Abstract
Phenazines are highly prevalent, natural bioactive substances secreted by microbes. However, their mode of action and potential involvement in shaping microbiomes remain elusive. Here we performed a comprehensive analysis of over 1.35 million bacterial genomes to identify phenazine-producing bacteria distributed across 193 species in 34 families. Analysis of rhizosphere microbiome and public rhizosphere metagenomic datasets revealed that phenazines could shape the microbial community by inhibiting Gram-positive bacteria, which was verified by pairwise interaction assays using Phenazine-1-carboxamide (PCN)-producing Pseudomonas chlororaphis. PCN induced DNA damage in Bacillus subtilis, a model Gram-positive target, where it directly bound to the bacterial topoisomerase IV, inhibiting its decatenation activity and leading to cell death. A two-species consortium of phenazine-producing Pseudomonas and resistant B. subtilis exhibited superior synergistic activity in preventing Fusarium crown rot in wheat plants. This work advances our understanding of a prevalent microbial interaction and its potential for biocontrol.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Amplicon sequencing data, genome sequences of PCN-resistant strains, and transcriptomic data of PCN-treated B. subtilis 3610 have been deposited in the Genome Sequence Archive of the Beijing Institute of Genomics (BIG) Data Center with accession numbers CRA028325, CRA028335 and CRA019862. The information on the phz+ phenazine biosynthesis gene cluster is available in the Science Data Bank at https://www.scidb.cn/s/EfmaIb (ref. 71). All data supporting the findings of this study are available in this article and its Supplementary Information. Source data are provided with this paper.
Code availability
The scripts employed in the ‘Identifier of Phenazine-Producing Bacteria (IPPB)’ pipeline are available in GitHub at https://github.com/xzliu919/Identifier-of-Phenazine-Producing-Bacteria-IPPB- (ref. 72).
References
Lee, J. Y., Tsolis, R. M. & Baumler, A. J. The microbiome and gut homeostasis. Science 377, eabp9960 (2022).
Argaw-Denboba, A. et al. Paternal microbiome perturbations impact offspring fitness. Nature 629, 652–659 (2024).
Duran, P. et al. Microbial interkingdom interactions in roots promote Arabidopsis survival. Cell 175, 973–983.e14 (2018).
Wilde, J., Slack, E. & Foster, K. R. Host control of the microbiome: mechanisms, evolution, and disease. Science 385, eadi3338 (2024).
Zhang, J. Y. et al. NRT1.1B is associated with root microbiota composition and nitrogen use in field-grown rice. Nat. Biotechnol. 37, 676–684 (2019).
Cordero, O. X. et al. Ecological populations of bacteria act as socially cohesive units of antibiotic production and resistance. Science 337, 1228–1231 (2012).
Goodrich, J. K., Davenport, E. R., Waters, J. L., Clark, A. G. & Ley, R. E. Cross-species comparisons of host genetic associations with the microbiome. Science 352, 532–535 (2016).
Lyng, M. & Kovacs, A. T. Frenemies of the soil: Bacillus and Pseudomonas interspecies interactions. Trends Microbiol. 31, 845–857 (2023).
Garcia-Bayona, L. & Comstock, L. E. Bacterial antagonism in host-associated microbial communities. Science 361, 1215 (2018).
Kehe, J. et al. Positive interactions are common among culturable bacteria. Sci. Adv. 7, eabi7159 (2021).
Dahlstrom, K. M., McRose, D. L. & Newman, D. K. Keystone metabolites of crop rhizosphere microbiomes. Curr. Biol. 30, R1131–R1137 (2020).
Krespach, M. K. C. et al. Streptomyces polyketides mediate bacteria–fungi interactions across soil environments. Nat. Microbiol. 8, 1348–1361 (2023).
Purtschert-Montenegro, G. et al. Pseudomonas putida mediates bacterial killing, biofilm invasion and biocontrol with a type IVB secretion system. Nat. Microbiol. 7, 1547–1557 (2022).
