Abstract
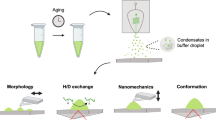
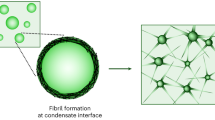
Biomolecular condensates regulate cellular physiology by sequestering and processing RNAs and proteins, yet how these processes are locally tuned within condensates remains unclear. Moreover, in neurodegenerative diseases such as amyotrophic lateral sclerosis, condensates undergo liquid-to-solid phase transitions, but capturing early intermediates in this process has been challenging. Here we present a surface multi-tethering approach to achieve intra-condensate single-molecule tracking of fluorescently labelled RNA and protein molecules within liquid-like condensates. Using RNA-binding protein fused-in-sarcoma as a model for condensates implicated in amyotrophic lateral sclerosis, we discover that RNA and protein diffusion is confined within distinct nanometre-scale domains, or nanodomains, which exhibit unique connectivity and chemical environments. The properties of these nanodomains are tunable by guest molecules. As condensates age, nanodomains reposition, facilitating fused-in-sarcoma fibrilization at the condensate surface, a process further enhanced by anti-amyotrophic lateral sclerosis drugs. Our findings demonstrate that nanodomain formation governs condensate function by modulating the residence time and spatial organization of constituent biomolecules, providing previously unattainable insights into condensate ageing and mechanisms underlying disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All the data that support the findings of this study are available within the article and its Supplementary Information. Source data are available at the Deep Blue Data Repository: https://doi.org/10.7302/eq18-p391.
Code availability
The source code developed for the data analysis is available via GitHub at https://github.com/walterlab-um/intra_condensate_SPT (ref. 62).
References
Mittag, T. & Pappu, R. V. A conceptual framework for understanding phase separation and addressing open questions and challenges. Mol. Cell 82, 2201–2214 (2022).
Pappu, R. V., Cohen, S. R., Dar, F., Farag, M. & Kar, M. Phase transitions of associative biomacromolecules. Chem. Rev. 123, 8945–8987 (2023).
Lyon, A. S., Peeples, W. B. & Rosen, M. K. A framework for understanding the functions of biomolecular condensates across scales. Nat. Rev. Mol. Cell Biol. 22, 215–235 (2021).
Alberti, S. & Hyman, A. A. Biomolecular condensates at the nexus of cellular stress, protein aggregation disease and ageing. Nat. Rev. Mol. Cell Biol. 22, 196–213 (2021).
Roden, C. & Gladfelter, A. S. RNA contributions to the form and function of biomolecular condensates. Nat. Rev. Mol. Cell Biol. 22, 183–195 (2021).
Ditlev, J. A., Case, L. B. & Rosen, M. K. Who’s in and who’s out—compositional control of biomolecular condensates. J. Mol. Biol. 430, 4666–4684 (2018).
Banani, S. F., Lee, H. O., Hyman, A. A. & Rosen, M. K. Biomolecular condensates: organizers of cellular biochemistry. Nat. Rev. Mol. Cell Biol. 18, 285–298 (2017).
Peeples, W. & Rosen, M. K. Mechanistic dissection of increased enzymatic rate in a phase-separated compartment. Nat. Chem. Biol. 17, 693–702 (2021).
Collins, M. J., Tomares, D. T., Nandana, V., Schrader, J. M. & Childers, W. S. RNase E biomolecular condensates stimulate PNPase activity. Sci. Rep. 13, 12937 (2023).
Portz, B., Lee, B. L. & Shorter, J. FUS and TDP-43 phases in health and disease. Trends Biochem. Sci. 46, 550–563 (2021).
Zuo, L. et al. Loci-specific phase separation of FET fusion oncoproteins promotes gene transcription. Nat. Commun. 12, 1491 (2021).
Levone, B. R. et al. FUS-dependent liquid-liquid phase separation is important for DNA repair initiation. J. Cell Biol. 220, e202008030 (2021).
Patel, A. et al. A liquid-to-solid phase transition of the ALS protein FUS accelerated by disease mutation. Cell 162, 1066–1077 (2015).
Wang, W. Y. et al. Interaction of FUS and HDAC1 regulates DNA damage response and repair in neurons. Nat. Neurosci. 16, 1383–1391 (2013).
Ishiguro, A., Katayama, A. & Ishihama, A. Different recognition modes of G-quadruplex RNA between two ALS/FTLD-linked proteins TDP-43 and FUS. FEBS Lett. 595, 310–323 (2021).
Ishiguro, A., Lu, J., Ozawa, D., Nagai, Y. & Ishihama, A. ALS-linked FUS mutations dysregulate G-quadruplex-dependent liquid-liquid phase separation and liquid-to-solid transition. J. Biol. Chem. 297, 101284 (2021).
