Abstract
Approximately 40% of the mammalian proteome undergoes N-terminal methionine excision and acetylation, mediated sequentially by methionine aminopeptidase (MetAP) and N-acetyltransferase A (NatA), respectively1. Both modifications are strictly cotranslational and essential in higher eukaryotic organisms1. The interaction, activity and regulation of these enzymes on translating ribosomes are poorly understood. Here we perform biochemical, structural and in vivo studies to demonstrate that the nascent polypeptide-associated complex2,3 (NAC) orchestrates the action of these enzymes. NAC assembles a multienzyme complex with MetAP1 and NatA early during translation and pre-positions the active sites of both enzymes for timely sequential processing of the nascent protein. NAC further releases the inhibitory interactions from the NatA regulatory protein huntingtin yeast two-hybrid protein K4,5 (HYPK) to activate NatA on the ribosome, enforcing cotranslational N-terminal acetylation. Our results provide a mechanistic model for the cotranslational processing of proteins in eukaryotic cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Models and electron microscopy maps were deposited at the PDB and EMDB with accession codes 9F1B, EMD-50124 (homogeneously refined cryo-EM map), EMD-50127 (locally refined cryo-EM map) and EMD-50130 (80S–NAC–NatA/E cryo-EM map) for the ternary complex, 9F1C, EMD-50125 (homogeneously refined cryo-EM map) and EMD-50128 (locally refined cryo-EM map) for the quaternary complex, and 9F1D, EMD-50126 (homogeneously refined cryo-EM map) and EMD-50129 (locally refined cryo-EM map) for the quaternary HYPK complex. Each EMDB entry holds either a homogeneously refined map or a locally refined map together with a set of lowpass-filtered or resampled maps presented throughout the manuscript as supplementary maps.
References
Aksnes, H., Drazic, A., Marie, M. & Arnesen, T. First things first: vital protein marks by N-terminal acetyltransferases. Trends Biochem. Sci 41, 746–760 (2016).
Wiedmann, B., Sakai, H., Davis, T. A. & Wiedmann, M. A protein complex required for signal-sequence-specific sorting and translocation. Nature 370, 434–440 (1994).
Gamerdinger, M. et al. Early scanning of nascent polypeptides inside the ribosomal tunnel by NAC. Mol. Cell 75, 996–1006.e8 (2019).
Arnesen, T. et al. The chaperone-like protein HYPK acts together with NatA in cotranslational N-terminal acetylation and prevention of Huntingtin aggregation. Mol. Cell. Biol. 30, 1898–1909 (2010).
Gottlieb, L. & Marmorstein, R. Structure of human NatA and its regulation by the huntingtin interacting protein HYPK. Structure 26, 925–935.e8 (2018).
Arnesen, T. et al. Proteomics analyses reveal the evolutionary conservation and divergence of N-terminal acetyltransferases from yeast and humans. Proc. Natl Acad. Sci. USA 106, 8157–8162 (2009).
Oh, J.-H., Hyun, J.-Y. & Varshavsky, A. Control of Hsp90 chaperone and its clients by N-terminal acetylation and the N-end rule pathway. Proc. Natl Acad. Sci. USA 114, E4370–E4379 (2017).
Monda, J. K. et al. Structural conservation of distinctive N-terminal acetylation-dependent interactions across a family of mammalian NEDD8 ligation enzymes. Structure 21, 42–53 (2013).
Scott, D. C. et al. Blocking an N-terminal acetylation-dependent protein interaction inhibits an E3 ligase. Nat. Chem. Biol. 13, 850–857 (2017).
Behnia, R., Panic, B., Whyte, J. R. C. & Munro, S. Targeting of the Arf-like GTPase Arl3p to the Golgi requires N-terminal acetylation and the membrane protein Sys1p. Nat. Cell Biol. 6, 405–413 (2004).
Hwang, C.-S., Shemorry, A. & Varshavsky, A. N-terminal acetylation of cellular proteins creates specific degradation signals. Science 327, 973–977 (2010).
