Abstract
Candida albicans is a common fungal member of the human microbiota but can also cause infections via expression of virulence factors associated with the yeast-to-hyphae transition. The evolutionary selection pressure to retain these pathogenic traits for a commensal microorganism remains unclear. Here we show that filamentation and hyphae-associated factors, including the toxin candidalysin, are crucial for colonization of the oral cavity, a major reservoir of C. albicans. Low-virulent strains of C. albicans expressed the candidalysin-encoding gene ECE1 transiently upon exposure to keratinocytes in vitro. In mice, ECE1 mutants were defective at accessing terminally differentiated oral epithelial layers where the fungus is protected from IL-17-mediated immune defence. Tight regulation of ECE1 expression prevented detrimental effects of candidalysin on the host. Our results suggest that hyphae-associated factors such as candidalysin govern not only pathogenicity, but also mucosal colonization through direct host interactions enabling C. albicans to create and maintain its niche in the oral mucosa.
Similar content being viewed by others
Main
Microorganisms colonizing human epithelial tissues are crucial for maintaining and modulating host barrier homeostasis and physiology. While microbiome research has primarily focused on bacteria, recent studies underscore the beneficial effects of commensal fungi for the host. Candida albicans can enhance resistance to infections via the induction of type 17 immunity1,2,3. Yet, C. albicans is also a pathobiont causing mucosal infections4,5. Furthermore, C. albicans can cause life-threatening systemic disease6 and is implicated in non-infectious chronic inflammatory disorders, including inflammatory bowel disease, hepatitis and airway allergies2,7,8,9. These opposing roles emphasize the importance of understanding how C. albicans maintains a balanced homeostatic relationship with its host.
Several studies suggest that the yeast morphology and repression of hyphae-associated genes are required for C. albicans gut colonization, while hyphal growth and associated factors are linked to pathogenicity3,10. Such a scenario would favour avirulent strains3. However, most clinical isolates of C. albicans retain considerable virulence potential11, raising the question why such traits have been conserved. One explanation is that hyphae may also play non-pathogenic roles during colonization12,13. In fact, hyphal formation and candidalysin—a hypha-associated peptide toxin encoded by the ECE1 gene and known to damage epithelial barriers and trigger inflammation14,15—have been shown to support gut colonization by inhibiting competing bacteria16.
Beyond the gut, C. albicans also inhabits the oral cavity, a potential primary reservoir for fungal colonization11,17. The oral mucosa, with its cornified stratified epithelium and continuous exposure to environmental stimuli, differs markedly from the gut and presents unique colonization challenges18. Unlike in the gut, where C. albicans is found in the lumen10,16, in the oral mucosa it resides within the uppermost layer of the stratified epithelium19,20. However, the fungal factors supporting stable colonization in this niche are not well defined.
Here we show that C. albicans relies on filamentation and candidalysin to establish and maintain oral colonization. In this process, candidalysin can breach the stratum corneum supporting growth and immune evasion without host adverse effects. These findings suggest that classical virulence traits have evolved not primarily for pathogenicity but to promote stable mucosal colonization.
Results
Filamentation and hyphae-associated genes are required for C. albicans oral colonization
The requirement of filamentation for the initiation of C. albicans oral colonization was tested by comparing the filamentation-competent wild type with yeast-locked mutant strains, which are unable to filament owing to the deletion of essential gene(s) of the yeast–hyphae transition, in a model of oral colonization in immunocompetent and not antibiotically treated mice. Three distinct yeast-locked mutant strains, SC5314efg1Δ/Δ (ref. 21), SC5314efg1/cph1Δ/Δ (ref. 22) and SC5314flo8Δ/Δ (ref. 23) showed a drastically reduced fungal burden compared with the parental strain24 by day 1 after fungal administration (Fig. 1a). We then tested the implication of hyphae-associated genes25 in this process known to act at the C. albicans–epithelial cell interface (Extended Data Fig. 1a,b). C. albicans mutants lacking the adhesin Hwp1 (SC5314hwp1Δ/Δ)26, the secreted aspartyl proteases 4–6 (SC5314sap456Δ/Δ)27 or the gene ECE1 encoding candidalysin (SC5314ece1Δ/Δ)14, but not strains lacking HYR1 (SC5314hyr1Δ/Δ)28 or SAP1–3 (SC5314sap123Δ/Δ)27, showed severely impaired host association (Fig. 1b–d and Extended Data Fig. 1c,d). The requirement for filamentation and hyphae-associated genes was evident as early as 8 h after fungal association (Fig. 1e–h and Extended Data Fig. 1e). Specific deletion of Ece1 peptide III-encoding sequences14, the precursor of candidalysin, confirmed that candidalysin was responsible for the observed effect of Ece1 (Fig. 1f). By 2 h, yeast cells adhered topically to the tongue epithelium, and by 8 h, individual filaments breached through the epithelial barrier (Fig. 1i). Penetration of the stratum corneum appeared to present a bottleneck with only few fungal cells succeeding. This was also reflected by the drastically reduced number of fungal cells recovered from the tongue tissue by 8 h, while thereafter, the fungus proliferated and reached high fungal loads by 24 h (Fig. 1j). The initial focal interaction of C. albicans with the epithelium was independent of filamentation (Fig. 1k) and formation of microcolonies, which were described as a specialized biofilm structure regulated by Sfl1 or Sfl2 (ref. 29; Extended Data Fig. 1f,g).
C57BL/6 wild-type (WT) mice were associated with C. albicans strain SC5314 (red symbols) or 101 (blue symbols) via sublingual administration. Gene deletion mutants (open symbols) were compared with their corresponding parental strain. CFU per gram of tongue tissue are shown at the indicated time points. a–h, Tongue CFU of SC5314efg1∆/∆, SC5314efg1/cph1∆/∆ and SC5314flo8∆/∆ (a, n = 6 per group; e, n = 8 per group; mean ± s.e.m.); SC5314ece1∆/∆ and SC5314ece1p3∆/∆ (b, n = 6 or 7 per group; f, n = 8 per group; mean ± s.e.m.); SC5314hwp1∆/∆ (c, g, n = 8 per group, mean ± s.e.m.); and SC5314sap4-6∆/∆ (d, h, n = 8 per group, mean ± s.e.m.) compared with the parental WT each at 1 day (a–d) or 8 h (e–h). i,j, PAS-stained tongue tissue sections (i) and tongue CFU at 2 h, 8 h and 24 h of colonization with SC5314WT or 101WT (j, n = 5–9 per group and time point, mean ± s.e.m.). The black arrows indicate fungal elements. k, Tongue CFU of SC5314efg1/cph1∆/∆ at 2 h; n = 8 per group; mean ± s.e.m. l–n, Tongue CFU of 101ece1∆/∆ and 101ece1p3∆/∆ (l; n = 9, 15 or 19 per group), 101hwp1∆/∆ (m; n = 4 or 8 per group, 2 different clones indicated by circles and triangles), 101sap4-6∆/∆ (n; n = 4 or 12 per group, 3 different clones indicated by circles, triangles and squares), all 101 background, at 8 h. Data presented as mean ± s.e.m. o, Tongue CFU of SC5314WT and 101WT at 8 h; n = 8 per group; mean ± s.e.m. Each data point represents an individual mouse. Data are pooled from at least 2 independent experiments. DL, detection limit; NS, not significant. The statistical significance of differences between groups was determined by ordinary one-way ANOVA (a, b, f, l, j), two-sided Mann–Whitney test (g) or unpaired two-tailed t-test (c, d, e, h, k, m, n, o), respectively.
To assess whether the process by which C. albicans associates with the tongue tissue was conserved among strains bearing a lower intrinsic capacity to filament and to express hyphae-associated genes than strain SC5314, we turned to low-virulent strain 101, originally isolated from a healthy child and bearing a low intrinsic capacity to cause epithelial cell damage19,20,30. The dynamics of filamentation, initial adhesion to and invasion of the stratum corneum and fungal load were remarkably similar between the two strains (Fig. 1i,j). In contrast to strain SC5314, which penetrated across the entire stratified epithelium of the murine tongue, strain 101 was confined to the stratum corneum, partially reverted back to the yeast and pseudohyphal morphology by 24 h and did not elicit tissue inflammation (Fig. 1i)19,20,31.
To further corroborate these findings and to test the implication of virulence determinants beyond filamentation in oral colonization, we deleted the hyphae-associated genes ECE1, HWP1 or SAP4-6 in strain 101 (101ece1Δ/Δ, 101hwp1Δ/Δ, 101sap456Δ/Δ). Consistent with results obtained with mutants of the high-virulent strain SC5314 (Fig. 1b,f), candidalysin, but not Hwp1 or Sap4–6, was an important strain-independent factor for the establishment of oral colonization (Fig. 1l–n). The selective dependence on Ece1 may explain the slightly inferior capacity of strain 101 compared with SC5314 to settle colonization by 8 h (Fig. 1o).
Low-virulent C. albicans acquires hallmarks of virulence for the establishment of oral colonization
The requirement of Ece1 and candidalysin for colonization by strain 101 appeared inconsistent with its low expression of ECE1 (ref. 19). We therefore investigated the ECE1 gene expression profile during the first 24 h of C. albicans–epithelial cell interaction in vitro, using human epidermal equivalents (HEEs) (Fig. 2a). The marked filamentation and epithelial invasion of both strains, SC5314 and 101, during the initial phase of colonization were largely comparable (Fig. 2a). ECE1 expression was induced massively in a strain-independent manner within 2 h (Fig. 2b) and rapidly declined thereafter in the case of strain 101, but not SC5314 (Fig. 2b). The acute and transient induction of ECE1 expression was recapitulated in TR146 oral keratinocytes and even in cell-free serum-containing medium, with even more transient expression kinetics for strain 101 despite persistent filamentation (Fig. 2c and Extended Data Fig. 2a,b). Similar gene expression profiles were also observed for the hyphae-associated genes HWP1 and SAP5 in strains SC5314 and 101 (Extended Data Fig. 2c,d), but not in the yeast-locked C. albicans SC5314efg1/cph1Δ/Δ mutant (Extended Data Fig. 2e). We also detected ECE1 expression in the tongue tissue colonized for 24 h by both strains (Fig. 2d). In agreement with the in vitro data, ECE1 mRNA levels were lower for strain 101 compared with SC5314, although differences were less dramatic than in vitro (Fig. 2d). Together, the highly dynamic and tight regulation of hyphae-associated genes, ECE1 in particular, may enable the establishment of colonization within hours of C. albicans–host cell interaction.