Kaur, T. et al. Molecular crowding tunes material states of ribonucleoprotein condensates. Biomolecules 9, 71 (2019).
Sumrall, E. R., Gao, G., Stakenas, S. & Walter, N. G. Surface-tethering enhances precision in measuring diffusion within 3D protein condensates. J. Mol. Biol. https://doi.org/10.1016/j.jmb.2025.169447 (2025).
Wu, T. et al. Single-fluorogen imaging reveals distinct environmental and structural features of biomolecular condensates. Nat. Phys. 21, 778–786 (2025).
Burnecki, K., Kepten, E., Garini, Y., Sikora, G. & Weron, A. Estimating the anomalous diffusion exponent for single particle tracking data with measurement errors—an alternative approach. Sci. Rep. 5, 11306 (2015).
Heckert, A., Dahal, L., Tjian, R. & Darzacq, X. Recovering mixtures of fast-diffusing states from short single-particle trajectories. Elife 11, e70169 (2022).
Gopal, A., Zhou, Z. H., Knobler, C. M. & Gelbart, W. M. Visualizing large RNA molecules in solution. RNA 18, 284–299 (2012).
Shen, Z. et al. Biological condensates form percolated networks with molecular motion properties distinctly different from dilute solutions. Elife 12, e81907 (2023).
Kamagata, K., Kusano, R., Kanbayashi, S., Banerjee, T. & Takahashi, H. Single-molecule characterization of target search dynamics of DNA-binding proteins in DNA-condensed droplets. Nucleic Acids Res. 51, 6654–6667 (2023).
Kamagata, K. et al. Structure-dependent recruitment and diffusion of guest proteins in liquid droplets of FUS. Sci. Rep. 12, 7101 (2022).
Kamagata, K. et al. Molecular principles of recruitment and dynamics of guest proteins in liquid droplets. Sci. Rep. 11, 19323 (2021).
Todorov, T. I., de Carmejane, O., Walter, N. G. & Morris, M. D. Capillary electrophoresis of RNA in dilute and semidilute polymer solutions. Electrophoresis 22, 2442–2447 (2001).
Holehouse, A. S., Das, R. K., Ahad, J. N., Richardson, M. O. G. & Pappu, R. V. CIDER: resources to analyze sequence-ensemble relationships of intrinsically disordered proteins. Biophys. J. 112, 16–21 (2017).
Manzo, C. & Garcia-Parajo, M. F. A review of progress in single particle tracking: from methods to biophysical insights. Rep. Prog. Phys. 78, 124601 (2015).
Lai, W.-J. C. et al. mRNAs and lncRNAs intrinsically form secondary structures with short end-to-end distances. Nat. Commun. 9, 4328 (2018).
Emmanouilidis, L. et al. A solid beta-sheet structure is formed at the surface of FUS droplets during aging. Nat. Chem. Biol. 20, 1044–1052 (2024).
He, C., Wu, C. Y., Li, W. & Xu, K. Multidimensional super-resolution microscopy unveils nanoscale surface aggregates in the aging of FUS condensates. J. Am. Chem. Soc. 145, 24240–24248 (2023).
Shen, Y. et al. The liquid-to-solid transition of FUS is promoted by the condensate surface. Proc. Natl Acad. Sci. USA 120, e2301366120 (2023).
Ausserwöger, H. et al. Biomolecular condensates sustain pH gradients at equilibrium through charge neutralisation. Preprint at bioRxiv https://doi.org/10.1101/2024.05.23.595321 (2024).
Hoffmann, C. et al. Electric potential at the interface of membraneless organelles gauged by graphene. Nano Lett. 23, 10796–10801 (2023).
Linsenmeier, M. et al. The interface of condensates of the hnRNPA1 low-complexity domain promotes formation of amyloid fibrils. Nat. Chem. 15, 1340–1349 (2023).
Deng, H., Gao, K. & Jankovic, J. The role of FUS gene variants in neurodegenerative diseases. Nat. Rev. Neurol. 10, 337–348 (2014).
Cruz, M. P. Edaravone (Radicava): a novel neuroprotective agent for the treatment of amyotrophic lateral sclerosis. P. T. 43, 25–28 (2018).
Albo, F., Pieri, M. & Zona, C. Modulation of AMPA receptors in spinal motor neurons by the neuroprotective agent riluzole. J. Neurosci. Res. 78, 200–207 (2004).
Ambadi Thody, S. et al. Small-molecule properties define partitioning into biomolecular condensates. Nat. Chem. 16, 1794–1802 (2024).
