Abstract
Acute lymphoblastic leukaemia (ALL) is the most common cancer of childhood. Here, we map emerging evidence suggesting that children with ALL at the time of diagnosis may have a delayed maturation of the gut microbiome compared with healthy children. This finding may be associated with early-life epidemiological factors previously identified as risk indicators for childhood ALL, including caesarean section birth, diminished breast feeding and paucity of social contacts. The consistently observed deficiency in short-chain fatty-acid-producing bacterial taxa in children with ALL has the potential to promote dysregulated immune responses and to, ultimately, increase the risk of transformation of preleukaemic clones in response to common infectious triggers. These data endorse the concept that a microbiome deficit in early life may contribute to the development of the major subtypes of childhood ALL and encourage the notion of risk-reducing microbiome-targeted intervention in the future.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
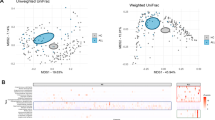
The primary data that support the findings presented in this Perspective article, including the results of our re-analysis of participant-level data of the study of Liu et al.71, are available as Supplementary figures.
References
Public Health England. Children, teenagers and young adults UK cancer statistics report 2021 1–30 (2021).
Pui, C. H. & Evans, W. E. A 50-year journey to cure childhood acute lymphoblastic leukemia. Semin. Hematol. 50, 185–196 (2013).
Kane, E. et al. Excess morbidity and mortality among survivors of childhood acute lymphoblastic leukaemia: 25 years of follow-up from the United Kingdom Childhood Cancer Study (UKCCS) population-based matched cohort. BMJ Open 12, e056216 (2022).
Greaves, M., Cazzaniga, V. & Ford, A. Can we prevent childhood leukaemia? Leukemia 35, 1258–1264 (2021).
Hauer, J., Fischer, U. & Borkhardt, A. Toward prevention of childhood ALL by early-life immune training. Blood 138, 1412–1428 (2021).
Greaves, M. F. Speculations on the cause of childhood acute lymphoblastic leukemia. Leukemia 2, 120–125 (1988).
Greaves, M. A causal mechanism for childhood acute lymphoblastic leukaemia. Nat. Rev. Cancer 18, 471–484 (2018).
Bach, J. The effect of infections on susceptibility to autoimmune and allergic diseases. N. Engl. J. Med. 347, 911–920 (2002).
Ford, A. M., Colman, S. & Greaves, M. Covert pre-leukaemic clones in healthy co-twins of patients with childhood acute lymphoblastic leukaemia. Leukemia 37, 47–52 (2023).
Wiemels, J. L. et al. Prenatal origin of acute lymphoblastic leukaemia in children. Lancet 354, 1499–1503 (1999).
Maia, A. T. et al. Prenatal origin of hyperdiploid acute lymphoblastic leukemia in identical twins. Leukemia 17, 2202–2206 (2003).
Taub, J. W. et al. High frequency of leukemic clones in newborn screening blood samples of children with B-precursor acute lymphoblastic leukemia. Blood 99, 2992–2996 (2002).
Swaminathan, S. et al. Mechanisms of clonal evolution in childhood acute lymphoblastic leukemia. Nat. Immunol. 16, 766–774 (2015).
Mori, H. et al. Chromosome translocations and covert leukemic clones are generated during normal fetal development. Proc. Natl Acad. Sci. USA 99, 8242–8247 (2002).
Tsuzuki, S., Seto, M., Greaves, M. & Enver, T. Modeling first-hit functions of the t(12;21) TEL-AML1 translocation in mice. Proc. Natl Acad. Sci. USA 22, 8443–8448 (2004).
Fidanza, M. et al. Inhibition of precursor B cell malignancy progression by toll-like receptor ligand-induced immune responses. Leukemia 30, 2116–2119 (2016).
Schäfer, D. et al. Five percent of healthy newborns have an ETV6-RUNX1 fusion as revealed by DNA-based GIPFEL screening. Blood 131, 821–826 (2018).
