Abstract
Asgard archaea played a key role in the origin of the eukaryotic cell, with extant genomes encoding relatives of diverse eukaryotic signature proteins (ESPs) involved in cellular organization. However, their often punctuated distribution and the absence of detectable homologues for many eukaryotic proteins limit our ability to reconstruct the cellular complexity of the Asgard archaeal ancestor of eukaryotes. Here we used de novo protein structure modelling and sequence similarity detection across an expanded Asgard archaeal genomic dataset to build a structural catalogue of the Asgard archaeal pangenome. We identified 908 ‘isomorphic’ ESPs—Asgard archaeal proteins with statistically enriched structural matches to eukaryotic proteins, often bridging deep sequence divergence. These isomorphic ESPs are enriched in information storage and processing roles and contain key components of the eukaryotic Vault (MVP) and Commander (COMMD) complexes, with potential roles in cellular compartmentalization and endosomal processing. These findings expand the repertoire of eukaryotic-like proteins in Asgard archaea and suggest a higher degree of eukaryote-like cellular complexity in the archaeal ancestor of eukaryotes.
Similar content being viewed by others
Main
The origin of the eukaryotic cell, with its complex and compartmentalized features, is regarded as the biggest evolutionary discontinuity since the advent of cellular life on Earth1,2. Yet, many key details regarding eukaryogenesis, the series of evolutionary events that led to the emergence of the eukaryotic cell from prokaryotic ancestors some 2 billion years ago3,4, remain elusive. The eukaryotic cell is the result of a symbiosis comprising an archaea-related host cell5,6 and a bacterial endosymbiont, the mitochondrial progenitor7,8. While the identity of the endosymbiont was traced back to the Alphaproteobacteria several decades ago9,10, the archaeal host remained elusive until recently. This changed with the discovery of Asgard archaea (phylum Asgardarchaeota11), which were shown to represent the closest prokaryotic relatives of the archaeal host cell from which eukaryotes evolved12,13,14,15. Analysis of Asgard archaeal genomes revealed the presence of numerous homologues of proteins previously deemed eukaryote-specific—so-called eukaryotic signature proteins (ESPs)16. Intriguingly, many of these ESPs represent fundamental building blocks of eukaryotic cellular complexity, including proteins essential for vesicular biogenesis and trafficking, as well as for the dynamic eukaryotic cytoskeleton. Recent work has indicated that several Asgard archaeal ESPs function similarly to their eukaryotic counterparts17,18,19,20, suggesting that Asgard archaea might display eukaryote-like cellular features beyond the dynamic actin cytoskeleton observed in the first enrichment cultures21,22. However, the detailed cellular characteristics and level of complexity of present-day Asgard archaea and the Asgard archaeal ancestor of eukaryotes remain unclear.
The definition, identification and characterization of ESPs are crucial for reconstructing the ancestral Asgard archaeal lineage and understanding its contributions to eukaryogenesis. Yet, the identification process is currently limited by several factors. Defining ESPs has proven challenging as increasingly sensitive homology search algorithms and improved sampling of genomic diversity across the tree of life have facilitated the discovery of ESP homologues in diverse prokaryotes13,15,23, including Asgard archaea12,13,15,23. While this has clarified the prokaryotic origins of many proteins in the last eukaryotic common ancestor, it has also reduced the set of strictly eukaryote-specific proteins. Therefore, a more relaxed definition of ESPs has been adopted, referring to proteins associated with conserved key eukaryotic processes6, or more specifically related to cellular complexity22. Furthermore, many eukaryotic proteins, especially those absent in common model organisms, remain poorly characterized. This, coupled with the limitations of sequence homology detection, makes it difficult to identify ESPs. Given the extensive divergence between present-day Asgard archaeal and eukaryotic proteins, reliable homology detection remains challenging. It becomes increasingly difficult to infer homology between two proteins with decreasing sequence similarity24. As the stem separating eukaryotes from their archaeal relatives represents one of the longest branches in the tree of life14,15, sequences from present-day Asgard archaea and eukaryotes have diverged extensively. Therefore, homology between these two groups might not even be detected, even when using sensitive methods24. However, protein structure is several times more conserved than protein sequence25, and structural information has been shown to increase the sensitivity of sequence homology inference26. Recent advances in de novo protein structure prediction using AlphaFold27 and related tools enable the large-scale generation of high-quality protein structure models. Combined with new methods to efficiently search large databases for similar structures28, it has become feasible to identify highly divergent homologues by using structural information29,30. This is particularly useful for non-model organisms, for which very few protein structures have been resolved. For example, the Protein Data Bank currently contains fewer than 50 Asgard archaeal protein structures (accessed on 31 March 2025).
Here, we explore these recent advances in protein structure prediction and comparison tools to expand the identification and characterization of ESPs in Asgard archaea beyond sequence similarity. By analysing an extended Asgard archaeal pangenome, we identified 908 new structure-based ‘isomorphic’ ESPs (iESPs), more than tripling the overall number of reported Asgard archaeal ESPs. Our structural catalogue of the Asgard archaeal pangenome reveals a marked increase of Asgard archaeal ESPs involved in information storage and processing, and in cellular processes and signalling, suggesting that the archaeal ancestor of eukaryotes was more eukaryote-like than was previously assumed.
Results
Structural modelling of the Asgard archaeal pangenome
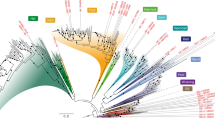
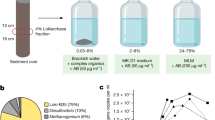
To generate structural models of representative proteins encoded by the Asgard archaeal pangenome, we analysed a diverse set comprising 936 Asgard archaeal draft genomes (Fig. 1a and Supplementary Data 1), including 497 metagenome-assembled genomes (MAGs) that were compiled and described in a recent study31. In addition to the previously sampled Asgard archaeal diversity11,15, this expanded dataset encompasses MAGs from Atabeyarchaeia32 and Ranarchaeia31, two additional deep-branching clades (Extended Data Fig. 1a). We grouped protein sequences encoded by these Asgard archaeal genomes by combining reference-based clustering into previously established Asgard archaeal clusters of orthologous genes (AsCOGs)23 with de novo gene clustering (Fig. 1b). This resulted in 96% of Asgard archaeal proteins grouped in 37,313 clusters of at least 5 proteins, including 22,609 de novo clusters (Fig. 1b). For computational feasibility, we selected one evolutionary representative protein sequence per cluster (Methods) to generate a high-quality structural model (Fig. 1c).
a, The number of Asgard archaeal draft genomes per group in the database used for pangenome-wide structural analyses (also see Extended Data Fig. 1a). Fill colour indicates publicly available genomes (grey) and newly added Asgard archaeal draft genomes (blue), respectively. b, Protein sequence clustering into existing Asgard archaeal COGs and de novo clustering with unassigned proteins. The x axis indicates the number of proteins and the y axis the number of respective clusters. Fill indicates protein sequences from publicly available genomes (grey) and added Asgard archaeal draft genomes (blue), respectively. c, Workflow for the pangenome-wide prediction of Asgard archaeal protein structures. d, Scatter plot depicting pLDDT scores of structure predictions of 100 randomly selected Prometheoarchaeum syntrophicum proteins computed with the default (x axis) and the Asgard archaea-enriched (y axis) ColabFold database, respectively. The diagonal black line indicates x = y, and the purple line indicates linear correlation fitted to the data. e, The distribution of average pLDDT scores of 37,223 predicted Asgard archaeal protein structures. MSA, multiple sequence alignment.
To determine an efficient and effective approach for de novo structure prediction, we modelled structures for 100 randomly selected proteins of the Asgard archaeon Prometheoarchaeum syntrophicum (Supplementary Data 2). As AlphaFold relies on homology information to predict protein structure, it tends to perform poorly if few homologues are found within its reference sequence database27. To solve this issue, we used ColabFold33, an accelerated AlphaFold workflow, with an expanded database containing all available Asgard archaeal protein sequences. In addition, we used ESMfold34, a prediction tool based on a protein language model that circumvents the time-consuming sequence homology search. We classified predictions as high quality if they had an average predicted local distance difference test (pLDDT) score of at least 80. We found that incorporating the Asgard archaeal proteins to the ColabFold homology search database led to better models for some proteins (Fig. 1d and Extended Data Fig. 1b). Overall, we obtained the most high-quality structure predictions when combining protein language model and sequence alignment-based techniques (Extended Data Fig. 1c). To optimize workflow efficiency, we predicted structures for each representative protein sequence using the fast ESMfold algorithm, and only if the average pLDDT score was below 80 did we employ the more time-consuming ColabFold method (Fig. 1c and Extended Data Fig. 1c,d). This approach resulted in 37,223 predicted structures with a median pLDDT of 82 (interquartile range (IQR) 71–86), covering 99.8% of all clusters (Fig. 1e).