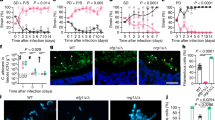
a–c, HEEs (a,b) and TR146 keratinocytes (c) were infected with C. albicans strain SC5314WT or 101WT for 2, 8 or 24 h, respectively, and analyzed by histology (a, PAS-stained HEE sections) and expression of ECE1 transcripts (b,c); data are presented as mean per group ± s.d. The black arrows indicate fungal elements. d,e, C57BL/6 WT mice were associated with SC5314WT or 101WT, and ECE1 transcript expression was quantified after 1 day (d, n = 8 or 10 per group, mean ± s.e.m.) or after 1 and 7 days (e, n = 3 per group, mean ± s.d.). f–h, C57BL/6 WT mice were associated with 101WT, a 50× reduced inoculum of 101WT (101WT l.d.), 101ece1Δ/Δ or 101ece1p3Δ/Δ and tongue CFU were enumerated on day 1 (f), day 7 (g) or day 28 (h), n = 8–31 per group; mean ± s.e.m. i, C57BL/6 WT mice were associated with 101tetOff-ECE1 and treated with doxycycline (Dox) to induce ECE1 deletion from day 21 to day 35, before tongue CFU were enumerated, n = 8 per group; mean ± s.e.m. j,k, Competitive colonization of C57BL/6 WT mice with 101WT_mCherry and 101ece1Δ/Δ at a 1:1 ratio for 3 days (j) or at a 1:50 ratio for 3, 7 and 28 days (k) before fluorescent and non-fluorescent tongue CFU were enumerated; n = 6 or 7 per group. Connected symbols are 101WT and 101ece1Δ/Δ CFU per individual animal (left) and fold differences between the two strains per animal (right). Data are presented as mean per group ± s.e.m. Data in a–c and e are from one representative of 3 independent experiments. Data in d and f–k are pooled from at least 2 independent experiments. The statistical significance of differences between groups was determined by two-way ANOVA (k (left)), repeated-measures two-way ANOVA (b,c), ordinary one-way ANOVA (f,g,h), Kruskal–Wallis test (k (right)), two-sided Mann–Whitney test (e), unpaired two-tailed t-test (d,i) or paired two-tailed t-test (j). rel. expr., relative expression.
Candidalysin is required for sustained oral colonization
Continuous ECE1 expression (Fig. 2e), which implies candidalysin’s necessity beyond initial niche establishment, prompted us to perform a kinetics experiment in colonized mice to evaluate the importance of Ece1 over time. Similarly to what we observed at 8 h (Fig. 1l), the tissue fungal burden of 101ece1Δ/Δ was at least 5× reduced at 24 h compared with the parental 101WT strain (Fig. 2f and Extended Data Fig. 2f). The difference was even more pronounced by day 7 and 28 (Fig. 2g,h). A similar colonization defect was also observed with 101ece1p3Δ/Δ (Fig. 2g,h), confirming the relevance of candidalysin in the Ece1-mediated process. When lowering the 101WT inoculum 50× relative to 101ece1Δ/Δ to achieve even colonization levels of both strains at day 1 (101WT low infection dose (l.d.); Fig. 2f), the colonization defect of 101ece1Δ/Δ persisted, while the fungal burden of 101WT caught up over time (Fig. 2g,h). We then engineered strain 101 to achieve doxycycline-inducible repression of ECE1 (101tetOff_ECE1; Extended Data Fig. 2g). Doxycycline administration from day 21, when colonization was established, resulted in almost complete clearance of the fungus by day 35 (Fig. 2i), while doxycycline did not affect colonization of 101WT (Extended Data Fig. 2h). To further address whether ECE1 provides a fitness advantage, we performed a series of competition experiments with a fluorescent variant of 101WT (101WT_mCherry (ref. 32), showing comparable colonization properties as 101WT; Extended Data Fig. 2i) and non-fluorescent 101ece1Δ/Δ. When co-colonizing mice with a 1:1 mixture, 101WT drastically outcompeted 101ece1Δ/Δ by day 3 (Fig. 2j). When using the two strains at a 1:50 ratio to eliminate the initial colonization difference, 101WT again outcompeted 101ece1Δ/Δ by day 7 and fully eliminated 101ece1Δ/Δ by day 28 (Fig. 2k), confirming the fundamental role of ECE1 not only to initiate but also to sustain fungal colonization. The predominant role of ECE1 over other hyphae-associated genes was confirmed by the observation that neither 101hwp1Δ/Δ nor 101sap4−6Δ/Δ showed a defect in oral colonization by day 7 (Extended Data Fig. 2j).
Candidalysin confers resistance to host antifungal responses
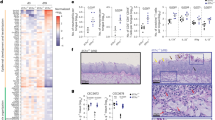
Contrary to the gut, where ECE1 provided a competitive advantage to C. albicans by impairing bacterial metabolic activity16, the fitness advantage provided by ECE1 in the murine oral mucosa appeared to be independent of antibiotically induced dysbiosis (Fig. 3a and Extended Data Fig. 3a,b). The ECE1-dependent competitive advantage (Fig. 2k) coincided with the manifestation of IL-17 antifungal immunity rising from day 3 after fungal administration (Fig. 3b)19,31, including IL-17 target genes as evidenced by RNA sequencing (RNAseq; Fig. 3c,d). The IL-17 pathway is critical for preventing C. albicans overgrowth, which manifests as mucocutaneous candidiasis in IL-17-deficient patients33. Therefore, we assessed whether ECE1 deficiency affects fungal colonization of the oral mucosa in IL-17 receptor-deficient mice. Surprisingly, the fitness of strain 101ece1Δ/Δ compared with that of 101WT (50× reduced inoculum) was restored in Il17rc−/− mice (Fig. 3e and Extended Data Fig. 3c). A co-colonization experiment further confirmed that the competitive fitness advantage of 101WT over 101ece1Δ/Δ was lost in the absence of IL-17 signalling (Fig. 3f). Together, these results indicate that C. albicans also requires Ece1 to resist the antifungal response and to sustain oral colonization independently of the oral microbiota.
a, Competitive colonization of C57BL/6 WT mice with 101WT_mCherry and 101ece1Δ/Δ at a 1:50 ratio for 7 days before fluorescent and non-fluorescent tongue CFU were enumerated. One group of mice was treated with ampicillin in the drinking water from day −4 and −3 until the end of the experiment. Tongue CFU of either strain were determined based on the presence or absence of fluorescence, n = 7 per group. Connected symbols are 101WT and 101ece1Δ/Δ CFU per individual animal (left) and fold differences between the two strains per animal (right) in a. Data are presented as mean per group ± s.e.m. b,c, Differentially regulated genes (log2(fold changes)) linked to antifungal immune response (b) and epidermal keratinization and antimicrobial activity (c) in the tongue of C57Bl/6 WT mice colonized with 101WT for the indicated period of time compared with naive mice. Data are extracted from a previously published RNAseq dataset31. d, Differential expression of the genes shown in c in Il17rc−/− compared with those in Il17rc+/− mice colonized with 101WT for 3 days. Data are from an RNAseq dataset deposited in the NCBI GEO repository with accession number GSE280210. e, Il17rc−/− mice were associated with 101WT (1/50 inoculum) or 101ece1Δ/Δ, and tongue CFU were enumerated on day 19; n = 4 or 12 per group; mean ± s.e.m. f, Competitive colonization of Il17rc−/− and Il17rc+/− mice with 101WT_mCherry and 101ece1Δ/Δ at a 1:50 ratio for 7 days; n = 7 per group. Fluorescent and non-fluorescent tongue CFU were enumerated and fold differences between the two strains per animal are shown as in a; data presented as mean per group ± s.e.m. Data in a, e and f are pooled from at least two independent experiments. Paired datapoints in competitive colonization experiments (a,f) are connected with a line. The statistical significance of differences between groups was determined by two-way ANOVA (a,f (left)), unpaired two-tailed t-test (a (right), e), two-sided Mann–Whitney test (f (right panel)).