Francis, S. S., Selvin, S., Yang, W., Buffler, P. A. & Wiemels, J. L. Unusual space–time patterning of the Fallon, Nevada leukemia cluster: evidence of an infectious etiology. Chem. Biol. Interact. 196, 102–109 (2012).
Cazzaniga, G. et al. Possible role of pandemic AH1N1 swine flu virus in a childhood leukemia cluster. Leukemia 31, 1819–1821 (2017).
Papaemmanuil, E. et al. RAG-mediated recombination is the predominant driver of oncogenic rearrangement in ETV6-RUNX1 acute lymphoblastic leukemia. Nat. Genet. 46, 116–125 (2014).
Marcotte, E. L. et al. Caesarean delivery and risk of childhood leukaemia: a pooled analysis from the Childhood Leukemia International Consortium (CLIC). Lancet Haematol. 3, e176–e185 (2016).
Sevelsted, A., Stokholm, J., Bønnelykke, K. & Bisgaard, H. Cesarean section chronic immune disorders. Pediatrics 135, e92–e98 (2015).
Amitay, E. L. & Keinan-Boker, L. Breastfeeding and childhood leukemia incidence: a meta-analysis and systematic review. JAMA Pediatr. 169, e151025 (2015).
Su, Q. et al. Breastfeeding and the risk of childhood cancer: a systematic review and dose-response meta-analysis. BMC Med. 19, 90 (2021).
Rudant, J. et al. Childhood acute lymphoblastic leukemia and indicators of early immune stimulation: a Childhood Leukemia International Consortium study. Am. J. Epidemiol. 181, 549–562 (2015).
Urayama, K. Y., Buffler, P. A., Gallagher, E. R., Ayoob, J. M. & Ma, X. A meta-analysis of the association between day-care attendance and childhood acute lymphoblastic leukaemia. Int. J. Epidemiol. 39, 718–732 (2010).
Kamper-Jørgensen, M. et al. Childcare in the first 2 years of life reduces the risk of childhood acute lymphoblastic leukemia. Leukemia 22, 189–193 (2008).
Shao, Y. et al. Stunted microbiota and opportunistic pathogen colonization in caesarean-section birth. Nature 574, 117–121 (2019).
Reyman, M. et al. Impact of delivery mode-associated gut microbiota dynamics on health in the first year of life. Nat. Commun. 10, 4997 (2019).
Stewart, C. J. et al. Temporal development of the gut microbiome in early childhood from the TEDDY study. Nature 562, 583–588 (2018).
Amir, A., Erez-granat, O., Braun, T., Sosnovski, K. & Hadar, R. Gut microbiome development in early childhood is affected by day care attendance. NPJ Biofilms Microbiomes 8, 2 (2022).
Olin, A. et al. Stereotypic immune system development in newborn children. Cell 174, 1277–1292.e14 (2018).
Stokholm, J. et al. Maturation of the gut microbiome and risk of asthma in childhood. Nat. Commun. 9, 141 (2018).
Depner, M. et al. Maturation of the gut microbiome during the first year of life contributes to the protective farm effect on childhood asthma. Nat. Med. 26, 1766–1775 (2020).
Stokholm, J. et al. Delivery mode and gut microbial changes correlate with an increased risk of childhood asthma. Sci. Transl. Med. 12, eaax9929 (2020).
Wen, Y., Jin, R. & Chen, H. Interactions between gut microbiota and acute childhood leukemia. Front. Microbiol. 10, 1300 (2019).
Cobaleda, C., Vicente-Duenas, C. & Sanchez-Garcia, I. An immune window of opportunity to prevent childhood B cell leukemia. Trends Immunol. 42, 371–374 (2021).
Ma, T., Chen, Y., Li, L.-J. & Zhang, L.-S. Opportunities and challenges for gut microbiota in acute leukemia. Front. Oncol. 11, 692951 (2021).
Uribe-Herranz, M., Klein-González, N., Rodríguez-Lobato, L. G., Juan, M. & de Larrea, C. F. Gut microbiota influence in hematological malignancies: from genesis to cure. Int. J. Mol. Sci. 22, 1026 (2021).
