportable impact metric. However, their use has not yet been widely adopted, possibly owing to associated costs with purchasing and maintaining DOIs, which can be prohibitive for many publicly funded research institutions. Potential measures to lower DOI costs could include large-scale agreements between research institutions and DOI providers. Other mechanisms of data citation have been discussed in the community but have also not been widely adopted12,13. Currently, data creators do not have any incentive or reward for releasing sequence data before an associated publication.
It is crucial to implement methodological and ethical guidelines that are based on the principles of good scientific practice and which are driven by the scientific community to facilitate appropriate use of public data. This need has been highlighted by recent conflicts between data creators and data consumers that played out over social media. Implementing and following guidelines for unpublished data usage by all scientists would create ‘safe spaces’ for data creators to publish their first analyses of data—particularly if they are delayed by resource, time or personnel constraints. The research topic also affects the expectations for open data. Research related to public health necessitates swift data release to counteract pandemics or identify zoonotic diseases. For example, in the event of a pandemic, there should be no data restriction on research related to the pandemic14. The general goal should be to promote open sharing of complete datasets as early and as widely as possible, across all institutions and individuals. This necessitates a technical framework that enhances the communication between data creators and data consumers regarding data reuse.
Here we propose a roadmap to enable equitable reuse of public microbiome data. This roadmap (1) addresses the lack of consensus in the field of microbiome research regarding public microbiome data use and reuse, (2) promotes communication between data consumers and data creators and (3) facilitates the rapid advancement of the microbiome field, including supporting the continued increases in data mining. To achieve the goal of this roadmap, we propose the introduction of a new machine-readable metadata tag, named DRI, containing Open Researcher and Contributor IDs (ORCIDs) of the data creators associated with data in public databases. The DRI will clearly indicate the point of contact for communication and if communication is desired by data creators. The ability to provide a point of contact for data reuse will lead to more rapid and complete data deposition. Following adoption by databases, authors and scientific journals would ideally integrate statements confirming that the best practices governed by DRI use were used in manuscripts and submission processes.
The roadmap is directly in line with the FAIR data principles, specifically contributing to FAIR principle R.1 in providing a machine-readable licence for data usage. This roadmap and its adoption by the scientific community (222 scientists as part of the Data Reuse Consortium—Supplementary Table 1—totalling 229 supporters, including the co-authors of this paper) will provide a citable resource regarding guidelines for public data reuse, will enable appropriate data reuse by data consumers and will reduce tension for data creators when submitting data. Ultimately, this roadmap outlines the expected practices for open data use for sequence data and represents a model for other biological data such as metabolomics or proteomics data.
Survey on data reuse
We created an anonymous survey with Google Forms, which was distributed to the international scientific community on 15 January 2024, to accumulate opinion data on a number of key topics related to this manuscript (Supplementary Box 1). Participation was voluntary and anonymous, and participants gave informed consent before participation. To ensure participant confidentiality, it was emphasized that signing the consensus statement could not be linked to survey responses. The University of Duisburg-Essen ethics committee evaluated the study and declared that the need for ethics approval has been waived. Questions included in the survey were formulated in a neutral fashion with the intent of not biasing responses, which were anonymized to ensure openness and transparency. This survey was online and open for responses over a total of 21 days. Efforts to ensure widespread awareness of this survey included actively advertising it across multiple social media platforms (X.com, LinkedIn, Bluesky.app) over the duration of the survey. To achieve this, 39 authors made use of their accounts across these platforms while also leveraging their working group and other institutional accounts. A blog advertising this survey was additionally hosted at the Springer Nature Research Communities blog15. Finally, a total of 78 microbiology institutions across the world were contacted to increase participation from the Global South and underrepresented segments of the global scientific community related to this survey. Raw data pertaining to the anonymous responses to this survey in TSV format are available at the Open Science Foundation16.