Barone, G. D. et al. Implications of bacteria–bacteria interactions within the plant microbiota for plant health and productivity. J. Zhejiang Univ. Sci. B 25, 1039–1054 (2024).
Mavrodi, D. V., Blankenfeldt, W. & Thomashow, L. S. Phenazine compounds in fluorescent Pseudomonas spp. biosynthesis and regulation. Annu. Rev. Phytopathol. 44, 417–445 (2006).
Dar, D., Thomashow, L. S., Weller, D. M. & Newman, D. K. Global landscape of phenazine biosynthesis and biodegradation reveals species-specific colonization patterns in agricultural soils and crop microbiomes. Elife 9, e59726 (2020).
Saunders, S. H. et al. Extracellular DNA promotes efficient extracellular electron transfer by pyocyanin in Pseudomonas aeruginosa biofilms. Cell 182, 919–932.e19 (2020).
McRose, D. L. & Newman, D. K. Redox-active antibiotics enhance phosphorus bioavailability. Science 371, 1033–1037 (2021).
McRose, D. L., Li, J. Y. & Newman, D. K. The chemical ecology of coumarins and phenazines affects iron acquisition by Pseudomonads. Proc. Natl Acad. Sci. USA 120, e2217951120 (2023).
Meirelles, L. A. & Newman, D. K. Phenazines and toxoflavin act as interspecies modulators of resilience to diverse antibiotics. Mol. Microbiol. 117, 1384–1404 (2022).
Moura-Alves, P. et al. AhR sensing of bacterial pigments regulates antibacterial defence. Nature 512, 387–392 (2014).
Guo, S. Q. et al. Microbial synthesis of antibacterial phenazine-1,6-dicarboxylic acid and the role of PhzG in Pseudomonas chlororaphis GP72AN. J. Agric. Food Chem. 68, 2373–2380 (2020).
Perry, E. K. & Newman, D. K. Prevalence and correlates of phenazine resistance in culturable bacteria from a dryland wheat field. Appl. Environ. Microbiol. 88, e0232021 (2022).
Kozdrój, J., Trevors, J. T. & van Elsas, J. D. Influence of introduced potential biocontrol agents on maize seedling growth and bacterial community structure in the rhizosphere. Soil Biol. Biochem. 36, 1775–1784 (2004).
Laursen, J. B. & Nielsen, J. Phenazine natural products: biosynthesis, synthetic analogues, and biological activity. Chem. Rev. 104, 1663–1685 (2004).
Chen, Y. et al. Wheat microbiome bacteria can reduce virulence of a plant pathogenic fungus by altering histone acetylation. Nat. Commun. 9, 3429 (2018).
Briard, B. et al. Pseudomonas aeruginosa manipulates redox and iron homeostasis of its microbiota partner Aspergillus fumigatus via phenazines. Sci. Rep. 5, 8820 (2015).
Hotter, V. et al. A polyyne toxin produced by an antagonistic bacterium blinds and lyses a Chlamydomonad alga. Proc. Natl Acad. Sci. USA 118, e2107695118 (2021).
Mavrodi, D. V. et al. Diversity and evolution of the phenazine biosynthesis pathway. Appl. Environ. Microbiol. 76, 866–879 (2010).
Imai, Y. et al. Evybactin is a DNA gyrase inhibitor that selectively kills Mycobacterium tuberculosis. Nat. Chem. Biol. 18, 1236–1244 (2022).
Perry, E. K., Meirelles, L. A. & Newman, D. K. From the soil to the clinic: the impact of microbial secondary metabolites on antibiotic tolerance and resistance. Nat. Rev. Microbiol. 20, 129–142 (2021).
Ran, H. M., Hassett, D. J. & Lau, G. W. Human targets of Pseudomonas aeruginosa pyocyanin. Proc. Natl Acad. Sci. USA 100, 14315–14320 (2003).
Vos, S. M., Tretter, E. M., Schmidt, B. H. & Berger, J. M. All tangled up: how cells direct, manage and exploit topoisomerase function. Nat. Rev. Mol. Cell Biol. 12, 827–841 (2011).
