Abstract
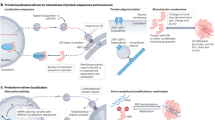
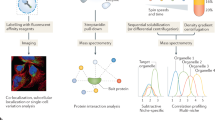
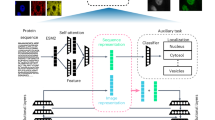
How proteins localize to specific compartments, function in coordination with other biomolecules and, ultimately, contribute to diverse cellular activities are crucial questions in cell biology. Complicating the answers to these questions are multilocalizing and multifunctional proteins, whose impact on the cell depends on both spatial and temporal contexts. Therefore, contextualizing protein functions based on their subcellular localization is necessary to fully understand cell behaviours. Recent advances in instrumentation and protein labelling techniques are rapidly increasing the availability of tools, technologies and applications that measure and control protein localization and compartment-specific function. In this Review, we first discuss microscopy, mass spectrometry-based correlation profiling and proximity labelling methods that assign localizations to proteins, ranging from cellular compartments to protein–protein interactions. We next examine the available tools for manipulating protein localization and measuring the effects of these manipulations, including localization tags and bifunctional molecules. For each technology, we assess the strengths and weaknesses that ultimately determine their usefulness. We conclude with an outlook on future technological advances in the field of spatial subcellular proteomics and their potential implications for cell biology and clinical applications.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Thul, P. J. et al. A subcellular map of the human proteome. Science 356, eaal3321 (2017).
Geladaki, A. et al. Combining LOPIT with differential ultracentrifugation for high-resolution spatial proteomics. Nat. Commun. 10, 331 (2019).
Sigaeva, A., Hutchings, C., Cesnik, A., Lilley, K. S. & Lundberg, E. Subcellular localization as a driver of protein function. Nat. Rev. Mol. Cell Biol. https://doi.org/10.1038/s41580-026-00947-3 (2026).
Christopher, J. A. et al. Subcellular proteomics. Nat. Rev. Methods Primers 1, 32 (2021).
Breckels, L. M. et al. Advances in spatial proteomics: mapping proteome architecture from protein complexes to subcellular localizations. Cell Chem. Biol. 31, 1665–1687 (2024).
Lundberg, E. & Borner, G. H. H. Spatial proteomics: a powerful discovery tool for cell biology. Nat. Rev. Mol. Cell Biol. 20, 285–302 (2019).
Schiff, L. et al. Integrating deep learning and unbiased automated high-content screening to identify complex disease signatures in human fibroblasts. Nat. Commun. 13, 1590 (2022).
Kapałczyńska, M. et al. 2D and 3D cell cultures—a comparison of different types of cancer cell cultures. Arch. Med. Sci. 14, 910–919 (2018).
Ayoubi, R. et al. Scaling of an antibody validation procedure enables quantification of antibody performance in major research applications. eLife 12, RP91645 (2023).
Berglund, L. et al. A genecentric Human Protein Atlas for expression profiles based on antibodies. Mol. Cell. Proteom. 7, 2019–2027 (2008).
Andersson, S. et al. Insufficient antibody validation challenges oestrogen receptor β research. Nat. Commun. 8, 15840 (2017).
Baker, M. Reproducibility crisis: blame it on the antibodies. Nature 521, 274–276 (2015).
Bradbury, A. & Plückthun, A. Reproducibility: standardize antibodies used in research. Nature 518, 27–29 (2015).
Schnell, U., Dijk, F., Sjollema, K. A. & Giepmans, B. N. G. Immunolabeling artifacts and the need for live-cell imaging. Nat. Methods 9, 152–158 (2012).
Gupta, A. et al. SubCell: vision foundation models for microscopy capture single-cell biology. Preprint at bioRxiv https://doi.org/10.1101/2024.12.06.627299 (2024).
Ouyang, W. et al. Analysis of the Human Protein Atlas image classification competition. Nat. Methods 16, 1254–1261 (2019).
Radtke, A. J. et al. IBEX: a versatile multiplex optical imaging approach for deep phenotyping and spatial analysis of cells in complex tissues. Proc. Natl Acad. Sci. USA 117, 33455–33465 (2020).
Radtke, A. J. et al. IBEX: an iterative immunolabeling and chemical bleaching method for high-content imaging of diverse tissues. Nat. Protoc. 17, 378–401 (2022).
Gut, G., Herrmann, M. D. & Pelkmans, L. Multiplexed protein maps link subcellular organization to cellular states. Science 361, eaar7042 (2018).
Hell, S. W. & Wichmann, J. Breaking the diffraction resolution limit by stimulated emission: stimulated-emission-depletion fluorescence microscopy. Opt. Lett. 19, 780–782 (1994).
Klar, T. A., Jakobs, S., Dyba, M., Egner, A. & Hell, S. W. Fluorescence microscopy with diffraction resolution barrier broken by stimulated emission. Proc. Natl Acad. Sci. USA 97, 8206–8210 (2000).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Rust, M. J., Bates, M. & Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 3, 793–795 (2006).
Gustafsson, M. G. Surpassing the lateral resolution limit by a factor of two using structured illumination microscopy. J. Microsc. 198, 82–87 (2000).
Kner, P., Chhun, B. B., Griffis, E. R., Winoto, L. & Gustafsson, M. G. L. Super-resolution video microscopy of live cells by structured illumination. Nat. Methods 6, 339–342 (2009).
