Abstract
Syntrophic microbial consortia can contribute substantially to the activity of anoxic ecosystems but are often too rare to allow the study of their in situ physiologies using traditional molecular methods. Here we combined bioorthogonal non-canonical amino acid tagging (BONCAT), stable isotope probing and metaproteomics to improve the recovery of proteins from active members and track isotope incorporation in an anaerobic digestion community. Click-chemistry-enabled cell sorting and direct protein pull-down coupled to metaproteomics improved recovery of isotopically labelled proteins during anaerobic acetate oxidation. BONCAT-enabled protein profiles revealed elevated activity and labelling of a rare and so-far uncharacterized syntrophic bacterium belonging to the family Natronincolaceae that expressed a previously hypothesized oxidative glycine pathway for syntrophic acetate oxidation. Stable-isotope-probing-informed metabolic modelling predicted that this organism accounted for a majority of acetate flux, suggesting that the oxidative glycine pathway is an important route for anaerobic carbon transformation and is probably central to community metabolism in natural and engineered ecosystems.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All sequencing reads from the time-series metagenomes, PacBio metagenomes, and the 466 high-quality MAGs are available through the NCBI BioProject identifier PRJNA1180090. Sequencing reads from the BONCAT mini-metagenomes are available through the NCBI BioProject identifier PRJNA1177472. Details of all metagenome samples, including individual NCBI accession numbers, can be found in Supplementary Table 1. NCBI accessions for individual MAGs from this study are provided in Supplementary Table 2. All proteomic data are available through the ProteomeXchange accession PXD057167. Additional data and metadata used in the analysis, including metabolomics, flow cytometry, metabolic modelling, metagenomics and genomic analyses can be found in the OSF repository at https://doi.org/10.17605/OSF.IO/U5VYC.
Code availability
Computational code used to analyse the metagenomic and metaproteomic data, conduct metabolic modelling and generate the figures shown in this paper are included in the Zenodo repository at https://doi.org/10.5281/zenodo.16903890 (ref. 108).
References
Sampara, P., Lawson, C. E., Scarborough, M. J. & Ziels, R. M. Advancing environmental biotechnology with microbial community modeling rooted in functional ‘omics. Curr. Opin. Biotechnol. 88, 103165 (2024).
Hatzenpichler, R., Krukenberg, V., Spietz, R. L. & Jay, Z. J. Next-generation physiology approaches to study microbiome function at single cell level. Nat. Rev. Microbiol. 18, 241–256 (2020).
Hua, Z.-S. et al. Ecological roles of dominant and rare prokaryotes in acid mine drainage revealed by metagenomics and metatranscriptomics. ISME J. 9, 1280–1294 (2015).
Jia, X., Dini-Andreote, F. & Salles, J. F. Unravelling the interplay of ecological processes structuring the bacterial rare biosphere. ISME Commun. 2, 96 (2022).
Dyksma, S., Jansen, L. & Gallert, C. Syntrophic acetate oxidation replaces acetoclastic methanogenesis during thermophilic digestion of biowaste. Microbiome 8, 105 (2020).
Lueders, T., Pommerenke, B. & Friedrich, M. W. Stable-isotope probing of microorganisms thriving at thermodynamic limits: syntrophic propionate oxidation in flooded soil. Appl. Environ. Microbiol. 70, 5778–5786 (2004).
Schink, B. Energetics of syntrophic cooperation in methanogenic degradation. Microbiol. Mol. Biol. Rev. 61, 262–280 (1997).
McInerney, M. J., Sieber, J. R. & Gunsalus, R. P. Syntrophy in anaerobic global carbon cycles. Curr. Opin. Biotechnol. 20, 623–632 (2009).
Westerholm, M., Dolfing, J. & Schnürer, A. Growth characteristics and thermodynamics of syntrophic acetate oxidizers. Environ. Sci. Technol. 53, 5512–5520 (2019).
Angelidaki, I. & Batstone, D. J. in Solid Waste Technology and Management (ed. Christensen, T. H.) 583–600 (John Wiley & Sons, 2010); https://doi.org/10.1002/9780470666883.ch37
Conrad, R. Contribution of hydrogen to methane production and control of hydrogen concentrations in methanogenic soils and sediments. FEMS Microbiol. Ecol. 28, 193–202 (1999).