Shemorry, A., Hwang, C.-S. & Varshavsky, A. Control of protein quality and stoichiometries by N-terminal acetylation and the N-end rule pathway. Mol. Cell 50, 540–551 (2013).
Gottlieb, L., Guo, L., Shorter, J. & Marmorstein, R. N-alpha-acetylation of Huntingtin protein increases its propensity to aggregate. J. Biol. Chem. 297, 101363 (2021).
Vinueza-Gavilanes, R. et al. N-terminal acetylation mutants affect alpha-synuclein stability, protein levels and neuronal toxicity. Neurobiol. Dis. 137, 104781 (2020).
Kang, L., Janowska, M. K., Moriarty, G. M. & Baum, J. Mechanistic insight into the relationship between N-terminal acetylation of α-synuclein and fibril formation rates by NMR and fluorescence. PLoS ONE 8, e75018 (2013).
Rope, A. F. et al. Using VAAST to identify an X-linked disorder resulting in lethality in male infants due to N-terminal acetyltransferase deficiency. Am. J. Hum. Genet. 89, 28–43 (2011).
Bader, I. et al. Severe syndromic ID and skewed X-inactivation in a girl with NAA10 dysfunction and a novel heterozygous de novo NAA10 p.(His16Pro) variant — a case report. BMC Med. Genet. 21, 153 (2020).
Lee, C.-F. et al. hNaa10p contributes to tumorigenesis by facilitating DNMT1-mediated tumor suppressor gene silencing. J. Clin. Invest. 120, 2920–2930 (2010).
Kim, S. M. et al. NAA10 as a new prognostic marker for cancer progression. Int. J. Mol. Sci. 21, E8010 (2020).
Bu, B. et al. N-terminal acetylation preserves α-synuclein from oligomerization by blocking intermolecular hydrogen bonds. ACS Chem. Neurosci. 8, 2145–2151 (2017).
Lima, V., de, A., do Nascimento, L. A., Eliezer, D. & Follmer, C. Role of Parkinson’s disease-linked mutations and N-terminal acetylation on the oligomerization of α-synuclein induced by 3,4-dihydroxyphenylacetaldehyde. ACS Chem. Neurosci. 10, 690–703 (2019).
Deng, S. & Marmorstein, R. Protein N-terminal acetylation: structural basis, mechanism, versatility, and regulation. Trends Biochem. Sci. 46, 15–27 (2021).
Gautschi, M. et al. The yeast Nα-acetyltransferase NatA is quantitatively anchored to the ribosome and interacts with nascent polypeptides. Mol. Cell. Biol. 23, 7403–7414 (2003).
Magin, R. S., Deng, S., Zhang, H., Cooperman, B. & Marmorstein, R. Probing the interaction between NatA and the ribosome for co-translational protein acetylation. PLoS ONE 12, e0186278 (2017).
Varland, S. & Arnesen, T. Investigating the functionality of a ribosome-binding mutant of NAA15 using Saccharomyces cerevisiae. BMC Res. Notes 11, 404 (2018).
Knorr, A. G. et al. Ribosome-NatA architecture reveals that rRNA expansion segments coordinate N-terminal acetylation. Nat. Struct. Mol. Biol. 26, 35–39 (2019).
Sandikci, A. et al. Dynamic enzyme docking to the ribosome coordinates N-terminal processing with polypeptide folding. Nat. Struct. Mol. Biol. 20, 843–850 (2013).
Mullen, J. R. et al. Identification and characterization of genes and mutants for an N-terminal acetyltransferase from yeast. EMBO J. 8, 2067–2075 (1989).
Deng, S., McTiernan, N., Wei, X., Arnesen, T. & Marmorstein, R. Molecular basis for N-terminal acetylation by human NatE and its modulation by HYPK. Nat. Commun. 11, 818 (2020).
Weyer, F. A. et al. Structural basis of HypK regulating N-terminal acetylation by the NatA complex. Nat. Commun. 8, 15726 (2017).