Tischbein, M. et al. The RNA-binding protein FUS/TLS undergoes calcium-mediated nuclear egress during excitotoxic stress and is required for GRIA2 mRNA processing. J. Biol. Chem. 294, 10194–10210 (2019).
Cataldi, R. et al. A dopamine metabolite stabilizes neurotoxic amyloid-β oligomers. Commun. Biol. 4, 19 (2021).
Chen, S. W. et al. Structure–toxicity relationship in intermediate fibrils from α-synuclein condensates. J. Am. Chem. Soc. 146, 10537–10549 (2024).
Eisenberg, D., Schwarz, E., Komaromy, M. & Wall, R. Analysis of membrane and surface protein sequences with the hydrophobic moment plot. J. Mol. Biol. 179, 125–142 (1984).
Gao, G. & Walter, N. G. Critical assessment of condensate boundaries in dual-color single particle tracking. J. Phys. Chem. B 127, 7694–7707 (2023).
Galvanetto, N. et al. Extreme dynamics in a biomolecular condensate. Nature 619, 876–883 (2023).
Custer, T. C. & Walter, N. G. In vitro labeling strategies for in cellulo fluorescence microscopy of single ribonucleoprotein machines. Protein Sci. 26, 1363–1379 (2017).
Schmidt, A., Gao, G., Little, S. R., Jalihal, A. P. & Walter, N. G. Following the messenger: recent innovations in live cell single molecule fluorescence imaging. Wiley Interdiscip. Rev. RNA 11, e1587 (2020).
Brito Querido, J. et al. Structure of a human 48S translational initiation complex. Science 369, 1220–1227 (2020).
Lin, Y., Protter, D. S. W., Rosen, M. K. & Parker, R. Formation and maturation of phase-separated liquid droplets by RNA-binding proteins. Mol. Cell 60, 208–219 (2015).
Yin, J. & Chen, X. Edaravone prevents high glucose-induced injury in retinal Müller cells through thioredoxin1 and the PGC-1α/NRF1/TFAM pathway. Pharm. Biol. 59, 1231–1242 (2021).
Sala, G. et al. Riluzole selective antioxidant effects in cell models expressing amyotrophic lateral sclerosis endophenotypes. Clin. Psychopharmacol. Neurosci. 17, 438–442 (2019).
Johnson-Buck, A. et al. Kinetic fingerprinting to identify and count single nucleic acids. Nat. Biotechnol. 33, 730–732 (2015).
Tokunaga, M., Imamoto, N. & Sakata-Sogawa, K. Highly inclined thin illumination enables clear single-molecule imaging in cells. Nat. Methods 5, 159–161 (2008).
Evangelidis, G. D. & Psarakis, E. Z. Parametric image alignment using enhanced correlation coefficient maximization. IEEE Trans. Pattern Anal. Mach. Intell. 30, 1858–1865 (2008).
Tinevez, J. Y. et al. TrackMate: an open and extensible platform for single-particle tracking. Methods 115, 80–90 (2017).
Michalet, X. & Berglund, A. J. Optimal diffusion coefficient estimation in single-particle tracking. Phys. Rev. E 85, 061916 (2012).
Berglund, A. J. Statistics of camera-based single-particle tracking. Phys. Rev. E 82, 011917 (2010).
Warren, S. C. et al. Rapid global fitting of large fluorescence lifetime imaging microscopy datasets. PLoS ONE 8, e70687 (2013).
Enderlein, J. & Erdmann, R. Fast fitting of multi-exponential decay curves. Opt. Commun. 134, 371–378 (1997).
Kohler, J., Hur, K. H. & Mueller, J. D. Autocorrelation function of finite-length data in fluorescence correlation spectroscopy. Biophys. J. 122, 241–253 (2023).
Gao, G. et al. intra_condensate_SPT. GitHub https://github.com/walterlab-um/intra_condensate_SPT (2025).
Acknowledgements
We sincerely thank S. Ray, Z. Chen, A. Decker, N. Rogers and X. Dai for their insightful discussions on developing the analysis pipelines for our SMT datasets. We appreciate help from D. Hoff at the Single Molecule Analysis in Real-Time (SMART) Center of Biophysics at the University of Michigan, for FLIM and FCS measurements and analysis, and E. Akbari and D. Pelekhov at the Nano-Systems Laboratory (NSL) of the Physics Department at the Ohio State University, for their help in using a LUMICKS C-Trap for condensate microrheology measurements. We much appreciate the invaluable feedback and proofreading efforts by Z. Chen, A. Chauvier, M. Jin, A. Johnson-Buck, S. Ma, Y. Zhu, and the University of Michigan U-M GPT as well as ChatGPT. We also thank L. Dai for his early joint efforts on protein purifications and M. Hijaz for efforts on quantifying the translatability of FL mRNAs. N.G.W. acknowledges funding from NIH grant R35 GM131922, a sub-award of NIH grant R01 NS097542, and Chan Zuckerberg Initiative (CZI) grant 2022-250725; whereas E.R.S. is thankful for an NSF GRFP fellowship DGE2241144.