Laponogov, I. et al. Structural basis of gate-DNA breakage and resealing by Type II topoisomerases. PLoS ONE 5, e11338 (2010).
Zhang, G., Wang, C., Sui, Z. H. & Feng, J. Insights into the evolutionary trajectories of fluoroquinolone resistance in Streptococcus pneumoniae. J. Antimicrob. Chemother. 70, 2499–2506 (2015).
Mostertz, J., Scharf, C., Hecker, M. & Homuth, G. Transcriptome and proteome analysis of Bacillus subtilis gene expression in response to superoxide and peroxide stress. Microbiology 150, 497–512 (2004).
Keren, I., Wu, Y., Inocencio, J., Mulcahy, L. R. & Lewis, K. Killing by bactericidal antibiotics does not depend on reactive oxygen species. Science 339, 1213–1216 (2013).
Liu, Y. Y. & Imlay, J. A. Cell death from antibiotics without the involvement of reactive oxygen species. Science 339, 1210–1213 (2013).
Yan, J. J. et al. Advances in phenazines over the past decade: review of their pharmacological activities, mechanisms of action, biosynthetic pathways and synthetic strategies. Mar. Drugs 19, 610 (2021).
Gonçalves, T. & Vasconcelos, U. Colour me blue: the history and the biotechnological potential of pyocyanin. Molecules 26, 927 (2021).
Dey, T. et al. Outcomes of clofazimine for the treatment of drug-resistant tuberculosis: a systematic review and meta-analysis. J. Antimicrob. Chemother. 68, 284–293 (2013).
Appelbaum, P. C. & Hunter, P. A. The fluoroquinolone antibacterials: past, present and future perspectives. Int. J. Antimicrob. Agents 16, 5–15 (2000).
Swanson, K. et al. Generative AI for designing and validating easily synthesizable and structurally novel antibiotics. Nat. Mach. Intell. 6, 338–353 (2024).
Su, P. et al. Microbiome homeostasis on rice leaves is regulated by a precursor molecule of lignin biosynthesis. Nat. Commun. 15, 23 (2024).
Netzker, T., Shepherdson, E. M. F., Zambri, M. P. & Elliot, M. A. Bacterial volatile compounds: functions in communication, cooperation, and competition. Annu. Rev. Microbiol. 74, 409–430 (2020).
He, Y. W. et al. DSF-family quorum sensing signal-mediated intraspecies, interspecies, and inter-kingdom communication. Trends Microbiol. 31, 36–50 (2023).
Molina-Santiago, C. et al. The extracellular matrix protects Bacillus subtilis colonies from Pseudomonas invasion and modulates plant co-colonization. Nat. Commun. 10, 1919 (2019).
Molina-Santiago, C. et al. Chemical interplay and complementary adaptative strategies toggle bacterial antagonism and co-existence. Cell Rep. 36, 109449 (2021).
Korgaonkar, A., Trivedi, U., Rumbaugh, K. P. & Whiteley, M. Community surveillance enhances Pseudomonas aeruginosa virulence during polymicrobial infection. Proc. Natl Acad. Sci. USA 110, 1059–1064 (2013).
Giddens, S. R., Feng, Y. & Mahanty, H. K. Characterization of a novel phenazine antibiotic gene cluster in Erwinia herbicola Eh1087. Mol. Microbiol. 45, 769–Eh1783 (2002).
Pierson, L. S. & Pierson, E. A. Metabolism and function of phenazines in bacteria: impacts on the behavior of bacteria in the environment and biotechnological processes. Appl. Microbiol. Biotechnol. 86, 1659–1670 (2010).
Mentel, M. et al. Of two make one: the biosynthesis of phenazines. ChemBioChem 10, 2295–2304 (2009).
Li, J., Wu, Q. & Zhu, X. Research progress of phenazine-1-carboxylic acid and its analogue. Chin. J. Org. Chem. 39, 2744–2758 (2019).
Madeira, F. et al. The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res. 47, W636–W641 (2019).