Candidalysin promotes oral colonization by enabling C. albicans to overcome the barrier of the stratum corneum
To understand the mechanism underlying Ece1-mediated colonization, we found, in line with the literature14, that ECE1 did not affect C. albicans adhesion to epithelial cells in culture, filamentation nor cellular invasion (Extended Data Fig. 4a–e). Candidalysin is known as a peptide toxin, which, at high concentrations, intercalates in host cell membranes, thereby inducing lytic cell death14,34,35. In contrast to strain SC5314 (ref. 34), strain 101 does not induce cellular damage in vitro (Fig. 4a and Extended Data Fig. 4f). At high fungal cell concentrations, strain 101 did, however, provoke host cell damage in spontaneously immortalized murine oral keratinocyte (IMOK) cells36 showing an increased sensitivity towards cell death induction compared with the more commonly used TR146 cell line. The response by IMOK cells was independent of ECE1, HWP1 or SAP4–6 factors (Extended Data Fig. 4g–j), which may be explained by the limited gene expression levels of strain 101 (Fig. 4b and Extended Data Fig. 4k). To assess the consequences of ECE1 under conditions more closely representing the situation in vivo in which expression levels by strain 101 are only slightly lower than in SC5314 (Fig. 4b), we aimed at overexpressing ECE1 in strain 101. Expression under control of the TDH3 or HWP1 promoters did not result in increased induction of cellular damage (Extended Data Fig. 4l)20. We then took advantage of strain 101tetOff_ECE1, which, in the absence of doxycycline, overexpressed ECE1 in vitro (but not in vivo) owing to the insertion of a strong promoter in strain 101 to levels comparable to those in strain SC5314 (Extended Data Fig. 4m,n). Compared with the parental strain 101WT, 101tetOff_ECE1 (-doxycycline) elicited enhanced lactate dehydrogenase (LDH) release from IMOK cells (Fig. 4a) and did so in a contact-dependent manner (Fig. 4c), as expected. Strain 101 is thus indeed capable of inducing cellular damage in an ECE1-dependent manner despite the complex regulation of candidalysin-mediated cellular damage beyond ECE1 expression34.
a, LDH release from IMOK cells in response to SC5314WT, SC5314ece1Δ/Δ 101WT and 101tetOff_ECE1 after 24 h of incubation (n = 3 or 6 per group). Each symbol represents an individual well with mean per group ± s.d. Data are from one representative of at least five independent experiments. b, Ratio of ECE1 expression by SC5314 versus 101 after 24 h of exposure to IMOK keratinocytes, HEEs or murine tongue tissue (in vivo). Each symbol represents an independent experiment (mean of 1–3 replicates each) with mean ± s.d. per group. c, LDH release from IMOK cells in response to 101WT or 101tetOff_ECE1 (1 × 105 yeast cells per well) in direct contact (cc) or transwell (TW) for 24 h (n = 3 or 4 per group). Each symbol represents an individual well with the mean per group ± s.e.m. Data are pooled from 2 independent experiments. d–f, C57Bl/6 WT mice were associated with 101WT or 101ece1Δ/Δ for 2 h; tongues were collected and incubated for 6 h (d,e) or 22 h (f) in a humid chamber at 37 °C, 5% CO2. Representative PAS-stained tongue tissue sections (d,f) and quantification of fungal foci outside the host, in the stratum corneum or in nucleated epithelial layers (e). Each symbol represents the percentage of all events per category and section from an individual mouse (>28 events per section); n = 3 animals per group with mean ± s.d. per group. g,h, Representative PAS-stained tongue tissue sections from C57Bl/6 WT mice (g) or Il17rc−/− mice (h) associated with 101WT or 101ece1Δ/Δ for 7 days (g) or 19 days (h). The yellow lines indicate the boundary between the stratum corneum and stratum granulosum. The black arrows indicate fungal elements. Data are pooled from at least 2 independent experiments. i–j, The tongue of C57Bl/6 mice was (+) or was not (−) mechanically disrupted before the association with 101ece1Δ/Δ Tongue CFU on day 1 (i) and representative histology sections of tongues at 2 h plus additional incubation in a humid chamber at 37 °C, 5% CO2, for 6 h (j); n = 4 pooled from 2 independent experiments with mean ± s.e.m. The statistical significance of differences between groups was determined by two-way ANOVA (a,c,e) or ordinary one-way ANOVA (b).
We therefore returned to our in vivo model to assess the mechanism by which candidalysin facilitates the establishment of C. albicans colonization in the complex environment of the stratified oral epithelium. Because too few fungal elements were retrievable at 8 h to observe invasion on tissue sections (Fig. 1i,j,o), we instead removed tongues 2 h after fungal association—minimizing loss to saliva and mechanical forces—and allowed ex vivo completion of the colonization process. Loss of ECE1 completely prevented C. albicans from invading the epithelium (Fig. 4d). Remarkably, fungal elements of both 101ece1Δ/Δ and SC5314ece1Δ/Δ strains remained peripheral despite extensive filamentation (Fig. 4d,e and Extended Data Fig. 4o). Extending the incubation period to 24 h made the critical role of Ece1 for invasion even more apparent (Fig. 4f). Ece1-dependent tissue invasion was not affected by pretreating mice with ampicillin (Extended Data Fig. 4p). The Ece1-mediated advantage for sustained colonization was also apparent at day 7, when the fungal burden of 101ece1Δ/Δ was low (Fig. 2g) and colonization foci were rare. In contrast to Ece1-competent C. albicans cells, which frequently cross the stratum corneum, reaching and breaching into the stratum granulosum, ECE1-deficient fungal cells were preferentially observed in the outer part of the stratum corneum (Fig. 4g). In Il17rc−/− mice, however, ECE1 is redundant for C. albicans to find its niche in the stratified epithelium (Fig. 4h). To provide further evidence for the proposed mode of action of Ece1 during oral colonization, we experimentally disrupted the cornified epithelial layer before fungal association. This enabled strain 101ece1Δ/Δ to overcome its defect in colonization and to establish its niche (Fig. 4i,j). These data show that Ece1 enables C. albicans to access a safe niche that prevents it from being shed from the oral epithelium, which exhibits a rapid turnover37.
Tight regulation of ECE1 expression ensures colonization by evading the inflammatory immune response
While the processes establishing C. albicans colonization of the oral mucosa are largely conserved between high- and low-virulent strains, the longer-term outcome of the fungus–host interaction differs greatly between the strains19,20. As such, strain SC5314 penetrates much deeper into the nucleated layers of the stratified epithelium than strain 101 (Figs. 1i and 4d,e, and Extended Data Fig. 3n). The in vivo phenotype of 101WT is reminiscent of SC5314eed1Δ/Δ, a mutant lacking the hyphal extension gene EED1 (ref. 38). As sustained colonization of the oral mucosa is supported by restrained filamentation and hyphae-associated gene expression20, we examined the colonization capacity of SC5314eed1Δ/Δ compared with that of SC5314WT and 101WT. While filamentation is initially induced in SC5314eed1Δ/Δ, and sufficient for the mutant to invade epithelial cells38, hyphal growth is not maintained. Once inside the epithelial cell, EED1 deficiency prevents the mutant from escaping and developing full virulence as it rapidly reverts to yeast cell growth38,39. Accordingly, ECE1 expression by SC5314eed1Δ/Δ was transient in response to epithelial cell contact in vitro (Fig. 5a) reminiscent of what we observed with strain 101WT (Fig. 2c). In vivo, again similar to strain 101WT, SC5314eed1Δ/Δ efficiently colonized the stratum corneum (Fig. 5b–d), reaching even higher fungal counts than SC5314WT on day 1, although it persists mainly in yeast morphology (Fig. 5c,d). Like 101WT, and in contrast to SC5314WT, SC5314eed1Δ/Δ did not elicit an inflammatory response (Fig. 5e,f and Extended Data Fig. 5a,b), emphasizing the relevance of restrained virulence for preventing immunopathology and hence promoting long-term colonization (Fig. 5g). Of note, ECE1 was even more repressed in SC5314eed1Δ/Δ than in 101WT (Fig. 5h). Consistent with the requirement of continuous moderate expression of ECE1 for persistence in the oral cavity, SC5314eed1Δ/Δ showed inferior capacity to sustain colonization than 101WT (Fig. 5g). This difference between 101WT and SC5314eed1Δ/Δ was even more pronounced in a competitive setting (Fig. 5i). Overexpression of EED1 in strain 101 (101OE_EED1) enhanced filamentation and ECE1 expression in vitro (Fig. 5j,k). Moreover, 101OE_EED1 elicited the recruitment of inflammatory cells to the site of fungal colonization in the oral mucosa, which resulted in fungal clearance by day 7 (Fig. 5l–n), reminiscent of what happens in the case of strain SC5314 (Fig. 5g). Inflammation foci characterized by infiltration of neutrophils in close proximity to fungal elements were significantly increased in the tongue of mice colonized with 101OE_EED1 compared with those with 101WT. The difference was even more pronounced when normalized to the fungal burden per tongue (Fig. 5n), given the low colonization efficiency of 101OE_EED1 due to its aggregating behaviour. These data illustrate how tight regulation of ECE1 expression via regulation of hyphal formation or maintenance curtails inflammation while promoting fungal fitness in the oral niche that facilitates fungal persistence in the mucosal tissue.
a, ECE1 expression by SC5314WT or SC5314eed1Δ/Δ exposed to TR146 keratinocytes for 0, 2, 4, 8 or 24 h. Each symbol represents the mean ± s.d. (n = 3 per group). b–h, C57Bl/6 WT mice associated with SC5314WT, SC5314eed1Δ/Δ or 101WT for 8 h (b), 1 day (c,e,f, h) or 7 days (g). Tongue CFU (b, c, g, n = 4 or 6 per group). Representative PAS-stained histology sections (d). Quantification of tongue neutrophils (CD45+Ly6G+Ly6C−) and inflammatory monocytes (CD45+Ly6G−Ly6C+) by flow cytometry (e, n = 4 or 6 per group). Cxcl2 and Il1b host transcripts (f) and ECE1, SAP5 and HWP1 fungal transcript (h) in the colonized tongue tissue (f,h, n = 6, 8 or 10 per group; g, n = 3 or 6 per group). Each symbol represents the mean per group ± s.d. (b) or mean per group ± s.e.m. (c,e,h). i, Competitive colonization of C57BL/6 WT mice with 101WT_mCherry and SC5314eed1Δ/Δ (1:1 ratio) for 7 days; n = 4 per group. Connected symbols are 101WT_mCherry and SC5314eed1Δ/Δ CFU per individual animal. j,k, SC5314WT, 101WT and 101OE_EED1 on spider agar at 37 °C, 5% CO2, for 5 days (j) or in serum-containing medium (med) for 24 h before quantification of the ECE1 transcript by RT-qPCR (k; n = 3 per group; mean ± s.d.). l–n, C57Bl/6 WT mice associated with 101WT or 101OE_EED1. Tongue CFU on day 1 and day 7 (l, n = 4 or 6 per group; mean ± s.d.). Representative PAS-stained histology sections on day 1 (m). Proportion of inflammatory foci (myeloid infiltrates) per fungal colonization site relative to the total number of CFU per tongue (n, n = 5 per group; mean ± s.e.m.). In d and m, the black arrows indicate fungal elements and the orange arrows indicate inflammatory infiltrates. Data are pooled from 2 to 3 independent experiments except for b, i and k. In h, data for SC5314WT and 101WT are the same as those shown in Fig. 2d. The statistical significance of differences between groups was determined by two-way ANOVA (a,l), ordinary one-way ANOVA (c,e,f,g,h), two-sided Mann–Whitney test (b), unpaired two-tailed t-test (n) or paired two-tailed t-test (i).
