Abstract
Understanding the mechanisms of brain function and dysfunction is at the core of the neuroscience mission. However, the field’s grasp of causal relationships between brain properties has been hindered by a focus on single modalities that neglects the complex interplay between the features found at different neural scales. Progress in neuroinformatics and the increasing availability of open datasets have helped overcome this limitation by facilitating the contextualization of brain maps against cellular, metabolic and network features. Despite the rapid uptake of data contextualization methods proposing that quantification of spatial similarity between brain maps may shed light on pathways of structure–function coupling, development and disease, their potential pitfalls have received little attention. In the context of neuroimaging research, these limitations include reliance on often small-sample and non-representative reference datasets, repeated use of the same brain maps across studies, and problems with intermodal and interindividual alignment. Applying data contextualization without considering these limitations can lead to circular reasoning, overfitting and correlational overreach, and limits the interpretation of findings to the properties of the source data. Here we provide a Roadmap of practical guidelines operating at the level of study design, analysis pipelines and interpretation of findings to encourage the development of best practices in data contextualization. A more informed use of brain map correlation approaches will improve mechanistic investigations and our understanding of causal relationships between brain properties.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
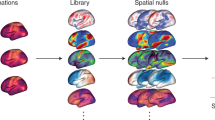
The data used to generate Fig. 1a are available as part of the MICA-MICs dataset (https://osf.io/j532r/)20. For subpanels displayed in Fig. 1b, Fig. 3a (bottom) and Fig. 3c, ultrahigh resolution 7 T MRI data are available as part of the MICA-PNI dataset (https://osf.io/mhq3f/)207. Histological data sampling cortical cyto-architecture are available via BigBrain25 (raw data: https://bigbrain.loris.ca) and BigBrainWarp (processed data: https://github.com/caseypaquola/BigBrainWarp)74. Gene expression data are provided by the Allen Human Brain Atlas (raw data: https://human.brain-map.org/)27. These data were processed with abagen, available via GitHub at https://github.com/rmarkello/abagen (ref. 96) and are made available in the ENIGMA toolbox via GitHub at https://github.com/MICA-MNI/ENIGMA (ref. 19). Data and analysis tools to reproduce Fig. 2 are available in neuromaps via GitHub at https://github.com/netneurolab/neuromaps (ref. 18).
References
Kitano, H. Systems biology: a brief overview. Science 295, 1662–1664 (2002).
Ross, L. N. & Bassett, D. S. Causation in neuroscience: keeping mechanism meaningful. Nat. Rev. Neurosci. 25, 81–90 (2024).
Kulkarni, S. & Bassett, D. S. Toward principles of brain network organization and function. Annu. Rev. Biophys. 54, 353–378 (2025).
Urai, A. E., Doiron, B., Leifer, A. M. & Churchland, A. K. Large-scale neural recordings call for new insights to link brain and behavior. Nat. Neurosci. 25, 11–19 (2022).
Stringer, C. et al. Spontaneous behaviors drive multidimensional, brainwide activity. Science 364, eaav7893 (2019).
O’Byrne, J. & Jerbi, K. How critical is brain criticality? Trends Neurosci. 45, 820–837 (2022).
Krakauer, J. W., Ghazanfar, A. A., Gomez-Marin, A., MacIver, M. A. & Poeppel, D. Neuroscience needs behavior: correcting a reductionist bias. Neuron 93, 480–490 (2017).
Brodmann, K. Vergleichende Lokalisationslehre der Grosshirnrinde in Ihren Prinzipien Dargestellt auf Grund des Zellenbaues (Barth, 1909).
Vogt, O. & Vogt, C. Zur anatomischen gliederung des cortex cerebri. J. Psychol. Neurol. 2, 160–180 (1903).
von Economo, C. F. & Koskinas, G. N. Die Cytoarchitektonik der Hirnrinde des Erwachsenen Menschen (Springer, 1925).
Nieuwenhuys, R. The myeloarchitectonic studies on the human cerebral cortex of the Vogt-Vogt school, and their significance for the interpretation of functional neuroimaging data. Brain Struct. Funct. 218, 303–352 (2013).
Vogt, C. & Vogt, O. Allgemeine Ergebnisse Unserer Hirnforschung Vol. 25 (Barth, 1919).
Pessoa, L. Understanding brain networks and brain organization. Phys. Life Rev. 11, 400–435 (2014).
Haak, K. V. & Beckmann, C. F. Understanding brain organisation in the face of functional heterogeneity and functional multiplicity. NeuroImage 220, 117061 (2020).
Edelman, G. M. & Gally, J. A. Degeneracy and complexity in biological systems. Proc. Natl Acad. Sci. USA 98, 13763–13768 (2001).
Passingham, R. E., Stephan, K. E. & Kötter, R. The anatomical basis of functional localization in the cortex. Nat. Rev. Neurosci. 3, 606–616 (2002).
DeKraker, J. et al. HippoMaps: multiscale cartography of human hippocampal organization. Nat. Methods 22, 2211–2222 (2025).
Markello, R. D. et al. Neuromaps: structural and functional interpretation of brain maps. Nat. Methods 19, 1472–1479 (2022).
Larivière, S. et al. The ENIGMA toolbox: multiscale neural contextualization of multisite neuroimaging datasets. Nat. Methods 18, 698–700 (2021).