Miklánková, P. et al. HYPK promotes the activity of the Nα-acetyltransferase A complex to determine proteostasis of nonAc-X2/N-degron-containing proteins. Sci. Adv. 8, eabn6153 (2022).
Gong, X. et al. OsHYPK-mediated protein N-terminal acetylation coordinates plant development and abiotic stress responses in rice. Mol. Plant 15, 740–754 (2022).
Jomaa, A. et al. Mechanism of signal sequence handover from NAC to SRP on ribosomes during ER-protein targeting. Science 375, 839–844 (2022).
Gamerdinger, M. et al. NAC controls cotranslational N-terminal methionine excision in eukaryotes. Science 380, 1238–1243 (2023).
Song, D., Peng, K., Palmer, B. E. & Lee, F. S. The ribosomal chaperone NACA recruits PHD2 to cotranslationally modify HIF-α. EMBO J. 41, e112059 (2022).
Hsieh, H.-H., Lee, J. H., Chandrasekar, S. & Shan, S.-O. A ribosome-associated chaperone enables substrate triage in a cotranslational protein targeting complex. Nat. Commun. 11, 5840 (2020).
Connell, E., Darios, F., Peak-Chew, S., Soloviev, M. & Davletov, B. N-terminal acetylation of the neuronal protein SNAP-25 is revealed by the SMI81 monoclonal antibody. Biochemistry 48, 9582–9589 (2009).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Mirdita, M. et al. ColabFold: making protein folding accessible to all. Nat. Methods 19, 679–682 (2022).
Garrabrant, T. et al. Small molecule inhibitors of methionine aminopeptidase type 2 (MetAP-2) fail to inhibit endothelial cell proliferation or formation of microvessels from rat aortic rings in vitro. Angiogenesis 7, 91–96 (2004).
Yang, C.-I., Hsieh, H.-H. & Shan, S.-O. Timing and specificity of cotranslational nascent protein modification in bacteria. Proc. Natl Acad. Sci. USA 116, 23050–23060 (2019).
Chen, X. et al. Three-dimensional structure of the complexin/SNARE complex. Neuron 33, 397–409 (2002).
Ingolia, N. T., Lareau, L. F. & Weissman, J. S. Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 147, 789–802 (2011).
Kulak, N. A., Pichler, G., Paron, I., Nagaraj, N. & Mann, M. Minimal, encapsulated proteomic-sample processing applied to copy-number estimation in eukaryotic cells. Nat. Methods 11, 319–324 (2014).
Minoia, M. et al. Chp1 is a dedicated chaperone at the ribosome that safeguards eEF1A biogenesis. Nat. Commun. 15, 1382 (2024).
Liszczak, G. et al. Molecular basis for N-terminal acetylation by the heterodimeric NatA complex. Nat. Struct. Mol. Biol. 20, 1098–1105 (2013).
Yin, J., Lin, A. J., Golan, D. E. & Walsh, C. T. Site-specific protein labeling by Sfp phosphopantetheinyl transferase. Nat. Protoc. 1, 280–285 (2006).
Sharma, A., Mariappan, M., Appathurai, S. & Hegde, R. S. in Protein Secretion, Vol. 619 (ed. Economou, A.) 339–363 (Humana Press, 2010).
Yin, J. et al. Genetically encoded short peptide tag for versatile protein labeling by Sfp phosphopantetheinyl transferase. Proc. Natl Acad. Sci. USA 102, 15815–15820 (2005).
Walker, K. W. & Bradshaw, R. A. Yeast methionine aminopeptidase I. Alteration of substrate specificity by site-directed mutagenesis. J. Biol. Chem. 274, 13403–13409 (1999).
Gottlieb, L. & Marmorstein, R. Biochemical and structural analysis of N-terminal acetyltransferases. Methods Enzymol. 626, 271–299 (2019).
Jarmoskaite, I., AlSadhan, I., Vaidyanathan, P. P. & Herschlag, D. How to measure and evaluate binding affinities. eLife 9, e57264 (2020).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D 75, 861–877 (2019).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Mello, C. & Fire, A. DNA transformation. Methods Cell Biol. 48, 451–482 (1995).