Mosbæk, F. et al. Identification of syntrophic acetate-oxidizing bacteria in anaerobic digesters by combined protein-based stable isotope probing and metagenomics. ISME J. 10, 2405–2418 (2016).
Kleiner, M. et al. Ultra-sensitive isotope probing to quantify activity and substrate assimilation in microbiomes. Microbiome 11, 24 (2023).
Hatzenpichler, R. et al. In situ visualization of newly synthesized proteins in environmental microbes using amino acid tagging and click chemistry. Environ. Microbiol. 16, 2568–2590 (2014).
Couradeau, E. et al. Probing the active fraction of soil microbiomes using BONCAT-FACS. Nat. Commun. 10, 2770 (2019).
Hellwig, P. et al. Tracing active members in microbial communities by BONCAT and click chemistry-based enrichment of newly synthesized proteins. ISME Commun. 4, ycae153 (2024).
Madill, M. B. W., Luo, Y., Sampara, P., Ziels, R. M. & Gilbert, J. A. Activity-based cell sorting reveals resistance of functionally degenerate Nitrospira during a press disturbance in nitrifying activated sludge. mSystems 6, e00712 (2021).
Scott, M. & Hwa, T. Shaping bacterial gene expression by physiological and proteome allocation constraints. Nat. Rev. Microbiol. 21, 327–342 (2023).
Lawson, C. E. et al. Rare taxa have potential to make metabolic contributions in enhanced biological phosphorus removal ecosystems. Environ. Microbiol. 17, 4979–4993 (2015).
Madsen, E. L. Identifying microorganisms responsible for ecologically significant biogeochemical processes. Nat. Rev. Microbiol. 3, 439–446 (2005).
Bowers, R. M., et al. Minimum information about a single amplified genome (MISAG) and a metagenome-assembled genome (MIMAG) of bacteria and archaea. Nat. Biotechnol. 35, 725–731 (2017).
Campanaro, S. et al. New insights from the biogas microbiome by comprehensive genome-resolved metagenomics of nearly 1600 species originating from multiple anaerobic digesters. Biotechnol. Biofuels 13, 25 (2020).
Mulat, D. G. et al. Quantifying contribution of synthrophic acetate oxidation to methane production in thermophilic anaerobic reactors by membrane inlet mass spectrometry. Environ. Sci. Technol. 48, 2505–2511 (2014).
Oren, A. in The Prokaryotes (eds Rosenberg, E. et al.) 297–306 (Springer, 2014); https://doi.org/10.1007/978-3-642-38954-2_277
Conklin, A., Stensel, H. D. & Ferguson, J. Growth kinetics and competition between Methanosarcina and Methanosaeta in mesophilic anaerobic digestion. Water Environ. Res. 78, 486–496 (2006).
Hao, L. et al. Novel syntrophic bacteria in full-scale anaerobic digesters revealed by genome-centric metatranscriptomics. ISME J. 14, 906–918 (2020).
Singh, A., Schnürer, A., Dolfing, J. & Westerholm, M. Syntrophic entanglements for propionate and acetate oxidation under thermophilic and high-ammonia conditions. ISME J. 17, 1966–1978 (2023).
Westerholm, M., Calusinska, M. & Dolfing, J. Syntrophic propionate-oxidizing bacteria in methanogenic systems. FEMS Microbiol. Rev. 46, fuab057 (2022).
Campanaro, S., Treu, L., Kougias, P. G., Luo, G. & Angelidaki, I. Metagenomic binning reveals the functional roles of core abundant microorganisms in twelve full-scale biogas plants. Water Res. 140, 123–134 (2018).
Sánchez-Andrea, I. et al. The reductive glycine pathway allows autotrophic growth of Desulfovibrio desulfuricans. Nat. Commun. 11, 5090 (2020).
Fink, C. et al. A shuttle-vector system allows heterologous gene expression in the thermophilic methanogen Methanothermobacter thermautotrophicus ΔH. mBio 12, e02766-21 (2021).
McDaniel, E. A. et al. Diverse electron carriers drive syntrophic interactions in an enriched anaerobic acetate-oxidizing consortium. ISME J. 17, 2326–2339 (2023).
Müller, V. & Hess, V. The minimum biological energy quantum. Front. Microbiol. 8, 2019 (2017).
Yi, Y. et al. Thermodynamic restrictions determine ammonia tolerance of methanogenic pathways in Methanosarcina barkeri. Water Res. 232, 119664 (2023).
