Abstract
Non-ribosomal peptide synthetases (NRPSs) are megaenzymes responsible for the biosynthesis of many clinically important natural products, from early modern medicines (penicillin, bacitracin) to current blockbuster drugs (cubicin, vancomycin) and newly approved therapeutics (rezafungin)1,2. The key chemical step in these biosyntheses is amide bond formation between aminoacyl building blocks, catalysed by the condensation (C) domain3. There has been much debate over the mechanism of this reaction3,4,5,6,7,8,9,10,11,12. NRPS condensation has been difficult to fully characterize because it is one of many successive reactions in the NRPS synthetic cycle and because the canonical substrates are each attached transiently as thioesters to mobile carrier domains, which are often both contained in the same very flexible protein as the C domain. Here we have produced a dimodular NRPS protein in two parts, modified each with appropriate non-hydrolysable substrate analogues13,14, assembled the two parts with protein ligation15, and solved the structures of the substrate- and product-bound states. The structures show the precise orientation of the megaenzyme preparing the nucleophilic attack of its key chemical step, and enable biochemical assays and quantum mechanical simulations to precisely interrogate the reaction. These data suggest that NRPS C domains use a concerted reaction mechanism, whereby the active-site histidine likely functions not as a general base, but as a crucial stabilizing hydrogen bond acceptor for the developing ammonium.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
X-ray crystal structures and associated diffraction data for this study are deposited under accession codes 9BE3 and 9BE4 in the Research Collaboratory for Structural Bioinformatics Protein Data Bank. Source data are provided with this paper.
References
Sieber, S. A. & Marahiel, M. A. Molecular mechanisms underlying nonribosomal peptide synthesis: Approaches to new antibiotics. Chem. Rev. 105, 715–738 (2005).
Hüttel, W. Echinocandins: structural diversity, biosynthesis, and development of antimycotics. Appl. Microbiol. Biotechnol. 105, 55–66 (2020).
Keating, T. A., Marshall, C. G., Walsh, C. T. & Keating, A. E. The structure of VibH represents nonribosomal peptide synthetase condensation, cyclization and epimerization domains. Nat. Struct. Biol. 9, 522–526 (2002).
Izore, T. et al. Structures of a non-ribosomal peptide synthetase condensation domain suggest the basis of substrate selectivity. Nat. Commun. 12, 2511 (2021).
Roche, E. D. & Walsh, C. T. Dissection of the EntF condensation domain boundary and active site residues in nonribosomal peptide synthesis. Biochemistry 42, 1334–1344 (2003).
Bergendahl, V., Linne, U. & Marahiel, M. A. Mutational analysis of the C-domain in nonribosomal peptide synthesis. Eur. J. Biochem. 269, 620–629 (2002).
Marshall, C. G., Hillson, N. J. & Walsh, C. T. Catalytic mapping of the vibriobactin biosynthetic enzyme VibF. Biochemistry 41, 244–250 (2002).
Zhong, L. et al. Engineering and elucidation of the lipoinitiation process in nonribosomal peptide biosynthesis. Nat. Commun. 12, 296 (2021).
Bloudoff, K., Alonzo, D. A. & Schmeing, T. M. Chemical probes allow structural insight into the condensation reaction of nonribosomal peptide synthetases. Cell Chem. Biol. 23, 331–339 (2016).
Samel, S. A., Schoenafinger, G., Knappe, T. A., Marahiel, M. A. & Essen, L. O. Structural and functional insights into a peptide bond-forming bidomain from a nonribosomal peptide synthetase. Structure 15, 781–792 (2007).
Kee, M.-J. C. Y. et al. Structural insights into the substrate-bound condensation domains of non-ribosomal peptide synthetase AmbB. Sci. Rep. 12, 5353 (2022).
Zhang, J. et al. Structural basis of nonribosomal peptide macrocyclization in fungi. Nat. Chem. Biol. 12, 1001–1003 (2016).
Reimer, J. M., Aloise, M. N., Harrison, P. M. & Schmeing, T. M. Synthetic cycle of the initiation module of a formylating nonribosomal peptide synthetase. Nature 529, 239–242 (2016).