McDonald, M., Mavrodi, D. V., Thomashow, L. S. & Floss, H. G. Phenazine biosynthesis in Pseudomonas fluorescens branchpoint from the primary shikimate biosynthetic pathway and role of phenazine-1,6-dicarboxylic acid. J. Am. Chem. Soc. 123, 9459–9460 (2001).
Shemesh, M. & Chai, Y. A combination of glycerol and manganese promotes biofilm formation in Bacillus subtilis via histidine kinase KinD signaling. J. Bacteriol. 195, 2747–2754 (2013).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013).
Rognes, T., Flouri, T., Nichols, B., Quince, C. & Mahé, F. VSEARCH: a versatile open source tool for metagenomics. PeerJ 4, e2584 (2016).
McMurdie, P. J. & Holmes, S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8, e61217 (2013).
Segata, N. et al. Microbial community function and biomarker discovery in the human microbiome. Genome Biol. 12, P47 (2011).
Chen, I. M. A. et al. IMG/M: integrated genome and metagenome comparative data analysis system. Nucleic Acids Res. 45, D507–D516 (2017).
Wood, D. E., Lu, J. & Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Biol. 20, 257 (2019).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60 (2014).
Gryczan, T. J., Contente, S. & Dubnau, D. Characterization of Staphylococcus aureus plasmids introduced by transformation into Bacillus subtilis. J. Bacteriol. 134, 318–329 (1978).
Chai, Y., Chu, F., Kolter, R. & Losick, R. Bistability and biofilm formation in Bacillus subtilis. Mol. Microbiol. 67, 254–263 (2008).
Ben-Yehuda, S., Rudner, D. Z. & Losick, R. RacA, a bacterial protein that anchors chromosomes to the cell poles. Science 299, 532–536 (2003).
Peters, J. M. et al. A comprehensive, CRISPR-based functional analysis of essential genes in bacteria. Cell 165, 1493–1506 (2016).
Methods for Dilution Antimicrobial Susceptibility Test for Bacteria that Grow Aerobically: Approved Standard CLSI Document M07-A9, 19th edn (CLSI, 2012).
Zhou, Y. & Chen, Y. phz+ strains data [DS/OL]. V1. Science Data Bank https://doi.org/10.57760/sciencedb.25989 (2025).
Liu, C. Identifier-of-Phenazine-Producing-Bacteria (IPPB). GitHub https://github.com/xzliu919/Identifier-of-Phenazine-Producing-Bacteria-IPPB- (2025).
Acknowledgements
We thank R. Losick (Harvard University, USA) for his suggestions during the revision of the paper; K. Lewis (Northeastern University, USA) for providing the E. coli MG1655, WO153 and ΔtolC mutant strains; and J. Feng (Chinese Academy of Sciences, China) for providing the S. pneumoniae R6WT and R6M-3 strains. This research was supported by the Natural Science Foundation of Zhejiang Province (LZ23C140004) to Y. Chen, National Natural Science Foundation of China U21A20219 to Z.M., National Key R&D Program of China 2022YFD1400100 to Y. Chen, National Natural Science Foundation of China U24A20808 to H.L., and ‘Pioneer’ and ‘Leading Goose’ R&D Program of Zhejiang 2023C02030 to Y. Chen.
Author information
Authors and Affiliations
Contributions
Y. Chen initiated, coordinated and supervised the project. Y.Z. performed and analysed most of the experiments. Y.Z., H.W. and J.S. performed MIC tests. C.L. collected and analysed the phz+ strains data. T.C. and W.A.W. collected and analysed the metagenome and metatranscriptome datasets. Y.H. performed ROS staining experiments. Y.Q. constructed the KD-ParE/C and KD-GyrA/B strains. L.L. and H.L. performed structural determination of phenazine-1-carboxylic acid. Y. Chen and Y.Z. wrote the paper. Y. Chen, Z.M., T.C., Y.B., Y. Chai and G.B. revised the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Wulf Blankenfeldt, Axel Brakhage and Ákos T Kovács for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 IPPB pipeline for identifying phz+ bacteria.