Royer, J. et al. An open MRI dataset for multiscale neuroscience. Sci. Data 9, 569 (2022).
Van Essen, D. C. et al. The WU-Minn human connectome project: an overview. NeuroImage 80, 62–79 (2013).
Frauscher, B. et al. Atlas of the normal intracranial electroencephalogram: neurophysiological awake activity in different cortical areas. Brain 141, 1130–1144 (2018).
Vaishnavi, S. N. et al. Regional aerobic glycolysis in the human brain. Proc. Natl Acad. Sci. USA 107, 17757–17762 (2010).
Hansen, J. Y. et al. Mapping neurotransmitter systems to the structural and functional organization of the human neocortex. Nat. Neurosci. 25, 1569–1581 (2022).
Amunts, K. et al. BigBrain: an ultrahigh-resolution 3D human brain model. Science 340, 1472–1475 (2013).
Alkemade, A. et al. A unified 3D map of microscopic architecture and MRI of the human brain. Sci. Adv. 8, eabj7892 (2022).
Hawrylycz, M. J. et al. An anatomically comprehensive atlas of the adult human brain transcriptome. Nature 489, 391–399 (2012).
Vos de Wael, R. et al. BrainSpace: a toolbox for the analysis of macroscale gradients in neuroimaging and connectomics datasets. Commun. Biol. 3, 103 (2020).
Gorgolewski, K. J. et al. BIDS apps: improving ease of use, accessibility, and reproducibility of neuroimaging data analysis methods. PLoS Comput. Biol. 13, e1005209 (2017).
Esteban, O. et al. fMRIPrep: a robust preprocessing pipeline for functional MRI. Nat. Methods 16, 111–116 (2019).
Markiewicz, C. J. et al. The openneuro resource for sharing of neuroscience data. eLife 10, e71774 (2021).
Halchenko, Y. O. et al. DataLad: distributed system for joint management of code, data, and their relationship. J. Open Source Softw. 6, 3262 (2021).
Hayashi, S. et al. brainlife.io: a decentralized and open-source cloud platform to support neuroscience research. Nat. Methods 21, 809–813 (2024).
Penny, W. D., Friston, K. J., Ashburner, J. T., Kiebel, S. J. & Nichols, T. E. Statistical Parametric Mapping: the Analysis of Functional Brain Images (Elsevier, 2011).
Goldblum, Z. et al. Pennsieve: a collaborative platform for translational neuroscience and beyond. Sci. Data 12, 1834 (2025).
Sydnor, V. J. et al. Neurodevelopment of the association cortices: patterns, mechanisms, and implications for psychopathology. Neuron 109, 2820–2846 (2021).
Wagstyl, K. & Raznahan, A. Converging cortical axes. Nat. Neurosci. 28, 8–10 (2025).
Saberi, A. et al. Adolescent maturation of cortical excitation-inhibition balance based on individualized biophysical network modeling. Sci. Adv. 11, eadr8164 (2025).
Salo, T. et al. NiMARE: neuroimaging meta-analysis research environment. NeuroLibre https://doi.org/10.52294/001c.87681 (2022).
Peraza, J. A. et al. Methods for decoding cortical gradients of functional connectivity. Imaging Neurosci. 2, 1–32 (2024).
Larivière, S. et al. BrainStat: a toolbox for brain-wide statistics and multimodal feature associations. NeuroImage 266, 119807 (2023).
Bazinet, V., Hansen, J. Y. & Misic, B. Towards a biologically annotated brain connectome. Nat. Rev. Neurosci. 24, 747–760 (2023).
Hansen, J. Y. et al. Mapping gene transcription and neurocognition across human neocortex. Nat. Hum. Behav. 5, 1240–1250 (2021).
Váša, F. & Mišić, B. Null models in network neuroscience. Nat. Rev. Neurosci. 23, 493–504 (2022).
Markello, R. D. & Misic, B. Comparing spatial null models for brain maps. NeuroImage 236, 118052 (2021).
Leech, R. et al. Variation in spatial dependencies across the cortical mantle discriminates the functional behaviour of primary and association cortex. Nat. Commun. 14, 5656 (2023).
Alexander-Bloch, A. F. et al. On testing for spatial correspondence between maps of human brain structure and function. NeuroImage 178, 540–551 (2018).
Delgado, J. M. R. Functional exploration of the brain with stereotaxic techniques. J. Neurosurg. 15, 269–280 (1958).
Talairach, J. & Tournoux, P. Co-Planar Stereotaxic Atlas of the Human Brain (Thieme, 1988).
Evans, A. C. et al. In 1993 IEEE Conference Record Nuclear Science Symposium and Medical Imaging Conference 1813–1817 (IEEE, 1993).
Fonov, V. et al. Unbiased average age-appropriate atlases for pediatric studies. NeuroImage 54, 313–327 (2011).
Laird, A. R. et al. ALE meta-analysis: controlling the false discovery rate and performing statistical contrasts. Hum. Brain Mapp. 25, 155–164 (2005).
Eickhoff, S. B. et al. Coordinate-based activation likelihood estimation meta-analysis of neuroimaging data: a random-effects approach based on empirical estimates of spatial uncertainty. Hum. Brain Mapp. 30, 2907–2926 (2009).
Eickhoff, S. B., Bzdok, D., Laird, A. R., Kurth, F. & Fox, P. T. Activation likelihood estimation meta-analysis revisited. NeuroImage 59, 2349–2361 (2012).