Westerholm, M., Roos, S. & Schnürer, A. Syntrophaceticus schinkii gen. nov., sp. nov., an anaerobic, syntrophic acetate-oxidizing bacterium isolated from a mesophilic anaerobic filter. FEMS Microbiol. Lett. 309, 100–104 (2010).
Manzoor, S., Bongcam-Rudloff, E., Schnürer, A. & Müller, B. Genome-guided analysis and whole transcriptome profiling of the mesophilic syntrophic acetate oxidising bacterium Syntrophaceticus schinkii. PLoS ONE 11, e0166520 (2016).
Nobu, M. K. et al. Catabolism and interactions of uncultured organisms shaped by eco-thermodynamics in methanogenic bioprocesses. Microbiome 8, 111 (2020).
Dueholm, M. K. D. et al. MiDAS 5: Global diversity of bacteria and archaea in anaerobic digesters. Nat. Commun. 15, 5361 (2024).
Westerholm, M., Müller, B., Singh, A., Karlsson Lindsjö, O. & Schnürer, A. Detection of novel syntrophic acetate-oxidizing bacteria from biogas processes by continuous acetate enrichment approaches. Microb. Biotechnol. 11, 680–693 (2018).
Hedlund, B. P. et al. SeqCode: a nomenclatural code for prokaryotes described from sequence data. Nat. Microbiol. 7, 1702–1708 (2022).
Kieft, B. et al. Genome-resolved correlation mapping links microbial community structure to metabolic interactions driving methane production from wastewater. Nat. Commun. 14, 5380 (2023).
Nobu, M. K. et al. Microbial dark matter ecogenomics reveals complex synergistic networks in a methanogenic bioreactor. ISME J. 9, 1710–1722 (2015).
Zhu, X. et al. Metabolic dependencies govern microbial syntrophies during methanogenesis in an anaerobic digestion ecosystem. Microbiome 8, 22 (2020).
Amann, R. & Fuchs, B. M. Single-cell identification in microbial communities by improved fluorescence in situ hybridization techniques. Nat. Rev. Microbiol. 6, 339–348 (2008).
Yilmaz, S., Haroon, M. F., Rabkin, B. A., Tyson, G. W. & Hugenholtz, P. Fixation-free fluorescence in situ hybridization for targeted enrichment of microbial populations. ISME J. 4, 1352–1356 (2010).
Harris, R. L. et al. FISH-TAMB, a fixation-free mRNA fluorescent labeling technique to target transcriptionally active members in microbial communities. Microb. Ecol. 84, 182–197 (2022).
APHA. Standard Methods for Examination of Water and Wastewater 22nd edn (American Public Health Association, 2012).
Anthonisen, A. C., Loehr, R. C., Prakasam, T. B. & Srinath, E. G. Inhibition of nitrification by ammonia and nitrous acid. J. Water Pollut. Control Fed. 48, 835–852 (1976).
Kolmogorov, M. et al. metaFlye: scalable long-read metagenome assembly using repeat graphs. Nat. Methods 17, 1103–1110 (2020).
Kalikar, S., Jain, C., Vasimuddin, M. & Misra, S. Accelerating minimap2 for long-read sequencing applications on modern CPUs. Nat. Comput. Sci. 2, 78–83 (2022).
Kang, D. D. et al. MetaBAT 2: an adaptive binning algorithm for robust and efficient genome reconstruction from metagenome assemblies. PeerJ 7, e7359 (2019).
Pan, S., Zhu, C., Zhao, X.-M. & Coelho, L. P. A deep siamese neural network improves metagenome-assembled genomes in microbiome datasets across different environments. Nat. Commun. 13, 2326 (2022).
Lamurias, A., Sereika, M., Albertsen, M., Hose, K. & Nielsen, T. D. Metagenomic binning with assembly graph embeddings. Bioinformatics 38, 4481–4487 (2022).
Sieber, C. M. K. et al. Recovery of genomes from metagenomes via a dereplication, aggregation and scoring strategy. Nat. Microbiol. 3, 836–843 (2018).
Chklovski, A., Parks, D. H., Woodcroft, B. J. & Tyson, G. W. CheckM2: a rapid, scalable and accurate tool for assessing microbial genome quality using machine learning. Nat. Methods 20, 1203–1212 (2023).