Discussion
Best known for its role as a pathogen causing superficial and systemic, often fatal, infections, C. albicans is primarily a harmless colonizer in most healthy individuals11,40. The mechanisms enabling stable, homeostatic colonization remained elusive13. We show that candidalysin—originally identified as a virulence factor—is essential for C. albicans to establish and sustain its niche in the oral epithelium, while excessive expression is avoided to prevent tissue damage and inflammation. The stratum corneum provides a safe space protected from direct exposure to antimicrobial peptides and IgA in the saliva41,42. These findings complement recent data showing that candidalysin targets commensal gut bacteria to overcome colonization resistance16 and highlight the pleiotropic functions of candidalysin in health and disease. Together, this reframes our understanding of C. albicans colonization in different tissue compartments and provides evidence for why fungal determinants have been conserved through evolution43 despite their pathogenic potential bearing a constant threat to the fungus–host equilibrium. As a pore-forming peptide, candidalysin agglomerates and incorporates in the host cell membrane when accumulating in the invasion pocket that is formed by C. albicans hyphae34,35, a process resulting in cell damage. Our data suggest that candidalysin acts via the same mechanism during homeostatic colonization of the oral mucosa to overcome the rigid barrier of the stratified epithelium, which consists of terminally differentiated transcriptionally inactive keratinocytes. Overexpression of ECE1 in strain 101 allowed demonstrating that this strain, despite its low virulence, can confer detectable cell damage via candidalysin. The continuous dependence of colonization on candidalysin suggests that the fungus is constantly repeating the cycle of tissue invasion to renew its niche and avoid being shed from the epithelium due to desquamation of the stratified oral epithelium44. IL-17 can accelerate tissue turnover and consequent cell shedding from stratum corneum by enhancing the proliferation rate of epithelial cells45. This may explain the competitive advantage of ECE1-competent versus deficient C. albicans in immunocompetent mice, which was not observed in the absence of a functional IL-17 pathway.
Desquamation acts as a host defence mechanism against microbial colonization as shown for Neisseria gonorrhoeae, which evades these host defense activities by suppressing exfoliation of the epithelium46. C. albicans may inhibit epithelial cell shedding by inducing interferon type 1 (refs. 47,48). In turn, some commensal bacteria can enhance shedding of epithelial cells and thereby reduce host cell damage induced by pathogenic C. albicans49.
Despite the fact that strains 101 and SC5314 greatly differ in their intrinsic host-damaging potential19, the initial process of oral colonization was unexpectedly well conserved in contrast to the longer-term outcome of colonization. Although originally reported to show only very limited capacity to filament and to lack expression of virulence factors19,20, we show that strain 101 does filament efficiently and robustly and expresses hyphae-associated genes at the host interface or when exposed to host-like conditions. The selective dependence on ECE1 for oral colonization by strain 101 may explain its shallower penetration depth in the stratified epithelium—compared with strain SC5314—to prevent access to deeper epithelial layers where induction of cell damage results in the production of antimicrobial effectors, release of inflammatory cues and rapid recruitment of inflammatory leukocytes, which in turn eliminate the fungus14,19,50,51. Whether in strain 101 the biological activity of candidalysin is regulated by additional mechanisms beyond ECE1 transcription remains to be determined34,52,53.
Restricted ECE1 expression is at least partially explained by high NRG1 expression20. As such, we previously showed that suppression of NRG1 in strain 101 enhances fungal virulence, translating in enhanced inflammation and more rapid fungal clearance in experimentally colonized mice20. In its role as a transcriptional repressor, Nrg1 also regulates filamentation by controlling EED1 (ref. 39). We found Eed1 to maintain ECE1 expression during hyphal extension validating the strong link between hyphae and hyphae-associated gene expression25. Consequently, EED1 deletion recapitulated features of homeostatic colonization in strain SC5314; however, this was less pronounced than in strain 101 owing to a substantial decrease of ECE1 expression and filamentation. Conversely, EED1 overexpression in strain 101 enhanced pathogenic traits. Together, this illustrates the importance of well-balanced virulence factor expression for evading inflammation while enabling sustained colonization. Our results further highlight the tissue-specific requirements for C. albicans colonization, as EED1 deficiency provides a fitness advantage in systemic organs54, which is not the case in the oral mucosa, where tonic virulence is a fitness benefit.
In summary, our findings attribute candidalysin additional functions beyond the previously known ones. In addition to serving as a classical virulence factor that causes host damage, and its function as an avirulence factor that stimulates a protective damage defense response in the host12, we show here that candidalysin is essential for successful long-term oral colonization of C. albicans by directly interacting with the host to create and maintain its niche in the oral mucosa without triggering tissue damage or inflammation. This property has probably contributed to the conservation of this toxin throughout C. albicans evolution55.
Limitations of this study
We show that candidalysin enables C. albicans to establish its niche in the oral mucosa in both high- and low-virulent isolates. Future studies with additional isolates will clarify conservation across the species regarding the ECE1-dependent mechanism.
We cannot exclude additional mechanisms by which candidalysin may facilitate oral colonization. The epithelial niche may, for instance, provide the fungus access to nutrients and hence contribute to its metabolic adaptation to the host56. Environmental cues to which C. albicans gets exposed in the niche may also modulate the fungal properties. Furthermore, we cannot exclude additional molecules that may support candidalysin’s function in facilitating oral colonization. We have not directly assessed these possibilities in this work.
Our study analysed the colonization process in the stratified epithelium of the tongue. It remains to be shown whether candidalysin is similarly involved in colonization of other stratified epithelia of the oral cavity and possibly the vaginal mucosa, while colonization of the gut is probably regulated differently given the distinct architecture of the epithelial lining in the gut.
Methods
Fungal strains
C. albicans strains used in this study are listed in Supplementary Table 1. All strains were maintained on YPD agar for short-term storage and in glycerol-supplemented medium at −80 °C for long-term storage. For fungal assays, C. albicans strains were first pre-cultured for 8 h in 5 ml YPD medium, then inoculated at OD600 = 0.1 in 10 ml of YPD medium and grown overnight, 15–18 h, at 30 °C and 180 rpm. At the end of the culture period, yeast cells were washed 3 times in PBS and their concentration was determined by spectrophotometry, whereby 1 OD600 = 107 yeast cells.
Generation of C. albicans mutants
The deletion mutant strain in the C. albicans 101 background was generated using CRISPR–Cas9 technology57. We used a CRISPR plasmid constitutively expressing C. albicans codon-optimized Cas9, locus-specific single guide RNA and a SAT1 resistance marker combined with a Flp-recombinase and FRT sites flanking the entire cassette together with a homology-directed repair template with 60 bp up- or downstream homology to the target locus and containing 20 bp overlap at their 3′ ends. Repair templates were generated by extension of DNA oligos using Taq polymerase (NEB). The guide RNAs for specific loci were identified using CRISPOR.org58. The corresponding DNA oligos were annealed and ligated into the guide RNA cloning site of the CRISPR plasmid sequence, and orientation was confirmed by Sanger sequencing. Then, 5 µg of linearized CRISPR plasmid and 3.5 µg of repair template were transformed into C. albicans using the polyethylene glycol (PEG3650)/lithium acetate (LiAc) method59 and selected on YPD containing 200 g ml−1 nourseothricin. Positive locus deletion was verified by colony PCR. Removal of the CRISPR cassette was achieved by induction of Flp recombination. The pNIM1R plasmid60 was used to generate a tet-repressible ECE1-expression mutant. The ECE1 open reading frame of C. albicans 101 was cloned into pNIM1R at SalI and BglII. The transformation cassette was excised with SacII and KpnI, and 5 µg was transformed into 101ece1Δ/Δ as described above. Correct integration into the ADH1 locus was confirmed via colony PCR.
For generating 1010E_EED1, the plasmid for EED1 overexpression was obtained by PCR amplifying EED1 on SC5314 genomic DNA using primers VB40 and VB41 and cloning it into pCRII-TOPO for sequencing. The EED1-bearing BstZI-NotI fragment was then inserted into pKS-PTDH3 cut with EcoRV and NotI. (pKS-PTDH3 results from the insertion of a PCR fragment, amplified on SC5314 genomic DNA with primers SZ11 and SZ12, in XhoI and EcoRV-cut pBluescript-KS(+)). The resulting plasmid was then cut with NsiI and ApaI and the EED1-bearing fragment ligated in the same sites of CIpSAT-PTDH3-GTW (CdE lab collection), yielding CIpSAT-PTDH3-EED1. This plasmid was linearized with StuI before the 101 strain was transformed using the lithium acetate protocol61.
A list of the oligonucleotide sequences for PCR and sequencing used in this study is provided in Supplementary Table 2.
