Abstract
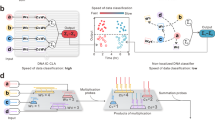
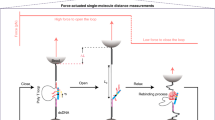
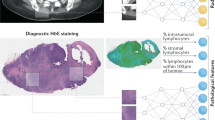
A molecular classification of diseases that accurately reflects clinical behaviour lays the foundation of precision medicine. The development of in silico classifiers coupled with molecular implementation based on DNA reactions marks a key advance in more powerful molecular classification, but it nevertheless remains a challenge to process multiple molecular datatypes. Here we introduce a DNA-encoded molecular classifier that can physically implement the computational classification of multidimensional molecular clinical data. To produce unified electrochemical sensing signals across heterogeneous molecular binding events, we exploit DNA-framework-based programmable atom-like nanoparticles with n valence to develop valence-encoded signal reporters that enable linearity in translating virtually any biomolecular binding events to signal gains. Multidimensional molecular information in computational classification is thus precisely assigned weights for bioanalysis. We demonstrate the implementation of a molecular classifier based on programmable atom-like nanoparticles to perform biomarker panel screening and analyse a panel of six biomarkers across three-dimensional datatypes for a near-deterministic molecular taxonomy of prostate cancer patients.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data that support the plots within this paper and other findings of this study are available from the corresponding author upon reasonable request. Furthermore, the miRNA, mRNA, PSA and SO data used in this study are available in ref. 47 and the National Center for Biotechnology Information database, https://www.ncbi.nlm.nih.gov/genome.
References
Collins, F. S. & Varmus, H. A new initiative on precision medicine. N. Engl. J. Med. 372, 793–795 (2015).
Thomasian, N. M., Kamel, I. R. & Bai, H. X. Machine intelligence in non-invasive endocrine cancer diagnostics. Nat. Rev. Endocrinol. 18, 81–95 (2022).
Vargas, A. J. & Harris, C. C. Biomarker development in the precision medicine era: lung cancer as a case study. Nat. Rev. Cancer 16, 525–537 (2016).
Nassiri, F. et al. Detection and discrimination of intracranial tumors using plasma cell-free DNA methylomes. Nat. Med. 26, 1044–1047 (2020).
Krzywinski, M. & Savig, E. Multidimensional data. Nat. Methods 10, 595 (2013).
Luo, Y. et al. A multidimensional precision medicine approach identifies an autism subtype characterized by dyslipidemia. Nat. Med. 26, 1375–1379 (2020).
Larance, M. & Lamond, A. I. Multidimensional proteomics for cell biology. Nat. Rev. Mol. Cell Biol. 16, 269–280 (2015).
Cohen, J. D. et al. Detection and localization of surgically resectable cancers with a multi-analyte blood test. Science 359, 926–930 (2018).
Berger, B., Peng, J. & Singh, M. Computational solutions for omics data. Nat. Rev. Genet. 14, 333–346 (2013).
Crichton, D. J. et al. Cancer biomarkers and big data: a planetary science approach. Cancer Cell 38, 757–760 (2020).
Liang, H. et al. Evaluation and accurate diagnoses of pediatric diseases using artificial intelligence. Nat. Med. 25, 433–438 (2019).
Kristensen, V. N. et al. Principles and methods of integrative genomic analyses in cancer. Nat. Rev. Cancer 14, 299–313 (2014).
Komori, T. The 2021 WHO classification of tumors, 5th edition, central nervous system tumors: the 10 basic principles. Brain Tumor Pathol. 39, 47–50 (2022).
Blanc, T., El Beheiry, M., Caporal, C., Masson, J. B. & Hajj, B. Genuage: visualize and analyze multidimensional single-molecule point cloud data in virtual reality. Nat. Methods 17, 1100–1102 (2020).
Adamcova, M. & Šimko, F. Multiplex biomarker approach to cardiovascular diseases. Acta Pharmacol. Sin. 39, 1068–1072 (2018).
Subramanian, I., Verma, S., Kumar, S., Jere, A. & Anamika, K. Multi-omics data integration, interpretation, and its application. Bioinf. Biol. Insights https://doi.org/10.1177/1177932219899051 (2020).