Annotation beyond the twilight zone of sequence similarity
Using sensitive sequence and structure-based annotation methods, we identified homologues for nearly half of the Asgard archaeal protein clusters (Fig. 2a). Structure-based searches enhanced the detection of homologues (Fig. 2b), particularly for clusters with high divergence, recovering significant hits in the SwissProt database for 47% of clusters (n = 17,309) versus 29% using sequence homology detection (n = 10,681). Of note, almost half of the protein representatives with both a highly confident (sequence-based) cluster of orthologous genes (COG) and structural hit displayed less than 20% sequence identity to their best structure hit (n = 4,263; median 18.6%, IQR 14.2–28.0%), falling below the ‘twilight zone’ of sequence identity (the zone between 20% and 35% sequence identity where homology becomes challenging to predict with regular algorithms)24. To illustrate the ability of our approach to annotate protein clusters even in cases of low sequence identity, we recovered the recently discovered distant Asgard archaeal homologue of Vps29 (ref. 15), a component of the eukaryotic retromer and retriever complexes, with sequence similarity searches (best structure hit amino-acid identity = 27.5%, HHsearch P = 99.8), as well as with local and global structural alignment (Foldseek E-value = 1.9 × E−20, DaliLite Z-score = 30; Fig. 2c). Extended analyses of these annotations, including domain-specific enrichment and sequence divergence patterns, are detailed in Supplementary Information (also see Supplementary Fig. 1).
a, Workflow to annotate Asgard archaeal proteins based on homology using sequence and structural similarity (also see Supplementary Fig. 1). b, Venn diagram depicting the number of clusters or cluster representing protein structures annotated using sequence homology detection with HHsearch against the COG/KOG database (orange) and structural similarity searches against AF2 SwissProt (violet), respectively. The intersection of both techniques is marked in pink. c, Structure prediction of Vps29 Asgard archaeal representative (left), its most similar SwissProt prediction (right; Cattle Vps29, Q3T0M0) and their overlay with the eukaryotic protein in violet (bottom). AA-identity, amino acid identity to best structure hit; P, hhsearch probability.
Expanding the repertoire of ESPs
Next, we used structure-based similarity searches to identify novel iESPs in Asgard archaea (Fig. 3a). We define an iESP as an Asgard archaeal protein structure that exhibits either exclusively eukaryotic hits, or a statistically significant overrepresentation of eukaryotic protein structures in (1) all hits or (2) the top 95% bit-score quantile of hits (Fig. 3b; Methods). This structure-based approach refines previous ESP classifications by incorporating a quantitative enrichment threshold rather than relying solely on presence/absence criteria. Unlike earlier definitions, which varied in their strictness or permissiveness, our method applies a standardized framework for assessing the overrepresentation of eukaryotic homologues for the investigated protein. This ensures that ESP identification remains systematic, biologically relevant and statistically justified.
a, Workflow to cluster protein structures and identify iESPs. b, Identification of Asgard archaeal iESPs based on structural similarity. c, Bar chart summarizing the clustering of previously described ESP and iESP protein structures into structural clusters, respectively. d, Sankey diagram displaying functional categories of newly identified iESP clusters and clusters containing previously established ESPs. Categories are inferred from the best SwissProt hits EggNOG annotation. ‘Multiple’ indicates an association of a structural cluster with multiple functional categories. e, Subgraph of protein structure similarity network, highlighting small GTPase (black outline) and Argonaute proteins. P, probability.
We identified 1,319 iESPs that have thus far not been identified as Asgard archaeal ESPs (Fig. 3b). Of note, we captured only 46% (611 proteins) of the 1,323 previously established Asgard archaeal ESPs, indicating that previous definitions for ESPs have been rather permissive (also see above; Fig. 3b and Supplementary Data 3). For example, 40 AsCOGs containing roadblock/LC7 domains were considered ESPs in a previous study, and Asgard archaeal proteins have been shown to form similar structures to their eukaryotic relatives35. However, only four Asgard archaeal roadblock/LC7 clusters (cog.000673, cog.000921, cog.006948 and cog.008459) are enriched in eukaryotes in our study. The marked change in coverage of previous ESPs is caused by our enrichment-based approach, which, rather than simply relying on sequence-based homology, is based on the overrepresentation of eukaryotic hits in structural similarity searches. Indeed, roadblock/LC7 domain (PF03259) containing proteins are common in prokaryotes with 24,892 and 2,494 such proteins encoded by bacterial and archaeal genomes, respectively, compared with 5,724 proteins in eukaryotes (Pfam database accessed 12 June 2024). While roadblock/LC7 domain proteins have important functions in eukaryotic cells, their widespread presence in prokaryotes suggests that previous studies may have overestimated the Asgard archaeal provenance of these proteins.
To reduce redundancy, and to obtain an overview of the structural connectivity within the (i)ESP landscape, we clustered the 37,223 predicted Asgard archaeal protein structures on the basis of their similarity, which we amalgamated into 19,775 structural clusters (Methods; Fig. 3a and Extended Data Fig. 2a). In total, the 1,319 newly identified iESPs and all 1,323 previously identified ESP protein structures are contained in 908 and 425 structural clusters (Fig. 3c), respectively, indicating that our structure-based approach more than triples the potential number of Asgard archaeal proteins that entered the eukaryotic stem lineage. A high-level functional assessment revealed remarkable differences between iESP and ESP structural clusters (Fig. 3d and Supplementary Data 3), despite the largely sparse distribution across Asgard archaeal genomes (Extended Data Figs. 3 and 4). For example, 64% of previously identified ESP clusters (336 of 425) have functions in cellular processing and signalling, including a hub of 59 clusters collectively encompassing 932 Asgard archaeal small GTPase protein representative structures (Fig. 3e), which are known to have undergone extensive duplication in both eukaryotes and Asgard archaea12,13,23,36,37. By contrast, only 28% of iESP clusters’ eukaryotic counterparts (258 of 908) are involved in cellular processing and signalling functions (when including clusters containing multiple functional categories). Among these, we identified a single cluster containing eight Argonaute-related Asgard archaeal iESPs (Extended Data Fig. 2). Argonautes are involved in DNA and RNA interference in prokaryotes and eukaryotes, respectively38. Recent studies indicate that some Asgard archaeal Argonautes appear to exhibit similar functions to their eukaryotic counterparts39,40. We obtained the best structural hits to eukaryotic AGO and PIWI proteins (Fig. 3e and Extended Data Fig. 2), illustrating their higher structural conservation despite their high level of sequence divergence38.
We also retrieved many iESP clusters specific to metabolism (Fig. 3d, n = 137), which was thus far poorly represented among previously found ESPs in Asgard archaea (n = 24; Extended Data Figs. 3 and 4). For example, we identified diverse iESPs, including best hits to proteins of the eukaryote-type mevalonate pathway (phosphomevalonate kinase, Swissprot accession: Q2KIU2), the oxygen-dependent degradation of prenylated proteins (PCYOX1, Q5R748), and reactive oxygen species defence (SOD1, P80566). As an outstanding feature, we identified many (n = 271) iESP clusters involved in information storage and processing functions, of which 169 are related to translation, ribosomal structure and biogenesis, a function in eukaryotes that is known to have an archaeal provenance41. iESPs identified within the latter functional category included best structural hits to eukaryotic elongation factor 1A lysine methyltransferase 1 (EEF1AKMT1, Q17QF2) and the malignant T-cell-amplified sequence 1 that is involved in translation re-initiation (MCT-1, Q2KIE4) (Supplementary Data 3). Altogether, our structure-based and functionally unbiased approach identified hundreds of new ESPs, bearing relevance for efforts to reconstruct the physiology and cell biological features of both extant Asgard archaea as well as the archaeal ancestor or eukaryotes.
iESPs indicate extended Asgard archaeal cellular complexity
The emergence of intricate cellular compartments has been a hallmark process of eukaryogenesis, yet the origins of many genes responsible for the formation of these compartments remain elusive42. To identify Asgard archaeal proteins potentially involved in cellular compartmentalization, we investigated iESPs with robust structural assignment but limited ‘twilight zone’ sequence similarity (Fig. 3d) and examined their relationship to their evolutionary eukaryotic counterparts. By using targeted sequence-based searches with iterative refinement guided by structural similarity, we could link several iESPs at the sequence level, after which we constructed multiple sequence alignments and performed phylogenetic analyses (Methods).