Oldenburg, M., Rüchel, N., Janssen, S., Borkhardt, A. & Gössling, K. L. The microbiome in childhood acute lymphoblastic leukemia. Cancers 13, 4947 (2021).
Masetti, R. et al. Gut microbiome in pediatric acute leukemia: from predisposition to cure. Blood Adv. 5, 4619–4629 (2021).
Arrieta, M. et al. The intestinal microbiome in early life: health and disease. Front. Immunol. 5, 427 (2014).
Derrien, M., Alvarez, A. S. & de Vos, W. M. The gut microbiota in the first decade of life. Trends Microbiol. 27, 997–1010 (2019).
Ferretti, P. et al. Mother-to-infant microbial transmission from different body sites shapes the developing infant gut microbiome. Cell Host Microbe 24, 133–145.e5 (2018).
Kirmiz, N., Robinson, R. C., Shah, I. M., Barile, D. & Mills, D. A. Milk glycans and their interaction with the infant-gut microbiota. Annu. Rev. Food Sci. Technol. 9, 429–450 (2018).
Bokulich, N. A. et al. Antibiotics, birth mode, and diet shape microbiome maturation during early life. Sci. Transl. Med. 8, 343ra82 (2016).
Matsuyama, M. et al. Breastfeeding: a key modulator of gut microbiota characteristics in late infancy. J. Dev. Orig. Health Dis. 10, 206–213 (2019).
Tsukuda, N. et al. Key bacterial taxa and metabolic pathways affecting gut short-chain fatty acid profiles in early life. ISME J. 15, 2574–2590 (2021).
Roswall, J. et al. Developmental trajectory of the healthy human gut microbiota during the first 5 years of life. Cell Host Microbe 29, 765–776.e3 (2021).
Odamaki, T. et al. Age-related changes in gut microbiota composition from newborn to centenarian: a cross-sectional study. BMC Microbiol. 16, 90 (2016).
Xiao, L., Wang, J., Zheng, J., Li, X. & Zhao, F. Deterministic transition of enterotypes shapes the infant gut microbiome at an early age. Genome Biol. 22, 243 (2021).
Hildebrand, F. et al. Dispersal strategies shape persistence and evolution of human gut bacteria. Cell Host Microbe 29, 1167–1176.e9 (2021).
Beller, L. et al. Successional stages in infant gut microbiota maturation. mBio 12, e01857-21 (2021).
Cox, L. M. et al. Altering the intestinal microbiota during a critical developmental window has lasting metabolic consequences. Cell 158, 705–721 (2014).
Mitchell, C. M. et al. Delivery mode affects stability of early infant gut microbiota. Cell Rep. Med. 1, 100156 (2020).
Niu, J. et al. Evolution of the gut microbiome in early childhood: a cross-sectional study of Chinese children. Front. Microbiol. 11, 439 (2020).
Azad, M. B. et al. Impact of maternal intrapartum antibiotics, method of birth and breastfeeding on gut microbiota during the first year of life: a prospective cohort study. Br. J. Obstetr. Gynecol. 123, 983–993 (2016).
Martin, R. et al. Early-life events, including mode of delivery and type of feeding, siblings and gender, shape the developing gut microbiota. PLoS ONE 11, e0158498 (2016).
Ho, N. T. et al. Meta-analysis of effects of exclusive breastfeeding on infant gut microbiota across populations. Nat. Commun. 9, 4169 (2018).
Bridgman, S. L. et al. Fecal short-chain fatty acid variations by breastfeeding status in infants at 4 months: differences in relative versus absolute concentrations. Front. Nutr. 4, 00011 (2017).
Bäckhed, F. et al. Dynamics and stabilization of the human gut microbiome during the first year of life. Cell Host Microbe 17, 690–703 (2015).