Yarkoni, T., Poldrack, R. A., Nichols, T. E., Van Essen, D. C. & Wager, T. D. Large-scale automated synthesis of human functional neuroimaging data. Nat. Methods 8, 665–670 (2011).
Fox, P. T. & Lancaster, J. L. Mapping context and content: the BrainMap model. Nat. Rev. Neurosci. 3, 319–321 (2002).
Tzourio-Mazoyer, N. et al. Automated anatomical labeling of activations in SPM using a macroscopic anatomical parcellation of the MNI MRI single-subject brain. NeuroImage 15, 273–289 (2002).
Eickhoff, S. B. et al. A new SPM toolbox for combining probabilistic cytoarchitectonic maps and functional imaging data. NeuroImage 25, 1325–1335 (2005).
Collins, D. L., Neelin, P., Peters, T. M. & Evans, A. C. Automatic 3D intersubject registration of MR volumetric data in standardized talairach space. J. Comput. Assist. Tomogr. 18, 192–205 (1994).
Collins, D.L., Peters, T.M. & Evans, A.C. in Proc. International Conference on Visualization in Biomedical Computing vol. 2359, 180–190 (SPIE, 1994).
Logothetis, N. K. What we can do and what we cannot do with fMRI. Nature 453, 869–878 (2008).
Raichle, M. E. A brief history of human brain mapping. Trends Neurosci. 32, 118–126 (2009).
Hansen, J. Y. et al. Local molecular and global connectomic contributions to cross-disorder cortical abnormalities. Nat. Commun. 13, 4682 (2022).
Alves, P. N., Forkel, S. J., Corbetta, M. & Thiebaut de Schotten, M. The subcortical and neurochemical organization of the ventral and dorsal attention networks. Commun. Biol. 5, 1343 (2022).
Wiesman, A. I. et al. Adverse and compensatory neurophysiological slowing in Parkinson’s disease. Prog. Neurobiol. 231, 102538 (2023).
Vo, A. et al. Network connectivity and local transcriptomic vulnerability underpin cortical atrophy progression in Parkinson’s disease. NeuroImage Clin. 40, 103523 (2023).
García-San-Martín, N. et al. Molecular and micro-architectural mapping of gray matter alterations in psychosis. Mol. Psychiatry 30, 1287–1296 (2024).
Liu, C. et al. Neural, genetic, and cognitive signatures of creativity. Commun. Biol. 7, 1324 (2024).
Cliff, A. D. & Ord, K. Spatial autocorrelation: a review of existing and new measures with applications. Econ. Geogr. 46, 269–292 (1970).
Legendre, P. Spatial autocorrelation: trouble or new paradigm? Ecology 74, 1659–1673 (1993).
Leech, R. et al. The impact of heterogeneous spatial autocorrelation on comparisons of brain maps. Preprint at bioRxiv https://doi.org/10.1101/2024.06.14.598987 (2024).
Taylor, P. N. et al. Normative brain mapping of interictal intracranial EEG to localize epileptogenic tissue. Brain 145, 939–949 (2022).
Janiukstyte, V. et al. Normative brain mapping using scalp EEG and potential clinical application. Sci. Rep. 13, 13442 (2023).
Paquola, C. et al. The BigBrainWarp toolbox for integration of BigBrain 3D histology with multimodal neuroimaging. eLife 10, e70119 (2021).
Valk, S. L. et al. Shaping brain structure: genetic and phylogenetic axes of macroscale organization of cortical thickness. Sci. Adv. 6, eabb3417 (2020).
Saberi, A. et al. The regional variation of laminar thickness in the human isocortex is related to cortical hierarchy and interregional connectivity. PLoS Biol. 21, e3002365 (2023).
Vos de Wael, R. et al. Anatomical and microstructural determinants of hippocampal subfield functional connectome embedding. Proc. Natl Acad. Sci. USA 115, 10154–10159 (2018).
Paquola, C. et al. Convergence of cortical types and functional motifs in the human mesiotemporal lobe. eLife 9, e60673 (2020).
Poppenk, J., Evensmoen, H. R., Moscovitch, M. & Nadel, L. Long-axis specialization of the human hippocampus. Trends Cogn. Sci. 17, 230–240 (2013).
Margulies, D. S. et al. Situating the default-mode network along a principal gradient of macroscale cortical organization. Proc. Natl Acad. Sci. USA 113, 12574–12579 (2016).
Park, B. -y et al. Signal diffusion along connectome gradients and inter-hub routing differentially contribute to dynamic human brain function. NeuroImage 224, 117429 (2021).
Dear, R. et al. Cortical gene expression architecture links healthy neurodevelopment to the imaging, transcriptomics and genetics of autism and schizophrenia. Nat. Neurosci. 27, 1075–1086 (2024).
Paquola, C. et al. Microstructural and functional gradients are increasingly dissociated in transmodal cortices. PLoS Biol. 17, e3000284 (2019).
Wagstyl, K. et al. BigBrain 3D atlas of cortical layers: cortical and laminar thickness gradients diverge in sensory and motor cortices. PLoS Biol. 18, e3000678 (2020).
Müller, E. J. et al. Core and matrix thalamic sub-populations relate to spatio-temporal cortical connectivity gradients. NeuroImage 222, 117224 (2020).