Keratinocyte cell culture
The human oral keratinocyte cell line TR146 (ref. 62; Sigma, reference number 10032305, previously used in the LeibundGut-Lab) was grown in DMEM medium supplemented with 10% FCS, penicillin and streptomycin. The murine oral keratinocyte cell line IMOK36 (obtained from L. Garrett-Sinha) was grown in CnT-07 Epithelial Proliferation Medium (CELLnTec). The human keratinocyte cell line N/TERT1 (ref. 63; obtained from H.-D. Beer) was grown in K-SFM (Gibco) supplemented with 0.005% (w/v) Bovine Pituitary Extract (Gibco) and 0.02% (v/v) Epidermal Growth Factor (Gibco). All cell lines were maintained at 37 °C, 5% CO2. For infection experiments, cells were seeded at 2 × 105 cells per well in 24-well tissue culture plates or at 2 × 104 cells per well in 96-well tissue culture plates, respectively, and grown to confluent monolayers for 2 days before infection as described below for the individual assays. In the case of TR146 cells, 1 day before the experiment, the DMEM medium was replaced with F12 medium (Hams’s Nutrient Mixture F12 medium (Sigma) supplemented with l-glutamine, penicillin, streptomycin and 1% FCS). In the case of the IMOK cells, the medium was renewed on the day of the infection.
HEEs
To generate HEEs64, 2 × 105 N/TERT1 cells per well were seeded in cell culture inserts in a 24-well plate containing 1 ml of CnT prime medium (CELLnTec) in the bottom of the well and incubated at 37 °C, 5% CO2 until confluent. The medium was replaced with 1 ml (bottom) and 100 μl (insert) of CnT Prime 3D Barrier medium (CELLnTec). After overnight incubation at 37 °C, 5% CO2, the medium was carefully removed from the insert to start the air–liquid-interface culture. The medium in the bottom well was replaced every 2–3 days over 3 weeks until HEEs were fully differentiated. For infection, 8 × 104 yeast cells in 100 μl of PBS were applied to HEEs. The excess liquid was removed after 30 min, and HEEs were incubated at 37 °C, 5% CO2, for 2, 8 or 24 h as indicated. To collect the 0 h timepoint, HEEs were centrifuged immediately after seeding C. albicans and excess liquid was removed. For histology, HEE samples were collected by detaching the tissue from the membrane and fixing in 4% PBS-buffered paraformaldehyde. For RNA isolation, HEEs were collected in TRI reagent (Sigma) and stored at −20 °C until further processing.
Adhesion assay
Adhesion of C. albicans to TR146 keratinocytes was determined as described previously34. Briefly, C. albicans cells were added to confluent TR146 cells in 24-well plates on 12-mm-diameter glass coverslips to a final concentration of 1 × 105 cells per well and incubated for 1 h at 37 °C, 5% CO2. Non-adherent C. albicans cells were removed by washing with PBS. Samples were fixed with 4% PBS-buffered paraformaldehyde for 15 min at room temperature and rinsed with PBS. Adherent C. albicans were stained with Calcoflour white (10 µg ml−1 in 100 mM TRIS∙HCl pH 9.5) for 30 min at room temperature. Samples were rinsed with Milli-Q-water and mounted with ProLong Gold Antifade Mountant (Invitrogen). Images were acquired with a Zeiss Axio Observer Z1 microscope. Adherence was measured by counting the number of C. albicans cells in a defined area.
Filamentation and cell invasion assay
To evaluate filamentation in serum-containing medium, 5 × 104 yeast cells were seeded in 0.5 ml F12 medium per well in a 24-well plate and incubated for 2 h at 37 °C, 5% CO2, before fixation with 4% PBS-buffered paraformaldehyde. To measure C. albicans filamentation on and invasion of keratinocytes, monolayers of IMOK cells in 24-well plates prepared as described above were infected with 1 × 105 yeast cells per well and incubated for 3.5 h at 37 °C, 5% CO2. Extracellular parts of fungal cells were then stained with 25 µl ml−1 ConA for 20 min at room temperature. After fixation with 4.2% formaldehyde for 20 min at room temperature, intracellular and extracellular fungal parts were stained with an FITC-conjugated polyclonal anti-C. albicans antibody (Meridian BioScience) overnight at 4 °C. Images were acquired with an EVOS FL Auto Microscope and analyzed with ImageJ software65. The percentage of invasive hyphae was calculated relative to the total number of hyphae per imaged field. The proportion of hyphae inside keratinocytes was determined by calculating the ratio of the length of green-only hyphae to the entire invasive filament length. Only individual (non-aggregated) filaments were assessed.
Cell damage assay
LDH release from TR146 and IMOK keratinocytes in response to C. albicans was performed as described66. Briefly, keratinocyte monolayers in 96-well plates prepared as described above were infected with the indicated number of yeast cells per well and incubated for 24 h at 37 °C, 5% CO2, before assessing LDH release using the LDH cytotoxicity kit (Roche) according to the manufacturer’s instructions. Control wells were incubated with medium only or with 1% Triton-X-100 to determine 0% and 100% damage, respectively.
Animals
Female WT C57BL/6j mice were purchased from Janvier Elevage. The Il17rc−/− mice (C57BL/6j background) were obtained from Amgen and bred at the Institute of Laboratory Animals Science (University of Zurich). All mice were kept in individually ventilated cages under specific pathogen-free conditions at 21–24 °C, 40–60% humidity and a standard light cycle (12 h:12 h) and were provided with unrestricted access to water and food (irradiated vitamin-fortified maintenance extrudate, Kliba Nafag 3435). Animals were used at 6–14 weeks of age in sex- and age-matched groups. Animals were allowed to acclimatize for 1 week after arrival in the animal experimentation unit of Laboratory Animals Science before the experiments started. Only animals in good health were included in experiments. Experiments with WT mice used a randomized design; experiments with genetically modified female and male Il7rc−/− and Il17rc+/− littermates were not done fully randomized owing to variable distribution of the different genotypes in each litter. Colonized and uncolonized animals were kept separately to avoid cross-contamination. Sample size was chosen using Fermi’s approximation and based on experience.
Ethics statement
All mouse experiments in this study were conducted in strict accordance with the guidelines of the Swiss Animals Protection Law and were performed under the protocols approved by the Veterinary Office of the Canton Zurich, Switzerland (license numbers ZH167/2018, ZH141/2021 and ZH186/2024). All efforts were made to minimize suffering and ensure the highest ethical and humane standards according to the replace, reduce, refine) principles67.
Oral colonization of mice with C. albicans
Mice were infected sublingually as described68, without immunosuppression or antibiotics unless stated explicitly, with 2.5 × 106 (normal infection dose) or with 5 × 104 (l.d.) C. albicans yeast cells. In competition experiments, C. albicans strains were mixed at a ratio of 1:1 (with 1.25 × 106 yeast cells of each strain) or 1:50 (with 5 × 104 101wt_mCherry and 2.5 × 106 yeast cells of the designated strain). In brief, mice were anaesthetized by injection of 100 mg kg−1 ketamine and 20 mg kg−1 xylazine in sterile saline intraperitoneally administered in two doses. C. albicans was administered by depositing a 2.5-mg cotton ball that was soaked in 100 µl yeast cell suspension under the tongue for 75–90 min. Mice were kept on a heating mat at 35–37 °C during the entire period of anaesthesia, administered with 10 ml kg−1 sterile saline to stabilize the circulation and treated with vitamin A ointment to avoid desiccation of the eyes. For experiments with 101tetOff_ECE1, mice were treated with doxycycline (2 mg ml−1) in 1% glucose drinking water for the indicated time. To induce dysbiosis, mice were treated with ampicillin (0.5 mg ml−1) added to the drinking water that was also supplemented with 1% glucose starting 3–4 days before fungal association until the end of the experiment. The drinking water of the control group was supplemented with 1% glucose.
In some experiments, mice were perfused 2 h after fungal association, and tongues were removed and directly placed into a 6-well plate that we converted into a humid chamber with moist tissues and incubated at 37 °C, 5% CO2, for 6–22 h, as indicated. Humidification was sufficient to maintain the integrity and moisture of the tissue. No ampicillin was added to the humid chamber.
In some experiments, the cornified layer of the stratified epithelium was experimentally disrupted by repeated shallow puncturing with a 26-G needle before fungal association.
Determination of fungal burden
Tongues were collected from euthanized animals and homogenized in sterile water supplemented with 0.05% NP40 in H2O and a 5-mm steel ball for 3 min at 25 Hz using a Tissue Lyzer (Qiagen). Serial dilutions were plated on YPD agar containing 100 µg ml−1 ampicillin, and colony-forming units (CFU) per gram tissue were enumerated. In the competition experiments, mCherry-positive colonies were identified using the IVIS Lumina III from Perkin Elmer.
Histology
HEE and tongue tissue were fixed in 4% PBS-buffered paraformaldehyde overnight and embedded in paraffin. Sagittal sections (9 µm) were stained with periodic acid–Schiff (PAS) reagent and counterstained with haematoxylin and mounted with Pertex (Biosystem) according to standard protocols. Images were acquired with a digital slide scanner (NanoZoomer 2.0-HT, Hamamatsu) and analyzed with NDP.view2. To determine the localization of C. albicans in the tongue tissue, accumulations of fungal elements spanning a maximum diameter of 20 μm were considered a unit. Histology images were scored blinded.