One of the eukaryotic complexes with a role in cell compartment biology and lacking a clear prokaryotic ancestry is the vault, the largest reported ribonucleoprotein complex conserved in diverse eukaryotes. This complex has been suggested to be involved in transport between cellular compartments, signal transmission, cellular stress protection and immune response43. Vaults are primarily composed of two symmetric cups, each consisting of 39 molecules of the major vault protein (MVP)44. While prokaryotic homologues of MVP have so far been described in only a few Bacteria45, we identified an Asgard archaeal protein structure with a reciprocal best hit to Xenopus laevis MVP (Q6PF69; Extended Data Fig. 5). In total, we found ten Asgard archaeal MVP homologues, half of which in our phylogenetic analysis affiliate with a clade including eukaryotic MVPs (Fig. 4a and Extended Data Fig. 5a). The representative Asgard archaeal MVP displays a predicted structure similar to the resolved rat MVP, including the cap helix, shoulder and repeat domains, even though the Asgard archaeal homologue contains only five instead of nine repeat domains present in the rat protein46 (Fig. 4b). While estimating multimeric stoichiometries remains a computationally challenging task in the absence of experimental data, here we used structural modelling to build a first model of the Asgard archaeal vault. Multimer structure modelling suggests a closed cup with ten Asgard archaeal MVP molecules (interface predicted template modelling score (ipTM) = 0.525, average pLDDT = 71.4; Extended Data Fig. 5). While the role of MVP homologues in Asgard archaea remains unknown, our findings support a prokaryotic—possibly Asgard archaeal—origin of eukaryotic MVP.
a–f, Asgard archaeal proteins related to eukaryotic MVPs (a–c) and COMMD-containing proteins (d–f). a, Phylogeny of prokaryotic and eukaryotic full-length MVPs. See Extended Data Fig. 5a for tree based only on the shoulder domain. b, Rat MVP complex46 next to Lokiarchaeial MVP (predicted structure) indicating the cap helix, shoulder and repeat domains (R). c, Biological assembly of the rat MVP cap (left) next to a multimer model of the Asgard archaeal homodecamer (right). d, Human COMMD2 next to Lokiarchaeial homologue indicating the HN and COMM domains. e, Phylogeny of prokaryotic and eukaryotic COMMD-containing proteins. f, Resolved human COMMD heterodecamer47 next to a multimer model of the Asgard archaeal homodecamer. g,h, Identification of Asgard archaeal iESPs of eukaryotic Ufm1 (g) and CINP (Hodarchaeales clade indicated with grey background) (h). Asgard archaeal query protein structure, best-scoring SwissProt target structural model and phylogenetic analysis of related protein sequences are indicated in the left, middle and right panel, respectively. Structural models exclude long terminal disordered regions. Additional data include Foldseek E-value, Dali Z-score, enrichment of eukaryotic structures (Fisher’s exact test, Bonferroni-corrected P value, ‘p-EukEnr’) and amino-acid identity to best structure hit (‘AA-identity’). Phylogenetic analyses highlight sequences for query and target structures, input MSA positions and substitution model. Scale bar, 1 amino acid substitution per position. Multimer model confidence measures (pLDDT, pTM and ipTM) are indicated. pTM, predicted template modelling score.
Another eukaryotic complex with an elusive origin is Commander. This complex is required for endosomal recycling of diverse transmembrane cargos and is composed of 16 subunits arranged into the CCC and retriever subcomplexes. While some retriever components have been reported in Asgard archaea before (Vps29, Fig. 2c; Vps35)47, the CCC (named after its components CCDC22, CCDC93 and COMMD) subunits, including the heterodecamer-forming COMMD proteins, thus far lacked prokaryotic homologues47. Our structure-based searches retrieved an Asgard archaeal iESP that displayed the characteristic COMMD protein structure, that is, an α-helical N-terminal (HN) and a C-terminal COMMD domain48, while displaying extremely low sequence identity (8.5%) (Fig. 4d). Subsequent sensitive HMM-based searches yielded homologues in diverse Asgard archaea (Lokiarchaeales, Helarchaeales and Heimdallarchaeia) and some other prokaryotes. In our phylogenetic analysis, eukaryotic COMMD proteins (COMMD1-10) form a near-monophyletic group (Fig. 4e), confirming that eukaryote-specific gene duplications gave rise to the COMMD heterodecamer47,49. While our phylogenetic analyses failed to resolve the origin of eukaryotic COMMD, multimer modelling of an Asgard archaeal homologue suggests that 8, 10 or 12 molecules may form a homomultimeric complex with high confidence (homomultimeric n = 10 in Fig. 4f; ipTM = 0.889, pLDDT = 88.4; see other homomultimers in Extended Data Fig. 5d,e).
In addition to homologues of eukaryotic proteins involved in cellular compartmentalization, we newly identified some proteins uniquely shared between eukaryotes and Asgard archaea. Despite limited sequence similarity, Ubiquitin fold modifier 1 (Ufm1) exhibits structural similarities to ubiquitin50 and is implicated in DNA damage and endoplasmic reticulum stress responses, although it has not been characterized extensively51. We identified Ufm1 homologues in nine of the major Asgard archaeal clades, but not in any other prokaryote (Fig. 4g), indicating an Asgard archaeal provenance of Ufm1 in eukaryotes. Similarly, no prokaryotic homologues have yet been reported for the cyclin-dependent kinase 2-interacting protein (CINP), a protein involved in DNA replication complex and DNA damage control52,53 that was recently also implicated in eukaryotic ribosome biogenesis54. Our sequence similarity searches revealed it is present in five major Asgard archaeal clades, but not in other prokaryotes. Phylogenetic analyses revealed that eukaryotic sequences are monophyletic and cluster with Hodarchaeal sequences with good support (Fig. 4h, UFBOOT: 99%), suggesting that eukaryotes inherited this protein from their Heimdallarchaeial ancestor15.
Discussion
This study leverages state-of-the-art structural prediction tools to uncover a broader spectrum of ESPs in Asgard archaea. Large-scale analyses of the protein structure universe are becoming powerful approaches to predicting the origins and functions of proteins beyond the capabilities of standard sequence-based homology searches55,56. Here, we explored the potential of these tools to gain insight into the archaeal provenance of the eukaryotic cell. By building and analysing a structural catalogue of the Asgard archaeal pangenome, we improved the annotation of Asgard archaeal proteins lacking significant sequence similarity. Our approach revealed many Asgard archaeal protein families, iESPs, that are structurally most similar to those of eukaryotes. As in previous studies that relied on sequence similarity searches to identify ESPs12,13,15,23, we identified iESPs involved in cellular processes and signalling, including many that participate in intracellular trafficking, secretion and vesicular transport. However, our extended analyses retrieved many iESPs involved in additional processes, such as information storage and processing. This observation is in line with the general conception that many eukaryotic proteins involved in translation, transcription, replication and DNA repair have an archaeal provenance57. Furthermore, we found that iESPs are also relatively enriched in metabolic functions, which contrasts with previous work indicating that metabolic functions in eukaryotes predominantly are of bacterial origin58,59. The underlying reason for this observation is unclear. Yet, in congruence with recent work showing that eukaryotic central carbon metabolic pathways are in part of Asgard archaeal origin60, these metabolic iESPs represent ancient homologues of eukaryotic proteins that have evolved beyond the limit of reliable sequence similarity detection. Given the scale of our dataset and the inclusion of high-confidence structure predictions independent of domain annotations, we anticipate that future studies will uncover novel domain architectures or previously uncharacterized folds among these proteins. Altogether, our analyses suggest that a thus far underappreciated fraction of the eukaryotic metabolic repertoire is of Asgard archaeal provenance. We point out that iESPs do not necessarily represent eukaryotic proteins that were directly inherited from Asgard archaea. Instead, they are Asgard proteins whose closest structural matches—often highly similar—are disproportionately found in eukaryotes. This pattern of enrichment suggests functional and evolutionary relevance, but not necessarily direct ancestry. Phylogenetic analyses to investigate the exact evolutionary relationship between iESPs and eukaryotic proteins are often hampered due to limited sequence similarity.
While several studies have revealed that some previously identified ESPs, such as small GTPases, actin homologues and several subunits of the endosomal sorting complex required for transport (ESCRT complex), are nearly universally distributed across Asgard archaeal genomes, many ESPs display a rather patchy distribution13,15,23. This patchiness is evident, for example, for Asgard archaeal homologues of adaptor proteins, Golgi-associated retrograde protein, homotypic fusion and protein sorting, and class C core vacuole/endosome tethering complexes15. A similar observation can be made for iESPs, which predominantly display patchy distribution patterns across Asgard archaeal taxa. These patchily distributed ESPs and iESPs probably represent ancient protein families that were already present in the Asgard archaeal lineage from which eukaryotes emerged, and were subject to multiple loss events or horizontal gene transfers among Asgard archaeal lineages. Overall, given their patchy distribution, combined with the evolutionary distance between present-day Asgard archaeal and eukaryotic proteins, it remains unclear to what extent Asgard archaeal iESPs are functionally equivalent to their eukaryotic counterparts. While structural conservation has been shown to be tightly linked to protein function, even at high levels of sequence divergence61, future studies are needed to corroborate the functions of Asgard archaeal iESPs and ESPs. Biochemical studies and high-resolution structural analyses will be crucial in determining whether these iESPs operate in cellular contexts analogous to their eukaryotic counterparts. Such efforts will provide deeper insights into the transitional features of eukaryotic common ancestors and refine our models of early eukaryotic evolution.