John, A. et al. A multimodal characterization of low-dimensional thalamocortical structural connectivity patterns. Commun. Biol. 8, 185 (2025).
Guell, X., Schmahmann, J. D., Gabrieli, J. D. E. & Ghosh, S. S. Functional gradients of the cerebellum. eLife 7, e36652 (2018).
King, M., Shahshahani, L., Ivry, R. B. & Diedrichsen, J. A task-general connectivity model reveals variation in convergence of cortical inputs to functional regions of the cerebellum. eLife 12, e81511 (2023).
Royer, J. et al. Myeloarchitecture gradients in the human insula: histological underpinnings and association to intrinsic functional connectivity. NeuroImage 216, 116859 (2020).
Katsumi, Y. et al. Correspondence of functional connectivity gradients across human isocortex, cerebellum, and hippocampus. Commun. Biol. 6, 401 (2023).
Dong, H.-M. et al. Ventral attention network connectivity is linked to cortical maturation and cognitive ability in childhood. Nat. Neurosci. 27, 2009–2020 (2024).
Dong, H.-M., Margulies, D. S., Zuo, X.-N. & Holmes, A. J. Shifting gradients of macroscale cortical organization mark the transition from childhood to adolescence. Proc. Natl Acad. Sci. USA 118, e2024448118 (2021).
Hong, S.-J. et al. Atypical functional connectome hierarchy in autism. Nat. Commun. 10, 1022 (2019).
Royer, J. et al. Cortical microstructural gradients capture memory network reorganization in temporal lobe epilepsy. Brain 146, 3923–3937 (2023).
Holmes, A. et al. Disruptions of hierarchical cortical organization in early psychosis and schizophrenia. Biol. Psychiatry Cogn. Neurosci. 8, 1240–1250 (2023).
Markello, R. D. et al. Standardizing workflows in imaging transcriptomics with the abagen toolbox. eLife 10, e72129 (2021).
Murphy, K., Bodurka, J. & Bandettini, P. A. How long to scan? The relationship between fMRI temporal signal to noise ratio and necessary scan duration. NeuroImage 34, 565–574 (2007).
Ojemann, J. G. et al. Anatomic localization and quantitative analysis of gradient refocused echo-planar fMRI susceptibility artifacts. NeuroImage 6, 156–167 (1997).
Esteban, O. et al. MRIQC: advancing the automatic prediction of image quality in MRI from unseen sites. PLoS ONE 12, e0184661 (2017).
Brett, M., Leff, A. P., Rorden, C. & Ashburner, J. Spatial normalization of brain images with focal lesions using cost function masking. NeuroImage 14, 486–500 (2001).
Glasser, M. F. et al. The human connectome project’s neuroimaging approach. Nat. Neurosci. 19, 1175–1187 (2016).
Amunts, K., Malikovic, A., Mohlberg, H., Schormann, T. & Zilles, K. Brodmann’s areas 17 and 18 brought into stereotaxic space — where and how variable? NeuroImage 11, 66–84 (2000).
Van Essen, D. C., Glasser, M. F., Dierker, D. L., Harwell, J. & Coalson, T. Parcellations and hemispheric asymmetries of human cerebral cortex analyzed on surface-based atlases. Cereb. Cortex 22, 2241–2262 (2012).
Reardon, P. K. et al. Normative brain size variation and brain shape diversity in humans. Science 360, 1222–1227 (2018).
Snyder, W. E. et al. A bimodal taxonomy of adult human brain sulcal morphology related to timing of fetal sulcation and trans-sulcal gene expression gradients. Neuron 112, 3396–3411 (2024).
Ono, M., Kubik, S. & Abernathey, C. D. Atlas of the Cerebral Sulci (Thieme, 1990).
Lyttelton, O., Boucher, M., Robbins, S. & Evans, A. An unbiased iterative group registration template for cortical surface analysis. NeuroImage 34, 1535–1544 (2007).
Coalson, T. S., Van Essen, D. C. & Glasser, M. F. The impact of traditional neuroimaging methods on the spatial localization of cortical areas. Proc. Natl Acad. Sci. USA 115, E6356–E6365 (2018).
Wandell, B. A. The neurobiological basis of seeing words. Ann. N. Y. Acad. Sci. 1224, 63–80 (2011).
Goldstone, R. L., Pestilli, F. & Börner, K. Self-portraits of the brain: cognitive science, data visualization, and communicating brain structure and function. Trends Cogn. Sci. 19, 462–474 (2015).
Ciantar, K. G. et al. Geometric effects of volume-to-surface mapping of fMRI data. Brain Struct. Funct. 227, 2457–2464 (2022).
Feilong, M., Jiahui, G., Gobbini, M. I. & Haxby, J. V. A cortical surface template for human neuroscience. Nat. Methods 21, 1736–1742 (2024).
Jeganathan, J. et al. Spurious correlations in surface-based functional brain imaging. Imaging Neurosci. https://doi.org/10.1162/imag_a_00478 (2025).
Robinson, E. C. et al. MSM: a new flexible framework for multimodal surface matching. NeuroImage 100, 414–426 (2014).
Haxby, J. V., Guntupalli, J. S., Nastase, S. A. & Feilong, M. Hyperalignment: modeling shared information encoded in idiosyncratic cortical topographies. eLife 9, e56601 (2020).