This resulted in responses from 306 scientists representing all continents (except Antarctica) with feedback on community interest in and likelihood of adopting the proposed roadmap (Supplementary Figs. 2–10 and Supplementary Tables 3–7). Survey questions were designed to enable quantitative analysis, namely, to appreciate the fraction of the community that agreed or disagreed with proposed aspects of the roadmap delineated in this manuscript. Raw data for this survey were imported into R 4.3.1 running in RStudio 2023.12.0 (Build 369) using the tidyverse ecosystem of data analysis packages to read TSV inputs (readr, version 2.1.5), filter (dplyr, version 1.1.4) and generate plots (ggplot2, version 3.5.1)17,18. For Fig. 1, survey data were processed using R, and visualizations in Fig. 1a–c were generated using ggplot2. Figure 1d was generated with a copyright-free image, coloured by normalizing height (in pixels) to percentage categories. Colours and formatting were manually edited. Sankey diagrams were generated via the ggsankey package (version 0.0.99999). Tables were generated with the kableExtra package (version 1.3.4)19. R code as well as system and package versions used for all data analysis in this manuscript are publicly available at a GitHub repository: https://github.com/GeoMicroSoares/DataUsage_Data_Analysis.
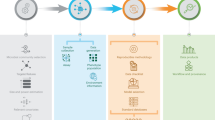
A 21-question survey examining the scientific community’s perceptions of data reuse was conducted in January 2024, with the survey distributed through social media, a blog and email to hundreds of scientists and two dozen scientific societies. A total of 306 respondents contributed to the survey. a–d, Responses were summarized for key questions and visualized using Adobe Illustrator, in which the first panel corresponds to questions 15–17 (a), the second panel to questions 20–21 (b), the third panel to questions 9–12 (c) and the fourth panel to questions 7–8 (d) (a descriptive analysis of responses to this survey and anonymous raw data to all questions are available in Supplementary Figs. 2–10 and Supplementary Tables 3–7). N/A indicates the question was left blank. Positive communication was defined as being contacted before data analysis or publication and asked for collaboration and opinion, or a positive answer when you requested data removal from a manuscript. Negative communication was defined as no contact before publication, or refusal to remove data from a manuscript upon request. The asterisk in a indicates that respondents agree or strongly agree that ‘Unauthorized use of my sequence data by other authors has had (or will have) negative impacts on my research programme and/or my mentees’. Respondents selected single-year intervals for the data presented in the hourglass image; years 4–6 and 7–9 were combined given very low proportions for all years except year 5. Notably, respondents did not agree on a time interval after which data should be made available in the absence of an available publication (Supplementary Fig. 7). As a result, our roadmap does not include a recommendation as to when publicly available data with a DRI but no publication can be reused without contacting the data creators (Fig. 2).
The initial DRI tag and the strategy for its implementation were defined by the Data Reuse Core Team before the survey. After the survey, the Data Reuse Core Team refined the roadmap and DRI strategy through conversations with members of the Data Reuse Consortium, of the Joint Genome Institute, of the European Nucleotide Archive and of the Genomic Standards Consortium, and the reviewers of this manuscript. Throughout the refinement process, the Data Reuse Core Team identified compromises between differing priorities and ensured that the DRI strategy was actionable within current database structures.
Microbiome data types most frequently reused include amplicon datasets of single genes, such as 16S rRNA or internal transcribed spacer (ITS) genes, as well as reads and assemblies of genomes, metagenomes and metatranscriptomes. These sequence data, while typically structured as machine-readable files, can come in various formats depending on the hosting repository.
Organizations hosting data usually have their own legal framework governing data download. In the case of the EMBL-EBI ENA database, for example, while records are typically available with no restrictions on reuse, the Terms of Use20 recognize that third parties may assert restrictions on reuse for a variety of reasons. The database is working towards more systematic implementation of categorizing reusability, probably under Creative Commons (CC0) licensing21. This puts the responsibility on the data consumer to ensure their action is indeed legal and covered by the copyrights that may be associated with the data they are accessing, even if the licensing information is not machine readable.