Nurk, S., Meleshko, D., Korobeynikov, A. & Pevzner, P. A. metaSPAdes: a new versatile metagenomic assembler. Genome Res. 27, 824–834 (2017).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Nawrocki, E. P. & Eddy, S. R. Infernal 1.1: 100-fold faster RNA homology searches. Bioinformatics 29, 2933–2935 (2013).
Olm, M. R., Brown, C. T., Brooks, B. & Banfield, J. F. dRep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication. ISME J. 11, 2864–2868 (2017).
Parks, D. H. et al. A complete domain-to-species taxonomy for bacteria and archaea. Nat. Biotechnol. 38, 1079–1086 (2020).
Chaumeil, P.-A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. GTDB-Tk v2: memory friendly classification with the genome taxonomy database. Bioinformatics 38, 5315–5316 (2022).
Shaw, J. & Yu, Y. W. Fast and robust metagenomic sequence comparison through sparse chaining with skani. Nat. Methods 20, 1661–1665 (2023).
Eren, A. M. et al. Community-led, integrated, reproducible multi-omics with anvi’o. Nat. Microbiol. 6, 3–6 (2020).
Hug, L. A. et al. A new view of the tree of life. Nat. Microbiol. 1, 16048 (2016).
Letunic, I. & Bork, P. Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res. 52, W78–W82 (2024).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinform. 11, 119 (2010).
Shaffer, M. et al. DRAM for distilling microbial metabolism to automate the curation of microbiome function. Nucleic Acids Res. 48, 8883–8900 (2020).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 45, D353–D361 (2017).
Mistry, J. et al. Pfam: the protein families database in 2021. Nucleic Acids Res. 49, D412–D419 (2021).
Sayers, E. W. et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 50, D20–D26 (2022).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152 (2012).
Vallenet, D. et al. MicroScope: an integrated platform for the annotation and exploration of microbial gene functions through genomic, pangenomic and metabolic comparative analysis. Nucleic Acids Res. https://doi.org/10.1093/nar/gkz926 (2019).
Karp, P. D. et al. Pathway Tools version 23.0 update: software for pathway/genome informatics and systems biology. Brief. Bioinform. 22, 109–126 (2021).
Ong, W. K., Midford, P. E. & Karp, P. D. Taxonomic weighting improves the accuracy of a gap-filling algorithm for metabolic models. Bioinformatics 36, 1823–1830 (2020).
Plugge, C. M. Anoxic media design, preparation, and considerations. Methods Enzymol. 397, 3–16 (2005).
Valero, D., Montes, J. A., Rico, J. L. & Rico, C. Influence of headspace pressure on methane production in biochemical methane potential (BMP) tests. Waste Manag. 48, 193–198 (2016).
Rinke, C. et al. Obtaining genomes from uncultivated environmental microorganisms using FACS–based single-cell genomics. Nat. Protoc. 9, 1038–1048 (2014).
Brent, R. P. Algorithms for Minimization without Derivatives. (Dover Publications, 2002).
Hatzenpichler, R. & Orphan, V. J. in Hydrocarbon and Lipid Microbiology Protocols (eds McGenity, T. J. et al.) 145–157 (Springer, 2015); https://doi.org/10.1007/8623_2015_61
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Xu, K. et al. Benchtop-compatible sample processing workflow for proteome profiling of <100 mammalian cells. Anal. Bioanal. Chem. 411, 4587–4596 (2019).
Nicora, C. D. et al. The MPLEx protocol for multi-omic analyses of soil samples. J. Vis. Exp. https://doi.org/10.3791/57343 (2018).
Kim, Y.-M. et al. Diel metabolomics analysis of a hot spring chlorophototrophic microbial mat leads to new hypotheses of community member metabolisms. Front. Microbiol. 6, 209 (2015).
Hiller, K. et al. MetaboliteDetector: comprehensive analysis tool for targeted and nontargeted GC/MS based metabolome analysis. Anal. Chem. 81, 3429–3439 (2009).
Dai, C. et al. quantms: a cloud-based pipeline for quantitative proteomics enables the reanalysis of public proteomics data. Nat. Methods https://doi.org/10.1038/s41592-024-02343-1 (2024).
Ewels, P. A. et al. The nf-core framework for community-curated bioinformatics pipelines. Nat. Biotechnol. 38, 276–278 (2020).