Dworetsky, A. et al. Two common and distinct forms of variation in human functional brain networks. Nat. Neurosci. 27, 1187–1198 (2024).
Bignardi, G. et al. Distinct genetic underpinnings of inter-individual differences in the sensorimotor-association axis of cortical organisation. Preprint at bioRxiv https://doi.org/10.1101/2023.07.13.548817 (2024).
Hettwer, M. D. et al. Coordinated cortical thickness alterations across six neurodevelopmental and psychiatric disorders. Nat. Commun. 13, 6851 (2022).
Larivière, S. et al. Connectome reorganization associated with temporal lobe pathology and its surgical resection. Brain 147, 2483–2495 (2024).
Serio, B. et al. Sex differences in functional cortical organization reflect differences in network topology rather than cortical morphometry. Nat. Commun. 15, 7714 (2024).
Bethlehem, R. A. et al. Dispersion of functional gradients across the adult lifespan. NeuroImage 222, 117299 (2020).
Larivière, S. et al. Multiscale structure–function gradients in the neonatal connectome. Cereb. Cortex 30, 47–58 (2020).
Park, B. -y et al. An expanding manifold in transmodal regions characterizes adolescent reconfiguration of structural connectome organization. eLife 10, e64694 (2021).
Paquola, C. et al. Shifts in myeloarchitecture characterise adolescent development of cortical gradients. eLife 8, e50482 (2019).
Bottenhorn, K. L., Corbett, J. D., Ahmadi, H. & Herting, M. M. Spatiotemporal patterns in cortical development: age, puberty, and individual variability from 9 to 13 years of age. J. Neurosci. 45, e1002242025 (2025).
Marquand, A. F., Rezek, I., Buitelaar, J. & Beckmann, C. F. Understanding heterogeneity in clinical cohorts using normative models: beyond case–control studies. Biol. Psychiatry 80, 552–561 (2016).
Rutherford, S. et al. The normative modeling framework for computational psychiatry. Nat. Protoc. 17, 1711–1734 (2022).
Gratton, C. et al. Defining individual-specific functional neuroanatomy for precision psychiatry. Biol. Psychiatry 88, 28–39 (2020).
Gordon, E. M. et al. Precision functional mapping of individual human brains. Neuron 95, 791–807 (2017).
Baum, G. L. et al. Development of structure–function coupling in human brain networks during youth. Proc. Natl Acad. Sci. USA 117, 771–778 (2020).
Medaglia, J. D. et al. Functional alignment with anatomical networks is associated with cognitive flexibility. Nat. Hum. Behav. 2, 156–164 (2018).
Griffa, A., Amico, E., Liégeois, R., Van De Ville, D. & Preti, M. G. Brain structure–function coupling provides signatures for task decoding and individual fingerprinting. NeuroImage 250, 118970 (2022).
Gu, Z., Jamison, K. W., Sabuncu, M. R. & Kuceyeski, A. Heritability and interindividual variability of regional structure–function coupling. Nat. Commun. 12, 4894 (2021).
Lotter, L. D. et al. Regional patterns of human cortex development correlate with underlying neurobiology. Nat. Commun. 15, 7987 (2024).
Shinn, M. et al. Functional brain networks reflect spatial and temporal autocorrelation. Nat. Neurosci. 26, 867–878 (2023).
Worsley, K. J., Taylor, J. E., Tomaiuolo, F. & Lerch, J. Unified univariate and multivariate random field theory. NeuroImage 23, S189–S195 (2004).
Worsley, K. J. Spatial smoothing of autocorrelations to control the degrees of freedom in fMRI analysis. NeuroImage 26, 635–641 (2005).
Váša, F. et al. Adolescent tuning of association cortex in human structural brain networks. Cereb. Cortex 28, 281–294 (2018).
Bazinet, V., Liu, Z.-Q. & Misic, B. The effect of spherical projection on spin tests for brain maps. Imaging Neurosci. 3, IMAG.a.118 (2025).
Burt, J. B., Helmer, M., Shinn, M., Anticevic, A. & Murray, J. D. Generative modeling of brain maps with spatial autocorrelation. NeuroImage 220, 117038 (2020).
Braga, R. M. & Buckner, R. L. Parallel interdigitated distributed networks within the individual estimated by intrinsic functional connectivity. Neuron 95, 457–471 (2017).
Yeo, B. T. T., Krienen, F. M., Chee, M. W. L. & Buckner, R. L. Estimates of segregation and overlap of functional connectivity networks in the human cerebral cortex. NeuroImage 88, 212–227 (2014).
Goldman-Rakic, P. S. Topography of cognition: parallel distributed networks in primate association cortex. Annu. Rev. Neurosci. 11, 137–156 (1988).
Paquola, C. et al. The architecture of the human default mode network explored through cytoarchitecture, wiring and signal flow. Nat. Neurosci. 28, 654–664 (2025).
Koussis, N. C. et al. Generation of surrogate brain maps preserving spatial autocorrelation through random rotation of geometric eigenmodes. Imaging Neurosci. https://doi.org/10.1162/IMAG.a.71 (2025).
Smallwood, J. et al. The default mode network in cognition: a topographical perspective. Nat. Rev. Neurosci. 22, 503–513 (2021).
Murphy, C. et al. Modes of operation: a topographic neural gradient supporting stimulus dependent and independent cognition. NeuroImage 186, 487–496 (2019).