a, Schematic representation of the Identifier of Phenazine-Producing Bacteria (IPPB) pipeline. The pipeline automates the search for homologous proteins of phenazine biosynthetic core proteins (PhzA/B, PhzC, PhzD, PhzE, PhzF, PhzG) within a strain’s genome. Protein identity thresholds are applied as indicated, and strains are classified as phz+ if all core proteins are identified and the corresponding genes are located within 20 kb of each other. Strains meeting these criteria are classified as phz+ bacterium, and included in the phylogenetic tree. Abbreviations: NGDC, National Geoscience Data Centre; IMG/M, Integrated Microbial Genomes & Microbiomes system. b, Schematic representation of 23 reference phenazine biosynthetic gene clusters. Core biosynthetic genes (phzA/B, phzC, phzD, phzE, phzF, phzG) are shown in color, while adjacent, non-core genes are depicted in grey.
Extended Data Fig. 2 Phylogenetic profiling and structural determination of phenazine-1-carboxylic acid produced in Spongiactinospora rosea LHW63015.
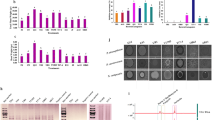
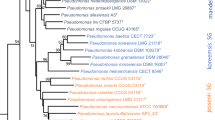
a, Maximum-likelihood phylogenetic tree of the phz+ bacterial genera based full-length 16S rRNA gene sequences. Phylum-level classifications are indicated. Bar plots on the right show the number of species within each phz+ genus and the total number of phz+ strains. b, 1H NMR (nuclear magnetic resonance) spectrum of phenazine-1-carboxylic acid (PCA) produced in S. rosea LHW63015. c, 13C NMR spectrum of PCA. d, DEPT135 NMR spectrum of PCA. e, X-ray ORTEP drawing for PCA.
Extended Data Fig. 3 phz+ bacteria drive shifts in soil microbiomes.
a, Number of ASVs in samples treated with phz+ ZJU60 and its phenazine mutant ΔphzA-H. b, Shannon diversity index in samples treated with phz+ ZJU60 and ΔphzA-H. c, Community composition in samples treated with phz+ ZJU60 and ΔphzA-H. d, Dynamics of wheat rhizosphere microbiota following soil irrigation with ZJU60 and ΔphzA–H. Soil samples were collected on days 3 and 7 after irrigation with ZJU60 and ΔphzA–H in wheat field microcosms. 16S rRNA gene fragment amplicon sequencing revealed distinct temporal changes in the composition and structure of the rhizosphere microbiomes. Data are presented as means ± s.d. of n = 5 biological replicates. For graphs shown in a, b and d, letters that are different from one another indicate that their means are statistically different using a two-sided ANOVA and Tukey’s HSD (P < 0.05).
Extended Data Fig. 4 Correlation analysis between the relative abundances of phz+ Pseudomonas species and other bacterial taxa within rhizosphere microbiomes.
a, Correlation analysis between the relative abundances (RA) of phz+ bacteria and Pseudomonas in rhizosphere microbiomes. b-d, Pairwise Spearman correlation between the abundance of the three most prevalent bacterial taxa at each taxonomic level and the abundance of phz+ Pseudomonas. The relative abundances of the analyzed taxa were obtained from shotgun-sequenced metagenome datasets and subsequently log-transformed. The Spearman correlation coefficient (R) and corresponding P value was calculated using the ggscatter function from the ggpubr package in R. The grey shaded area indicates the 95% confidence interval of the regression line, shown in blue.
Extended Data Fig. 5 Correlation analysis between the phzA/BDEFG gene transcripts and the abundances of bacteria assigned to the class Bacilli and to the genus Pseudomonas.
a, Correlation analysis between phzA/BDEFG gene transcripts (log-transformed counts per million, CPM) and the relative abundances (RA) of bacteria assigned to the class Bacilli. b, Correlation analysis between the phzA/BDEFG gene transcripts and the relative abundance of bacteria assigned to the genus Pseudomonas. Metatranscriptomic datasets were used to determine both gene expression and the relative abundances of the target taxa. The Spearman correlation coefficient (R) and corresponding P value (P < 0.05) were calculated using the ggscatter function from the ggpubr package in R. The grey shaded area indicates the 95% confidence interval of the regression line, shown in blue.