Quantification of tongue neutrophils and monocytes by flow cytometry
Tongues were collected from euthanized and perfused mice, cut into fine pieces and digested with DNase I (200 μg ml−1, Roche) and Collagenase IV (4.8 mg ml−1, Invitrogen) in PBS for 45 min at 37 °C. Single-cell suspensions were passed through a 70-μm strainer and stained in ice-cold PBS supplemented with 1% FCS, 5 mM EDTA and 0.02% NaN3 with LIVE/DEAD Fixable Near-IR Stain (Life Technologies, 1/1,000 dilution) and fluorochrome-conjugated antibodies against mouse CD45.2-AlexaFluor 700 (clone 104, Biolegend 109822, used at 1/200 dilution), CD11b-PE-Cy7 (clone M1/70, Biolegend 101216, used at 1/200 dilution), CD3-PE-Cy5 (clone 145-2C11, Biolegend 100310, used at 1/250 dilution), Ly6G-Pacific Blue (clone 1A8, Biolegend 127612, used at 1/250 dilution), Ly6C-FITC (clone AL-21, BD Biosciences 553104, used at 1/200 dilution), CD64-APC (clone X54-5/7.1, Biolegend 139306, used at 1/100 dilution) and F4/80-PE (clone BM8, Biolegend 123109, used at 1/100 dilution). Cells were fixed in 4.2% formaldehyde before acquisition on a SP6800 Spectral Analyzer (Sony). Data analysis was performed using FlowJo software v10 (FlowJo LLC). The gating of the flow cytometric data was performed according to the guidelines for the use of flow cytometry and cell sorting in immunological studies69 including pre-gating on viable and single cells for analysis. A defined number of counting beads (BD Bioscience, Calibrite Beads), which were added to the samples before flow cytometric acquisition, were used to quantify absolute cell numbers.
Isolation of fungal RNA from infected keratinocytes and HEEs
Monolayers of TR146 or IMOK cells in 24-well plates prepared as described above were infected with 1 × 105 yeast cells per well. After 0, 2, 4 and 24 h of incubation at 37 °C, 5% CO2, the medium was removed and cells were frozen on liquid nitrogen and stored at −20 °C until further processing. C. albicans in serum-containing medium was manipulated the same way, except that no keratinocytes were seeded. Infected HEEs (see above) were homogenized in Tri Reagent with a 5-mm steel ball using a Tissue Lyzer (Qiagen) for 6 min at 25 Hz. RNA was then isolated with the RNeasy Mini Kit (Qiagen). Briefly, to break up fungal cell walls, samples were homogenized with 0.5-mm acid-washed glass beads (Sigma) using a Tissue Lyzer (Qiagen) for two cycles of 2 min at 30 Hz interrupted by 30-s cooling periods on ice. Cell debris was removed by centrifugation. One volume of 75% ethanol was admixed, and each sample was transferred to an RNeasy spin column. The loaded columns were washed using RW1 buffer and RPE buffer. RNA was eluted in 30 μl of RNAse-free water.
RNA isolation from colonized tongue tissue
Total RNA (including fungal RNA) from colonized murine tongues was isolated with TRI Reagent. The tissue was homogenized in TRI Reagent with a 0.5-mm steel ball using a Tissue Lyzer for 6 min at 25 Hz. Fungal cell walls were broken up with acid-washed glass beads (0.5 mm) using a Tissue Lyzer for two cycles of 2 min at 30 Hz interrupted by a 30-s resting period on ice. RNA isolation was then pursued according to the manufacturer’s instructions.
RT-qPCR
cDNA was generated using RevertAid reverse transcriptase (Thermo Fisher). Quantitative PCR was performed using SYBR green (Roche) and an Applied Biosystems 7500 Fast instrument (Thermo Fisher Scientific). The primers used in this study to measure the expression of C. albicans and murine genes are listed in Supplementary Tables 3 and 4, respectively. All qPCRs were performed in duplicates. The relative expression (rel. expr.) of each C. albicans gene was determined after normalization to ACT1 transcript levels while the rel. expr. of each murine gene was normalized to ACTB transcript levels. Data were analyzed using ABI7500 software v2.3.
RNAseq data analysis
To determine differentially expressed genes in the colonized tongues of C57Bl/6 WT mice over time, we explored a published dataset available via National Center for Biotechnology Information (NCBI) BioProject accession number PRJNA491801 (ref. 31). To determine IL-17-dependent regulation of selected genes of interest, we explored an RNAseq dataset from Il17rc−/− and Il17rc+/− littermate control animals colonized with strain 101WT for 3 days (NCBI Gene Expression Omnibus (GEO) repository, accession number GSE280210) using Illumina bcl2fastq Conversion and SUSHI app of the Functional Genomics Centre Zurich. Heat maps were generated using GraphPad prism v10.
Statistical analysis
No statistical methods were used to predetermine sample sizes, but our sample sizes are similar to those reported in previous publications19,20,30. Data collection and analysis were not performed blind to the conditions of the experiments, unless specified. All statistical analyses were done with GraphPad Prism software v10. Normality was always calculated using the D’Agostino and Pearson test, Anderson–Darling test, Shapiro–Wilk test and Kolmogorov–Smirnov test. If all samples passed at least one normality test, they were considered to have normal distribution. The statistical test was applied depending on the normality result, variables and number of groups. Unpaired or paired two-tailed t-test or Mann–Whitney test was used for parametric or non-parametric datasets, respectively, when two groups were compared. One-way ANOVA with Sidák multiple comparisons was used when comparing more than two groups. Two-way ANOVA with Sidák or Dunnet’s multiple-comparison or Fisher’s LSD test was used when comparing more than two sets of variables. Correction for repeated measures was applied for time course experiments. Comparison of matched values was conducted for competition experiments. Data presented on a logarithmic scale underwent log transformation before statistical evaluation. For datasets containing suspected outliers, an outlier analysis was performed.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this Article.
Data availability
The data that support the findings of this study are publicly available via Zenodo at https://doi.org/10.5281/zenodo.16734135 (ref. 70). RNAseq datasets used in this study are available at NCBI (BioProject PRJNA491801 and GEO repository GSE280210).
References
Leonardi, I. et al. Mucosal fungi promote gut barrier function and social behavior via type 17 immunity. Cell 185, 831–846.e14 (2022).
Shao, T. Y. et al. Commensal Candida albicans positively calibrates systemic Th17 immunological responses. Cell Host Microbe 25, 404–417.e6 (2019).
Tso, G. H. W. et al. Experimental evolution of a fungal pathogen into a gut symbiont. Science 362, 589–595 (2018).
Lu, S. Y. Oral candidosis: pathophysiology and best practice for diagnosis, classification, and successful management. J. Fungi https://doi.org/10.3390/jof7070555 (2021).
Denning, D. W., Kneale, M., Sobel, J. D. & Rautemaa-Richardson, R. Global burden of recurrent vulvovaginal candidiasis: a systematic review. Lancet Infect. Dis. 18, e339–e347 (2018).
Denning, D. W. Global incidence and mortality of severe fungal disease. Lancet Infect. Dis. https://doi.org/10.1016/S1473-3099(23)00692-8 (2024).
Li, X. V., Leonardi, I. & Iliev, I. D. Gut mycobiota in immunity and inflammatory disease. Immunity 50, 1365–1379 (2019).
Zeng, S. et al. Candida albicans-specific Th17 cell-mediated response contributes to alcohol-associated liver disease. Cell Host Microbe 31, 389–404.e7 (2023).
Bacher, P. et al. Human anti-fungal Th17 immunity and pathology rely on cross-reactivity against Candida albicans. Cell 176, 1340–1355.e15 (2019).
Witchley, J. N. et al. Candida albicans morphogenesis programs control the balance between gut commensalism and invasive infection. Cell Host Microbe 25, 432–443.e6 (2019).
Anderson, F. M. et al. Candida albicans selection for human commensalism results in substantial within-host diversity without decreasing fitness for invasive disease. PLoS Biol. 21, e3001822 (2023).
Siscar-Lewin, S., Hube, B. & Brunke, S. Emergence and evolution of virulence in human pathogenic fungi. Trends Microbiol. 30, 693–704 (2022).
Fróis-Martins, R., Lagler, J. & LeibundGut-Landmann, S. Candida albicans virulence traits in commensalism and disease. Curr. Clin. Microbiol. Rep. https://doi.org/10.1007/s40588-024-00235-8 (2024).
Moyes, D. L. et al. Candidalysin is a fungal peptide toxin critical for mucosal infection. Nature 532, 64–68 (2016).
Allert, S. et al. Candida albicans-induced epithelial damage mediates translocation through intestinal barriers. mBio https://doi.org/10.1128/mBio.00915-18 (2018).
Liang, S. H. et al. The hyphal-specific toxin candidalysin promotes fungal gut commensalism. Nature 627, 620–627 (2024).
Zollner-Schwetz, I. et al. Oral and intestinal Candida colonization in patients undergoing hematopoietic stem-cell transplantation. J. Infect. Dis. 198, 150–153 (2008).
Gaffen, S. L. & Moutsopoulos, N. M. Regulation of host–microbe interactions at oral mucosal barriers by type 17 immunity. Sci. Immunol. https://doi.org/10.1126/sciimmunol.aau4594 (2020).
Schonherr, F. A. et al. The intraspecies diversity of C. albicans triggers qualitatively and temporally distinct host responses that determine the balance between commensalism and pathogenicity. Mucosal Immunol. 10, 1335–1350 (2017).
Lemberg, C. et al. Candida albicans commensalism in the oral mucosa is favoured by limited virulence and metabolic adaptation. PLoS Pathog. 18, e1010012 (2022).
Setiadi, E. R., Doedt, T., Cottier, F., Noffz, C. & Ernst, J. F. Transcriptional response of Candida albicans to hypoxia: linkage of oxygen sensing and Efg1p-regulatory networks. J. Mol. Biol. 361, 399–411 (2006).
Wartenberg, A. et al. Microevolution of Candida albicans in macrophages restores filamentation in a nonfilamentous mutant. PLoS Genet. 10, e1004824 (2014).
Cao, F. et al. The Flo8 transcription factor is essential for hyphal development and virulence in Candida albicans. Mol. Biol. Cell 17, 295–307 (2006).