Mill, R. D., Ito, T. & Cole, M. W. From connectome to cognition: the search for mechanism in human functional brain networks. NeuroImage 160, 124–139 (2017).
Zhang, X.-H. et al. The cell-type underpinnings of the human functional cortical connectome. Nat. Neurosci. 28, 150–160 (2025).
Romero-Garcia, R. et al. Structural covariance networks are coupled to expression of genes enriched in supragranular layers of the human cortex. NeuroImage 171, 256–267 (2018).
Pedersen, R. et al. Dopamine D1-receptor organization contributes to functional brain architecture. J. Neurosci. 44, e0621232024 (2024).
Ricard, J. A. et al. A shared spatial topography links the functional connectome correlates of cocaine use disorder and dopamine D2/3 receptor densities. Commun. Biol. 7, 1178 (2024).
Tremblay, C. et al. Brain atrophy progression in parkinson’s disease is shaped by connectivity and local vulnerability. Brain Commun. 3, fcab269 (2021).
Grosz, M. P., Rohrer, J. M. & Thoemmes, F. The taboo against explicit causal inference in nonexperimental psychology. Perspect. Psychol. Sci. 15, 1243–1255 (2020).
Alvarez-Vargas, D. et al. Hedges, mottes, and baileys: causally ambiguous statistical language can increase perceived study quality and policy relevance. PLoS ONE 18, e0286403 (2023).
Mesulam, M.-M. From sensation to cognition. Brain 121, 1013–1052 (1998).
Kerr, N. L. HARKing: hypothesizing after the results are known. Pers. Soc. Psychol. Rev. 2, 196–217 (1998).
Baryshnikova, A. Systematic functional annotation and visualization of biological networks. Cell Syst. 2, 412–421 (2016).
Reimand, J. et al. Pathway enrichment analysis and visualization of omics data using g:profiler, GSEA, cytoscape and enrichmentMap. Nat. Protoc. 14, 482–517 (2019).
Chen, P. B. & Flint, J. What connectomics can learn from genomics. PLoS Genet. 17, e1009692 (2021).
Eickhoff, S. B., Yeo, B. T. & Genon, S. Imaging-based parcellations of the human brain. Nat. Rev. Neurosci. 19, 672–686 (2018).
Gong, Z.-Q. & Zuo, X.-N. Dark brain energy: toward an integrative model of spontaneous slow oscillations. Phys. Life Rev. 52, 278–297 (2025).
Hernán, M. A. The C-word: scientific euphemisms do not improve causal inference from observational data. Am. J. Public Health 108, 616–619 (2018).
Larsen, B. & Luna, B. Adolescence as a neurobiological critical period for the development of higher-order cognition. Neurosci. Biobehav. Rev. 94, 179–195 (2018).
Toyoizumi, T. et al. A theory of the transition to critical period plasticity: inhibition selectively suppresses spontaneous activity. Neuron 80, 51–63 (2013).
Takesian, A. E. & Hensch, T. K. in Changing Brains: Applying Brain Plasticity to Advance and Recover Human Ability (eds Merzenich, M. M., Nahum, M. & Van Vleet, T. M.) 3–34 (Elsevier, 2013).
Grydeland, H. et al. Waves of maturation and senescence in micro-structural mri markers of human cortical myelination over the lifespan. Cereb. Cortex 29, 1369–1381 (2018).
Sotiras, A. et al. Patterns of coordinated cortical remodeling during adolescence and their associations with functional specialization and evolutionary expansion. Proc. Natl Acad. Sci. USA 114, 3527–3532 (2017).
Whitaker, K. J. et al. Adolescence is associated with genomically patterned consolidation of the hubs of the human brain connectome. Proc. Natl Acad. Sci. USA 113, 9105–9110 (2016).
Baum, G. L. et al. Modular segregation of structural brain networks supports the development of executive function in youth. Curr. Biol. 27, 1561–1572 (2017).
Valk, S. L. et al. Structural plasticity of the social brain: differential change after socio-affective and cognitive mental training. Sci. Adv. 3, e1700489 (2017).
Monaghan, A. et al. Canonical neurodevelopmental trajectories of structural and functional manifolds. eLife https://doi.org/10.7554/eLife.103097.2 (2025).
Reid, A. T. et al. Advancing functional connectivity research from association to causation. Nat. Neurosci. 22, 1751–1760 (2019).
Frässle, S. et al. Regression DCM for fMRI. NeuroImage 155, 406–421 (2017).
Friston, K. J., Kahan, J., Biswal, B. & Razi, A. A DCM for resting state fMRI. NeuroImage 94, 396–407 (2014).
Rabuffo, G. et al. Mapping global brain reconfigurations following local targeted manipulations. Proc. Natl Acad. Sci. USA 122, e2405706122 (2025).
Rocchi, F. et al. Increased fMRI connectivity upon chemogenetic inhibition of the mouse prefrontal cortex. Nat. Commun. 13, 1056 (2022).
Sanz-Leon, P., Knock, S. A., Spiegler, A. & Jirsa, V. K. Mathematical framework for large-scale brain network modeling in the virtual brain. NeuroImage 111, 385–430 (2015).
Ritter, P., Schirner, M., McIntosh, A. R. & Jirsa, V. K. The virtual brain integrates computational modeling and multimodal neuroimaging. Brain Connect. 3, 121–145 (2013).