Extended Data Fig. 6 Antimicrobial activity of Pseudomonas chlororaphis ZJU60 against representative strains isolated from the wheat microbiome.
The antimicrobial activity of ZJU60 was assessed on LBGM agar medium against 70 bacterial species. One representative strain from each species was selected for testing. ZJU60 was inoculated on the top of a lawn of the individual test strains, and inhibition zones were indicated by dashed circles after 72 h of incubation. Scale bar = 5 mm. Detailed information on the tested strains can be found in Supplementary Table 1.
Extended Data Fig. 7 PCN is the primary antibacterial compound produced by P. chlororaphis ZJU60.
a, b, Time-resolved interactions between P. chlororaphis ZJU60 or ΔphzA-H and B. subtilis 3610-GFP over 24, 48, and 72 h in LBGM agar medium (left panel). The log(CFU·mL⁻¹·cm⁻²) of ZJU60 or ΔphzA-H and 3610 in the interaction areas at different time points (24, 48, 72 h) is shown in the right panel. Cells from the three sections of the interaction zone were collected, sonicated, fully re-suspended, diluted, and plated to determine the CFU of each species. Scale bar = 8 mm. c, Time-resolved interactions between ΔphzA-H (supplemented with 50 µg/mL PCN) and B. subtilis 3610 over 24, 48, and 72 h in LBGM medium. d, Growth curves of B. subtilis 3610 with ZJU60 supernatant, ΔphzA-H supernatant, and ΔphzA-H supernatant supplemented with 50 µg/mL PCN. Data for graphs in a, b and d are presented as means ± s.d. For a and b, n = 3 biological replicates. For d, n = 6 biological replicates.
Extended Data Fig. 8 Morphological changes in four PCN-sensitive bacterial strains following PCN treatment.
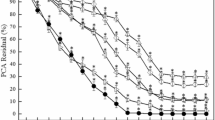
Differential interference contrast (DIC) images (left) showing cell morphology of four PCN-sensitive bacterial strains in the absence and presence of PCN. Scale bar = 5 μm. Quantification of cell length from images (right). The graphs represent results from one representative experiment. Statistical differences were analyzed by a two-sided Student’s t-test. Data are presented as means ± s.d. of n = 50 individual cells. Experiments were independently repeated three times with similar results.
Extended Data Fig. 9 ROS is not the primary contributor to DNA damage induced by PCN.
a, Fluorescence staining of reactive oxygen species (ROS) signals following PCN treatment of B. subtilis 3610. Scale bar = 10 μm. b, Quantification of fluorescence intensity from panel a. Data are presented as mean ± s.d. of n = 78 individual cells. Differences between groups were analyzed using a two-sided Student’s t-test. c, Analysis of the expression of ROS response-related mutant genes following PCN treatment. d, Growth curves for B. subtilis 3610 and ROS-scavenging mutants, with or without treatment with 15 μg/mL PCN. Data are presented as means ± s.d. of n = 3 biological replicates.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–6 and Tables 1 and 2.
Supplementary Data 1 (download XLSX )
Supplementary Data 1–10.
Supplementary Data 2 (download XLSX )
Source data for Supplementary Fig. 3.
Source data
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download PDF )
Unprocessed western blots and gels.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download PDF )
Unprocessed gels.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhou, Y., Wang, H., Sun, J. et al. Phenazines contribute to microbiome dynamics by targeting topoisomerase IV. Nat Microbiol 10, 2396–2411 (2025). https://doi.org/10.1038/s41564-025-02118-0
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41564-025-02118-0
This article is cited by
-
Ecological function of phenazine in soil
Nature Microbiology (2025)