Reimer, J. M. et al. Structures of a dimodular nonribosomal peptide synthetase reveal conformational flexibility. Science 366, 6466 (2019).
Yang, R. et al. Engineering a catalytically efficient recombinant protein ligase. J. Am. Chem. Soc. 139, 5351–5358 (2017).
Quadri, L. E. et al. Characterization of Sfp, a Bacillus subtilis phosphopantetheinyl transferase for peptidyl carrier protein domains in peptide synthetases. Biochemistry 37, 1585–1595 (1998).
Crosby, J. & Crump, M. P. The structural role of the carrier protein—active controller or passive carrier. Nat. Prod. Rep. 29, 1111–1137 (2012).
Bloudoff, K. & Schmeing, T. M. Structural and functional aspects of the nonribosomal peptide synthetase condensation domain superfamily: discovery, dissection and diversity. Biochim. Biophys. Acta 1865, 1587–1604 (2017).
Maruyama, C. et al. A stand-alone adenylation domain forms amide bonds in streptothricin biosynthesis. Nat. Chem. Biol. 8, 791–797 (2012).
Fortinez, C. M. et al. Structures and function of a tailoring oxidase in complex with a nonribosomal peptide synthetase module. Nat. Commun. 13, 548548 (2022).
Wang, J. et al. Catalytic trajectory of a dimeric nonribosomal peptide synthetase subunit with an inserted epimerase domain. Nat. Commun. 13, 592 (2022).
Katsuyama, Y. et al. Structural and functional analyses of the tridomain-nonribosomal peptide synthetase FmoA3 for 4-methyloxazoline ring formation. Angew. Chem. Int. Ed. Engl. 60, 14554–14562 (2021).
Bloudoff, K., Rodionov, D. & Schmeing, T. M. Crystal structures of the first condensation domain of CDA synthetase suggest conformational changes during the synthetic cycle of nonribosomal peptide synthetases. J. Mol. Biol. 425, 3137–3150 (2013).
Chang, C. Y. et al. Structural insights into the free-standing condensation enzyme SgcC5 catalyzing ester-bond formation in the biosynthesis of the enediyne antitumor antibiotic C-1027. Biochemistry 57, 3278–3288 (2018).
Drake, E. J. et al. Structures of two distinct conformations of holo-non-ribosomal peptide synthetases. Nature 529, 235–238 (2016).
Tanovic, A., Samel, S. A., Essen, L. O. & Marahiel, M. A. Crystal structure of the termination module of a nonribosomal peptide synthetase. Science 321, 659–663 (2008).
Miller, B. R., Drake, E. J., Shi, C., Aldrich, C. C. & Gulick, A. M. Structures of a nonribosomal peptide synthetase module bound to MbtH-like proteins support a highly dynamic domain architecture. J. Biol. Chem. 291, 22559–22571 (2016).
Kreitler, D. F., Gemmell, E. M., Schaffer, J. E., Wencewicz, T. A. & Gulick, A. M. The structural basis of N-acyl-α-amino-β-lactone formation catalyzed by a nonribosomal peptide synthetase. Nat. Commun. 10, 3432 (2019).
Tarry, M. J., Haque, A. S., Bui, K. H. & Schmeing, T. M. X-ray crystallography and electron microscopy of cross- and multi-module nonribosomal peptide synthetase proteins reveal a flexible architecture. Structure 25, 783–793 (2017).
Ho, Y. T. C. et al. Exploring the selectivity and engineering potential of an NRPS condensation domain involved in the biosynthesis of the thermophilic siderophore fuscachelin. Front. Catal. 3, 1184959 (2023).
Wheadon, M. J. & Townsend, C. A. Accurate substrate-like probes for trapping late-stage intermediates in nonribosomal peptide synthetase condensation domains. ACS Chem. Biol. 17, 2046–2053 (2022).
Worthington, A. S. & Burkart, M. D. One-pot chemo-enzymatic synthesis of reporter-modified proteins. Org. Biomol. Chem. 4, 44–46 (2006).
Mao, H., Hart, S. A., Schink, A. & Pollok, B. A. Sortase-mediated protein ligation: a new method for protein engineering. J. Am. Chem. Soc. 126, 2670–2671 (2004).