Hess, S. T., Girirajan, T. P. K. & Mason, M. D. Ultra-high resolution imaging by fluorescence photoactivation localization microscopy. Biophys. J. 91, 4258–4272 (2006).
Balzarotti, F. et al. Nanometer resolution imaging and tracking of fluorescent molecules with minimal photon fluxes. Science 355, 606–612 (2016).
Wirth, J. O. et al. MINFLUX dissects the unimpeded walking of kinesin-1. Science 379, 1004–1010 (2023).
Gwosch, K. C. et al. MINFLUX nanoscopy delivers 3D multicolor nanometer resolution in cells. Nat. Methods 17, 217–224 (2020).
Sahl, S. J. et al. Direct optical measurement of intramolecular distances with angstrom precision. Science 386, 180–187 (2024).
Schermelleh, L. et al. Super-resolution microscopy demystified. Nat. Cell Biol. 21, 72–84 (2019).
Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature 425, 737–741 (2003).
Huh, W.-K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003).
Gavin, A.-C. et al. Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature 415, 141–147 (2002).
Cho, N. H. et al. OpenCell: endogenous tagging for the cartography of human cellular organization. Science 375, eabi6983 (2022).
Davidi, D. et al. Assessment of GFP tag position on protein localization and growth fitness in yeast. J. Mol. Biol. 431, 636–641 (2019).
Reicher, A. et al. Pooled multicolour tagging for visualizing subcellular protein dynamics. Nat. Cell Biol. 26, 745–756 (2024).
Los, G. V. et al. HaloTag: a novel protein labeling technology for cell imaging and protein analysis. ACS Chem. Biol. 3, 373–382 (2008).
Keppler, A. et al. A general method for the covalent labeling of fusion proteins with small molecules in vivo. Nat. Biotechnol. 21, 86–89 (2003).
Gautier, A. et al. An engineered protein tag for multiprotein labeling in living cells. Chem. Biol. 15, 128–136 (2008).
Erdmann, R. S. et al. Labeling strategies matter for super-resolution microscopy: a comparison between halotags and snap-tags. Cell Chem. Biol. 26, 584–592.e6 (2019).
Götzke, H. et al. The ALFA-tag is a highly versatile tool for nanobody-based bioscience applications. Nat. Commun. 10, 4403 (2019).
Fredriksson, S. et al. Protein detection using proximity-dependent DNA ligation assays. Nat. Biotechnol. 20, 473–477 (2002).
Söderberg, O. et al. Direct observation of individual endogenous protein complexes in situ by proximity ligation. Nat. Methods 3, 995–1000 (2006).
Hansen, J. N. et al. Intrinsic heterogeneity of primary cilia revealed through spatial proteomics. Cell 188, 6804–6824.e16 (2025).
Karlsson, F. et al. Molecular pixelation: spatial proteomics of single cells by sequencing. Nat. Methods 21, 1044–1052 (2024).
Wu, D. et al. Profiling surface proteins on individual exosomes using a proximity barcoding assay. Nat. Commun. 10, 3854 (2019).
Weinstein, J. A., Regev, A. & Zhang, F. DNA microscopy: optics-free spatio-genetic imaging by a stand-alone chemical reaction. Cell 178, 229–241.e16 (2019).
Faulk, W. P. & Taylor, G. M. An immunocolloid method for the electron microscope. Immunochemistry 8, 1081–1083 (1971).
Petralia, R. S. & Wang, Y.-X. Review of post-embedding immunogold methods for the study of neuronal structures. Front. Neuroanat. 15, 763427 (2021).
Polishchuk, R. S. et al. Correlative light-electron microscopy reveals the tubular-saccular ultrastructure of carriers operating between Golgi apparatus and plasma membrane. J. Cell Biol. 148, 45–58 (2000).
Watanabe, S. et al. Protein localization in electron micrographs using fluorescence nanoscopy. Nat. Methods 8, 80–84 (2011).
Bykov, Y. S. et al. High-throughput ultrastructure screening using electron microscopy and fluorescent barcoding. J. Cell Biol. 218, 2797–2811 (2019).
Fung, H. K. H. et al. Genetically encoded multimeric tags for subcellular protein localization in cryo-EM. Nat. Methods 20, 1900–1908 (2023).
Greenblatt, J. F., Alberts, B. M. & Krogan, N. J. Discovery and significance of protein–protein interactions in health and disease. Cell 187, 6501–6517 (2024).
Hein, M. Y. et al. A human interactome in three quantitative dimensions organized by stoichiometries and abundances. Cell 163, 712–723 (2015).
Drew, K. et al. Integration of over 9,000 mass spectrometry experiments builds a global map of human protein complexes. Mol. Syst. Biol. 13, 932 (2017).
Yamanaka, T., Nishiyama, R., Shimogori, T. et al. Proteomics-based approach identifies altered ER domain properties by ALS-linked VAPB mutation. Sci. Rep. 10, 7610 (2020).
Abu-Remaileh, M. et al. Lysosomal metabolomics reveals V-ATPase- and mTOR-dependent regulation of amino acid efflux from lysosomes. Science 358, 807–813 (2017).