Gillum, A. M., Tsay, E. Y. & Kirsch, D. R. Isolation of the Candida albicans gene for orotidine-5′-phosphate decarboxylase by complementation of S. cerevisiae ura3 and E. coli pyrF mutations. Mol. Gen. Genet. 198, 179–182 (1984).
Martin, R. et al. A core filamentation response network in Candida albicans is restricted to eight genes. PLoS ONE 8, e58613 (2013).
Staab, J. F., Bradway, S. D., Fidel, P. L. & Sundstrom, P. Adhesive and mammalian transglutaminase substrate properties of Candida albicans Hwp1. Science 283, 1535–1538 (1999).
Lermann, U. et al. Secreted aspartic proteases are not required for invasion of reconstituted human epithelia by Candida albicans. Microbiology. https://doi.org/10.1099/mic.0.2008/022525-0 (2008).
Bailey, D. A., Feldmann, P. J., Bovey, M., Gow, N. A. & Brown, A. J. The Candida albicans HYR1 gene, which is activated in response to hyphal development, belongs to a gene family encoding yeast cell wall proteins. J. Bacteriol. 178, 5353–5360 (1996).
McCall, A. D., Kumar, R. & Edgerton, M. Candida albicans Sfl1/Sfl2 regulatory network drives the formation of pathogenic microcolonies. PLoS Pathog. 14, e1007316 (2018).
Frois-Martins, R. et al. Genome-guided manipulation of regulators of morphogenesis in a C. albicans strain with low virulence is not sufficient to trigger a high-virulence phenotype. Preprint at bioRxiv https://doi.org/10.1101/2025.07.16.665085 (2025).
Kirchner, F. R. et al. Persistence of Candida albicans in the oral mucosa induces a curbed inflammatory host response that is independent of immunosuppression. Front. Immunol. 10, 330 (2019).
Kirchner, F. R. & LeibundGut-Landmann, S. Tissue-resident memory Th17 cells maintain stable fungal commensalism in the oral mucosa. Mucosal Immunol. 14, 455–467 (2021).
Tangye, S. G. & Puel, A. The Th17/IL-17 axis and host defense against fungal infections. J. Allergy Clin. Immunol. Pract. 11, 1624–1634 (2023).
Mogavero, S. et al. Candidalysin delivery to the invasion pocket is critical for host epithelial damage induced by Candida albicans. Cell Microbiol. 23, e13378 (2021).
Russell, C. M. et al. The Candida albicans virulence factor candidalysin polymerizes in solution to form membrane pores and damage epithelial cells. eLife https://doi.org/10.7554/eLife.75490 (2022).
Parikh, N., Nagarajan, P., Sei-ichi, M., Sinha, S. & Garrett-Sinha, L. A. Isolation and characterization of an immortalized oral keratinocyte cell line of mouse origin. Arch. Oral. Biol. 53, 1091–1100 (2008).
Cutright, D. E. & Bauer, H. Cell renewal in the oral mucosa and skin of the rat. I. Turnover time. Oral. Surg. Oral. Med. Oral. Pathol. 23, 249–259 (1967).
Zakikhany, K. et al. In vivo transcript profiling of Candida albicans identifies a gene essential for interepithelial dissemination. Cell. Microbiol. 9, 2938–2954 (2007).
Martin, R. et al. The Candida albicans-specific gene EED1 encodes a key regulator of hyphal extension. PLoS ONE 6, e18394 (2011).
Delavy, M. et al. Unveiling Candida albicans intestinal carriage in healthy volunteers: the role of micro- and mycobiota, diet, host genetics and immune response. Gut Microbes 15, 2287618 (2023).
Lindh, E. et al. Autoimmunity and cystatin SA1 deficiency behind chronic mucocutaneous candidiasis in autoimmune polyendocrine syndrome type 1. J. Autoimmun. 42, 1–6 (2013).
Conti, H. R. et al. New mechanism of oral immunity to mucosal candidiasis in hyper-IgE syndrome. Mucosal Immunol. 4, 448–455 (2011).
Richardson, J. P. et al. Candidalysins are a new family of cytolytic fungal peptide toxins. mBio 13, e0351021 (2022).
Deo, P. N. & Deshmukh, R. Pathophysiology of keratinization. J. Oral. Maxillofac. Pathol. 22, 86–91 (2018).
Rabeony, H. et al. Inhibition of keratinocyte differentiation by the synergistic effect of IL-17A, IL-22, IL-1α, TNFα and oncostatin M. PLoS ONE 9, e101937 (2014).
Muenzner, P. & Hauck, C. R. Neisseria gonorrhoeae blocks epithelial exfoliation by nitric-oxide-mediated metabolic cross talk to promote colonization in mice. Cell Host Microbe 27, 793–808.e5 (2020).
Pekmezovic, M. et al. Candida pathogens induce protective mitochondria-associated type I interferon signalling and a damage-driven response in vaginal epithelial cells. Nat. Microbiol. 6, 643–657 (2021).
Sala, A. et al. A new phenotype in Candida–epithelial cell interaction distinguishes colonization- versus vulvovaginal candidiasis-associated strains. mBio 14, e0010723 (2023).
Graf, K. et al. Keeping Candida commensal: how lactobacilli antagonize pathogenicity of Candida albicans in an in vitro gut model. Dis. Model Mech. https://doi.org/10.1242/dmm.039719 (2019).
Verma, A. H. et al. Oral epithelial cells orchestrate innate type 17 responses to Candida albicans through the virulence factor candidalysin. Sci. Immunol. https://doi.org/10.1126/sciimmunol.aam8834 (2017).
Trautwein-Weidner, K., Gladiator, A., Nur, S., Diethelm, P. & LeibundGut-Landmann, S. IL-17-mediated antifungal defense in the oral mucosa is independent of neutrophils. Mucosal Immunol. 8, 221–231 (2015).
Richardson, J. P. et al. Processing of Candida albicans Ece1p is critical for candidalysin maturation and fungal virulence. mBio https://doi.org/10.1128/mBio.02178-17 (2018).
Muller, R. et al. Secretion of the fungal toxin candidalysin is dependent on conserved precursor peptide sequences. Nat. Microbiol. 9, 669–683 (2024).
Dunker, C. et al. Rapid proliferation due to better metabolic adaptation results in full virulence of a filament-deficient Candida albicans strain. Nat. Commun. 12, 3899 (2021).
Konwar, A. et al. Insights into the evolution of candidalysin and recent developments. Arch. Microbiol. 207, 206 (2025).
Sprague, J. L. et al. Candida albicans translocation through the intestinal epithelial barrier is promoted by fungal zinc acquisition and limited by NFκB-mediated barrier protection. PLoS Pathog. 20, e1012031 (2024).
Vyas, V. K., Barrasa, M. I. & Fink, G. R. A Candida albicans CRISPR system permits genetic engineering of essential genes and gene families. Sci. Adv. 1, e1500248 (2015).
Concordet, J. P. & Haeussler, M. CRISPOR: intuitive guide selection for CRISPR/Cas9 genome editing experiments and screens. Nucleic Acids Res. 46, W242–W245 (2018).
Walther, A. & Wendland, J. An improved transformation protocol for the human fungal pathogen Candida albicans. Curr. Genet. 42, 339–343 (2003).
Prieto, D., Roman, E., Correia, I. & Pla, J. The HOG pathway is critical for the colonization of the mouse gastrointestinal tract by Candida albicans. PLoS ONE 9, e87128 (2014).
Huppert, F. A. & Piercy, M. Recognition memory in amnesic patients: effect of temporal context and familiarity of material. Cortex 12, 3–20 (1976).
Rupniak, H. T. et al. Characteristics of four new human cell lines derived from squamous cell carcinomas of the head and neck. J. Natl Cancer Inst. 75, 621–635 (1985).
Dickson, M. A. et al. Human keratinocytes that express hTERT and also bypass a p16(INK4a)-enforced mechanism that limits life span become immortal yet retain normal growth and differentiation characteristics. Mol. Cell. Biol. 20, 1436–1447 (2000).
Smits, J. P. H. et al. Immortalized N/TERT keratinocytes as an alternative cell source in 3D human epidermal models. Sci. Rep. 7, 11838 (2017).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Wachtler, B., Wilson, D., Haedicke, K., Dalle, F. & Hube, B. From attachment to damage: defined genes of Candida albicans mediate adhesion, invasion and damage during interaction with oral epithelial cells. PLoS ONE 6, e17046 (2011).
Blache, D., Martin, G. B. & Maloney, S. K. Towards ethically improved animal experimentation in the study of animal reproduction. Reprod. Domest. Anim. 43, 8–14 (2008).
Solis, N. V. & Filler, S. G. Mouse model of oropharyngeal candidiasis. Nat. Protoc. 7, 637–642 (2012).
Cossarizza, A. et al. Guidelines for the use of flow cytometry and cell sorting in immunological studies (second edition). Eur. J. Immunol. 49, 1457–1973 (2019).
Fróis Martins, R. & LeibundGut-Landmann, S. Dynamic expression of candidalysin facilitates oral colonization of Candida albicans in mice. Zenodo https://doi.org/10.5281/zenodo.16734135 (2025).