Methods
Genome dataset selection
Dataset assembly
To construct a representative initial dataset, we retrieved all publicly available Asgard genomes from the National Center for Biotechnology Information (NCBI)62 up to 6 October 2022. This collection also included the recently published Asgard archaeal MAGs from refs. 31,32. To ensure data quality, MAGs were evaluated using CheckM v1.2.1 (ref. 63). Those MAGs with estimated completeness below 50% and estimated contamination exceeding 10% were identified as low-quality and consequently excluded from the initial dataset. Taxonomic classification of the initial dataset was conducted using GTDB-Tk v2.3.2 (ref. 64) with default parameters. The final dataset comprised 936 genomes (Supplementary Data 1) covering all known Asgard archaeal lineages. Gene prediction was performed using Prokka v1.14.6 (ref. 65) (options ‘--metagenome --kingdom Archaea’).
Phylogenomic inference of the species tree
To obtain an adequate outgroup dataset for inferring the phylogenetic relationships among the different Asgard archaeal lineages, we downloaded genus-level representatives of other archaeal lineages from the Genome Taxonomy Database (GTDB), release 214 (ref. 66). We based our selection on genome quality score (GQS), defined as GQS = completeness (%) − 5 × contamination (%), as described in ref. 67. In cases where two genomes had equal GQS, a random selection was made between the two. The final outgroup dataset included 311 genus-level representatives classified as members of the Thermoproteota (excluding Korarchaeia, to avoid artefacts derived from their uncertain affiliation68 and their strong thermophilic compositions15), Methanobacteria B and Hadarchaeota lineages.
To infer the species tree, we performed phylogenomic analysis based on 47 non-ribosomal proteins, which were selected from a set of 200 markers previously identified as core archaeal proteins69 (Supplementary Data 1). Homologous sequences within the final genome dataset were recruited using PSI-BLAST70 v2.10.0+ (‘-evalue 1e-10’). All recruited sequences per taxon per protein marker were selected, aligned using MAFFT L-INS-i71 v7.453, followed by trimming with trimAl72 v1.4.rev22 (‘-gt 0.5’) and removal of sequences with more than 60% gaps. We constructed the individual protein phylogenies using IQ-TREE73 v2.1.3, incorporating model selection from ModelFinder74. The best-fitting model was selected among the combination of the LG, Q.pfam and WAG models by adding the mixture model C20 with rate heterogeneity (+R4 or +G4) (‘-mset LG+C20,Q.pfam+C20,WAG+C20 -mrate G4,R4 -mfreq ""’). We assessed branch robustness for each marker with 1,000 ultrafast bootstraps75 and Shimodaira–Hasegawa-like approximate likelihood ratio tests (SH-aLRT)76. From the resulting phylogenies, we removed sequences indicative of contamination, paralogy or horizontal gene transfer events and realigned and trimmed the remaining sequences as described above. The curated alignments were then concatenated into a supermatrix containing 1,244 sequences. To mitigate effects related to compositional bias, we performed heterogeneous site removal using χ2 trimming77 where the 50% most heterogeneous sites were removed, resulting in an alignment of 8,068 amino acid positions. We inferred a species phylogeny for the χ2-trimmed alignment using ModelFinder within IQ-TREE v2.1.3 to select among the LG + C10, Q.pfam + C10 and WAG + C10 models and rate heterogeneity components (+R4 or +G4). A posterior mean site frequency (PMSF) approximation78 of the best-fitting model (WAG + C10 + R4) using the resulting tree was then employed to reconstruct a final tree with 100 non-parametric bootstrap pseudoreplicates.
Clustering and selection of representative protein sequences
The dataset of 936 Asgard archaeal genomes comprised 2.68 million proteins. We assigned AsCOG domains23 to 2.1 million Asgard archaeal proteins according to the best hit to an AsCOG member using MMseqs2 (ref. 79) with ‘-e 0.001’ and ‘-s 9’ and at least 80% of the best hit had to be covered. Unassigned proteins (0.6 million) or protein fragments (0.2 million) of at least 60 amino acids were clustered de novo using MMseqs2 (ref. 80) v14.7e284 at 20% sequence identity and a coverage of 50%. We built sequence profiles for 14,467 (2,084,964 represented proteins) AsCOGs and 22,846 (448,812 represented proteins) de novo clusters with at least 5 members. To select an evolutionary representative sequence per cluster, we searched members of the 37,313 clusters with at least five members against their respective cluster profile using MMseqs2 (‘mmseqs search’), ranked them based on their bit-score and selected the highest-ranked sequence per cluster as the representative sequence79.
Protein structure prediction
Supplementing the ColabFold database with Asgard archaeal proteins
Protein structure prediction using AlphaFold2 (ref. 27) has been shown to generally perform poorly if few sequences can be aligned to the target sequence27. We therefore wanted to evaluate whether adding our Asgard archaeal protein dataset to the ‘genetic search’ workflow of ColabFold33, an accelerated adaptation of AlphaFold2, would increase overall prediction quality. To this end, we implemented a version of the ‘genetic search’ workflow of ColabFold that queries the Asgard archaeal protein dataset (‘enriched’) in addition to the default databases (‘default’). For the enriched workflow, we added a third MMseqs2 sequence search step against the Asgard archaeal protein database as after the searches against the two default ColabFold databases with the same parameters.
Comparing performance of structure prediction algorithms for an Asgard archaeon
To evaluate performance of different structure prediction algorithms as well as the ColabFold ‘default’ versus the ‘enriched’ database, we created a test set of 100 Asgard archaeal proteins (Supplementary Data 2). We downloaded 100 randomly selected proteins of a reference Asgard archaeal proteome, P. syntrophicum, from UniProt (Supplementary Data 2; Proteome ID: UP000321408; accessed on 17 January 2023). We first predicted structural models from the primary sequences using the protein language model based ESMfold v2.0.0 (ref. 34) with option ‘-r 12’. To measure the quality of predictions, we used the average pLDDT score, ranging from low to high confidence (0–100). We considered predictions with an average pLDDT ≥80 as high-quality, as a compromise between the suggested pLDDT ≥90 for ‘high accuracy’ and pLDDT ≥70 for ‘general correct backbone’ according to ref. 27. Second, we generated multiple sequence alignments with the ‘genetic search’ module of ColabFold v1.3.0 (ref. 33) with default and enriched database, respectively. We then ran the ColabFold prediction workflow on each alignment using the default ‘exhaustive’ setting and a premature stopping rule (‘early-stop’) designed to reduce computation time; specifically, the algorithm terminates if a pLDDT of at least 85 is reached or if the first prediction yields a pLDDT below 50 (‘--stop-at-score 85 --stop-at-score-below 50’). The ‘genetic search’ module was run on a computer equipped with two AMD EPYC 7H12 processors (64 cores each, 2.6 GHz, 280 W) and 1 TiB of memory, whereas the ‘prediction’ module was run on a system with four NVIDIA A100 graphics processing units (40 GiB HBM2 memory each).
Protein structure prediction workflow
Based on the highest ratio of high-quality proteins and lowest computational resource demands for our 100 test proteins, we opted for a hybrid approach of using protein language model- and multiple sequence alignment-based prediction algorithms. We first used ESMfold v2.0.0 (ref. 34) with ‘-r 12’ to calculate structural models for each representative Asgard archaeal protein. Second, structures with an average pLDDT <80 in ESMfold were predicted again using ColabFold v1.3.0 (ref. 33) with the enriched database and the ‘early-stop’ settings. Large proteins that could not be folded with ESMfold v2.0.0 and ColabFold v1.3.0 because of exceeding memory demands were attempted to be folded with ColabFold v1.5.2.
Structure similarity searches
Best structural hit annotation
We searched Asgard archaeal structures reciprocally against SwissProt predicted structures (downloaded 8 July 2022) using FoldSeek v6.29e2557 (ref. 28) ‘foldseek search’ with ‘--max-seqs 10000’. To ensure robustness in structural comparisons, we use the default local structural alignment via Foldseek rather than relying on global fold similarity (for example, TM-score). This mitigates potential biases introduced by differences between ColabFold and ESMFold models, as functionally relevant local motifs remain detectable regardless of global conformational variations. We retained the highest bit-score non-overlapping hits along the query sequence to accommodate fusion proteins and checked for reciprocal best hits. We mapped the annotation of the SwissProt best hits to each query protein. As described above, but unidirectionally, we searched Asgard archaeal structures against the Protein Data Bank and UniProt50 databases (downloaded 9 February 2023).