Chen, W. W., Freinkman, E. & Sabatini, D. M. Rapid immunopurification of mitochondria for metabolite profiling and absolute quantification of matrix metabolites. Nat. Protoc. 12, 2215–2231 (2017).
Park, H. et al. Spatial snapshots of amyloid precursor protein intramembrane processing via early endosome proteomics. Nat. Commun. 13, 6112 (2022).
Fasimoye, R. et al. Golgi-IP, a tool for multimodal analysis of Golgi molecular content. Proc. Natl Acad. Sci. USA 120, e2219953120 (2023).
Steiner, F. A., Talbert, P. B., Kasinathan, S., Deal, R. B. & Henikoff, S. Cell-type-specific nuclei purification from whole animals for genome-wide expression and chromatin profiling. Genome Res. 22, 766–777 (2012).
Hein, M. Y. et al. Global organelle profiling reveals subcellular localization and remodeling at proteome scale. Cell 188, 1137–1155.e20 (2025).
Zhong, B. et al. In vivo cross-linking-based affinity purification and mass spectrometry for targeting intracellular protein–protein interactions. Anal. Chim. Acta 1265, 341273 (2023).
Dunkley, T. P. J., Watson, R., Griffin, J. L., Dupree, P. & Lilley, K. S. Localization of organelle proteins by isotope tagging (LOPIT). Mol. Cell Proteom. 3, 1128–1134 (2004).
Christoforou, A. et al. A draft map of the mouse pluripotent stem cell spatial proteome. Nat. Commun. 7, 9992 (2016).
Borner, G. H. H. Organellar maps through proteomic profiling — a conceptual guide. Mol. Cell. Proteom. 19, 1076–1087 (2020).
Orre, L. M. et al. SubCellBarCode: proteome-wide mapping of protein localization and relocalization. Mol. Cell 73, 166–182.e7 (2019).
Villanueva, E. et al. System-wide analysis of RNA and protein subcellular localization dynamics. Nat. Methods 21, 60–71 (2024).
Kennedy, M. A., Hofstadter, W. A. & Cristea, I. M. TRANSPIRE: a computational pipeline to elucidate intracellular protein movements from spatial proteomics data sets. J. Am. Soc. Mass. Spectrom. 31, 1422–1439 (2020).
Martinez-Val, A. et al. Spatial-proteomics reveals phospho-signaling dynamics at subcellular resolution. Nat. Commun. 12, 7113 (2021).
Currie, J. et al. Simultaneous proteome localization and turnover analysis reveals spatiotemporal features of protein homeostasis disruptions. Nat. Commun. 15, 2207 (2024).
Liao, P.-C., Bergamini, C., Fato, R., Pon, L. A. & Pallotti, F. Isolation of mitochondria from cells and tissues. Methods Cell Biol. 155, 3–31 (2020).
Katholnig, K., Poglitsch, M., Hengstschläger, M. & Weichhart, T. in The Nucleus (ed. Hancock, R.) 15–23 (Springer, 2015).
Chhoy, P., Brown, C. W., Amante, J. J. & Mercurio, A. M. Protocol for the separation of extracellular vesicles by ultracentrifugation from in vitro cell culture models. STAR Protoc. 2, 100303 (2021).
Crook, O. M., Smith, T., Elzek, M. & Lilley, K. S. Moving profiling spatial proteomics beyond discrete classification. Proteomics 20, 1900392 (2020).
Armingol, E., Officer, A., Harismendy, O. & Lewis, N. E. Deciphering cell–cell interactions and communication from gene expression. Nat. Rev. Genet. 22, 71–88 (2021).
Branon, T. C. et al. Efficient proximity labeling in living cells and organisms with TurboID. Nat. Biotechnol. 36, 880–887 (2018).
Kubitz, L. et al. Engineering of ultraID, a compact and hyperactive enzyme for proximity-dependent biotinylation in living cells. Commun. Biol. 5, 1–14 (2022).
Lam, S. S. et al. Directed evolution of APEX2 for electron microscopy and proximity labeling. Nat. Methods 12, 51–54 (2015).
Roux, K. J., Kim, D. I., Raida, M. & Burke, B. A promiscuous biotin ligase fusion protein identifies proximal and interacting proteins in mammalian cells. J. Cell Biol. 196, 801–810 (2012).
Choi-Rhee, E., Schulman, H. & Cronan, J. E. Promiscuous protein biotinylation by Escherichia coli biotin protein ligase. Protein Sci. 13, 3043–3050 (2004).
Kim, D. I. et al. An improved smaller biotin ligase for BioID proximity labeling. Mol. Biol. Cell 27, 1188–1196 (2016).
Xiong, Z. et al. In vivo proteomic mapping through GFP-directed proximity-dependent biotin labelling in zebrafish. eLife 10, e64631 (2021).
Mair, A., Xu, S.-L., Branon, T. C., Ting, A. Y. & Bergmann, D. C. Proximity labeling of protein complexes and cell-type-specific organellar proteomes in Arabidopsis enabled by TurboID. eLife 8, e47864 (2019).
Zhang, Y. et al. TurboID-based proximity labeling reveals that UBR7 is a regulator of N NLR immune receptor-mediated immunity. Nat. Commun. 10, 3252 (2019).
Shi, W. et al. Combination of in vivo proximity labeling and co-immunoprecipitation identifies the host target network of a tumor-inducing effector in the fungal maize pathogen Ustilago maydis. J. Exp. Bot. 74, 4736–4750 (2023).