Chikunova, A. & Ubbink, M. The roles of highly conserved, non-catalytic residues in class A β-lactamases. Protein Sci. 31, e4328 (2022).
Zhang, F. et al. Modulating the pH activity profiles of phenylalanine ammonia lyase from Anabaena variabilis by modification of center-near surface residues. Appl. Biochem. Biotechnol. 183, 699–711 (2017).
Elfstrom, L. T. & Widersten, M. Catalysis of potato epoxide hydrolase, StEH1. Biochem. J. 390, 633–640 (2005).
Tantillo, D. J., Chen, J. & Houk, K. N. Theozymes and compuzymes: theoretical models for biological catalysis. Curr. Opin. Chem. Biol. 2, 743–750 (1998).
Folger, I. B. et al. High-throughput reprogramming of an NRPS condensation domain. Nat. Chem. Biol. 20, 761–769 (2024).
Shi, C., Miller, B. R., Alexander, E. M., Gullick, A. M. & Aldrich, C. C. Design, synthesis, and biophysical evaluation of mechanism-based probes for condensation domains of nonribosomal peptide synthetases. ACS Chem. Biol. 15, 1813–1819 (2020).
Ulrich, V. & Cryle, M. J. SNaPe: a versatile method to generate multiplexed protein fusions using synthetic linker peptides for in vitro applications. J. Pept. Sci. 23, 16–27 (2017).
Kosol, S. et al. Structural basis for chain release from the enacyloxin polyketide synthase. Nat. Chem. 11, 913–923 (2019).
Patel, K. D., MacDonald, M. R., Ahmed, S. F., Singh, J. & Gulick, A. M. Structural advances toward understanding the catalytic activity and conformational dynamics of modular nonribosomal peptide synthetases. Nat. Prod. Rep. 40, 1550–1582 (2023).
Haslinger, K., Peschke, M., Brieke, C., Maximowitsch, E. & Cryle, M. J. X-domain of peptide synthetases recruits oxygenases crucial for glycopeptide biosynthesis. Nature 521, 105–109 (2015).
Williams, A. Concerted mechanisms of acyl group transfer-reactions in solution. Accounts Chem. Res. 22, 387–392 (1989).
Guthrie, J. P. & Pike, D. C. Hydration of acylimidazoles: tetrahedral intermediates in acylimidazole hydrolysis and nucleophilic-attack by imidazole on esters. The question of concerted mechanisms for acyl transfers. Can. J. Chem. 65, 1951–1969 (1987).
Guthrie, J. P. Concerted mechanism for alcoholysis of esters—an examination of the requirements. J. Am. Chem. Soc. 113, 3941–3949 (1991).
Hengge, A. C. & Hess, R. A. Concerted or stepwise mechanisms for acyl transfer-reactions of p-nitrophenyl acetate—transition-state structures from isotope effects. J. Am. Chem. Soc. 116, 11256–11263 (1994).
Blake, J. F. & Jorgensen, W. L. Ab initio study of the displacement reactions of chloride ion with formyl and acetyl chloride. J. Am. Chem. Soc. 109, 3856–3861 (1987).
Kwan, E. E., Zeng, Y. W., Besser, H. A. & Jacobsen, E. N. Concerted nucleophilic aromatic substitutions. Nat. Chem. 10, 917–923 (2018).
Bürgi, H. B., Dunitz, J. D., Lehn, J. M. & Wipff, G. Stereochemistry of reaction paths at carbonyl centres. Tetrahedron 30, 1563–1572 (1974).
Kessler, N., Schuhmann, H., Morneweg, S., Linne, U. & Marahiel, M. A. The linear pentadecapeptide gramicidin is assembled by four multimodular nonribosomal peptide synthetases that comprise 16 modules with 56 catalytic domains. J. Biol. Chem. 279, 7413–7419 (2004).
Chalut, C., Botella, L., de Sousa-D’Auria, C., Houssin, C. & Guilhot, C. The nonredundant roles of two 4′-phosphopantetheinyl transferases in vital processes of Mycobacteria. Proc. Natl Acad. Sci. USA 103, 8511–8516 (2006).