Montaner, J. et al. Multilevel omics for the discovery of biomarkers and therapeutic targets for stroke. Nat. Rev. Neurol. 16, 247–264 (2020).
Tarazona, S., Arzalluz-Luque, A. & Conesa, A. Undisclosed, unmet and neglected challenges in multi-omics studies. Nat. Comput. Sci. 1, 395–402 (2021).
Tarazona, S. et al. Harmonization of quality metrics and power calculation in multi-omic studies. Nat. Commun. 11, 3092 (2020).
Lopez de Maturana, E. et al. Challenges in the integration of omics and non-omics data. Genes 10, 238 (2019).
Benenson, Y., Gil, B., Ben-Dor, U., Adar, R. & Shapiro, E. An autonomous molecular computer for logical control of gene expression. Nature 429, 423–429 (2004).
Seelig, G., Soloveichik, D., Zhang, D. Y. & Winfree, E. Enzyme-free nucleic acid logic circuits. Science 314, 1585–1588 (2006).
Lopez, R., Wang, R. & Seelig, G. A molecular multi-gene classifier for disease diagnostics. Nat. Chem. 10, 746–754 (2018).
Zhang, C. et al. Cancer diagnosis with DNA molecular computation. Nat. Nanotechnol. 15, 709–715 (2020).
Yao, G. et al. Meta-DNA structures. Nat. Chem. 12, 1067–1075 (2020).
Yao, G. et al. Programming nanoparticle valence bonds with single-stranded DNA encoders. Nat. Mater. 19, 781–788 (2020).
Li, J. et al. Encoding quantized fluorescence states with fractal DNA frameworks. Nat. Commun. 11, 2185 (2020).
Wiraja, C. et al. Framework nucleic acids as programmable carrier for transdermal drug delivery. Nat. Commun. 10, 1147 (2019).
Zhang, T. et al. Design, fabrication and applications of tetrahedral DNA nanostructure-based multifunctional complexes in drug delivery and biomedical treatment. Nat. Protoc. 15, 2728–2757 (2020).
Song, P. et al. Programming bulk enzyme heterojunctions for biosensor development with tetrahedral DNA framework. Nat. Commun. 11, 838 (2020).
Lin, M. et al. Programmable engineering of a biosensing interface with tetrahedral DNA nanostructures for ultrasensitive DNA detection. Angew. Chem. Int. Ed. 54, 2151–2155 (2015).
Woehrstein, J. B. et al. 100-nm metafluorophores with digitally tunable optical properties self-assembled from DNA. Sci. Adv. 3, e1602128 (2017).
Ulbrich, M. H. & Isacoff, E. Y. Subunit counting in membrane-bound proteins. Nat. Methods 4, 319–321 (2007).
Hearty, S., Leonard, P. & O’Kennedy, R. Barcodes check out prostate cancer. Nat. Nanotechnol. 5, 9–10 (2010).
Hill, H. D. & Mirkin, C. A. The bio-barcode assay for the detection of protein and nucleic acid targets using DTT-induced ligand exchange. Nat. Protoc. 1, 324–336 (2006).
Nam, J.-M., Thaxton, C. S. & Mirkin, C. A. Nanoparticle-based bio-bar codes for the ultrasensitive detection of proteins. Science 301, 1884–1886 (2003).
Zebda, A. et al. Mediatorless high-power glucose biofuel cells based on compressed carbon nanotube-enzyme electrodes. Nat. Commun. 2, 370 (2011).
de Jong, O. G. et al. A CRISPR-Cas9-based reporter system for single-cell detection of extracellular vesicle-mediated functional transfer of RNA. Nat. Commun. 11, 1113 (2020).
Zhao, Z. et al. Nanocaged enzymes with enhanced catalytic activity and increased stability against protease digestion. Nat. Commun. 7, 10619 (2016).
He, L. et al. Transducing complex biomolecular interactions by temperature-output artificial DNA signaling networks. J. Am. Chem. Soc. 142, 14234–14239 (2020).
Li, H., Brouwer, C. R. & Luo, W. A universal deep neural network for in-depth cleaning of single-cell RNA-Seq data. Nat. Commun. 13, 1901 (2022).