EggNOG annotation of SwissProt best hits
Proteins representing the best SwissProt hits were mapped against EggNOG v5 (ref. 81) with the emapper user interface (http://eggnog-mapper.embl.de/) with default parameters, and we extracted root non-supervised orthologous group (NOG) and eukaryotic NOG identifiers and functional categories.
Identification eukaryotic hit enriched structures
For each Asgard archaeal predicted structure, we collected the best 10,000 hits of predicted UniProt50 structures (downloaded 9 February 2023), which contains proteins from all domains of life, ensuring that our ESP identification pipeline inherently considers homologues across bacteria, archaea and eukaryotes. Per Asgard archaeal protein representative, we performed a one-tailed Fisher’s exact test with the function ‘fisher.test’ and the ‘alternative=less’ parameter with Bonferroni correction with the function ‘p.adjust’ in R v4.2.1 (ref. 82) on the domain-level taxonomy of hit UniProt proteins to test for a statistical enrichment in eukaryotic sequences. To test for eukaryotic enrichment in only the most similar proteins, we also performed the same statistical test using only the top 5% bit-score percentile of the hits. Structures with an enrichment in hits to eukaryotic proteins were classified as candidate isomorphic (i)ESPs, that is, proteins that look structurally similar to proteins that are overrepresented in eukaryotes. We clustered all Asgard archaeal structures with Foldseek ‘foldseek cluster’ into clusters of isomorphic protein structures and identified structural clusters uniquely added with iESPs.
NCBI COG and KOG annotation of gene families
We created multiple sequence alignments for each Asgard archaeal protein cluster using FAMSA v2.2.2 (ref. 83) with ‘-refine_mode on’. We performed profile–profile searches with the HHsuite3 (ref. 84) program HHsearch v3.3.0 with parameters ‘-glob -M 50’ against the profile COG–eukaryotic orthologous groups (KOG) database (ftp://ftp.tuebingen.mpg.de/pub/protevo/toolkit/databases/hhsuite_dbs/COG_KOG.tar.gz)85.
Mapping of ESPs described by Eme et al. (2023)
To identify conserved protein domains in the proteomes of the Asgard archaeal dataset, we used InterProScan v5.57-90.0 (ref. 86) with default parameters and using hidden Markov models (HMM) from the databases AntiFam v7.0 (ref. 87), CDD v3.18 (ref. 88), Coils v2.2.1 (ref. 89), Gene3D v4.3.0 (ref. 90), MMobiDBLite v2.0 (ref. 91), PANTHER v15.0 (ref. 92), Pfam v35.0 (ref. 93), PIRSF v3.10 (ref. 94), PRINTS v42.0 (ref. 95), SFLD v4 (ref. 96), SMART v7.1 (ref. 97), SUPERFAMILY v1.75 (ref. 98) and TIGRFAM v15.0 (ref. 99).
We then identified the AsCOG and de novo cluster protein domains containing at least 80% of the length of a Pfam or Interpro domains reported as ESPs15.
Phylogenetic inferences of iESPs
iESP selection
To illustrate how iESP confer information about the origins of eukaryotic functions and their proteins, we selected several iESPs for phylogenetic analysis, based on the following criteria: the Asgard archaeal query structure is well covered (>80% of protein length) by its alignment to its best structure hit; the best (eukaryotic) structure hit reciprocally has the Asgard archaeal query structure as its best hit; eukaryotic structures are overrepresented among the hits (Fig. 3b); the eukaryotic hit structures are consistent (are evidently homologous to one another); they comprise eukaryote-relevant functions; neither the query nor the hit appears to embody particularly complex evolutionary histories (for example, they do not contain repeat domains or highly composite multidomain architectures); and the Asgard archaeal query is unlikely to represent contamination, as it is found in more than one Asgard archaeal taxon. Finally, we required that the candidates lack a well-scoring sequence-based hit to eukaryotic sequences, as determined by HHsearch; consequently, they fall into the ‘twilight zone’ of sequence homology (Fig. 3c).
Establishing remote sequence similarity between iESP and eukaryotic structure hits
Subsequently, we found that the iESPs, although divergent, retain sequence signals that connect them to the eukaryotic proteins they match structurally. For this, we sought to gradually expand the homologue set of the iESP via manually supervised, iterative HMM searches. In each round, we checked the newly hit proteins before adding them to the multiple sequence alignment, as we ensured these are genuine homologues by inspecting both their sequences and (predicted) protein structures. We executed these profile HMM-based searches using online tools (HHpred and HMMer web server) as well as local hmmsearches onto our local databases (see description below). Note that, in addition to eukaryotic and Asgard archaeal sequences, we included bacterial and other archaeal sequences in the search database, as they may also have homologues that could help link the iESP and related Asgard archaeal sequences to their eukaryotic structural hits.
Selecting homologues for phylogenetic inference
We made use of three sequence datasets for retrieving sequences for the phylogenetic analysis. First, we subsampled our in-house Asgard archaeal set, including only a single representative protein set per species. This representative for a given species was selected based on the quality of the predicted proteomes, as reflected by their predicted completeness and contamination, measured by CheckM63. Note that ‘species’ here signifies groups of genomes that can be clustered at the 95% average nucleotide identity level. Second, we used a subsampled version of an in-house eukaryotic dataset100, including 25 eukaryotic taxa of all of the major eukaryotic groups, taking the taxon with the best, most complete, predicted proteome quality, as measured by BUSCO101. Third, we used a subsampled version of GTDB (r207)66, of which first the Asgard archaea were removed, and then we selected the best assembly for each family, which was also based on the CheckM quality parameters. Using the final, most inclusive yet accurate profile HMM obtained, and our manually determined bit-score cut-offs (described above), we employed hmmsearch onto these three datasets and retrieved all sequences meeting the cut-off. Because COMMD and CINPL comprised virtually full-length hits, both at the structural comparisons as well as in our sequence similarity searches, we extracted the entire protein sequence of each hit protein. For Ufm1, we observed that some hits in our sequence searches were not full-length, and others contained multiple hit regions; in these cases, we extracted only the protein segment corresponding to the best-scoring hit. For the MVP, in addition to the smaller full-length phylogeny (Fig. 4a), we performed a broader phylogenetic analysis of the shoulder domain only, which is a type of Band 7 domain found in many prokaryotic and eukaryotic proteins45,102, and which are united in the SPFH (for stomatins, prohibitins, flotillins and HflK/C) family ‘clan’ (https://www.ebi.ac.uk/interpro/set/pfam/CL0433/entry/pfam/).
Phylogenetic analysis and annotation of the phylogeny
For each family, we inferred gene trees using multiple sequences alignments generated by MAFFT (v7.505, mode L-INS-i)71 and the web server of PROMALS3D103. For the latter, we used the default options, except for detecting and using homologues with three-dimensional structures (included DaliLite v5 (ref. 104)), pairwise alignments between input three-dimensional structures (included DaliLite) and aligning sequences within groups in the first alignment stage (PROMALS instead of MAFFT). We supplemented PROMALS3D with predicted protein structures from diverse sequences in the AlphaFold Protein Structure Database, as well as with structures from our own predictions (described above), including those of the iESPs and, where available, other Asgard archaeal homologues. Before inferring the gene trees, we trimmed the multiple sequence alignment using BMGE v1.12 (settings: ‘-m BLOSUM30 --h 0.6 -g 0.7 -b 3’)105, which selects good-quality aligned positions. However, in some cases (for example, COMMD MAFFT alignment), this produced very short alignments, prompting us to switch to trimAl (v1.4.1, mode ‘gappyout’)72. For phylogenetic inference in a maximum-likelihood framework, we used IQ-TREE v.2.0.3 (settings ‘-B 1000 -m MFP -mset LG,JTT,Q.pfam,WAG,LG+C20,LG+C40,LG+C60,LG+C20+R+F,LG+C40+R+F,LG+C60+R+F,WAG+C20,WAG+C40,WAG+C60,WAG+C20+R+F,WAG+C40+R+F,WAG+C60+R+F,JTT+C20,JTT+C40,JTT+C60,JTT+C20+R+F,JTT+C40+R+F,JTT+C60+R+F,Q.pfam+C20,Q.pfam+C40,Q.pfam+C60,Q.pfam+C20+R+F,Q.pfam+C40+R+F,Q.pfam+C60+R+F’)73 to first select the best evolutionary model using ModelFinder74 and then infer a phylogeny with 1,000 ultrafast bootstraps75. For each iESP/family, we subsequently selected the phylogeny displaying the most informative and probably accurate tree, which entailed post-hoc selecting the alignment algorithm (MAFFT-L-INS-i versus PROMALS3D) (based on ultrafast bootstrap support values at key branches, and monophyly of expected monophyletic sequence groups). We coloured the branches in the tree according to the species group the sequences belong to: Eukaryota, Asgard archaea, Archaea (other) and Bacteria. We also annotated the eukaryotic clades with the names of their proteins, specifically labelling each clade reflecting a single gene in the last eukaryotic common ancestor. Trees were visualized using iTOL106.