Nogacka, A. et al. Impact of intrapartum antimicrobial prophylaxis upon the intestinal microbiota and the prevalence of antibiotic resistance genes in vaginally delivered full-term neonates. Microbiome 5, 93 (2017).
Prescott, S. et al. Impact of intrapartum antibiotic prophylaxis on offspring microbiota. Front. Pediatr. 9, 754013 (2021).
Yassour, M. et al. Natural history of the infant gut microbiome and impact of antibiotic treatments on strain-level diversity and stability. Sci. Transl. Med. 8, 1173–1178 (2016).
McDonnell, L. et al. Association between antibiotics and gut microbiome dysbiosis in children: systematic review and meta-analysis. Gut Microbes 13, 1870402 (2021).
Laursen, M. F. et al. Having older siblings is associated with gut microbiota development during early childhood. BMC Microbiol. 15, 154 (2015).
De Filippo, C. et al. Diet, environments, and gut microbiota. A preliminary investigation in children living in rural and urban Burkina Faso and Italy. Front. Microbiol. 8, 1979 (2017).
Borbet, T. C. et al. Influence of the early-life gut microbiota on the immune responses to an inhaled allergen. Mucosal Immunol. 15, 1000–1011 (2022).
Hakim, H. et al. Gut microbiome composition predicts infection risk during chemotherapy in children with acute lymphoblastic leukemia. Clin. Infect. Dis. 67, 541–548 (2018).
Liu, X. et al. Pediatric acute lymphoblastic leukemia patients exhibit distinctive alterations in the gut microbiota. Front. Cell Infect. Microbiol. 10, 558799 (2020).
de Pietri, S. et al. Gastrointestinal toxicity during induction treatment for childhood acute lymphoblastic leukemia: the impact of the gut microbiota. Int. J. Cancer 147, 1953–1962 (2020).
Gao, X. et al. A new insight into acute lymphoblastic leukemia in children: influences of changed intestinal microfloras. BMC Pediatr. 20, 290 (2020).
Rajagopala, S. V. et al. Persistent gut microbial dysbiosis in children with acute lymphoblastic leukemia (ALL) during chemotherapy. Microb. Ecol. 79, 1034–1043 (2020).
Bai, L., Zhou, P., Li, D. & Ju, X. Changes in the gastrointestinal microbiota of children with acute lymphoblastic leukaemia and its association with antibiotics in the short term. J. Med. Microbiol. 66, 1297–1307 (2017).
Chua, L. L. et al. Temporal changes in gut microbiota profile in children with acute lymphoblastic leukemia prior to commencement-, during-, and post-cessation of chemotherapy. BMC Cancer 20, 151 (2020).
Koh, A., De Vadder, F., Kovatcheva-Datchary, P. & Bäckhed, F. From dietary fiber to host physiology: short-chain fatty acids as key bacterial metabolites. Cell 165, 1332–1345 (2016).
Kim, M. & Kim, C. H. Regulation of humoral immunity by gut microbial products. Gut Microbes 8, 392–399 (2017).
Yu, B., Wang, L. & Chu, Y. Gut microbiota shape B cell in health and disease settings. J. Leukoc. Biol. 110, 271–281 (2021).
Dunn, K. A. et al. Antibiotic and antifungal use in pediatric leukemia and lymphoma patients are associated with increasing opportunistic pathogens and decreasing bacteria responsible for activities that enhance colonic defense. Front. Cell. Infect. Microbiol. 12, 924707 (2022).
Gensollen, T., Iyer, S. S., Kasper, D. L., Blumberg, R. S. & Medical, H. How colonization by microbiota in early life shapes the immune system. Science 352, 539–544 (2016).
Zhang, X., Zhivaki, D. & Lo-Man, R. Unique aspects of the perinatal immune system. Nat. Rev. Immunol. 17, 495–507 (2017).
Rooks, M. G. & Garrett, W. S. Gut microbiota, metabolites and host immunity. Nat. Rev. Immunol. 16, 341–352 (2016).