Demirtaş, M. et al. Hierarchical heterogeneity across human cortex shapes large-scale neural dynamics. Neuron 101, 1181–1194 (2019).
Sanz Leon, P. et al. The virtual brain: a simulator of primate brain network dynamics. Front. Neuroinform. https://doi.org/10.3389/fninf.2013.00010 (2013).
Zuo, X.-N., Xu, T. & Milham, M. P. Harnessing reliability for neuroscience research. Nat. Hum. Behav. 3, 768–771 (2019).
Xu, T., Kiar, G., Zuo, X.-N., Vogelstein, J. T. & Milham, M. P. Challenges in measuring individual differences of brain function. Imaging Neurosci. https://doi.org/10.1162/imag_a_00430 (2025).
Wilkinson, M. D. et al. The FAIR guiding principles for scientific data management and stewardship. Sci. Data 3, 160018 (2016).
Gorgolewski, K. J. et al. The brain imaging data structure, a format for organizing and describing outputs of neuroimaging experiments. Sci. Data 3, 160044 (2016).
Poldrack, R. A. et al. The past, present, and future of the brain imaging data structure (BIDS). Imaging Neurosci. 2, 1–19 (2024).
Zuo, X.-N. et al. An open science resource for establishing reliability and reproducibility in functional connectomics. Sci. Data 1, 140049 (2014).
Dhamala, E. et al. Considering the interconnected nature of social identities in neuroimaging research. Nat. Neurosci. 28, 222–233 (2025).
Besharati, S. & Akinyemi, R. Accelerating African neuroscience to provide an equitable framework using perspectives from West and Southern Africa. Nat. Commun. 14, 8107 (2023).
Henrich, J., Heine, S. J. & Norenzayan, A. Most people are not WEIRD. Nature 466, 29 (2010).
Liu, S. et al. Chinese color nest project : an accelerated longitudinal brain-mind cohort. Dev. Cogn. Neurosci. 52, 101020 (2021).
Zhang, Y. et al. The consortium on vulnerability to externalizing disorders and addictions (c-VEDA): an accelerated longitudinal cohort of children and adolescents in India. Mol. Psychiatry 25, 1618–1630 (2020).
Salum, G. A. et al. Cohort profile: brazilian high-risk cohort for mental health conditions (BHRC). Int. J. Epidemiol. 54, dyaf192 (2025).
Cooley, C. Z. et al. A portable scanner for magnetic resonance imaging of the brain. Nat. Biomed. Eng. 5, 229–239 (2021).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Gopinath, K. et al. Synthetic data in generalizable, learning-based neuroimaging. Imaging Neurosci. 2, 1–22 (2024).
Kropp, J. O., Schiffer, C., Amunts, K. & Dickscheid, T. in 2024 IEEE International Symposium on Biomedical Imaging (ISBI) 1–5 (IEEE, 2024).
Vaden, K. I., Gebregziabher, M., Dyslexia Data, C. & Eckert, M. A. Fully synthetic neuroimaging data for replication and exploration. NeuroImage 223, 117284 (2020).
Steeg, K. et al. Re-identification of anonymised MRI head images with publicly available software: investigation of the current risk to patient privacy. eClinicalMedicine 78, 102930 (2024).
Dai, Y. et al. Leveraging deep learning to enhance MRI for brain disorders. Preprint at medRxiv https://doi.org/10.1101/2025.02.10.25321126 (2025).
Gicquel, M. et al. Converting T1-weighted MRI from 3T to 7T quality using deep learning. Preprint at https://arxiv.org/abs/2507.13782 (2025).
Iglesias, J. E. et al. Quantitative brain morphometry of portable low-field-strength MRI using super-resolution machine learning. Radiology 306, e220522 (2022).
Lau, V. et al. Pushing the limits of low-cost ultra-low-field MRI by dual-acquisition deep learning 3D superresolution. Magn. Reson. Med. 90, 400–416 (2023).
Dai, X. et al. Multimodal MRI synthesis using unified generative adversarial networks. Med. Phys. 47, 6343–6354 (2020).
Royer, J. et al. Human cortical dynamics reflect graded contributions of local geometry and network topography. Nat. Commun. 17, 1002 (2025).
Wagstyl, K. et al. Transcriptional cartography integrates multiscale biology of the human cortex. eLife 12, RP86933 (2024).
Cabalo, D. G. et al. Multimodal precision MRI of the individual human brain at ultra-high fields. Sci. Data 12, 526 (2025).
Cruces, R. R. et al. Micapipe: a pipeline for multimodal neuroimaging and connectome analysis. NeuroImage 263, 119612 (2022).
Tustison, N. J. & Avants, B. B. Explicit B-spline regularization in diffeomorphic image registration. Front. Neuroinform. 7, 39 (2013).
Fischl, B., Sereno, M. I. & Dale, A. M. Cortical surface-based analysis: II: inflation, flattening, and a surface-based coordinate system. NeuroImage 9, 195–207 (1999).
Dale, A. M., Fischl, B. & Sereno, M. I. Cortical surface-based analysis: I. Segmentation and surface reconstruction. NeuroImage 9, 179–194 (1999).
Henschel, L. et al. FastSurfer — a fast and accurate deep learning based neuroimaging pipeline. NeuroImage 219, 117012 (2020).