Frøkjær-Jensen, C. et al. Random and targeted transgene insertion in Caenorhabditis elegans using a modified Mos1 transposon. Nat. Methods 11, 529–534 (2014).
Redemann, S. et al. Codon adaptation-based control of protein expression in C. elegans. Nat. Methods 8, 250–252 (2011).
Ketting, R. F., Tijsterman, M. & Plasterk, R. H. A. Introduction of double-stranded RNA in C. elegans by feeding. CSH Protoc. 2006, pdb.prot4317 (2006).
Acknowledgements
We thank R. Marmorstein and L. Gottlieb for expression plasmids for NatA/E and HYPK; M. Leibundgut and M. Jaskolowski for advice on model building, cryo-EM data processing and feedback on the manuscript; the ETH Scientific Center for Optical and Electron Microscopy (ScopeM) for technical support; L. Hang, G. Hunaeus and R. Schloemer for technical assistance; and members of the Shan laboratory for feedback on the manuscript. This work was supported by National Institutes of Health grant R35 GM136321 and National Science Foundation grant 2219287 to S.-o.S., the German Science Foundation grants SFB969/A01/A07 and 537004599 to E.D. and M.G., and the Swiss National Science Foundation (SNSF grant 310030_212308 to N.B.), the National Center of Excellence in Research RNA & Disease Program of the SNSF (grant 51NF40-205601 to N.B.), and the European Research Council (Synergy Grant 101072047 CoTransComplex to N.B.).
Author information
Authors and Affiliations
Contributions
A.M.L. and S.-o.S. conceived the project. A.M.L., S.-o.S., D.Y., N.B., M.G. and E.D. designed research. A.M.L. and S.C. performed biochemical experiments and analysed data, D.Y., A.S. and N.B. performed cryo-EM studies. M.G. and L.R. performed studies in C. elegans and human cells. S.-o.S., N.B. and E.D supervised the project. A.M.L. and D.Y. wrote the manuscript. S.-o.S., N.B., E.D. and M.G. revised and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewer(s) for their contribution to the peer review of this work. Peer review reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 FRET of NatA with RNC and with NAC on the ribosome.
(a-c) Scheme of the nascent chain constructs used in this work. Grey depicts the ribosome exit tunnel up to the PTC. The nascent chain for FRET between RNC and NatA (a) contains an N-terminal 3xFLAG tag, SUMO, and residues 1–53 of the RpL4 nascent chain with Met1 mutated to serine. An amber codon at residue 12 allows for fluorescence labeling using BODIPY-FL (BDP). After FLAG affinity purification, SUMO cleavage by Ulp1 generates a 53 amino acid (aa) long nascent chain with a defined N-terminus recognized by NatA. Nascent chains for enzymatic assays in RRL are shown in (b). Constructs to measure iMet excision (upper) contained varying lengths of RpL4 with a single iMet, which was radioactively labeled with [35S] and detected via autoradiography. Constructs to measure Nt-acetylation (lower) contained residues 2–12 from SNAP25, which is specifically detected by the SMI antibody upon Nt-acetylation (SMI81 epitope), followed by varying lengths of the RpL4 nascent chain. The nascent chain for measurement of Nt-acetylation reconstituted with purified RNC and proteins is shown in (c) and contains an N-terminal 3xFLAG tag, SUMO, the SMI81 epitope, a GSGS linker, and residues 1-90 of the RpL4 nascent chain. After purification of RNC, SUMO cleavage by Ulp1 generates a defined N-terminus for acetylation. (d) Purified proteins (as indicated) were visualized on a 12.5% SDS-PAGE gel and Coomassie Blue staining. The dashed line indicates that the two lanes were from the same gel. Similar protein quality was observed from 2, 1, 3, and 2 independent preparations of NatA, Naa50, MetAP1, and HYPK, respectively. (e) Steady-state kinetics of WT human NatA expressed and purified from S. cerevisiae. Reactions