Grüning, B., et al.; The Bioconda Team. Bioconda: sustainable and comprehensive software distribution for the life sciences. Nat. Methods 15, 475–476 (2018).
Da Veiga Leprevost, F. et al. BioContainers: an open-source and community-driven framework for software standardization. Bioinformatics 33, 2580–2582 (2017).
Pfeuffer, J. et al. OpenMS 3 enables reproducible analysis of large-scale mass spectrometry data. Nat. Methods 21, 365–367 (2024).
Di Tommaso, P. et al. Nextflow enables reproducible computational workflows. Nat. Biotechnol. 35, 316–319 (2017).
Kim, S. & Pevzner, P. A. MS-GF+ makes progress towards a universal database search tool for proteomics. Nat. Commun. 5, 5277 (2014).
Eng, J. K., Jahan, T. A. & Hoopmann, M. R. Comet: an open-source MS/MS sequence database search tool. Proteomics 13, 22–24 (2013).
Elias, J. E. & Gygi, S. P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
Käll, L., Canterbury, J. D., Weston, J., Noble, W. S. & MacCoss, M. J. Semi-supervised learning for peptide identification from shotgun proteomics datasets. Nat. Methods 4, 923–925 (2007).
R Core Team R: A Language and Environment for Statistical Computing (R Core Team, 2022).
Wickham, H. et al. Welcome to the tidyverse. J. Open Source Softw. 4, 1686 (2019).
Delogu, F. et al. Integration of absolute multi-omics reveals dynamic protein-to-RNA ratios and metabolic interplay within mixed-domain microbiomes. Nat. Commun. 11, 4708 (2020).
Wiśniewski, J. R. et al. Extensive quantitative remodeling of the proteome between normal colon tissue and adenocarcinoma. Mol. Syst. Biol. 8, 611 (2012).
Wiśniewski, J. R. & Rakus, D. Multi-enzyme digestion FASP and the ‘Total Protein Approach’-based absolute quantification of the Escherichia coli proteome. J. Proteom. 109, 322–331 (2014).
Zhang, X. et al. Metaproteomics reveals associations between microbiome and intestinal extracellular vesicle proteins in pediatric inflammatory bowel disease. Nat. Commun. 9, 2873 (2018).
Li, L. et al. Revealing proteome-level functional redundancy in the human gut microbiome using ultra-deep metaproteomics. Nat. Commun. 14, 3428 (2023).
Sachsenberg, T. et al. MetaProSIP: automated inference of stable isotope incorporation rates in proteins for functional metaproteomics. J. Proteome Res. 14, 619–627 (2015).
Berthold, M. R. et al. KNIME - the Konstanz information miner: version 2.0 and beyond. ACM SIGKDD Explor. 11, 26–31 (2009).
Stahl, D. A. & Amann, R. Development and application of nucleic acid probes in bacterial systematics. In Sequencing and Hybridization Techniques in Bacterial Systematics (eds Stackebrandt, E. & Goodfellow, M.) 205–248 (John Wiley and Sons, 1991).
Meier, H., Amann, R., Ludwig, W. & Schleifer, K. H. Specific oligonucleotide probes for in situ detection of a major group of Gram-positive bacteria with low DNA G+C content. Syst. Appl. Microbiol. 22, 186–196 (1999).
Ebrahim, A., Lerman, J. A., Palsson, B. O. & Hyduke, D. R. COBRApy: constraints-based reconstruction and analysis for Python. BMC Syst. Biol. 7, 74 (2013).
Holzhütter, H.-G. The principle of flux minimization and its application to estimate stationary fluxes in metabolic networks. Eur. J. Biochem. 271, 2905–2922 (2004).
Ziels, R. Activity-targeted metaproteomics uncovers rare syntrophic bacteria central to anaerobic community metabolism. Zenodo https://doi.org/10.5281/zenodo.16903890 (2025).