Planer, J. D. et al. Development of the gut microbiota and mucosal IgA responses in twins and gnotobiotic mice. Nature 534, 263–266 (2016).
Turroni, F. et al. The infant gut microbiome as a microbial organ influencing host well-being. Ital. J. Pediatr. 46, 16 (2020).
Li, H. et al. Mucosal or systemic microbiota exposures shape the B cell repertoire. Nature 584, 274–278 (2020).
New, J. S. et al. Neonatal exposure to commensal-bacteria-derived antigens directs polysaccharide-specific B-1 B cell repertoire development. Immunity 53, 172–186.e6 (2020).
Chen, H. et al. BCR selection and affinity maturation in Peyer’s patch germinal centres. Nature 582, 421–425 (2020).
Wesemann, D. R. et al. Microbial colonization influences early B-lineage development in the gut lamina propria. Nature 501, 112–115 (2013).
Hapfelmeier, S. et al. Reversible microbial colonization of germ-free mice reveals the dynamics of IgA immune responses. Science 328, 1705–1709 (2010).
Sefik, E. et al. Individual intestinal symbionts induce a distinct population of RORγ+ regulatory T cells. Science 349, 993–997 (2015).
Cahenzli, J., Köller, Y., Wyss, M., Geuking, M. B. & McCoy, K. D. Intestinal microbial diversity during early-life colonization shapes long-term IgE levels. Cell Host Microbe 14, 559–570 (2013).
Round, J. L. et al. The Toll-like receptor 2 pathway establishes colonization by a commensal of the human microbiota. Science 332, 974–977 (2011).
Yang, C. et al. Fecal IgA levels are determined by strain-level differences in bacteroides ovatus and are modifiable by gut microbiota manipulation. Cell Host Microbe 27, 467–475.e6 (2020).
Pabst, O., Cerovic, V. & Hornef, M. Secretory IgA in the coordination of establishment and maintenance of the microbiota. Trends Immunol. 37, 287–296 (2016).
Marcotte, E. et al. Cesarean delivery and risk of childhood leukemia: findings from the Childhood Leukemia International Consortium (CLIC). Cancer Res. 75, LB-194 (2015).
Zachariassen, L. F. et al. Cesarean section induces microbiota-regulated immune disturbances in C57BL/6 mice. J. Immunol. 202, 142–150 (2019).
Busi, S. B. et al. Persistence of birth mode-dependent effects on gut microbiome composition, immune system stimulation and antimicrobial resistance during the first year of life. ISME Commun. 1, 8 (2021).
Ramanan, D. et al. An immunologic mode of multigenerational transmission governs a gut Treg setpoint. Cell 181, 1276–1290.e13 (2020).
van den Elsen, L. W. J., Garssen, J., Burcelin, R. & Verhasselt, V. Shaping the gut microbiota by breastfeeding: the gateway to allergy prevention? Front. Pediatr. 7, 47 (2019).
Lundell, A.-C. et al. Infant B cell memory differentiation and early gut bacterial colonization. J. Immunol. 188, 4315–4322 (2012).
Wood, H. et al. Breastfeeding promotes early neonatal regulatory T-cell expansion and immune tolerance of non-inherited maternal antigens. Allergy 76, 2447–2460 (2021).
Henrick, B. M. et al. Bifidobacteria-mediated immune system imprinting early in life. Cell 184, 3884–3898.e11 (2021).
Vatanen, T. et al. Variation in microbiome LPS immunogenicity contributes to autoimmunity in humans. Cell 165, 842–853 (2016).
Zmora, N., Suez, J. & Elinav, E. You are what you eat: diet, health and the gut microbiota. Nat. Rev. Gastroenterol. Hepatol. 16, 35–56 (2019).
Arpaia, N. et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 504, 451–455 (2013).
Kaisar, M. M. M., Pelgrom, L. R., van der Ham, A. J., Yazdanbakhsh, M. & Everts, B. Butyrate conditions human dendritic cells to prime type 1 regulatory T cells via both histone deacetylase inhibition and G protein-coupled receptor 109A signaling. Front. Immunol. 8, 1429 (2017).