The current state of data policies across online repositories of biological data, including EMBL-EBI, is largely reflective of a now outdated statement in the TOR. Here data users are advised to contact data creators to ‘discuss publication plans’, a demanding and often unfeasible approach given today’s rate of data production and the availability of new data analysis pipelines. The TOR, from 2009, and the FLA, from 2003, are the agreements guiding the field to date, although their contents are often not well known by microbiome scientists across all career stages (Supplementary Figs. 4–6). The FLA specifies, among other things, that “sequence assemblies of 2 kb or greater by large-scale sequencing efforts” must be rapidly released. The language around the scale of data reflects the outdated guidance, and the emphasis on large-scale data creators is no longer in line with the prevalence of independent labs as data creators today. Interestingly, the conflict between data sharing and publishing a first analysis was acknowledged in the FLA, but it does not provide guidelines for navigating this concern. The TOR elaborated further on the conflict between “data [creators] and data users”, stating that, in the author’s experience, conflicts have rarely arisen.
The TOR lists a set of conditions (scale, utility, reference data, community acceptance) to consider for prepublication data sharing that are of limited relevance for today’s data landscape3. Currently, individual research groups can contribute substantial datasets, both in terms of size and scientific value, and the idea of individualized, private agreements on data sharing, as suggested by the TOR, is no longer viable. In response to this need, repositories have developed their own policies (for example, the EMBL-EBI licence information). Data distributors recognize that some public datasets, although hosted by their respective services, carry additional restrictions that are currently neither easily visible nor machine readable. Given the ease of accessibility and sheer volume of sequence data, it has become impractical and, in some cases, impossible for a data consumer to verify and to comply with recommendations and restrictions.
Identifying conflicts of interest between data consumers and data creators
The current scale of open sequence data, including massive open datasets such as the Tara Oceans (7.2 Tbp of metagenomic data) and Integrative Human Microbiome (1.3 Tbp of metagenomic data as of 2019) projects, has made meta-analyses drawing on public data a powerful avenue to explore microbial systems22,23. Use of public data is now routine; close to 80% of respondents to our poll on data usage identified themselves as both data creators and data consumers (Box 1 and Supplementary Table 7). Access to public data has unequivocally improved the depth of the science conducted. However, it is often difficult to assess whether data reuse follows the expectations of communication and collaboration outlined in the TOR. There are many widely used software tools that include use of secondarily accessed data for which no primary publication exists (for example, the Genome Taxonomy Database, GToTree)24,25. Identification of the publication status associated with specific data in many repositories is not straightforward. As more governments begin to require data deposition on short or immediate timelines, there is a growing tension between data creators and data consumers around public data use. Clarifying and facilitating data reuse is therefore in the best interest of the community.
The first step is to identify roots of the potential conflict of interest between data creators and data consumers, as established following discussions between the authors of this manuscript as well as within their academic networks. These are discussed in detail below.
Disconnect between efforts of data creation and ease of reuse
One source of conflict is a disconnect between the efforts expended by data creators in generating sequence data and associated metadata, and the ease of reuse and limited or inconsistent acknowledgement of data origins by consumers. Creators sometimes feel that their monetary, time and intellectual investments to design and conduct sample collection and experiments; obtain permission for, plan, fund and carry out research expeditions; process samples; and deposit data and metadata are not adequately acknowledged or are potentially ignored by data consumers. Data creators must obtain legal documents (for example, sampling permits, visas) and follow international agreements (for example, the Convention on Biological Diversity, Nagoya Protocol), and manage the risks that come with certain fieldwork (for example, treacherous terrain, wilderness areas, areas with high criminal activity). In addition, they must secure funding for custom-design vehicles and instrumentation needed for sample collection (for example, drill ships, research vessels, submarines, buoys, remote samplers) and maintain research sites in hard-to-access areas (for example, polar regions or the Amazon). Unbeknownst to data consumers, the original data creators may be bound by restrictive agreements on appropriate or ethical data use if research was conducted in a national park, on private land or land owned by Indigenous nations, and/or for samples obtained from human specimens or biobanks26. There are currently limited rewards for data creators when their data are reused, and data creators have little incentive to make detailed metadata available. Systems for reporting and incentivizing data deposition have been proposed but are not yet the norm11,27.