Woodcroft, B. J. et al. Comprehensive taxonomic identification of microbial species in metagenomic data using SingleM and Sandpiper. Nat. Biotechnol. https://doi.org/10.1038/s41587-025-02738-1 (2025).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Acknowledgements
This work was performed under the auspices of the Natural Sciences and Engineering Research Council of Canada (grant number CRDPJ 543962-19 and RGPIN-2018-04585, both to R.M.Z.), Genome British Columbia (grant number SIP027 to R.M.Z. and S.J.H.), the Canada Foundation for Innovation (grant number 37513 to R.M.Z.), and the US Department of Energy (DOE) Joint Genome Institute (JGI) and EMSL Facilities Integrating Collaborations for User Science (JGI-EMSL FICUS) programme (10.46936/fics.proj.2020.51515/60000205 to R.M.Z.) and used resources at the JGI (https://ror.org/04xm1d337) and EMSL (https://ror.org/04rc0xn13) under contract numbers DE-AC02-05CH11231 (JGI) and DE-AC05-76RL01830 (EMSL). A portion of this research was supported under the EMSL user project award https://doi.org/10.46936/intm.proj.2022.60473/60008546 to V.S.L., H.M.O. and R.M.Z. for developing technologies for and using capabilities operating at the EMSL, a DOE Office of Science User Facility, sponsored by the Biological and Environmental Research programme under contract number DE-AC05-76RL01830. The LABGeM (CEA/Genoscope and CNRS UMR8030), the France Génomique and French Bioinformatics Institute national infrastructures (funded as part of Investissement d’Avenir programme managed by Agence Nationale pour la Recherche, contracts ANR-10-INBS-09 and ANR-11-INBS-0013) are acknowledged for support within the MicroScope annotation platform. We thank C. Nicora for her advice and guidance with sample milling. We thank R. Moore and P. Lalli for performing proteomics runs. We thank M. Schmid for help with confocal microscopy. We thank M. Kazak and F. Crozier for assisting with sample collection from the full-scale bioenergy facility.
Author information
Authors and Affiliations
Contributions
R.M.Z. and S.J.H. conceived the research. S.F. and R.M.Z. processed the metagenomic and metaproteomic data and analysed the results. S.F. and E.A.M. performed the microcosm incubation experiment. M.M. and K.W. performed the time-series sample collection and advised on the microcosm incubation. M.K.D.D. performed the global distribution analysis based on 16S rRNA gene amplicon sequencing data from the MiDAS project. R.R.M. and D.G. designed and performed the mini-metagenome FACS protocol. W.C. performed the FACS MP protocol. L.J.G., C.J.K., L.H.G., H.M.O. and V.S.L. designed and performed the BONCAT protein pull-down assay and MS. H.M.O performed protein extraction for the bulk MP. J.T. and N.M. performed the bulk metabolomics extraction and untargeted metabolomics analysis by GC–MS. S.M.W. performed MS for the bulk and BONCAT–FACS samples. S.B.L. performed IRMS analysis. N.T. and L.P.-T. performed the µPOTS method. M.L. advised on the project formation. R.M.Z. and N.T. advised on the proteomic data analysis. S.F. and R.M.Z. drafted the paper. All authors discussed the results and commented on the paper.
Corresponding author
Ethics declarations
Competing interests
S.J.H. is a co-founder of Koonkie Inc., a bioinformatics consulting company that designs and provides scalable algorithmic and data analytics solutions in the cloud. Koonkie Inc. was not involved in any aspect of this research. The other authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Metabolic fluxes predicted for the five most 13C labeled members at 24 h.
Model fluxes are based on consumption of one mol of acetate, and thus represent the stoichiometry across the metabolic networks at the discrete time point of 24 h of the SIP incubation. Values were predicted with parsimonious flux balance analysis while constraining each organism’s ATP production to be proportional to the mass of 13C labeled proteins at 24 h. Ack = Acetate kinase, Acs = Acetyl-CoA synthetase, CODH = carbon monoxide dehydrogenase, Dld = Dihydrolipoyl dehydrogenase, Ech = Energy-converting hydrogenase, Fdh = Formate dehydrogenase, Fhs = Formate-tetrahydrofolate ligase, FolD = Methylenetetrahydrofolate cyclohydrolase, Fpo = F420H2 dehydrogenase, Frh = F420-reducing hydrogenase, Ftr = Formylmethanofuran:tetrahyromethanopterin formyltransferase, Fwd = Formylmethanofuran dehydrogenase, GcvH = Glycine cleavage system H protein, GcvP = Glycine cleavage system P protein, GcvT = Glycine cleavage system T protein, GlyA = Glycine hydroxymethyltransferase, Grd = Glycine reductase complex, Hdr = heterodisulfide reductase, Hyd = Bifurcating [FeFe] hydrogenase, Mch = Methenyltetrahydromethanopterin cyclohydrolase, Mcr = Methyl-CoM reductase, Mer = 5,10-Methylenetetrahydromethanopterin reductase, Met = 5,10-Methylenetetrahydrofolate reductase, Mtd = F420-dependent methylenetetrahydromethanopterin dehydrogenase, Mtr = 5-Tetrahydromethanopterin:CoM-S-methyltransferase, Mvh = F420-nonreducing hydrogenase, Por = Pyruvate:ferredoxin oxidoreductase, Pta = Phosphate acetyltransferase, Sda = Serine deaminase, Rnf = Na+-translocating ferredoxin:NAD+ oxidoreductase complex, Vho = F420-nonreducing hydrogenase.