contained the indicated concentrations of the H4 peptide substrate (SGRGKGGKGLGKGGAKRHR), 20 µM [14C]-acetyl-CoA, 80 µM [12C]-acetyl-CoA, and 10 nM NatA. The progress of the reaction was monitored in the linear range (<10% of product formation). The concentration dependence of initial rates was fit to the Michaelis-Menten equation to obtain the steady-state parameters. (f) The position of the N-terminal ybbR tag and the site of acceptor dye (TMR) label are shown on the human NatA structure (PDB: 6C9M) using a dotted line. Naa15 and Naa10 are colored in dark and light green, respectively. The figure was generated with Pymol v.2.5.5. (g) Fluorescence emission spectra of RNCBDP with the 53 aa long RpL4 nascent chain (green), NatATMR (purple), RNCBDP with NatATMR (red), and RNCBDP•NatATMR chased with excess unlabeled NatA (black). Where indicated, the reactions contained 1 nM RNCBDP, 100 nM NatATMR, and 1 µM unlabeled NatA. (h, i) Fluorescence emission spectra measuring FRET between NatABDP and NACTMR in the absence (h) or presence (i) of 5 nM unlabeled RNC with a 53 aa long RpL4 nascent chain. 5 nM NatABDP + 5 nM unlabeled NAC are shown in green, 5 nM NatABDP + 5 nM NACTMR in red, 5 nM NACTMR alone in purple, 5 nM NatABDP + 5 nM NACTMR + 250 nM unlabeled NAC in black, and buffer only in brown.
Extended Data Fig. 2 Ribosome binding of NatA in human cells depends on NAC.
(a) Ribosome association of NatA after knockdown of NACα in human HEK293T cells. Total and ribosomal pellet fractions were analyzed by immunoblotting. ns, nonsense siRNA control. Representative immunoblot is shown. Experiment was repeated three times. (b) Ribosome association of NatA after knockdown of NACα in human HEK293T cells using siRNAs targeting the endogenous NACα mRNA in the 3′ UTR. NACα expression was restored in knockdown cells by transient expression FLAG-tagged NACα variants from plasmids containing a different 3′ UTR. Two NACα variants carrying point mutations in the NACα UBA-Naa15 binding interface (mtNAC-UBA-1 and -2; see Extended Data Table 2) and a deletion mutant lacking the entire UBA domain (∆UBA; see Extended Data Table 2) were analyzed. ns, nonsense siRNA control. Representative immunoblot is shown. Experiment was repeated three times. (c) Ribosome association of FLAG-tagged Naa15 variants in human HEK293T cells in the endogenous Naa15 knockdown background. A mutant variant carrying three point mutations in the hydrophobic NACα UBA binding interface (L73A/L77A/W83A) was compared with wildtype Naa15. Total and ribosomal pellet fractions were analyzed by immunoblotting. Representative immunoblot is shown. Experiment was repeated three times.
Extended Data Fig. 3 Local resolution estimates, orientation distribution and refinement statistics of cryo-EM maps.
Each row shows, from the left to right: an overview and slice-through of the cryo-EM map, both filtered and coloured according to estimated local resolution, a viewing direction distribution heatmap, and Fourier shell correlations (between unmasked or masked half maps – blue and purple curves, respectively; between the map and the model after the final round of refinement – orange curves). The local resolution estimation and filtering was done in CryoSPARC. Maps were coloured on the same scale from 2.5 to 10 Å. The viewing direction distribution for each map was taken from corresponding CryoSPARC refinement outputs. For locally refined maps, only areas within refinement masks were coloured. The FSC curves for maps and map vs. model were obtained from outputs of map refinements in CryoSPARC and real-space refinements of models in Phenix, respectively.
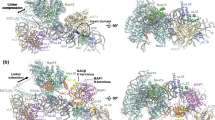
Extended Data Fig. 4 Cryo-EM structure and model of the RNCRpL4-NAC-NatA/E ternary complex.