Timely deposition contrasts with lengthy multi-omics analyses
Both data creators and consumers generally share ideals of open science and rapid advancement of science. However, conflicts arise from disagreements in the timing of sharing data and prioritization of access. Data creators must balance long trainee timelines with publication of datasets intended for multiple research questions. Publishing a first paper on a large dataset and depositing the full data may make additional research projects associated with that dataset vulnerable to scooping. Results from our poll suggest that 53% of researchers are concerned about negative impacts on their research programme and/or mentees from unauthorized data reuse (Fig. 1 and Supplementary Figs. 7–9). As a result, partial or raw datasets or datasets lacking key metadata are deposited in place of more polished, complete datasets with full metadata to guide interpretation of genome data. In the absence of open data, data consumers are frequently unable to access contextual information (for example, physicochemical parameters, geolocations) of the field site that are essential to accurately interpret the data, to the detriment of downstream analyses. These issues are exacerbated by public sequence databases lacking (links to) the associated metadata and a general lack of familiarity of many data consumers with the literature on, or the environmental context of, a specific system.
Perceived threats to research and career goals
Duplication of effort and the potential for lowered impact or difficulties publishing replicated results are a loss for both creators and consumers. For data creators, raw data underlying published scientific results must be made public to meet expectations for reproducibility. However, unrestricted access to public data can compromise permits, site access agreements and research ethics board approvals, all of which can negatively impact the data creators’ and their mentees’ ongoing research. For data consumers, even unintentional reuse of restricted data can slow research progress while appropriate permissions are sought, delay publications while data are removed and, in extreme cases, lead to paper retractions. The perceived threat is that unauthorized data reuse can also negatively impact planned research directions, funded research goals, acquisition of new funds or career perspectives of early career researchers for both creators and consumers. A lack of formal structure for data reuse causes tension for data creators and consumers alike.
A roadmap to reduce tension between data creators and data consumers
There are multiple potential avenues for mitigating the three conflicts of interest discussed above, yet not every approach is suitable or can be realized. For instance, funding agencies have the power to set rules for data release and data reuse in principle. However, besides the differentiation between private and taxpayer-funded agencies, funders usually have diverging agendas that not only differ across political borders but are also heterogeneous within a single country. To address the current tension(s) between data creators and data consumers and to update the existing agreements from more than 15 years ago, we propose a comprehensive roadmap for data reuse (Fig. 2). We recommend following this roadmap except in cases in which institutions or funding agencies have a different policy for data reuse in place or there is a restricting licence associated with the dataset itself. This roadmap was developed with the aim of minimizing friction between data creators and data consumers, while promoting open science, and involves the introduction of a new machine-readable DRI metadata tag for facilitating communication between data creators, generators and consumers.
This flow diagram is applicable unless there is a differing policy in place by the institution using the data consumer or there is a restricting licence associated with the dataset itself. aDRI can be updated or added. bManuscripts on a preprint server or in a peer-reviewed journal are included in these recommendations. cRepositories could more easily update publication status.
Transparent, equitable and ethical use of public data necessitates clear labelling of its usability by data distributors. Although we initially considered a system that places limitations on free data use, following consultations with the Genomic Standards Consortium and polling 306 microbiome scientists, we have converged on an approach that focuses on both simplicity and openness while achieving nearly all the desired effects. The DRI tag would attach ORCIDs of the data creators to deposited data, signalling that data creators wish to be contacted (for example, via email) before data use28. This way, the DRI tag also provides stable contact information to allow data consumers to easily reach data creators. ORCIDs are both free and ubiquitous and, most importantly, are already used internally by the INSDC community. The absence of a DRI tag would signal that the authors of the dataset agree to its reuse without the need to be further involved or contacted.