Furusawa, Y. et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504, 446–450 (2013).
Smith, P. M. et al. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 341, 569–573 (2013).
Cait, A. et al. Microbiome-driven allergic lung inflammation is ameliorated by short-chain fatty acids. Mucosal Immunol. 11, 785–795 (2018).
Rosser, E. C. et al. Regulatory B cells are induced by gut microbiota-driven interleukin-1β and interleukin-6 production. Nat. Med. 20, 1334–1339 (2014).
Sanchez, H. N. et al. B cell-intrinsic epigenetic modulation of antibody responses by dietary fiber-derived short-chain fatty acids. Nat. Commun. 11, 60 (2020).
Makki, K., Deehan, E. C., Walter, J. & Bäckhed, F. The impact of dietary fiber on gut microbiota in host health and disease. Cell Host Microbe 23, 705–715 (2018).
Corte, V. et al. Microbiota derived short chain fatty acids, propionate and butyrate, contribute to modulate the inflammatory response in chronic kidney disease. Nephrol. Dial. Transplant. 35, gfaa140.MO046 (2020).
Stražar, M. et al. The influence of the gut microbiome on BCG-induced trained immunity. Genome Biol. 22, 275 (2021).
Nakajima, A. et al. Maternal high fiber diet during pregnancy and lactation influences regulatory T cell differentiation in offspring in mice. J. Immunol. 199, 3516–3524 (2017).
Kwan, M. L. et al. Maternal diet and risk of childhood acute lymphoblastic leukemia. Public Health Rep. 124, 503–514 (2009).
Heiss, C. N. et al. The gut microbiota regulates hypothalamic inflammation and leptin sensitivity in Western diet-fed mice via a GLP-1R-dependent mechanism. Cell Rep. 35, 109163 (2021).
Lu, Z. et al. Fasting selectively blocks development of acute lymphoblastic leukemia via leptin-receptor upregulation. Nat. Med. 23, 79–90 (2017).
Blaser, M., Nomura, A., Lee, J., Stemmerman, G. & Perez-Perez GI. Early-life family structure and microbially induced cancer risk. PLoS ONE 4, e100 (2007).
Wellbrock, M. et al. 28-year incidence and time trends of childhood leukaemia in former East Germany compared to West Germany after German reunification: a study from the German Childhood Cancer Registry. Cancer Epidemiol. 73, 101968 (2021).
Steliarova-Foucher, E. et al. Changing geographical patterns and trends in cancer incidence in children and adolescents in Europe, 1991–2010 (automated childhood cancer information system): a population-based study. Lancet Oncol. 19, 1159–1169 (2018).
Linet, M. S. et al. International long-term trends and recent patterns in the incidence of leukemias and lymphomas among children and adolescents ages 0–19 years. Int. J. Cancer 138, 1862–1874 (2016).
Zhou, L. et al. Faecalibacterium prausnitzii produces butyrate to maintain Th17/Treg balance and to ameliorate colorectal colitis by inhibiting histone deacetylase 1. Inflamm. Bowel Dis. 24, 1926–1940 (2018).
Zhang, M. et al. Faecalibacterium prausnitzii produces butyrate to decrease c-Myc-related metabolism and Th17 differentiation by inhibiting histone deacetylase 3. Int. Immunol. 31, 499–514 (2019).
Ponsonby, A. et al. Household size, T regulatory cell development, and early allergic disease: a birth cohort study. Pediatr. Allergy Immunol. 33, e13810 (2022).
Roslund, M. I. et al. Biodiversity intervention enhances immune regulation and health-associated commensal microbiota among daycare children. Sci. Adv. 6, eaba2578 (2020).
Chang, J. S. et al. Profound deficit of IL10 at birth in children who develop childhood acute lymphoblastic leukemia. Cancer Epidemiol. Biomark. Prev. 20, 1736–1740 (2011).