(a) Front view of the cryo-EM map of RNCRpL4 bound with NAC and NatA/E. The map shows the NatA/E complex bound to the large ribosomal subunit next to the globular domain of NAC. The map was lowpass-filtered to the estimated local resolution. The black outline shows the same map lowpass-filtered to 8 Å resolution. (b) Detailed view of the ternary complex model depicted in (A) showing NatA/E and NAC on the 60 S ribosomal subunit. The yellow dotted line represents the flexible linker between NACα helix H2 and the UBA domain. NatA/E contacts the ribosome through its Naa15 auxiliary subunit via patches of positively charged residues within the basic helix of Naa15 and N-terminal helices that bind to the backbone of ribosomal RNA. The NACα C-terminal tail forms a bipartite contact with Naa15 with its UBA domain and H2. (c-e) Details of cryo-EM maps of the ternary complex showing a segment of Naa15 in contact with the NACα UBA domain (c), the NACα-H2 density connecting to the globular domain of NAC (d), and the N-terminal ribosome anchor of NACβ (e). The locally refined map (c), the homogeneously refined map lowpass filtered to 6 Å resolution (d) and the homogeneously refined map (e) are displayed as semi-transparent surfaces. (f-h) ColabFold-predicted dimer model of the Naa15-NACα complex. The AlphaFold model is coloured by chain (f), with Naa15 in green and NACα in yellow, and by the pLDDT confidence score (g). (h) shows the predicted Aligned Error (PAE) plot for the predicted model, with Naa15 as chain A and NACα as chain B. The chain residues are marked along the x and y axis, and the heatmap indicates the estimated position error (in Å) for residue x when predicted and true structures are aligned on residue y. Low PAE for residue pairs from NACα UBA and Naa15 (marked on the plot with *) and NACα H2 and Naa15 (marked as **) indicate that the relative positions of corresponding domains are well-defined in the AlphaFold prediction.
Extended Data Fig. 5 Mammals and yeast employ distinct modes of NatA/E recruitment to ribosomes.
Side (a, b) and top (c, d) views of the mammalian ternary RNCRpL4-NAC-NatA/E complex (a, c) and those of the yeast RNC-NatA/E complex (b, d; PDB# 6HD5). The ribosomes are shown in surface representation, bound factors and selected expansion segments in cartoon. The NAC heterodimer is shown in yellow and orange, mammalian NatA/E in hues of green, rRNA expansion segments coordinating NatA/E in blue, and yeast NatA/E in hues of pink. While the N-terminal helical domain of Naa15 contacts the ribosome near the exit tunnel in both mammalian and yeast systems, all other structural elements that mediate NatA recruitment and positioning on the ribosome are distinct in the two organisms. Firstly, ribosome-bound NAC captures and helps position NatA/E in mammals, whereas in yeast, the rRNA extension Es27 is proposed to act as a protein recruitment hub for NatA in place of NAC. Secondly, the second catalytic subunit Naa50 mediates an additional contact with the rRNA extionsion Es7a on the yeast ribosome, whereas Naa50 is not involved in ribosome contact and hence does not contribute to the ribosome affinity of mammalian NatA/E as shown in Fig. 1. Thirdly, the locations of the NatA/E complex at the ribosome exit site are distinct in the two organisms. Finally, the ribosome binding site of mammalian NAC heavily overlaps with that of yeast NatA/E, suggesting that yeast NAC antagonizes rather than facilitates the ribosome recruitment of NatA/E. These differences, together with the absence of the NatA regulator HYPK in S. cerevisiae, suggest that the ribosome recruitment mechanisms for protein biogenesis factors are distinct between yeast and higher eukaryotic organisms.
Extended Data Fig. 6 Equilibrium titrations to measure the binding of NatA to RNC•NAC with the indicated NAC or NatA variants.
(a-c) Titrations contained 1 nM RNCBDP, indicated concentrations of WT NatATMR or NatATMR variants (a, b), and 50 nM NAC WT or NAC variants (c). (d) Close-up view of the interaction between Naa15 TPR and the NACα UBA domain. Mutated residues are colored as in (b) and (c).