Acknowledgements
We thank H.-D. Beer for support with establishing HEE cultures, the staff of the Laboratory Animal Service Center of University of Zürich for support with animal husbandry, the staff of the Laboratory for Animal Model Pathology of University of Zürich for support with histology studies, the Center for Clinical Studies (ZKS) for access to equipment, and members of the LeibundGut lab, Jakob Sprague and Lydia Kaspar for helpful advice and discussions. This study was supported by the Swiss National Science Foundation (grant number CRSII5_173863 to S.L.L., D.S. and C.d’E.), the European Union’s Horizon 2020 research and innovation programme under the Marie Sklodowska-Curie action, Innovative Training Network: FunHoMic (grant number 812969 to S.L.L., B.H. and C.d’E.), the Novartis Foundation for Medical-Biological Research (grant number 22C224 to S.L.L.), the Uniscientia Foundation (grant number 206-2023 to S.L.L. and J.L.) and a UZH Candoc grant (to R.F.-M.). B.H. and T.B.S. were supported by the German Research Foundation (Deutsche Forschungsgemeinschaft (DFG)) within the Cluster of Excellence ‘Balance of the Microverse’, under Germany’s Excellence Strategy, EXC 2051, project ID 390713860; B.H. was further supported by the Collaborative Research Centre/Transregio 124 ‘FungiNet’ (DFG project number 210879364, projects C1 and C2) of the German Research Foundation. Work in the laboratory of C.d’E. is supported by the Agence Nationale de Recherche (ANR-10-LABX-62-IBEID).
Author information
Authors and Affiliations
Contributions
R.F.-M. and S.L.-L. designed the study and wrote the paper. R.F.-M. performed most of the experiments and analyzed the data. J.L., K.M.d.S.V., S. Mertens and T.B.S. performed selected experiments. M.S. and I.K. helped with the experiments. T.B.S., O.E., S. Mogavero and B.H. generated C. albicans mutants, provided advice and edited the paper. N.S., S.B.-B. and C.d’E. constructed the 101OE_EED1 mutant. D.S. generated fluorescent variants of C. albicans mutants and ECE1 overexpression mutants. S.L.-L. oversaw the study design and data analysis and acquired funding. All authors discussed the results and commented on the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Jigar Desai, Anna Dongari-Bagtzoglo and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Filamentation and hyphae-associated genes are required for C. albicans oral colonization.
a–b. SC51314efg1∆/∆, SC51314efg1/cph1∆/∆, SC51314flo8∆/∆ and the parental SC5314WT strain were incubated for 2 hours in serum containing medium and assessed for filamentation (a, scale bar, 100 μm) and expression of ECE1, HWP1 and SAP5 transcripts (b); n = 3 / group; mean ± SD. c–g. C57BL/6 WT mice were associated with C. albicans strain SC51314hyr1∆/∆ (c), SC51314sap1-3∆/∆ (d), SC51314 efg1/cph1∆/∆ and SC51314 efg1/cph1∆/∆_pENO_ECE1 (e), or SC51314sfl1∆/∆, and SC51314sfl2∆/∆ (f–g) in comparison to their parental SC5314WT strain for 8 hours and analyzed for tongue CFU (c–f) and histology (g, PAS-stained tongues sections, scale bar, 100 μm; black arrows indicate fungal elements); n = 4 / group. DL, detection limit. Each data point represents an independent biological sample (a, b) or an individual mouse (c–f) with mean/group ± SD per group. The statistical significance of differences between groups was determined by unpaired two-tailed t-test (c, d) or ordinary One-way ANOVA (b, e and f).
Extended Data Fig. 2 Low virulent C. albicans acquires hallmarks of virulence for the establishment of oral colonization.
a. ECE1 expression by SC5314WT and 101WT incubated in serum containing medium for 0, 2, 4 and 24 h. b. CFW staining of C. albicans strain 101 grown on IMOK cells or in serum-containing medium for 4 h, 8 h or 24 h. Scale bars, 100 μm. c–d SAP5 (c), and HWP1 expression (d) by SC5314WT and 101WT incubated in serum containing medium (left panels in c–d), on TR146 keratinocytes (middle panels in c–d) or on human epidermal equivalents (HEE, right panels in c-d) for 0, 2, 4 and 24 hours as indicated. e. ECE1 (top), SAP5 (middle), and HWP1 expression (bottom) by SC5314WT and SC5314efg1/cph1∆/∆ incubated in serum-containing medium for 0, 2, 4 and 24 hours. Each symbol from a–e represents the mean ± SD of triplicate wells. Data are representative of at least 2 independent experiments. f. Tongue CFU of C57Bl/6 WT mice associated with 3 different clones of 101ece1∆/∆ (n = 6 / group ± SEM) at 24 hours. g. Loss of ECE1 expression by 101tetOff_ECE1 upon doxycycline treatment (1 µg/ml) for 24 hours in serum containing medium; n = 3 / group with mean ± SD. h. Tongue CFU of C57Bl/6 WT mice that were associated with 101WT for 35 days and treated or not with doxycycline from day 21 to the end of the experiment; n = 4 / group with mean ± SD. i. Competitive colonization of 101WT and 101WT_mCherry at a 1:1 ratio for 7 days before enumerating fluorescent and non-fluorescent tongue CFU; n = 4. Connected symbols are 101WT and 101WT_mCherry CFUs per individual animal. j. C57BL/6 WT mice were associated with strain 101hwp1∆/∆ (n = 6, 2 independent clones indicated by circles and triangles) or 101sap4-6∆/∆ (n = 9, 3 independent clones indicated by circles, triangles and squares) in comparison to the parental strain 101 (n = 4) and tongue CFU (mean ± SD) were enumerated on day 7. The statistical significance of differences between groups was determined by Two-way ANOVA (a, c–e), One-way ANOVA (f, j), unpaired two-tailed t-test (g, h), paired two-tailed t-test (i).
Extended Data Fig. 3 Candidalysin confers resistance to host antifungal responses.
a–b. C57Bl/6 mice were or were not treated with ampicillin from day -4/-3 prior to fungal association. Tongue bacteria growing on YPD agar was assessed at day 7; n = 7 pooled from 2 independent experiments with mean ± SEM. c. Il17rc+/− mice were associated with strain 101WT or strain 101ece1∆/∆ and tongue CFU were enumerated on day 19. n = 4 or 9 with mean ± SD. The statistical significance of differences between groups was determined by paired two-tailed t-test (b) or unpaired two-tailed t-test (c).
Extended Data Fig. 4 Candidalysin promotes oral colonization by enabling C. albicans to overcome the barrier of the stratum corneum.
a. Adhesion of C. albicans 101WT and 101ece1∆/∆ to TR146 keratinocytes at 1 hour post infection. b–e. C. albicans 101WT and 101ece1∆/∆ were incubated on IMOK cells for 3.5 hours and stained with concanavalin A (extracellular parts of C. albicans, red) and with FITC-labelled anti-C. albicans (intra- and extracellular parts, green). Total filament length (b), % of invading fungi (c, % of fungal elements that are at least partially stained green in e) and % of fungal elements inside IMOK cells (d, proportion of the stained green part of fungal element within all, red and green stain, of the respective invasive filament) were quantified. Representative images are shown in (e). Scale bars, 100 μm. In A, each symbol represents the number of adherent fungal cells in a defined area with mean ± SD per group. In b and d, each symbol represents the measurement of an individual filament with mean / group ± SD. In c, each symbol is the percentage of all invading filaments per image with mean / group ± SD. Data in b-e are representative of two independent experiments. f. LDH release from TR146 keratinocytes in response to SC5314WT or 101WT (2 x 104 yeast cells / well) at 24 hours (n = 4 / group ± SD). Each symbol represents an individual well. g–J. LDH release from IMOK cells in response to SC5314WT and 101WT (g), 101WT and 101ece1∆/∆ (h) or 101WT and 101hwp1∆/∆ (2 different clones indicated by circles and triangles) (i), or 101WT and 101sap4-6∆/∆ (3 different clones indicated by circles, triangles and squares) (j) at the indicated yeast cell numbers per well for 24 hours with mean ± SD. In g–j each symbol represent the mean (3-6 technical replicates). g-j are data pooled from 2 independent experiments. k. Relative ECE1 expression by SC5314 and 101 after 24 hours of exposure to IMOK keratinocytes or murine tongue tissue (in vivo) underlying the data shown in Fig. 3b, the mean ± SEM is indicated. Each symbol represents an individual well (IMOK) or an individual mouse (in vivo). l. LDH release from IMOK cells in response to 101WT, 101TDH3-ECE1 and 101HWP1-ECE1 at 24 hours (n = 3 / group). Each symbol represents an individual well, the mean / group ± SD is indicated. Each symbol represents an individual well. m–n. Relative ECE1 expression by SC5314 WT, 101WT and 101ECE1-OE exposed to IMOK cells for 24 hours (m, n = 3 /group ± SD) or in the tongue tissue after 1 day or 7 days after colonization (n, n = 3 /group ± SD). While in M each symbol represents an individual well, in N each symbol represents an individual mouse. o. Representative PAS-stained tongue tissue sections of C57Bl/6 WT mice that were associated with SC5314WT or SC5314ece1∆/∆ for 2 hours followed by 6 hours ex vivo incubation of the tissue as in Fig. 3. p. Representative PAS-stained tongue tissue sections of ampicillin-pretreated C57Bl/6 WT mice that were associated with 101WT and 101ece1∆/∆ for 2 hours followed by 6 hours ex vivo incubation of the tissue as in Fig. 3. Scale bars in o and p, 100μm (inserts: 25μm). Black arrows indicate fungal elements. The statistical significance of differences between groups was determined by unpaired two-tailed t-test (a, c, f), two-sided Mann-Whitney test (b and d) or Two-way ANOVA (g–k) or ordinary One-way ANOVA (m).
Extended Data Fig. 5 Tight regulation of ECE1 expression ensures commensalism by evading the inflammatory immune response.
a–b. Gating strategy used to quantify neutrophils and monocytes in the colonized tongue (a) and representative plots for each fungal strain (b) shown in Fig. 4e.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fróis-Martins, R., Lagler, J., Schille, T.B. et al. Dynamic expression of candidalysin facilitates oral colonization of Candida albicans in mice. Nat Microbiol 10, 2472–2485 (2025). https://doi.org/10.1038/s41564-025-02122-4
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41564-025-02122-4