Author information
Authors and Affiliations
Contributions
G.G.: conceptualization, investigation, formal analysis, methodology, software, visualization and writing (original draft). E.R.S.: conceptualization, investigation, formal analysis, methodology, software, visualization and writing (original draft). N.G.W.: conceptualization, funding acquisition, methodology, project administration, supervision and writing (final draft).
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Nanotechnology thanks Dragomir Milovanovic and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Representative SMT trajectory reconstruction images of FUS and FL mRNA.
Representative reconstruction images of dual-color SMT-trajectory localization data from FUS (cyan) and FL mRNA (red). Color bars are in the unit of number of single-molecule localizations per pixel. All scale bars are 1 µm.
Extended Data Fig. 2 Representative SMT trajectory reconstruction images of FUS and miRNA-21.
Representative reconstruction images of dual-color SMT-trajectory localization data from FUS (cyan) and miRNA (red). Color bars are in the unit of number of single-molecule localizations per pixel. All scale bars are 1 µm.
Extended Data Fig. 3 Nanodomains exhibit extremely slow dynamics while providing distinct chemical environments as measured by fluorescence lifetimes.
a | Time-lapse SMT-PAINT reconstructions of FUS nanodomains within the indicated zoomed-in regions. Diffusive motions of the underlying nanodomains reach a smaller localization error (\({\sigma }_{{nanodmain}}=\frac{{\sigma }_{{SMLM}}}{\sqrt{{N}_{{trajectories}}}}\)) than SMT trajectories (\({\sigma }_{{SMLM}}=\frac{{\sigma }_{{diffraction}}}{\sqrt{{N}_{{photon}}}}\)) and are depicted by orange trajectories, noted with the velocity calculated from their mean step sizes. b | Fluorescence lifetime imaging (FLIM) of Alexa Fluor 488-labeled FUS. All scale bars are 1 µm.
Extended Data Fig. 4 Effect of aging and macromolecular crowding on the intra-condensate diffusion profile of mRNA molecules.
a │ Fractions of the three types of SMT trajectories, immobile (I.), confined (C.), and normal (N.) diffusion, of mRNA molecules within FUS condensates at varying times after reconstitution of FUS condensates with or without the crowding reagent Dextran T-500 at a final concentration of 10% (w/v). Error bars are SEM from at least three biological replicates. b │ Distribution of Dapp calculated from normal diffusion trajectories by SA analysis, comparing the impact of aging with or without macromolecular crowding from Dextran T-500 as indicated. Vertical dotted lines indicate the peak positions.
Extended Data Fig. 5 Effect of total RNA background and macromolecular crowding on the intra-condensate diffusion profile of mRNA molecules.
a │ Fractions of the three types of SMT trajectories, immobile (I.), confined (C.), and normal (N.) diffusion, of mRNA molecules within FUS condensates with or without 50 ng/μL HeLa cell total RNA with or without the crowding reagent Dextran T-500 at a final concentration of 10% (w/v). Error bars are SEM from at least three biological replicates. b │ Distribution of Dapp calculated from normal diffusion trajectories by SA analysis, comparing the impact of total RNA background with or without macromolecular crowding from Dextran T-500 as indicated. Vertical dotted lines indicate the peak positions.
Extended Data Fig. 6 Bright field images of condensates in a representative FOV undergoing aging.
Four fields of view shown for FUS condensates over 24 h. For each FOV, condensates that are liquid at time point 0 h are observed to have surface fibril features at 24 h.
Extended Data Fig. 7 Small molecule drugs impact condensate aging.
FUS condensates under different small molecule drug treatment imaged in bright field at the 24-h time point. All scale bars are 1 µm.
Extended Data Fig. 8 Translational and rotational motions of untethered condensates.
a-b│ Condensates can undergo coalescence (a) and translational (arrows in a) or rotational (arrows in b) motions in the absence of FUS-biotin molecular tethers. The position of condensate (b) and the angular position of three RNA molecules (b) in the current and previous frame are shown in white and yellow respectively. All scale bars are 1 µm.
Supplementary information
Supplementary Information (download PDF )
Supplementary Discussion, Table 1, Figs. 1–19 and Methods.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gao, G., Sumrall, E.R. & Walter, N.G. Nanoscale domains govern local diffusion and ageing within fused-in-sarcoma condensates. Nat. Nanotechnol. 21, 249–258 (2026). https://doi.org/10.1038/s41565-025-02077-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41565-025-02077-x