Extended Data Fig. 7 HYPK forms an ultra-stable complex with NatA.
(a) Fluorescence emission spectra of 5 nM NatABDP with (red) and without (green) 50 nM HYPKTMR. (b) Equilibrium titrations to measure the binding of NatABDP to HYPKTMR. Data are shown as mean ± SD, with n = 3 independent measurements. (c) FRET between NatA-HYPK was not changed by increasing concentrations of the 80 S ribosome. (d) Kinetics of dissociation of NatA-HYPK. The line is a fit of the data to a single exponential function assuming FRET = 0 at the end of the reaction. (e) Summary of the dissociation rate constant of the NatA-HYPK complex, in the absence and presence of the additional Naa50 catalytic subunit or by increasing the concentration of the ribosome high-salt wash fraction (HSW). Individual data points for independent measurements are shown.
Extended Data Fig. 8 Structural modeling and cryoEM analysis of HYPK bound to NatA on the RNC-NAC complex.
(a-d) HYPK H2 in inhibitory conformation would clash with bound H2 of NACα. In (a), the model of HYPK from NatA-HYPK complex (PDB 6C95) is superimposed on the model of NatA/E from the quaternary RNC-NAC-NatA/E-MetAP1 complex. Models were aligned by Naa15, only HYPK from the isolated NatA-HYPK complex is displayed. (b) shows the model of NAC-NatA/E from the quaternary RNC-NAC-NatA/E-MetAP1 complex. (c) shows the model of HYPK from NatA-HYPK complex (PDB 6C95) superimposed on the model of NAC-NatA/E from the quaternary RNC-NAC-NatA/E-MetAP1 complex. The binding site of HYPK H2 in its inhibitory conformation on Naa15 in part overlaps with that of H2 of NACα, when NatA/E is recruited to the ribosome. (d) shows a model of NatA/E-HYPK-NAC from the quaternary complex with HYPK. The N terminus of HYPK is remodelled upon recruitment to the ribosome by NAC. The flexible linker connecting NACα H2 and UBA and displaced HYPK N terminus are shown as yellow and purple dotted lines, respectively. (e-j) Comparison between cryo-EM maps of quaternary complexes with and without HYPK. e-g, details of the locally refined cryo-EM map of the quaternary complex showing the front view of the quaternary complex (e), a segment of Naa15 with NACα UBA (f), and MetAP1 (g). h-j, details of the locally refined cryo-EM map of the quaternary complex with HYPK showing the front view of the complex (h) and a segment of Naa15 with NACα UBA and HYPK UBA (i). A detail of a homogeneously refined map, lowpass filtered to 6 Å resolution, shows how NACα-H2 connects to the globular domain of NAC (j). The maps are displayed as solid surfaces (e, h) or as semi-transparent surfaces superimposed on models of the complexes (f-g, i-j). Colors are the same as in Fig. 1e, with MetAP1 in blue-grey and HYPK in purple.
Supplementary information
Supplementary Information (download PDF )
This file contains Supplementary Tables 2 and 3, and Supplementary Figs. 2 and 3.
Supplementary Figure 1 (download JPG )
Uncropped gels, western blots, and autoradiographs.
Supplementary Table 1 (download XLSX )
Original data points for all figures.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lentzsch, A.M., Yudin, D., Gamerdinger, M. et al. NAC guides a ribosomal multienzyme complex for nascent protein processing. Nature 633, 718–724 (2024). https://doi.org/10.1038/s41586-024-07846-7
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-024-07846-7
This article is cited by
-
NatA engages in multi-factor complexes at the ribosomal polypeptide tunnel exit
Nature Communications (2026)
-
Structural basis of co-translational N-myristoylation in humans
Nature Communications (2026)
-
Illuminating the impact of N-terminal acetylation: from protein to physiology
Nature Communications (2025)
-
Engineering a membrane protein chaperone to ameliorate the proteotoxicity of mutant huntingtin
Nature Communications (2025)
-
NAC couples protein synthesis with nascent polypeptide myristoylation on the ribosome
The EMBO Journal (2025)