Hollstein, L. S., Schmitt, K., Valerius, O., Stahlhut, G. & Pöggeler, S. Establishment of in vivo proximity labeling with biotin using TurboID in the filamentous fungus Sordaria macrospora. Sci. Rep. 12, 17727 (2022).
Go, C. D. et al. A proximity-dependent biotinylation map of a human cell. Nature 595, 120–124 (2021).
Qin, W. et al. Dynamic mapping of proteome trafficking within and between living cells by TransitID. Cell 186, 3307–3324.e30 (2023).
Lee, A. et al. OrthoID: profiling dynamic proteomes through time and space using mutually orthogonal chemical tools. Nat. Commun. 15, 1851 (2024).
Martell, J. D. et al. A split horseradish peroxidase for the detection of intercellular protein–protein interactions and sensitive visualization of synapses. Nat. Biotechnol. 34, 774–780 (2016).
Han, Y. et al. Directed evolution of split APEX2 peroxidase. ACS Chem. Biol. 14, 619–635 (2019).
Bezawork-Geleta, A. et al. Proximity proteomics reveals a mechanism of fatty acid transfer at lipid droplet–mitochondria–endoplasmic reticulum contact sites. Nat. Commun. 16, 2135 (2025).
Cho, K. F. et al. Split-TurboID enables contact-dependent proximity labeling in cells. Proc. Natl Acad. Sci. USA 117, 12143–12154 (2020).
Kwak, C. et al. Contact-ID, a tool for profiling organelle contact sites, reveals regulatory proteins of mitochondrial-associated membrane formation. Proc. Natl Acad. Sci. USA 117, 12109–12120 (2020).
Lee, S.-Y. et al. Engineered allostery in light-regulated LOV-Turbo enables precise spatiotemporal control of proximity labeling in living cells. Nat. Methods 20, 908–917 (2023).
Lee, S.-Y. et al. APEX fingerprinting reveals the subcellular localization of proteins of interest. Cell Rep. 15, 1837–1847 (2016).
Zhang, H. et al. Mass spectrometry imaging for spatially resolved multi-omics molecular mapping. npj Imaging 2, 1–15 (2024).
Caprioli, R. M., Farmer, T. B. & Gile, J. Molecular imaging of biological samples: localization of peptides and proteins using MALDI-TOF MS. Anal. Chem. 69, 4751–4760 (1997).
Zavalin, A., Yang, J., Hayden, K., Vestal, M. & Caprioli, R. M. Tissue protein imaging at 1 μm laser spot diameter for high spatial resolution and high imaging speed using transmission geometry MALDI TOF MS. Anal. Bioanal. Chem. 407, 2337–2342 (2015).
Kompauer, M., Heiles, S. & Spengler, B. Atmospheric pressure MALDI mass spectrometry imaging of tissues and cells at 1.4-μm lateral resolution. Nat. Methods 14, 90–96 (2017).
Grgic, A., Cuypers, E., Dubois, L. J., Ellis, S. R. & Heeren, R. M. A. MALDI MSI protocol for spatial bottom-up proteomics at single-cell resolution. J. Proteome Res. 23, 5372–5379 (2024).
Keren, L. et al. MIBI-TOF: a multiplexed imaging platform relates cellular phenotypes and tissue structure. Sci. Adv. 5, eaax5851 (2019).
Jean Beltran, P. M., Mathias, R. A. & Cristea, I. M. A portrait of the human organelle proteome in space and time during cytomegalovirus infection. Cell Syst. 3, 361–373.e6 (2016).
Mund, A. et al. Deep visual proteomics defines single-cell identity and heterogeneity. Nat. Biotechnol. 40, 1231–1240 (2022).
Gordon, A. & Gousset, K. Utilization of laser capture microdissection coupled to mass spectrometry to uncover the proteome of cellular protrusions. Methods Mol. Biol. 2259, 25–45 (2021).
Dong, Z. et al. Spatial proteomics of single cells and organelles on tissue slides using filter-aided expansion proteomics. Nat. Commun. 15, 9378 (2024).
Chen, Y.-D. et al. Microscopy-guided subcellular proteomic discovery by high-speed ultra-content photo-biotinylation. Preprint at bioRxiv https://doi.org/10.1101/2023.12.27.573388 (2023).
Qin, Y. et al. A multi-scale map of cell structure fusing protein images and interactions. Nature 600, 536–542 (2021).
Schaffer, L. V. et al. Multimodal cell maps as a foundation for structural and functional genomics. Nature https://doi.org/10.1038/s41586-025-08878-3 (2025).
Fischer, S. N. et al. hu.MAP3.0: atlas of human protein complexes by integration of >25,000 proteomic experiments. Mol. Syst. Biol. 21, 911–943 (2025).
Wang, G., Xue, M.-Q., Shen, H.-B. & Xu, Y.-Y. Learning protein subcellular localization multi-view patterns from heterogeneous data of imaging, sequence and networks. Brief. Bioinform. 23, bbab539 (2022).
Cesnik, A. et al. Mapping the multiscale proteomic organization of cellular and disease phenotypes. Annu. Rev. Biomed. Data Sci. 7, 369–389 (2024).