The DRI will consist of a tag with one or more associated ORCIDs identifying the data creators. In computer science notation, the DRI tag will have the following structure:
DRI = {ORCID1, ORCID2, …}
We note that, implicitly, data consumers are expected to acknowledge or cite any data they use for their scientific work. For the new ethical use of data with DRI, we expect data consumers to follow the approach summarized in Fig. 2.
Sequencing datasets published in public databases (for example, in Genbank) have tags, attributes and fields that indicate which publications are connected to the respective datasets. Examples of such tags are the following: (1) for GenBank entries, ‘REFERENCE’, ‘REFERENCE/AUTHORS’, ‘REFERENCE/TITLE’ and ‘REFERENCE/JOURNAL’; (2) for BioSample, ‘reference for biomaterial’; and (3) for BioProject, ‘Publications’. The content of these tags is input initially and/or updated by the data submitter. In theory, they can be updated later either manually by the submitter or automatically by the system. Large scientific literature databases such as PubMed (https://pubmed.ncbi.nlm.nih.gov/) and Europe PMC (https://europepmc.org/) actively monitor published scientific articles and index listings of sequencing accession numbers, generating crucial linkage information that can help address updating such information in public sequence databases. However, as of 6 November 2024, only 147,632 and 78,430 publications for PubMed and Europe PMC, respectively, have had 196,632 and 91,440 sequence accession numbers assigned. Linkage information on literature and sequence data stored in these databases contrasts poorly with the scale of exponentially growing data hosted in the NCBI SRA, amounting to 9 million accession numbers as of 28 March 2023 (12 petabytes of sequence data). Coordination between publication and sequencing databases could in theory be improved if both database types use ORCID associated with their entries. In our roadmap, the DRI, which will contain the ORCID of at least one of the data creators (typically the corresponding creators, that is, the project leader), could address these issues. The content of this tag would be input or updated solely by the data submitter, preferably during the initial data deposition.
The presence of a DRI tag indicates that data creators prefer to be contacted if a data consumer reuses their data, especially if the respective data have no associated publication. The intentions behind this preference can be manifold, including a willingness to share additional metadata or datasets, or the preference to collaborate to help protect early career researchers’ (for example, PhD students) ability to finish their studies and graduate. Including one or several ORCID(s) with the DRI will provide a stable point of contact, bolster transparency in science and adherence to the FAIR principles (findable, accessible, interoperable and reusable)6 and also facilitate science through the exchange of metadata and increased collaboration. There have been instances in which such collaboration with data creators, for example, provision of additional metadata that are not publicly available, has strengthened the content of research studies29,30. This should be the norm rather than the exception. In other cases, communication with data creators has allowed proper citation of datasets used, and acknowledgement of funding that supported critical datasets, thus providing some benefit to the data creator for their data reuse29,31.
In the long run, the DRI tag would help facilitate automatic updates of the publication-associated tags. For example, a GenBank sequence entry could be updated as follows: if the GenBank entry or its corresponding BioProject and BioSample entries are mentioned in a PubMed-indexed publication together with the ORCID(s) found in the DRI, then the new publication will be automatically added to the respective publication tags. The proposed metadata tag (that is, the DRI) will substantially reduce the amount of time needed to clarify the status of any public dataset, enabling automated rules for dataset screening and reducing tension between data creators and data consumers in the future. The DRI thus bridges a gap generated by the FAIR principles, for those who are invested in making their data open access, but where it is equitable for the creator to maintain some control over its reuse.
Ideally, DRIs would propagate automatically within databases (for example, from datasets of sequencing reads to the assembled genomes) and across other online databases of -omics sequence data for a given data creator. In the absence of automatic connections, data consumers can manually add DRIs to downstream data depositions (for example, metagenome assembled genomes) via a custom metadata field. These can be the original DRIs from the original data creator or, following conversation with the data creator, new DRIs connected to the data consumer.