Visual representation of protein structures
Structural models were either visualized in ChimeraX v1.6.1 (Fig. 4b–f)107 or in R with the ‘r3dmol’ package v0.1.2 (Fig. 4g,h) (https://github.com/swsoyee/r3dmol)108.
Statistics and reproducibility
No statistical method was used to predetermine sample size. No data were excluded from the analyses. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment. For benchmarking structure prediction methods (Extended Data Fig. 1b–d), 100 proteins were randomly sampled from the proteome of P. syntrophicum (UniProt ID: UP000321408; Supplementary Data 2). Each protein was evaluated once per prediction condition; no technical replicates were performed. This sample size was selected to provide a representative yet computationally feasible comparison.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Asgard archaeal genome data were obtained from NCBI GenBank (https://www.ncbi.nlm.nih.gov/nucleotide), and identifiers can be found in Supplementary Data 1. Archaeal outgroup proteomes were downloaded from Genome Taxonomy Database (GTDB) release 214 (https://data.gtdb.ecogenomic.org/releases/release214/214.0/genomic_files_reps/gtdb_proteins_aa_reps_r214.tar.gz) and proteins for gene phylogenies from release 207 (https://data.gtdb.ecogenomic.org/releases/release207/207.0/genomic_files_reps/gtdb_proteins_aa_reps_r207.tar.gz). The Asgard protein database and all predicted structures, original multiple sequence alignments and IQ-TREE outputs are available via figshare at https://doi.org/10.6084/m9.figshare.26057632 (ref. 109). The uncollapsed phylogenies can be found via the iTOL106 website at https://itol.embl.de/tree/62145192210399341699888333 (CINP; Fig. 4a), https://itol.embl.de/tree/13722425212199811699868285 (COMMD; Fig. 4c) and https://itol.embl.de/tree/62145192210319901699902102 (Ufm1; Fig. 4d).
Code availability
Custom code is available via GitHub at https://github.com/stephkoest/structural_genomics.
References
Stanier, R. Y., Doudoroff, M. & Adelberg, E. A. The Microbial World (Prentice-Hall, 1963).
Vosseberg, J. et al. The emerging view on the origin and early evolution of eukaryotic cells. Nature 633, 295–305 (2024).
Betts, H. C. et al. Integrated genomic and fossil evidence illuminates life’s early evolution and eukaryote origin. Nat. Ecol. Evol. 2, 1556–1562 (2018).
Mahendrarajah, T. A. et al. ATP synthase evolution on a cross-braced dated tree of life. Nat. Commun. 14, 7456 (2023).
Cox, C. J., Foster, P. G., Hirt, R. P., Harris, S. R. & Embley, T. M. The archaebacterial origin of eukaryotes. Proc. Natl Acad. Sci. USA 105, 20356–20361 (2008).
Eme, L., Spang, A., Lombard, J., Stairs, C. W. & Ettema, T. J. G. Archaea and the origin of eukaryotes. Nat. Rev. Microbiol. 15, 711–723 (2017).
Roger, A. J., Muñoz-Gómez, S. A. & Kamikawa, R. The origin and diversification of mitochondria. Curr. Biol. 27, R1177–R1192 (2017).
Martijn, J. & Ettema, T. J. G. From archaeon to eukaryote: the evolutionary dark ages of the eukaryotic cell. Biochem. Soc. Trans. 41, 451–457 (2013).
Schwartz, R. M. & Dayhoff, M. O. Origins of prokaryotes, eukaryotes, mitochondria, and chloroplasts: a perspective is derived from protein and nucleic acid sequence data. Science 199, 395–403 (1978).
Yang, D., Oyaizu, Y., Oyaizu, H., Olsen, G. J. & Woese, C. R. Mitochondrial origins. Proc. Natl Acad. Sci. USA 82, 4443–4447 (1985).
Tamarit, D. et al. Description of Asgardarchaeum abyssi gen. nov. spec. nov., a novel species within the class Asgardarchaeia and phylum Asgardarchaeota in accordance with the SeqCode. Syst. Appl. Microbiol. 47, 126525 (2024).
Spang, A. et al. Complex archaea that bridge the gap between prokaryotes and eukaryotes. Nature 521, 173–179 (2015).
Zaremba-Niedzwiedzka, K. et al. Asgard archaea illuminate the origin of eukaryotic cellular complexity. Nature 541, 353–358 (2017).
Williams, T. A., Cox, C. J., Foster, P. G., Szöllősi, G. J. & Embley, T. M. Phylogenomics provides robust support for a two-domains tree of life. Nat. Ecol. Evol. 4, 138–147 (2020).
Eme, L. et al. Inference and reconstruction of the heimdallarchaeial ancestry of eukaryotes. Nature 618, 992–999 (2023).
Hartman, H. & Fedorov, A. The origin of the eukaryotic cell: a genomic investigation. Proc. Natl Acad. Sci. USA 99, 1420–1425 (2002).
Akıl, C. & Robinson, R. C. Genomes of Asgard archaea encode profilins that regulate actin. Nature 562, 439–443 (2018).
Akıl, C. et al. Insights into the evolution of regulated actin dynamics via characterization of primitive gelsolin/cofilin proteins from Asgard archaea. Proc. Natl Acad. Sci. USA 117, 19904–19913 (2020).
Survery, S. et al. Heimdallarchaea encodes profilin with eukaryotic-like actin regulation and polyproline binding. Commun. Biol. 4, 1024 (2021).
Hatano, T. et al. Asgard archaea shed light on the evolutionary origins of the eukaryotic ubiquitin-ESCRT machinery. Nat. Commun. 13, 3398 (2022).
Imachi, H. et al. Isolation of an archaeon at the prokaryote–eukaryote interface. Nature 577, 519–525 (2020).
Rodrigues-Oliveira, T. et al. Actin cytoskeleton and complex cell architecture in an Asgard archaeon. Nature 613, 332–339 (2023).
Liu, Y. et al. Expanded diversity of Asgard archaea and their relationships with eukaryotes. Nature 593, 553–557 (2021).
Rost, B. Twilight zone of protein sequence alignments. Protein Eng. 12, 85–94 (1999).
Illergård, K., Ardell, D. H. & Elofsson, A. Structure is three to ten times more conserved than sequence—a study of structural response in protein cores. Proteins 77, 499–508 (2009).
Vanni, C. et al. Unifying the known and unknown microbial coding sequence space. eLife 11, e67667 (2022).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
van Kempen, M. et al. Fast and accurate protein structure search with Foldseek. Nat. Biotechnol. 42, 243–246 (2024).
Ruperti, F. et al. Cross-phyla protein annotation by structural prediction and alignment. Genome Biol. 24, 113 (2023).
Seong, K. & Krasileva, K. V. Prediction of effector protein structures from fungal phytopathogens enables evolutionary analyses. Nat. Microbiol. 8, 174–187 (2023).
K.E. Appler et al. Oxygen metabolism in descendants of the archaeal-eukaryotic ancestor. Nature https://doi.org/10.1038/s41586-026-10128-z (2026).
Valentin-Alvarado, L. E. et al. Asgard archaea modulate potential methanogenesis substrates in wetland soil. Nat. Commun. 15, 6384 (2024).
Mirdita, M. et al. ColabFold: making protein folding accessible to all. Nat. Methods 19, 679–682 (2022).
Lin, Z. et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science 379, 1123–1130 (2023).
Tran, L. T., Akıl, C., Senju, Y. & Robinson, R. C. The eukaryotic-like characteristics of small GTPase, roadblock and TRAPPC3 proteins from Asgard archaea. Commun. Biol. 7, 273 (2024).
Klinger, C. M., Spang, A., Dacks, J. B. & Ettema, T. J. G. Tracing the archaeal origins of eukaryotic membrane-trafficking system building blocks. Mol. Biol. Evol. 33, 1528–1541 (2016).
Vosseberg, J. et al. Timing the origin of eukaryotic cellular complexity with ancient duplications. Nat. Ecol. Evol. 5, 92–100 (2021).
Swarts, D. C. et al. The evolutionary journey of Argonaute proteins. Nat. Struct. Mol. Biol. 21, 743–753 (2014).
Bastiaanssen, C. et al. RNA-guided RNA silencing by an Asgard archaeal Argonaute. Nat. Commun. 15, 5499 (2024).
Leão, P. et al. Asgard archaea defense systems and their roles in the origin of eukaryotic immunity. Nat. Commun. 15, 6386 (2024).