Zhang, T. et al. ENO1 suppresses cancer cell ferroptosis by degrading the mRNA of iron regulatory protein 1. Nat. Cancer 3, 75–89 (2022).
Claude, A. Fractionation of mammalian liver cells by differential centrifugation; problems, methods, and preparation of extract. J. Exp. Med. 84, 51–59 (1946).
de Duve, C., Pressman, B. C., Gianetto, R., Wattiaux, R. & Appelmans, F. Tissue fractionation studies. 6. Intracellular distribution patterns of enzymes in rat-liver tissue. Biochem. J. 60, 604–617 (1955).
De Robertis, E., De Lores Arnaiz, G. R., Alberici, M., Butcher, R. W. & Sutherland, E. W. Subcellular distribution of adenyl cyclase and cyclic phosphodiesterase in rat brain cortex. J. Biol. Chem. 242, 3487–3493 (1967).
Wolff, J. & Jones, A. B. The purification of bovine thyroid plasma membranes and the properties of membrane-bound adenyl cyclase. J. Biol. Chem. 246, 3939–3947 (1971).
Zhang, J., Campbell, R. E., Ting, A. Y. & Tsien, R. Y. Creating new fluorescent probes for cell biology. Nat. Rev. Mol. Cell Biol. 3, 906–918 (2002).
Dixon, A. S. et al. NanoLuc complementation reporter optimized for accurate measurement of protein interactions in cells. ACS Chem. Biol. 11, 400–408 (2016).
Pfleger, K. D. G. & Eidne, K. A. Illuminating insights into protein–protein interactions using bioluminescence resonance energy transfer (BRET). Nat. Methods 3, 165–174 (2006).
Namkung, Y. et al. Monitoring G protein-coupled receptor and β-arrestin trafficking in live cells using enhanced bystander BRET. Nat. Commun. 7, 12178 (2016).
Suzuki, J., Kanemaru, K. & Iino, M. Genetically encoded fluorescent indicators for organellar calcium imaging. Biophys. J. 111, 1119–1131 (2016).
Massengill, C. I. et al. Sensitive genetically encoded sensors for population and subcellular imaging of cAMP in vivo. Nat. Methods 19, 1461–1471 (2022).
Kostyuk, A. I. et al. In vivo imaging with genetically encoded redox biosensors. Int. J. Mol. Sci. 21, 8164 (2020).
Lyons, A. C., Mehta, S. & Zhang, J. Fluorescent biosensors illuminate the spatial regulation of cell signaling across scales. Biochemical J. 480, 1693–1717 (2023).
Kwon, Y. et al. Non-canonical β-adrenergic activation of ERK at endosomes. Nature 611, 173–179 (2022).
Gest, A. M. M. et al. Molecular spies in action: genetically encoded fluorescent biosensors light up cellular signals. Chem. Rev. 124, 12573–12660 (2024).
Yeh, H.-W. & Ai, H.-W. Development and applications of bioluminescent and chemiluminescent reporters and biosensors. Annu. Rev. Anal. Chem. 12, 129–150 (2019).
Arce, M. M. et al. Central control of dynamic gene circuits governs T cell rest and activation. Nature 637, 930–939 (2025).
Kramer, N. J. et al. CRISPR–Cas9 screens in human cells and primary neurons identify modifiers of C9ORF72 dipeptide-repeat-protein toxicity. Nat. Genet. 50, 603–612 (2018).
Feldman, D. et al. Optical pooled screens in human cells. Cell 179, 787–799.e17 (2019).
Funk, L. et al. The phenotypic landscape of essential human genes. Cell 185, 4634–4653.e22 (2022).
Lacoste, J. et al. Pervasive mislocalization of pathogenic coding variants underlying human disorders. Cell 187, 6725–6741.e13 (2024).
Chen, W. & Choi, J. Molecular circuits for genomic recording of cellular events. Trends Genet. 41, 647–659 (2025).
Barnea, G. et al. The genetic design of signaling cascades to record receptor activation. Proc. Natl Acad. Sci. USA 105, 64–69 (2008).
Coukos, R. et al. An engineered transcriptional reporter of protein localization identifies regulators of mitochondrial and ER membrane protein trafficking in high-throughput CRISPRi screens. eLife 10, e69142 (2021).
Reich, N. C. STATs get their move on. JAKSTAT 2, e27080 (2013).
Hofweber, M. & Dormann, D. Friend or foe—post-translational modifications as regulators of phase separation and RNP granule dynamics. J. Biol. Chem. 294, 7137–7150 (2019).
Chen, B., Sun, Y., Niu, J., Jarugumilli, G. K. & Wu, X. Protein lipidation in cell signaling and diseases: function, regulation, and therapeutic opportunities. Cell Chem. Biol. 25, 817–831 (2018).
Cao, X. et al. A phosphorylation-controlled switch confers cell cycle-dependent protein relocalization. Nat. Cell Biol. 26, 1804–1816 (2024).
Kliche, J. et al. Large-scale phosphomimetic screening identifies phospho-modulated motif-based protein interactions. Mol. Syst. Biol. 19, e11164 (2023).
Sakamoto, K. M. et al. PROTACs: chimeric molecules that target proteins to the Skp1–Cullin-F box complex for ubiquitination and degradation. Proc. Natl Acad. Sci. USA 98, 8554–8559 (2001).