Extended Data Fig. 2 Metabolic fluxes predicted for the five most 13C labeled members at 72 h.
Model fluxes are based on consumption of one mol of acetate, and thus represent the stoichiometry across the metabolic networks at the discrete time point of 72 h of the SIP incubation. Values were predicted with parsimonious flux balance analysis while constraining each organism’s ATP production to be proportional to the mass of 13C labeled proteins at 72 h. Ack = Acetate kinase, Acs = Acetyl-CoA synthetase, CODH = carbon monoxide dehydrogenase, Dld = Dihydrolipoyl dehydrogenase, Ech = Energy-converting hydrogenase, Fdh = Formate dehydrogenase, Fhs = Formate-tetrahydrofolate ligase, FolD = Methylenetetrahydrofolate cyclohydrolase, Fpo = F420H2 dehydrogenase, Frh = F420-reducing hydrogenase, Ftr = Formylmethanofuran:tetrahyromethanopterin formyltransferase, Fwd = Formylmethanofuran dehydrogenase, GcvH = Glycine cleavage system H protein, GcvP = Glycine cleavage system P protein, GcvT = Glycine cleavage system T protein, GlyA = Glycine hydroxymethyltransferase, Grd = Glycine reductase complex, Hdr = heterodisulfide reductase, Hyd = Bifurcating [FeFe] hydrogenase, Mch = Methenyltetrahydromethanopterin cyclohydrolase, Mcr = Methyl-CoM reductase, Mer = 5,10-Methylenetetrahydromethanopterin reductase, Met = 5,10-Methylenetetrahydrofolate reductase, Mtd = F420-dependent methylenetetrahydromethanopterin dehydrogenase, Mtr = 5-Tetrahydromethanopterin:CoM-S-methyltransferase, Mvh = F420-nonreducing hydrogenase, Por = Pyruvate:ferredoxin oxidoreductase, Pta = Phosphate acetyltransferase, Sda = Serine deaminase, Rnf = Na+-translocating ferredoxin:NAD+ oxidoreductase complex, Vho = F420-nonreducing hydrogenase.
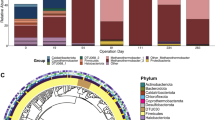
Extended Data Fig. 3 Global distribution of Natronincolaceae_JAAZJW01_1 (that is, Syntrophacetatiphaga salishiae).
(A) Relative abundance in publicly available metagenomes on NCBI (database generated on 20 Feb, 2025) based on Sandpiper 1.0.1.109 metagenomic profiling, in which Syntrophacetatiphaga salishiae was detected at over 0.05% relative abundance (n = 72 anaerobic digester metagenomes, n = 9 landfill metagenomes). (B, C) Maximum observed relative abundance across anaerobic digesters organized by (B) country and (C) substrate type and temperature based on 16S rRNA gene amplicon sequencing data from the MiDAS global survey of anaerobic digesters38. V4 region amplicon sequence variants (ASVs) representing Natronincolaceae_JAAZJW01_1 (>97% sequence identity) were identified by mapping the ASVs against the 16S rRNA genes from the MAG using usearch110 with the command -usearch_global -maxrejects 0 -maxaccepts 0 -id 0 -top_hit_only.
Supplementary information
Supplementary Information (download PDF )
Supplementary Text 1 and Figs. 1–17.
Supplementary Tables (download XLSX )
Supplementary Tables 1–13.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Friedline, S., McDaniel, E.A., Scarborough, M. et al. Activity-targeted metaproteomics uncovers rare syntrophic bacteria central to anaerobic community metabolism. Nat Microbiol 10, 2749–2767 (2025). https://doi.org/10.1038/s41564-025-02146-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41564-025-02146-w