Outlook
In this Consensus Statement, we propose a roadmap to facilitate equitable data reuse in the microbiome field. The implementation of machine-readable labels reflecting contact information of data creators will permit efficient reuse of data, accelerate scientific discoveries and hopefully lead data creators to more readily share their valuable datasets with the scientific public at an earlier stage. Using the standards devised by the GSC32, complete metadata are expected to be made available by data creators. Here we propose the addition of a DRI tag to GSC metadata. Enabling equitable use of public microbiome data will rely on close collaborative work between the data creators, data distributors and data consumers. We envision that the GSC will play a pivotal role in helping all parties (that is, data distributors) implement a DRI tag within submission systems for microbiome data to public repositories, as well as in socializing the new approach to data mining (that is, the flowchart and algorithm given above). We note that even tangible incentives (for example, MG-RAST granting priority access to computing for data made public33) have not alleviated data creators’ hesitancy to make their data available. However, while we do not anticipate the DRI will achieve a complete resolution to this challenge, we do think it is an essential first step in the right direction.
The full adoption of a DRI tag for microbiome data in public repositories will ultimately require broad support from the scientific community, data distributors, and journals and publishing houses. As an encouraging first step, the ENA has independently implemented an ORCID metadata category, allowing data creators to attach identifying information to their submissions. Propagation of this practice to other major databases will set the stage for the DRI to be used to screen for data availability. In the meantime, data creators are encouraged to apply DRI metadata tags to their datasets. This will allow data consumers to connect with data creators to discuss data reuse. We, the Data Reuse Core Team and Data Reuse Consortium, propose that scientific manuscripts partially or exclusively making use of public data should include a written statement by the authors confirming that they have complied with these guidelines for public data use. This statement would include protocols for data download and use of analysis tools, or reproducible workflows describing how the tag was incorporated in the workflow or, in case of a missing DRI, how authors adhered to the roadmap outlined in Fig. 2.
We are aware that the implementation of the DRI could substantially impact the timeline from analysis to publication by increasing the workload (for example, needing to identify email addresses of data creators). This is especially true for projects accessing many datasets (for example, mining single genes for phylogenetic trees). We are confident that, with time, automated and standardized informatic tools will become available that will lower the administrative burden of following the DRI guidelines.
At the same time, the high proportion of participants who stated that they would respect the DRI when reusing data (266 participants (96.73%); Fig. 3) suggests that data creators will be able to more freely and more frequently share their data in the future. This would facilitate adoption of the FAIR principles in science and hugely benefit the scientific community in the long run. Moreover, by fostering collaborations, the scientific best practices, and the roadmap for data sharing in microbiome research as introduced here, will enable research by lower-resourced laboratories, reducing financial bias in scientific progress. However, we do not expect that the recommendations in this roadmap will be applied retroactively to datasets already deposited in public databases before the implementation of the DRI.
Flow height (response category, in colour) per stage (survey question, x-axis) is proportional to number of responses. See Supplementary Table 2 for respondent numbers in each category. PI, principal investigator.
The scale of nucleic acid sequence data required early pioneers to establish public databases across political borders and to confront data sharing considerations early. Other omics technologies are maturing (for example, proteomics), and scientists are recognizing the need to establish data mining approaches34. We propose that this roadmap for equitable reuse of public sequence data should be expanded to other fields including but not limited to proteomics, lipidomics, metabolomics, phenomics, microscopy and spectroscopy as data mining becomes routine with these types of data.
Change history
05 November 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41564-025-02212-3
References
The Wellcome Trust Sharing Data from Large-Scale Biological Research Projects: A System of Tripartite Responsibility (National Human Genome Research Institute, 2003).