Koonin, E. V. & Yutin, N. The dispersed archaeal eukaryome and the complex archaeal ancestor of eukaryotes. Cold Spring Harb. Perspect. Biol. 6, a016188 (2014).
Prokopchuk, G. et al. Lessons from the deep: mechanisms behind diversification of eukaryotic protein complexes. Biol. Rev. Camb. Philos. Soc. 98, 1910–1927 (2023).
Berger, W., Steiner, E., Grusch, M., Elbling, L. & Micksche, M. Vaults and the major vault protein: novel roles in signal pathway regulation and immunity. Cell. Mol. Life Sci. 66, 43–61 (2009).
Frascotti, G. et al. The Vault nanoparticle: a gigantic ribonucleoprotein assembly involved in diverse physiological and pathological phenomena and an ideal nanovector for drug delivery and therapy. Cancers 13, 707 (2021).
Daly, T. K., Sutherland-Smith, A. J. & Penny, D. In silico resurrection of the major vault protein suggests it is ancestral in modern eukaryotes. Genome Biol. Evol. 5, 1567–1583 (2013).
Casañas, A. et al. New features of vault architecture and dynamics revealed by novel refinement using the deformable elastic network approach. Acta Crystallogr. D 69, 1054–1061 (2013).
Healy, M. D. et al. Structure of the endosomal Commander complex linked to Ritscher–Schinzel syndrome. Cell 186, 2219–2237 (2023).
Healy, M. D. et al. Structural insights into the architecture and membrane interactions of the conserved COMMD proteins. eLife 7, e35898 (2018).
Laulumaa, S., Kumpula, E.-P., Huiskonen, J. T. & Varjosalo, M. Structure and interactions of the endogenous human Commander complex. Nat. Struct. Mol. Biol. 31, 925–938 (2024).
Komatsu, M. et al. A novel protein-conjugating system for Ufm1, a ubiquitin-fold modifier. EMBO J. 23, 1977–1986 (2004).
Zhou, X. et al. UFMylation: a ubiquitin-like modification. Trends Biochem. Sci 49, 52–67 (2024).
Lovejoy, C. A. et al. Functional genomic screens identify CINP as a genome maintenance protein. Proc. Natl Acad. Sci. USA 106, 19304–19309 (2009).
Grishina, I. & Lattes, B. A novel Cdk2 interactor is phosphorylated by Cdc7 and associates with components of the replication complexes. Cell Cycle 4, 1120–1126 (2005).
Ni, C. et al. Labeling of heterochronic ribosomes reveals C1ORF109 and SPATA5 control a late step in human ribosome assembly. Cell Rep. 38, 110597 (2022).
Durairaj, J. et al. Uncovering new families and folds in the natural protein universe. Nature 622, 646–653 (2023).
Barrio-Hernandez, I. et al. Clustering predicted structures at the scale of the known protein universe. Nature 622, 637–645 (2023).
Yutin, N., Makarova, K. S., Mekhedov, S. L., Wolf, Y. I. & Koonin, E. V. The deep archaeal roots of eukaryotes. Mol. Biol. Evol. 25, 1619–1630 (2008).
Rivera, M. C. & Lake, J. A. The ring of life provides evidence for a genome fusion origin of eukaryotes. Nature 431, 152–155 (2004).
Brueckner, J. & Martin, W. F. Bacterial genes outnumber archaeal genes in eukaryotic genomes. Genome Biol. Evol. 12, 282–292 (2020).
Molina, C. S., Williams, T. A., Snel, B. & Spang, A. Chimeric origins and dynamic evolution of central carbon metabolism in eukaryotes. Nat. Ecol. Evol. 9, 613–627 (2025).
Friedberg, I. & Margalit, H. Persistently conserved positions in structurally similar, sequence dissimilar proteins: roles in preserving protein fold and function. Protein Sci. 11, 350–360 (2002).
Sayers, E. W. et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 50, D20–D26 (2022).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Chaumeil, P.-A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. GTDB-Tk v2: memory friendly classification with the genome taxonomy database. Bioinformatics 38, 5315–5316 (2022).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Rinke, C. et al. A standardized archaeal taxonomy for the Genome Taxonomy Database. Nat. Microbiol. 6, 946–959 (2021).
Parks, D. H. et al. Recovery of nearly 8,000 metagenome-assembled genomes substantially expands the tree of life. Nat. Microbiol. 2, 1533–1542 (2017).
Tahon, G. et al. Phylogenomics and ancestral reconstruction of Korarchaeota reveals genomic adaptation to habitat switching. Preprint at bioRxiv https://www.biorxiv.org/content/10.1101/2023.09.28.559970v2 (2023).
Petitjean, C., Deschamps, P., López-García, P., Moreira, D. & Brochier-Armanet, C. Extending the conserved phylogenetic core of archaea disentangles the evolution of the third domain of life. Mol. Biol. Evol. 32, 1242–1254 (2015).
Schäffer, A. A. et al. Improving the accuracy of PSI-BLAST protein database searches with composition-based statistics and other refinements. Nucleic Acids Res. 29, 2994–3005 (2001).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., von Haeseler, A. & Jermiin, L. S. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat. Methods 14, 587–589 (2017).
Hoang, D. T., Chernomor, O., von Haeseler, A., Minh, B. Q. & Vinh, L. S. UFBoot2: improving the ultrafast bootstrap approximation. Mol. Biol. Evol. 35, 518–522 (2018).
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010).
Viklund, J., Ettema, T. J. G. & Andersson, S. G. E. Independent genome reduction and phylogenetic reclassification of the oceanic SAR11 clade. Mol. Biol. Evol. 29, 599–615 (2012).
Wang, H.-C., Minh, B. Q., Susko, E. & Roger, A. J. Modeling site heterogeneity with posterior mean site frequency profiles accelerates accurate phylogenomic estimation. Syst. Biol. 67, 216–235 (2018).
Steinegger, M. & Söding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 35, 1026–1028 (2017).
Steinegger, M. & Söding, J. Clustering huge protein sequence sets in linear time. Nat. Commun. 9, 2542 (2018).
Huerta-Cepas, J. et al. eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res. 47, D309–D314 (2019).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2021).
Deorowicz, S., Debudaj-Grabysz, A. & Gudyś, A. FAMSA: fast and accurate multiple sequence alignment of huge protein families. Sci. Rep. 6, 33964 (2016).
Steinegger, M. et al. HH-suite3 for fast remote homology detection and deep protein annotation. BMC Bioinformatics 20, 473 (2019).
Biegert, A., Mayer, C., Remmert, M., Söding, J. & Lupas, A. N. The MPI Bioinformatics Toolkit for protein sequence analysis. Nucleic Acids Res. 34, W335–W339 (2006).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014).
Eberhardt, R. Y. et al. AntiFam: a tool to help identify spurious ORFs in protein annotation. Database 2012, bas003 (2012).
Lu, S. et al. CDD/SPARCLE: the conserved domain database in 2020. Nucleic Acids Res. 48, D265–D268 (2020).
Linding, R. et al. Protein disorder prediction: implications for structural proteomics. Structure 11, 1453–1459 (2003).
Lees, J. et al. Gene3D: a domain-based resource for comparative genomics, functional annotation and protein network analysis. Nucleic Acids Res. 40, D465–D471 (2012).
Necci, M., Piovesan, D., Dosztányi, Z. & Tosatto, S. C. E. MobiDB-lite: fast and highly specific consensus prediction of intrinsic disorder in proteins. Bioinformatics 33, 1402–1404 (2017).
Thomas, P. D. et al. PANTHER: making genome-scale phylogenetics accessible to all. Protein Sci. 31, 8–22 (2022).
Mistry, J. et al. Pfam: the protein families database in 2021. Nucleic Acids Res. 49, D412–D419 (2021).
Wu, C. H. et al. PIRSF: family classification system at the Protein Information Resource. Nucleic Acids Res. 32, D112–114 (2004).
Attwood, T. K. The PRINTS database: a resource for identification of protein families. Brief. Bioinform. 3, 252–263 (2002).
Akiva, E. et al. The Structure–Function Linkage Database. Nucleic Acids Res. 42, D521–D530 (2014).
Letunic, I., Doerks, T. & Bork, P. SMART 7: recent updates to the protein domain annotation resource. Nucleic Acids Res. 40, D302–D305 (2012).
Wilson, D. et al. SUPERFAMILY—sophisticated comparative genomics, data mining, visualization and phylogeny. Nucleic Acids Res. 37, D380–D386 (2009).
Haft, D. H., Selengut, J. D. & White, O. The TIGRFAMs database of protein families. Nucleic Acids Res. 31, 371–373 (2003).
de Potter, B., Raas, M. W. D., Seidl, M. F., Verrijzer, C. P. & Snel, B. Uncoupled evolution of the Polycomb system and deep origin of non-canonical PRC1. Commun. Biol. 6, 1144 (2023).