Lin, M. et al. Electrochemical detection of nucleic acids, proteins, small molecules and cells using a DNA-nanostructure-based universal biosensing platform. Nat. Protoc. 11, 1244–1263 (2016).
Gorog, D. A. et al. Current and novel biomarkers of thrombotic risk in COVID-19: a Consensus Statement from the International COVID-19 Thrombosis Biomarkers Colloquium. Nat. Rev. Cardiol. 19, 475–495 (2022).
Schwarzenbach, H., Hoon, D. S. B. & Pantel, K. Cell-free nucleic acids as biomarkers in cancer patients. Nat. Rev. Cancer 11, 426–437 (2011).
Xiao, B. et al. Plasma microRNA panel is a novel biomarker for focal segmental glomerulosclerosis and associated with podocyte apoptosis. Cell Death Dis. 9, 533 (2018).
Bhanvadia, R. R. et al. MEIS1 and MEIS2 expression and prostate cancer progression: a role for HOXB13 binding partners in metastatic disease. Clin. Cancer Res. 24, 3668–3680 (2018).
Kumar, D., Gupta, A., Mandhani, A. & Sankhwar, S. N. Metabolomics-derived prostate cancer biomarkers: fact or fiction? J. Proteome Res. 14, 1455–1464 (2015).
Rajakumar, T. et al. A blood-based miRNA signature with prognostic value for overall survival in advanced stage non-small cell lung cancer treated with immunotherapy. npj Precis. Oncol. 6, 19 (2022).
Nassiri, F. et al. A clinically applicable integrative molecular classification of meningiomas. Nature 597, 119–125 (2021).
Li, F. et al. Ultrafast DNA sensors with DNA framework-bridged hybridization reactions. J. Am. Chem. Soc. 142, 9975–9981 (2020).
Acknowledgements
This work was financially supported by the National Natural Science Foundation of China (T2188102, 22025404, 22001168); National Key R&D Program of China (2021YFF1200300); China National Postdoctoral Program for Innovative Talents (BX2021190) by the China Postdoctoral Science Foundation; Innovative Research Team of High-Level Local Universities in Shanghai (SHSMU-ZLCX20212602); 2022 Shanghai ‘Science and Technology Innovation Action Plan’ Fundamental Research Project (22JC1401202); Shanghai Jiao Tong University Scientific and Technological Innovation Funds (21X010202096) and Shanghai Municipal Health Commission (2022JC027).
Author information
Authors and Affiliations
Contributions
X.Z., C.F. and F.Y. conceived the study. F.Y., H.Z. and S.L. performed the experiments. F.Y performed the TIRFM imaging and nucleic acid information translation. H.Z. performed the TEM imaging and SO information translation. S.L. performed the AFM imaging and PSA information translation. J. Shen performed the target screen and data training. B.D. and W.X. provided samples and analysed the clinical data. F.Y., H.Z. and S.L. performed the clinical sample detection. F.Y., J. Shi, M.L., X.M., F.L. and J.L. carried out the assays and analysed the results. X.Z. and C.F. directed the research. X.Z., C.F. and F.Y. wrote the paper. X.Z., C.F. and X.Y. supervised the project. All authors read the paper and provided comments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Nanotechnology thanks Hao Yan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–41, Tables 1–23, Discussion, Notes and References.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yin, F., Zhao, H., Lu, S. et al. DNA-framework-based multidimensional molecular classifiers for cancer diagnosis. Nat. Nanotechnol. 18, 677–686 (2023). https://doi.org/10.1038/s41565-023-01348-9
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41565-023-01348-9
This article is cited by
-
Enabling global-scale nucleic acid repositories through versatile, scalable biochemical selection from room-temperature archives
Nature Communications (2026)
-
Polymerase-based DNA reactions for molecularly computing cancerous diagnostic valences of multiple miRNAs
Journal of Nanobiotechnology (2025)
-
Hierarchical core–shell DNA condensates enable programmable information storage and encryption
Nature Communications (2025)
-
A biochemical sensor with continuous extended stability in vivo
Nature Biomedical Engineering (2025)
-
DNA-based biocomputing circuits and their biomedical applications
Nature Reviews Bioengineering (2025)