Report of the International Strategy Meeting on Human Genome Sequencing held at the Princess Hotel, Southampton, Bermuda, on 25th–28th February 1996 (unpublished manuscript, 1996); http://hdl.handle.net/10161/7715
Toronto International Data Release Workshop Authors. Prepublication data sharing. Nature 461, 168–170 (2009).
Parties to the Convention on Biological Diversity. Nagoya Protocol on Access to Genetic Resources and the Fair and Equitable Sharing of Benefits Arising from their Utilization to the Convention on Biological Diversity 234–249 (Official Journal of the European Union, 2014); https://eur-lex.europa.eu/eli/agree_prot/2014/283/oj
GenBank and WGS Statistics (National Center for Biotechnology Information, accessed February 2024); https://www.ncbi.nlm.nih.gov/genbank/statistics/
Wilkinson, M. D. et al. The FAIR Guiding Principles for scientific data management and stewardship. Sci. Data 3, 160018 (2016).
Data Management. NIH Grants & Funding https://sharing.nih.gov/data-management-and-sharing-policy/data-management (2025).
Womack, R. P. Research data in core journals in biology, chemistry, mathematics, and physics. PLoS ONE 10, e0143460 (2015).
Zuiderwijk, A. & Spiers, H. Sharing and re-using open data: a case study of motivations in astrophysics. Int. J. Inf. Manag. 49, 228–241 (2019).
Uhlir, P. Developing Data Attribution and Citation Practices and Standards: Summary of an International Workshop (National Academies, 2012).
Amann, R. I. et al. Toward unrestricted use of public genomic data. Science 363, 350–352 (2019).
Borgman, C. L. in Theories of Informetrics and Scholarly Communication (ed. Sugimoto, C. R.) 93–116 (De Gruyter, 2016).
Cousijn, H., Feeney, P., Lowenberg, D., Presani, E. & Simons, N. Bringing citations and usage metrics together make data count 18, 9 (2019).
Rourke, M., Eccleston-Turner, M., Phelan, A. & Gostin, L. Policy opportunities to enhance sharing for pandemic research. Science 368, 716–718 (2020).
Hug, L. Contribution needed for developing a new community standard for reusing sequencing data. Springer Nature Research Communities https://communities.springernature.com/posts/contribution-needed-for-developing-a-new-community-standard-for-reusing-sequencing-data (2024).
Soares, A. R. Data Usage Manuscript - Data Deposition (OSFHOME, 2025); https://osf.io/skw4a/
Wickham, H. et al. Welcome to the Tidyverse. J. Open Source Softw. 4, 1686 (2019).
Wickham, H. et al. dplyr: a grammar of data manipulation. R package version 3.1 (2023).
Zhu, H. et al. kableExtra: construct complex table with ‘kable’ and pipe syntax. R package version 3.1 (2024).
EMBL-EBI Terms of Use. EMBL-EBI https://www.ebi.ac.uk/about/terms-of-use/ (2025).
Licensing of EMBL-EBI data resources. EMBL-EBI https://www.ebi.ac.uk/licencing/ (2025).
Sunagawa, S. et al. Tara Oceans: towards global ocean ecosystems biology. Nat. Rev. Microbiol. 18, 428–445 (2020).
Proctor, L. M. et al. The Integrative Human Microbiome Project. Nature 569, 641–648 (2019).
Lee, M. D. GToTree: a user-friendly workflow for phylogenomics. Bioinformatics 35, 4162–4164 (2019).
Chaumeil, P.-A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. GTDB-Tk: a toolkit to classify genomes with the Genome Taxonomy Database. Bioinformatics 36, 1925–1927 (2020).
Jennings, L. et al. Applying the ‘CARE Principles for Indigenous Data Governance’ to ecology and biodiversity research. Nat. Ecol. Evol. 7, 1547–1551 (2023).
Westoby, M., Falster, D. S. & Schrader, J. Motivating data contributions via a distinct career currency. Proc. R. Soc. B 288, 20202830 (2021).