Manni, M., Berkeley, M. R., Seppey, M., Simão, F. A. & Zdobnov, E. M. BUSCO update: novel and streamlined workflows along with broader and deeper phylogenetic coverage for scoring of eukaryotic, prokaryotic, and viral genomes. Mol. Biol. Evol. 38, 4647–4654 (2021).
Sokolskyi, T. H. Bacterial Major Vault Protein homologs shed new light on origins of the enigmatic organelle. Preprint at bioRxiv https://doi.org/10.1101/872010 (2019).
Pei, J. & Grishin, N. V. PROMALS3D: multiple protein sequence alignment enhanced with evolutionary and three-dimensional structural information. Methods Mol. Biol. 1079, 263–271 (2014).
Holm, L. Dali server: structural unification of protein families. Nucleic Acids Res. 50, W210–W215 (2022).
Criscuolo, A. & Gribaldo, S. BMGE (Block Mapping and Gathering with Entropy): a new software for selection of phylogenetic informative regions from multiple sequence alignments. BMC Evol. Biol. 10, 210 (2010).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
Meng, E. C. et al. UCSF ChimeraX: tools for structure building and analysis. Protein Sci. 32, e4792 (2023).
Rego, N. & Koes, D. 3Dmol.js: molecular visualization with WebGL. Bioinformatics 31, 1322–1324 (2015).
Dataset for: Prediction of eukaryotic cellular complexity in Asgard archaea using structural modelling. figshare https://doi.org/10.6084/m9.figshare.26057632 (2026).
Acknowledgements
We thank F. Homa and V. de Jager for technical assistance and SURF (www.surf.nl) for supporting the use of the National Supercomputer Snellius, facilitated through a grant from the Dutch Research Council (NWO-2021.059, T.J.G.E.). This work was supported by the European Research Council Consolidator and Advanced Grants 817834 and 101142180, respectively (T.J.G.E.), the Dutch Research Council VI.C.192.016 (T.J.G.E.) and VI.Veni.212.099 (J.J.E.v.H.), the Volkswagen Foundation Grant 96725 (T.J.G.E.) and the Simons Foundation as part of the Moore-Simons Project on the Origin of the Eukaryotic Cell (Grant 73592LPI; https://doi.org/10.46714/735925LPI) (T.J.G.E. and B.J.B.). Computational resources were provided by the SURF Cooperative, grant no. EINF-2953.
Author information
Authors and Affiliations
Contributions
S.K. and T.J.G.E. conceptualized the study. S.K. led orthology assignment, protein modelling and sequence homology searches, with support from J.J.E.v.H. Structural genomics analyses were performed by S.K. and J.J.E.v.H. Genome data generation and curation were carried out by K.E.A., B.J.B. and V.D.A., while phylogenetic analyses were conducted by J.J.E.v.H., S.K. and K.P. Data interpretation involved S.K., J.J.E.v.H., K.P., K.E.A., D.T. and T.J.G.E. Supervision was provided by T.J.G.E. S.K., J.J.E.v.H. and T.J.G.E. wrote the original draft, and all authors (S.K., J.J.E.v.H., K.P., D.T., K.E.A., V.D.A., B.J.B. and T.J.G.E.) contributed to reviewing and editing the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Damien Devos, Robert Robinson and Rui Zhao for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Asgard archaeal phylogenomic tree and comparison of structural prediction algorithms.
a, Maximum-likelihood phylogenetic tree of 935 Asgard archaeal genomes, using Euryarchaeota and Thermoproteota archaeal representatives as outgroup. The tree is based on 47 concatenated non-ribosomal proteins (8,068 sites and 1,244 taxa), using IQ-TREE under the WAG+C10+R4 model. PMSF approximated non-parametric bootstrap support ≥70 is indicated on branches. Scalebar represents the average expected substitutions per site. b-d, Evaluation of different aspects of the structure prediction workflow in the following panels were performed on a set of 100 proteins of Prometheoarchaeum syntrophicum (n = 100; see Methods). b, Number of aligned reference sequences (x-axis) and average structure model pLDDT (y-axis) with default (blue) and enriched (purple) database. Each dot represents a structure prediction for one of the 100 randomly selected proteins from Prometheoarchaeum syntrophicum. Lines show linear regression between pLDDT and the number of aligned sequences with shaded 95% confidence intervals. c, Number of high-quality structure predictions (pLDDT ≥80) based on different predictions strategies. d, Inference times of ColabFold prediction modules with different inference strategies, including the default setting and database, or the enriched database with either default settings or an early stop criterion (see Methods). Boxes represent the interquartile range (IQR), with the centre line showing the median. Whiskers extend to the most extreme data points within 1.5×IQR from the box. Outliers are shown as individual points.
Extended Data Fig. 2 Analyses of the Asgard archaeal protein structure similarity network.
a, Subgraph complementing the protein structure similarity network depicted in Fig. 3, once again highlighting Argonaute proteins. b, Distribution across Asgard archaeal groups of eight Asgard archaeal Argonaute-related iESPs contained in a single structural cluster.
Extended Data Fig. 3 Distribution of ESPs and iESPs in Asgard archaea.
Complementing Fig. 3d, this heatmap displays the presence of ESPs (green) and iESPs (purple) across Asgard archaeal genomes. Genomes (y-axis) are grouped by taxonomy, and structural clusters (x-axis) are sorted by conservation across genomes and functional categories.
Extended Data Fig. 4 Distribution of ESPs and iESPs across Asgard archaea by functional category of best structural hit.
Presence of eukaryotic signature proteins (ESPs) and isomorphic ESPs (iESPs) across Asgard archaeal genomes grouped by taxonomy (y-axis). Structural clusters are ordered by functional category (x-axis). Each column represents a distinct structural cluster, categorized based on predicted functional annotations. Functional categories (x-axis labels) follow COG annotations, reflecting major biological processes, including information storage and processing, cellular processes and signaling, and metabolism. Asgard archaeal genomes are grouped into taxonomic lineages (abbreviation on y-axis). Black lines demarcate major Asgard archaeal clades. ESPs (green) and iESPs (purple) show distinct patterns of conservation across taxonomic groups and functional categories. This extended dataset builds on the high-level summary in Fig. 3 and Extended Data Fig. 3, providing deeper resolution into functional distributions of ESPs and iESPs. Functional categories follow COG annotations and are labeled by their letter codes, including: J, Translation, ribosomal structure and biogenesis; A, RNA processing and modification; K, Transcription; L, Replication, recombination and repair; B, Chromatin structure and dynamics; D, Cell cycle control, cell division, chromosome partitioning; T, Signal transduction mechanisms; M, Cell wall/membrane/envelope biogenesis; Z, Cytoskeleton; W, Extracellular structures; U, Intracellular trafficking, secretion, and vesicular transport; O, Posttranslational modification, protein turnover, chaperones; C, Energy production and conversion; G, Carbohydrate transport and metabolism; E, Amino acid transport and metabolism; F, Nucleotide transport and metabolism; H, Coenzyme transport and metabolism; I, Lipid transport and metabolism; P, Inorganic ion transport and metabolism; Q, Secondary metabolites biosynthesis, transport and catabolism; and Multiple, for clusters assigned to more than one category.
Extended Data Fig. 5 Phylogenetic and structural analyses of iESPs.
a, Protein domain phylogeny based on Band 7, MVP and related shoulder domains. The depicted phylogenetic tree is based on 90 aligned positions and was generated under the LG+C60+R7 model (see Methods). b, ipTM score of Asgard archaeal MVP homopolymers modeled with different numbers of subunits with local optima highlighted. c, Multimer model of Lokiarchaeial MVP with different number of subunits. d, ipTM score of Asgard archaeal COMMD homopolymers modeled with different numbers of subunits with local optima highlighted. e, Homo-multimer model of Lokiarchaeial COMMD-containing protein with different number of subunits.
Supplementary information
Supplementary Information (download PDF )
Supplementary Discussion and Fig. 1.
Supplementary Data 1–3 (download XLSX )
Supplementary Data 1. Spread sheet with genome information of the outgroup genomes used for Extended Data Fig. 1a and the dataset of 936 Asgard archaeal draft genomes. Supplementary Data 2. Spread sheet of UniProt protein IDs and annotations of sampled Prometheoarchaeum syntrophicum. Supplementary Data 3. Spread sheet including the annotation of structures in the of ESPs and iESP structural clusters, as well as ESP and iESP proteins in Prometheoarchaeum syntrophicum.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Köstlbacher, S., van Hooff, J.J.E., Panagiotou, K. et al. Prediction of eukaryotic cellular complexity in Asgard archaea using structural modelling. Nat Microbiol 11, 747–758 (2026). https://doi.org/10.1038/s41564-026-02273-y
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41564-026-02273-y