Fitch, B. et al. Decreased IL-10 accelerates B-cell leukemia/lymphoma in a mouse model of pediatric lymphoid leukemia. Blood Adv. 6, 854–865 (2021).
Harper, A. et al. Viral infections, the microbiome, and probiotics. Front. Cell Infect. Microbiol. 10, 596166 (2021).
Erttmann, S. F. et al. The gut microbiota prime systemic antiviral immunity via the cGAS-STING-IFN-I axis. Immunity 55, 847–861.e10 (2022).
Wirusanti, N. I., Baldridge, M. T. & Harris, V. C. Microbiota regulation of viral infections through interferon signaling. Trends Microbiol. 30, 778–792 (2022).
Haak, B. W. et al. Impact of gut colonization with butyrate-producing microbiota on respiratory viral infection following allo-HCT. Blood 131, 2978–2986 (2018).
Brown, J. A. et al. Gut microbiota-derived metabolites confer protection against SARS-CoV-2 infection. Gut Microbes 14, 2105609 (2022).
Albrich, W. C. et al. A high-risk gut microbiota configuration associates with fatal hyperinflammatory immune and metabolic responses to SARS-CoV-2. Gut Microbes 14, 2073131 (2022).
Huda, M. N. et al. Stool microbiota and vaccine responses of infants. Pediatrics 134, 3937 (2014).
de Jong, S. E., Olin, A. & Pulendran, B. The impact of the microbiome on immunity to vaccination in humans. Cell Host Microbe 28, 169–179 (2020).
Trompette, A. et al. Dietary fiber confers protection against flu by shaping Ly6c− patrolling monocyte hematopoiesis and CD8+ T cell metabolism. Immunity 48, 992–1005.e8 (2018).
Vicente-Dueñas, C. et al. An intact gut microbiome protects genetically predisposed mice against leukemia. Blood 136, 2003–2017 (2020).
Wang, R. et al. Gut microbiota regulates acute myeloid leukaemia via alteration of intestinal barrier function mediated by butyrate. Nat. Commun. 13, 2522 (2022).
Beneforti, L. et al. Pro-inflammatory cytokines favor the emergence of ETV6-RUNX1-positive pre-leukemic cells in a model of mesenchymal niche. Br. J. Haematol. 190, 262–273 (2020).
Dander, E., Palmi, C., D’amico, G. & Cazzaniga, G. The bone marrow niche in B-cell acute lymphoblastic leukemia: the role of microenvironment from pre-leukemia to overt leukemia. Int. J. Mol. Sci. 22, 4426 (2021).
Xiao, E. et al. Microbiota regulates bone marrow mesenchymal stem cell lineage differentiation and immunomodulation. Stem Cell Res. Ther. 8, 213 (2017).
Marcos-Zambrano, L. J. et al. Applications of machine learning in human microbiome studies: a review on feature selection, biomarker identification, disease prediction and treatment. Front. Microbiol. 12, 634511 (2021).
McCoubrey, L. E., Elbadawi, M., Orlu, M., Gaisford, S. & Basit, A. W. Harnessing machine learning for development of microbiome therapeutics. Gut Microbes 13, 1872323 (2021).
Mirzayi, C. et al. Reporting guidelines for human microbiome research: the STORMS checklist. Nat. Med. 27, 1885–1892 (2021).
Blaser, M. J. The theory of disappearing microbiota and the epidemics of chronic diseases. Nat. Rev. Immunol. 17, 461–463 (2017).
Panigrahi, P. et al. A randomized synbiotic trial to prevent sepsis among infants in rural India. Nature 548, 407–412 (2017).
Korpela, K. et al. Probiotic supplementation restores normal microbiota composition and function in antibiotic-treated and in caesarean-born infants. Microbiome 6, 182 (2018).
Durack, J. et al. Delayed gut microbiota development in high-risk for asthma infants is temporarily modifiable by Lactobacillus supplementation. Nat. Commun. 9, 707 (2018).