Credit where credit is due. Nature 462, 825–825 (2009).
Buessecker, S. et al. An essential role for tungsten in the ecology and evolution of a previously uncultivated lineage of anaerobic, thermophilic Archaea. Nat. Commun. 13, 3773 (2022).
McKay, L. J. et al. Co-occurring genomic capacity for anaerobic methane and dissimilatory sulfur metabolisms discovered in the Korarchaeota. Nat. Microbiol. 4, 614–622 (2019).
Viljakainen, V. R. & Hug, L. A. The phylogenetic and global distribution of bacterial polyhydroxyalkanoate bioplastic-degrading genes. Environ. Microbiol. 23, 1717–1731 (2021).
Field, D. et al. The Genomic Standards Consortium. PLoS Biol. 9, e1001088 (2011).
Meyer, F. et al. The metagenomics RAST server—a public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinform. 9, 386 (2008).
Vaudel, M. et al. Exploring the potential of public proteomics data. Proteomics 16, 214–225 (2016).
Acknowledgements
The Data Reuse Core Team thanks the numerous participants of our survey who chose not to be listed as authors on this manuscript for their valuable perspectives and the many people who discussed this endeavour with us on multiple occasions. We thank L. Rothe for consultations on data visualizations. A.J.P. acknowledges funding by the German Research Foundation (DFG), CRC 1439/1 and CRC 1439/2, project number 426547801 (project INF). L.A.H. acknowledges support from the Canada Research Chairs. R.H. acknowledges support from the US National Science Foundation (OCE-2049445). C. Moraru acknowledges funding by the German Research Foundation (DFG), Priority Program SPP 2330, project number MO 3498/2-1. F.M. acknowledges support from the German Federal Ministry of Education and Research (BMBF project number 01ZZ2013).
Author information
Authors and Affiliations
Consortia
Contributions
A.J.P. was invited to write this manuscript and formed the ‘Data Reuse Core Team’ via invitation. All members of this team (A.J.P., L.A.H., A.R.S., C. Moraru, F.M. and R.H.) contributed equally to this manuscript. A.J.P., F.M. and A.R.S. led discussions with JGI, ENA and GSC. L.A.H. and A.R.S. performed the analysis and visualization of survey response data. A.H. supervised the construction of the scientific survey, ensuring the quality and impartiality of questions. The ‘Data Reuse Consortium’ provided support and feedback to this manuscript.
Corresponding author
Ethics declarations
Competing interests
R.H. was a member (2021–2024) of the User Executive Committee of the US Department of Energy’s (DOE) Joint Genome Institute. A.P. is an affiliate scientist at JGI and sits on the Prokaryotic Advisory Committee. All opinions expressed in this paper are the authors’ and do not necessarily reflect the policies and views of the DOE. R.K. is a scientific advisory board member and consultant for BiomeSense, Inc., has equity and receives income. He is a scientific advisory board member and has equity in GenCirq. He has equity in and acts as a consultant for Cybele. He is a co-founder of Biota, Inc. and has equity. He is a co-founder of Micronoma and has equity and is a scientific advisory board member. He is a board member of Microbiota Vault, Inc. He is a board member of N=1 IBS advisory board and receives income. He is a Senior Visiting Fellow of HKUST Jockey Club Institute for Advanced Study. The terms of these arrangements have been reviewed and approved by the University of California, San Diego in accordance with its conflict of interest policies.
Peer review
Peer review information
Nature Microbiology thanks Marnix Medema and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1–7, Supplementary Box 1 and Supplementary Figs. 1–10.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hug, L.A., Hatzenpichler, R., Moraru, C. et al. A roadmap for equitable reuse of public microbiome data. Nat Microbiol 10, 2384–2395 (2025). https://doi.org/10.1038/s41564-025-02116-2
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41564-025-02116-2
This article is cited by
-
Give credit where credit is due, also for omics data
EMBO Reports (2026)