French, L. & Paus, T. A FreeSurfer view of the cortical transcriptome generated from the Allen Human Brain atlas. Front. Neurosci. 9, 323 (2015).
van den Akker, O. R. et al. Preregistration in practice: a comparison of preregistered and non-preregistered studies in psychology. Behav. Res. Methods 56, 5424–5433 (2024).
Scheel, A. M., Schijen, M. R. M. J. & Lakens, D. An excess of positive results: comparing the standard psychology literature with registered reports. Adv. Methods Pract. Psychol. Sci. 4, 25152459211007467 (2021).
Acknowledgements
J.R. received support from the Canadian Institutes of Health Research (CIHR Fellowship, PhD) and a Banting Postdoctoral Fellowship from the National Science and Engineering Research Council of Canada (NSERC). C.P. is supported by the Deutsche Forschungsgemeinschaft (DFG Emmy Noether Programme 524408221). S.L. acknowledges research support from the Centre de Recherche du CHUS, Molson Foundation and the National Science and Engineering Research Council of Canada (NSERC Discovery RGPIN-2025-06138). B.C.B. acknowledges research support from the National Science and Engineering Research Council of Canada (NSERC RGPIN-2025-05932), CIHR (FDN-154298, PJT-174995, PJT-191853 and PJT-203761), SickKids Foundation (NI17-039), Helmholtz International BigBrain Analytics and Learning Laboratory (HIBALL), HBHL, Brain Canada Foundation, FRQS, Tier-2 Canada Research Chairs Program and The Centre for Excellence in Epilepsy at the Neuro (CEEN).
Author information
Authors and Affiliations
Contributions
J.R. and B.C.B. researched data for the article. J.R., B.C.B, C.P., S.L. and R.L. contributed substantially to the discussion of the content and wrote the article. All authors reviewed and/or edited the manuscript before submission.
Corresponding authors
Ethics declarations
Competing interests
B.C.B. is co-founder of BrainScores Inc. The other authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Neuroscience thanks Mike Hawrylycz, Franco Pestilli and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Glossary
- Brain cartography
-
Analogous to geographic cartography mapping terrain, brain cartography refers to efforts to deliver a detailed mapping of the brain’s anatomy and function.
- Brain map
-
As the product of brain cartography, brain maps render structural or functional properties of the brain across its anatomical landscape.
- Effective connectivity
-
In the context of neuroimaging, effective connectivity aims to capture the directed, causal influence that one brain region exerts over another. This is distinct from the statistical association captured by functional connectivity, which describes the temporal co-variance or correlation between neural signals without regard to directionality. Effective connectivity models the direction and magnitude of information flow and is therefore model dependent: estimates are derived by fitting a generative or causal model to observed data and inferring the underlying circuit architecture that best explains the measured activity. Common analytical approaches include dynamic causal modelling and Granger causality analysis.
- Functional connectivity
-
Statistical relationships (such as correlated or coherent fluctuations) between neural activity time series from different regions, commonly estimated with functional MRI, electroencephalography or magnetoencephalography; these relationships do not imply direct structural connections or causality.
- HARKing
-
A questionable research practice consisting of hypothesizing after the results are known.
- Interpolation
-
When using sparsely sampled data (for example, transcriptomic samples or intracranial electroencephalography recordings), values in an unsampled location may be estimated by generalizing from spatially neighbouring data points. This interpolation can be necessary in certain analytical pipelines when sparse data is contextualized against more densely sampled brain maps (for example, collected using MRI).
- Multivariate dominance analysis
-
A statistical method used to determine the relative importance of multiple predictor variables in explaining variance in one or more outcome variables within a multivariate regression framework.
- Multiple imputation
-
Multiple imputation is a statistical procedure for handling missing or withheld data, notably used in psychometrics and survey data analysis. It replaces each missing value with a set of m plausible values drawn from a predictive distribution conditioned on the observed data. Extensions to neuroimaging-focused applications have notably aimed to replace sensitive identifying features (for example, combinations of demographic variables that uniquely identify a participant) with statistically plausible synthetic values. Another example application has focused on generating multiple plausible completions of partially incomplete feature sets (for example, missing longitudinal timepoints).
- Sensory–association axis
-
A topographic gradient of brain organization recapitulated in several data modalities that spans from primary sensory areas to higher-order association areas, reflecting how input from the external environment is processed and integrated into abstract mental representations.
- Spatial autocorrelation
-
A property of brain maps by which data points that are proximally located in space are non-independent; that is, a correlation between spatial proximity and measurement similarity of data points making up a given brain map is observed.
- Spatial normalization
-
The process of transforming individual brain images into a common anatomical space or template, so that brain structures are aligned across different participants or scans. This allows for comparisons across individuals by accounting for differences in brain size, shape and orientation. Spatial normalization may involve linear (translations, scaling or rotations) and nonlinear (warping) transformations to match input images to a standard brain template reference.
- Stereotactic space
-
A standard three-dimensional coordinate system to precisely locate or refer to a given area of the brain.
- Structural connectivity
-
The brain’s physical network of anatomical links between regions, typically quantified in humans in vivo by mapping white matter pathways with diffusion MRI and tractography.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Royer, J., Paquola, C., Larivière, S. et al. Opportunities and pitfalls of data contextualization in neuroimaging. Nat. Rev. Neurosci. (2026). https://doi.org/10.1038/s41583-026-01038-0
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41583-026-01038-0