Wang, Z., Che, S. & Yu, Z. PROTAC: novel degradable approach for different targets to treat breast cancer. Eur. J. Pharm. Sci. 198, 106793 (2024).
Shen, J., Zhou, G. & Wang, W. Chemogenetic tools in focus: proximity, conformation, and sterics. Chemistry–Methods 4, e202300051 (2024).
Stanton, B. Z., Chory, E. J. & Crabtree, G. R. Chemically induced proximity in biology and medicine. Science 359, eaao5902 (2018).
Banaszynski, L. A., Liu, C. W. & Wandless, T. J. Characterization of the FKBP·rapamycin·FRB ternary complex. J. Am. Chem. Soc. 127, 4715–4721 (2005).
Bayle, J. H. et al. Rapamycin analogs with differential binding specificity permit orthogonal control of protein activity. Chem. Biol. 13, 99–107 (2006).
Putyrski, M. & Schultz, C. Protein translocation as a tool: the current rapamycin story. FEBS Lett. 586, 2097–2105 (2012).
Irannejad, R. et al. Functional selectivity of GPCR-directed drug action through location bias. Nat. Chem. Biol. 13, 799–806 (2017).
Miyamoto, T. et al. Rapid and orthogonal logic gating with a gibberellin-induced dimerization system. Nat. Chem. Biol. 8, 465–470 (2012).
Liang, F.-S., Ho, W. Q. & Crabtree, G. R. Engineering the ABA plant stress pathway for regulation of induced proximity. Sci. Signal. 4, rs2 (2011).
Park, S.-Y. et al. An orthogonalized PYR1-based CID module with reprogrammable ligand-binding specificity. Nat. Chem. Biol. 20, 103–110 (2024).
Ziegler, M. J. et al. Mandipropamid as a chemical inducer of proximity for in vivo applications. Nat. Chem. Biol. 18, 64–69 (2022).
Wang, T. et al. Repurposing salicylic acid as a versatile inducer of proximity. Nat. Chem. Biol. 21, 1444–1456 (2025).
Pöschko, P. et al. Photoactivatable plant hormone-based chemical inducers of proximity for in vivo applications. ACS Chem. Biol. 20, 332–339 (2025).
Levskaya, A., Weiner, O. D., Lim, W. A. & Voigt, C. A. Spatiotemporal control of cell signalling using a light-switchable protein interaction. Nature 461, 997–1001 (2009).
Kennedy, M. J. et al. Rapid blue-light-mediated induction of protein interactions in living cells. Nat. Methods 7, 973–975 (2010).
Guntas, G. et al. Engineering an improved light-induced dimer (iLID) for controlling the localization and activity of signaling proteins. Proc. Natl Acad. Sci. USA 112, 112–117 (2015).
Benedetti, L. et al. Optimized Vivid-derived Magnets photodimerizers for subcellular optogenetics in mammalian cells. eLife 9, e63230 (2020).
Benedetti, L. et al. Light-activated protein interaction with high spatial subcellular confinement. Proc. Natl Acad. Sci. USA 115, E2238–E2245 (2018).
Uda, Y. et al. Efficient synthesis of phycocyanobilin in mammalian cells for optogenetic control of cell signaling. Proc. Natl Acad. Sci. USA 114, 11962–11967 (2017).
Kögler, A. C. et al. Extremely rapid and reversible optogenetic perturbation of nuclear proteins in living embryos. Dev. Cell 56, 2348–2363.e8 (2021).
Yumerefendi, H. et al. Control of protein activity and cell fate specification via light-mediated nuclear translocation. PLoS One 10, e0128443 (2015).
He, L. et al. Optical control of membrane tethering and interorganellar communication at nanoscales. Chem. Sci. 8, 5275–5281 (2017).
Glantz, S. T. et al. Directly light-regulated binding of RGS-LOV photoreceptors to anionic membrane phospholipids. Proc. Natl Acad. Sci. USA 115, E7720–E7727 (2018).
Di Ventura, B. & Kuhlman, B. Go in! Go out! Inducible control of nuclear localization. Curr. Opin. Chem. Biol. 34, 62–71 (2016).
Phua, S. C., Pohlmeyer, C. & Inoue, T. Rapidly relocating molecules between organelles to manipulate small GTPase activity. ACS Chem. Biol. 7, 1950–1955 (2012).
Yang, X., Jost, A. P.-T., Weiner, O. D. & Tang, C. A light-inducible organelle-targeting system for dynamically activating and inactivating signaling in budding yeast. Mol. Biol. Cell 24, 2419–2430 (2013).
Tei, R. & Baskin, J. M. Spatiotemporal control of phosphatidic acid signaling with optogenetic, engineered phospholipase Ds. J. Cell Biol. 219, e201907013 (2020).
Ong, Q. et al. Spatiotemporal control of subcellular O-GlcNAc signaling using Opto-OGT. Nat. Chem. Biol. 21, 300–308 (2025).
Yu, D. et al. Optogenetic activation of intracellular antibodies for direct modulation of endogenous proteins. Nat. Methods 16, 1095–1100 (2019).
Gil, A. A. et al. Optogenetic control of protein binding using light-switchable nanobodies. Nat. Commun. 11, 4044 (2020).
Ng, C. S. C., Liu, A., Cui, B. & Banik, S. M. Targeted protein relocalization via protein transport coupling. Nature 633, 941–951 (2024).