Acknowledgements
The authors acknowledge support from the Cancer Research UK (CRM 171X), The Children’s Cancer and Leukaemia Group (CCLGA2019.02), The Royal Marsden Cancer Charity, the Wood family-in memory of Artemis and The Institute for Cancer Research, London.
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Cancer thanks Martin Blaser, Stephen Sallan and Josef Vormoor for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Embase: https://www.embase.com/search/quick
MEDLINE: https://pubmed.ncbi.nlm.nih.gov/
Supplementary information
Glossary
- α-Diversity
-
A measure of the number of different taxa (richness) and/or the degree of evenness in their relative abundance within a single sample.
- β-Diversity
-
A measure of the degree of similarity or distance between the composition of the microbial communities of two samples.
- Activation-induced cytidine deaminase
-
(AID). An enzyme required for somatic hypermutation and class-switch recombination of immunoglobulin genes during B cell maturation and immune response.
- Area under the receiver operating characteristic curve
-
(AUC). An aggregate measure of the performance of a predictive model across all possible classification thresholds.
- Bray–Curtis dissimilarity
-
A measure of β-diversity that quantifies the degree of dissimilarity in the composition of the microbial communities of two samples.
- Chao1
-
A measure of α-diversity that takes into account the number of different taxa (richness) within a sample.
- Human milk oligosaccharides
-
(HMOs). Human milk oligosaccharides are unconjugated complex glycans that have a central role in the development of the gut microbiome–immune system axis.
- High hyperdiploidy
-
A genetic aberration characterized by chromosomal gains (>51 chromosomes) that is commonly found in preleukaemic clones of childhood B cell precursor ALL.
- Inverse Simpson index
-
A measure of α-diversity that takes into account both richness and evenness within a sample, giving more weight to common taxa.
- Leptin
-
A hormone produced by adipose tissue that has a central role in the regulation of energy balance and has widespread effects in multiple organ systems, including haematopoietic cells.
- Linear discriminant analysis of effect size
-
(LEfSe). Determines the taxa most likely to explain differences between study groups. It uses standard statistical tests to detect taxa with significant difference in relative abundance between the groups, as well as additional tests to assess the biological significance and relevance of these taxa.
- Microorganism-associated molecular patterns
-
(MAMPs). Molecular structures conserved among classes of microorganisms that can be recognized by pattern recognition receptors to elicit immune responses.
- Neonatal Guthrie cards
-
Samples of dried blood routinely collected after birth via heel prick for the purpose of universal screening for genetic conditions.
- Peyer’s patches
-
Gut-associated lymphoid tissue found in the small intestine that forms the interface of the gut microbiome-mediated immune system priming.
- Shannon diversity index
-
A measure of α-diversity that takes into account both richness and evenness of taxa within a sample, giving more weight to rare taxa.
- Short-chain fatty acids
-
(SCFAs). Metabolites produced by gut commensals through the fermentation of non-digestible fibre.
- Weighted Unifrac distance
-
A measure of β-diversity that incorporates abundance information and places more weight to common species. By contrast, the unweighted Unifrac distance is a measure of β-diversity that takes into account the presence and absence of taxa.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Peppas, I., Ford, A.M., Furness, C.L. et al. Gut microbiome immaturity and childhood acute lymphoblastic leukaemia. Nat Rev Cancer 23, 565–576 (2023). https://doi.org/10.1038/s41568-023-00584-4
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41568-023-00584-4
This article is cited by
-
Harnessing the microbiome for cancer therapy
Nature Reviews Microbiology (2026)
-
LC-MS analysis of serum lipidomic and metabolomic signatures in pediatric patients with acute lymphoblastic leukemia
Italian Journal of Pediatrics (2025)
-
Disruption and adaptation: infant gut microbiota’s dynamic response to SARS-CoV-2 infection
Microbiome (2025)
-
A reconceptualized framework for human microbiome transmission in early life
Nature Communications (2025)
-
Distinct functional and compositional properties in the gut microbiome of children with acute lymphoblastic leukaemia identified by shotgun metagenomics
Scientific Reports (2025)