Keber, F. C., Nguyen, T., Mariossi, A., Brangwynne, C. P. & Wühr, M. Evidence for widespread cytoplasmic structuring into mesoscale condensates. Nat. Cell Biol. 26, 346–352 (2024).
Till, N. A., Ramanathan, M. & Bertozzi, C. R. Induced proximity at the cell surface. Nat. Biotechnol. 43, 702–711 (2025).
Cheng, S. H. et al. Defective intracellular transport and processing of CFTR is the molecular basis of most cystic fibrosis. Cell 63, 827–834 (1990).
Riordan, J. R. et al. Identification of the cystic fibrosis gene: cloning and characterization of complementary DNA. Science 245, 1066–1073 (1989).
Rommens, J. M. et al. Identification of the cystic fibrosis gene: chromosome walking and jumping. Science 245, 1059–1065 (1989).
Atkinson, R. et al. TDP-43 mislocalization drives neurofilament changes in a novel model of TDP-43 proteinopathy. Dis. Model. Mech. 14, dmm047548 (2021).
Neumann, M. et al. Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science 314, 130–133 (2006).
Ling, S.-C., Polymenidou, M. & Cleveland, D. W. Converging mechanisms in ALS and FTD: disrupted RNA and protein homeostasis. Neuron 79, 416–438 (2013).
Walkup, W. G. IV et al. A model for regulation by SynGAP-α1 of binding of synaptic proteins to PDZ-domain ‘slots’ in the postsynaptic density. eLife 5, e16813 (2016).
Gou, G. et al. SynGAP splice variants display heterogeneous spatio-temporal expression and subcellular distribution in the developing mammalian brain. J. Neurochem. 154, 618–634 (2020).
Meili, F. et al. Multi-parametric analysis of 57 SYNGAP1 variants reveal impacts on GTPase signaling, localization, and protein stability. Am. J. Hum. Genet. 108, 148–162 (2021).
Kornienko, J. et al. Mislocalization of pathogenic RBM20 variants in dilated cardiomyopathy is caused by loss-of-interaction with Transportin-3. Nat. Commun. 14, 4312 (2023).
Schraivogel, D. et al. High-speed fluorescence image-enabled cell sorting. Science 375, 315–320 (2022).
Snyder, L. B. et al. Preclinical evaluation of bavdegalutamide (ARV-110), a novel PROteolysis TArgeting Chimera androgen receptor degrader. Mol. Cancer Ther. 24, 511–522 (2025).
Sarott, R. C. et al. Relocalizing transcriptional kinases to activate apoptosis. Science 386, 6717 (2024).
Vargas, M. V. et al. Psychedelics promote neuroplasticity through the activation of intracellular 5-HT2A receptors. Science 379, 700–706 (2023).
Jiang, Y., Wang, D., Wang, W. & Xu, D. Computational methods for protein localization prediction. Comput. Struct. Biotechnol. J. 19, 5834–5844 (2021).
Senior, A. W. et al. Improved protein structure prediction using potentials from deep learning. Nature 577, 706–710 (2020).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Lin, Z. et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science 379, 1123–1130 (2023).
Hayes, T. et al. Simulating 500 million years of evolution with a language model. Science 387, 850–858 (2025).
Madani, A. et al. Large language models generate functional protein sequences across diverse families. Nat. Biotechnol. 41, 1099–1106 (2023).
Nguyen, E. et al. Sequence modeling and design from molecular to genome scale with Evo. Science 386, eado9336 (2024).
Brixi, G. et al. Genome modeling and design across all domains of life with Evo 2. Preprint at bioRxiv https://doi.org/10.1101/2025.02.18.638918 (2025).
Lu, J. et al. Types of nuclear localization signals and mechanisms of protein import into the nucleus. Cell Commun. Signal. 19, 60 (2021).
Blobel, G. & Dobberstein, B. Transfer of proteins across membranes. I. Presence of proteolytically processed and unprocessed nascent immunoglobulin light chains on membrane-bound ribosomes of murine myeloma. J. Cell Biol. 67, 835–851 (1975).
Caras, I. W., Weddell, G. N., Davitz, M. A., Nussenzweig, V. & Martin, D. W. Jr. Signal for attachment of a phospholipid membrane anchor in decay accelerating factor science. Science 238, 1280–1283 (1987).
Hurt, E. C., Pesold-Hurt, B. & Schatz, G. The amino-terminal region of an imported mitochondrial precursor polypeptide can direct cytoplasmic dihydrofolate reductase into the mitochondrial matrix. EMBO J. 3, 3149–3156 (1984).
Dunn, C. D. & Paavilainen, V. O. Wherever I may roam: organellar protein targeting and evolvability. Curr. Opin. Genet. Dev. 58–59, 9–16 (2019).
Jiang, Y. et al. MULocDeep: a deep-learning framework for protein subcellular and suborganellar localization prediction with residue-level interpretation. Comput. Struct. Biotechnol. J. 19, 4825–4839 (2021).
Thumuluri, V., Almagro Armenteros, J. J., Johansen, A. R., Nielsen, H. & Winther, O. DeepLoc 2.0: multi-label subcellular localization prediction using protein language models. Nucleic Acids Res. 50, W228–W234 (2022).
Stärk, H., Dallago, C., Heinzinger, M. & Rost, B. Light attention predicts protein location from the language of life. Bioinforma. Adv. 1, vbab035 (2021).
Kilgore, H. R. et al. Protein codes promote selective subcellular compartmentalization. Science 387, 1095–1101 (2025).
Khwaja, E., Song, Y. S., Agarunov, A. & Huang, B. CELLE-2: translating proteins to pictures and back with a bidirectional text-to-image transformer. Adv. Neural Inf. Process. Syst. 36, 4899–4914 (2023).
Khwaja, E., Song, Y. S. & Huang, B. CELL-E: a text-to-image transformer for protein localization prediction. Preprint at bioRxiv https://doi.org/10.1101/2022.05.27.493774 (2024).
Gupta, A., Wefers, Z., Ahmed, N., Zhang, X. & Lundberg, E. A comprehensive benchmark of sequence-based subcellular location predictors for human proteins. Figshare https://doi.org/10.6084/m9.figshare.31282576 (2026).
Kobayashi, H., Cheveralls, K. C., Leonetti, M. D. & Royer, L. A. Self-supervised deep learning encodes high-resolution features of protein subcellular localization. Nat. Methods 19, 995–1003 (2022).
Le, T. et al. Analysis of the Human Protein Atlas weakly supervised single-cell classification competition. Nat. Methods 19, 1221–1229 (2022).
Doron, M. et al. Unbiased single-cell morphology with self-supervised vision transformers. Preprint at bioRxiv https://doi.org/10.1101/2023.06.16.545359 (2023).
Das, S., Vera, M., Gandin, V., Singer, R. H. & Tutucci, E. Intracellular mRNA transport and localized translation. Nat. Rev. Mol. Cell Biol. 22, 483–504 (2021).
Krahmer, N. et al. Organellar proteomics and phospho-proteomics reveal subcellular reorganization in diet-induced hepatic steatosis. Dev. Cell 47, 205–221.e7 (2018).
Aebersold, R. et al. How many human proteoforms are there? Nat. Chem. Biol. 14, 206–214 (2018).
Dittmann, K., Mayer, C. & Rodemann, H. P. Nuclear EGFR as novel therapeutic target: insights into nuclear translocation and function. Strahlenther. Onkol. 186, 1–6 (2010).
Thiele, A., Nagamine, Y., Hauschildt, S. & Clevers, H. AU-rich elements and alternative splicing in the β-catenin 3′UTR can influence the human β-catenin mRNA stability. Exp. Cell Res. 312, 2367–2378 (2006).
Kjer-Hansen, P., Phan, T. G. & Weatheritt, R. J. Protein isoform-centric therapeutics: expanding targets and increasing specificity. Nat. Rev. Drug. Discov. 23, 759–779 (2024).
Sheynkman, G. M., Shortreed, M. R., Frey, B. L. & Smith, L. M. Discovery and mass spectrometric analysis of novel splice-junction peptides using RNA-seq. Mol. Cell Proteom. 12, 2341–2353 (2013).
Lee, K., Cui, Y., Lee, L. P. & Irudayaraj, J. Quantitative imaging of single mRNA splice variants in living cells. Nat. Nanotechnol. 9, 474–480 (2014).
Acknowledgements
The authors thank A. Sigaeva, C. Hutchings, A. Cesnik, S.-J. Tsai and K. S. Lilley for comments and feedback during the writing of the manuscript. E.L. was supported by the Wallenberg Foundation (2021.0346), Erling Persson Foundation, Göran Gustafsson Foundation, Schmidt Futures, the Bridge2AI Program (National Institutes of Health (NIH) Common Fund; OT2 OD032742), the Cancer Cell Map Initiative (National Cancer Institute (NCI) Center for Cancer Systems Biology; U54 CA274502), Stanford Institute for Human-Centered AI, Chan Zuckerberg Initiative, Phil & Penny Knight Initiative for Brain Resilience at the Wu Tsai Neurosciences Institute, Danaher and Param Hansa Philanthropies. R.T. was supported by the Life Sciences Research Foundation Fellowship (sponsored by Astellas Pharma). A.T. was supported by the Chan Zuckerberg Biohub – San Francisco and the Stanford Bio-X.
Author information
Authors and Affiliations
Contributions
W.L., R.T. and A.M. researched data for the article. E.L., W.L. and R.T. contributed substantially to discussion of the content. W.L., R.T. and A.M. wrote the article. E.M. and A.T. reviewed and/or edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
E.L. is an adviser for Element Biosciences, Nautilus Biotechnology, Cartography Biosciences, Genbio.ai and Pixelgen Technologies. A.T. is a scientific adviser to Third Rock Ventures. The terms of these arrangements have been reviewed and approved by Stanford University in accordance with its conflict of interest policies.
Peer review
Peer review information
Nature Reviews Molecular Cell Biology thanks Edward Lau, who co-reviewed with Matthew Juber; Mikko Taipale; and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Ensembl: https://www.ensembl.org/index.html
Human Protein Atlas: https://www.proteinatlas.org/
UniProt: https://www.uniprot.org/
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Leineweber, W., Tei, R., Mäkiniemi, A. et al. Technologies to measure and modulate protein subcellular localization. Nat Rev Mol Cell Biol (2026). https://doi.org/10.1038/s41580-026-00957-1
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41580-026-00957-1