Pfeifer, B. A., Admiraal, S. J., Gramajo, H., Cane, D. E. & Khosla, C. Biosynthesis of complex polyketides in a metabolically engineered strain of E. coli. Science 291, 1790–1792 (2001).
Phan, J. et al. Structural basis for the substrate specificity of tobacco etch virus protease. J. Biol. Chem. 277, 50564–50572 (2002).
Nazi, I., Koteva, K. P. & Wright, G. D. One-pot chemoenzymatic preparation of coenzyme A analogues. Anal. Biochem. 324, 100–105 (2004).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods. Enzymol. 276, 307–326 (1997).
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Karplus, P. A. & Diederichs, K. Linking crystallographic model and data quality. Science 336, 1030–1033 (2012).
Bulut, H. et al. Crystal structures of receptors involved in small molecule transport across membranes. Eur. J. Cell Biol. 91, 318–325 (2012).
Berman, H. M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Bloudoff, K., Fage, C. D., Marahiel, M. A. & Schmeing, T. M. Structural and mutational analysis of the nonribosomal peptide synthetase heterocyclization domain provides insight into catalysis. Proc. Natl Acad. Sci. USA 114, 95–100 (2017).
Rausch, C., Hoof, I., Weber, T., Wohlleben, W. & Huson, D. H. Phylogenetic analysis of condensation domains in NRPS sheds light on their functional evolution. BMC Evol. Biol. 7, 78 (2007).
Schafer, A., Horn, H. & Ahlrichs, R. Fully optimized contracted Gaussian-basis sets for atoms Li to Kr. J. Chem. Phys. 97, 2571–2577 (1992).
Scalmani, G. & Frisch, M. J. Continuous surface charge polarizable continuum models of solvation. I. General formalism. J. Chem. Phys. 132, 114110 (2010).
Maier, J. A. et al. ff14SB: Improving the accuracy of protein side chain and backbone parameters from ff99SB. J. Chem. Theory Comput. 11, 3696–3713 (2015).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Wang, J. M., Wolf, R. M., Caldwell, J. W., Kollman, P. A. & Case, D. A. Development and testing of a general amber force field. J. Comput. Chem. 25, 1157–1174 (2004).
Bayly, C. I., Cieplak, P., Cornell, W. D. & Kollman, P. A. A well-behaved electrostatic potential based method using charge restraints for deriving atomic charges—the Resp model. J. Phys. Chem. 97, 10269–10280 (1993).
Besler, B. H., Merz, K. M. & Kollman, P. A. Atomic charges derived from semiempirical methods. J. Comput. Chem. 11, 431–439 (1990).
Singh, U. C. & Kollman, P. A. An approach to computing electrostatic charges for molecules. J. Comput. Chem. 5, 129–145 (1984).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald—an N⋅log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Götz, A. W. et al. Routine microsecond molecular dynamics simulations with AMBER on GPUs. 1. Generalized born. J. Chem. Theory Comput. 8, 1542–1555 (2012).
Reimer, J. M., Haque, A. S., Tarry, M. J. & Schmeing, T. M. Piecing together nonribosomal peptide synthesis. Curr. Opin. Struct. Biol. 49, 104–113 (2018).
Reimer, J. M., Aloise, M. N., Powell, H. R. & Schmeing, T. M. Manipulation of an existing crystal form unexpectedly results in interwoven packing networks with pseudo-translational symmetry. Acta Crystallogr. D 72, 1130–1136 (2016).
Chen, W.-H., Li, K., Guntaka, N. S. & Bruner, S. D. Interdomain and intermodule organization in epimerization domain containing nonribosomal peptide synthetases. ACS Chem. Biol. 11, 2293–2303 (2016).
Liu, Y., Zheng, T. & Bruner, S. D. Structural basis for phosphopantetheinyl carrier domain interactions in the terminal module of nonribosomal peptide synthetases. Chem. Biol. 18, 1482–1488 (2011).
Gahloth, D. et al. Structures of carboxylic acid reductase reveal domain dynamics underlying catalysis. Nat. Chem. Biol. 13, 975–981 (2017).
Deshpande, S., Altermann, E., Sarojini, V., Lott, J. S. & Lee, T. V. Structural characterization of a PCP–R di-domain from an archaeal non-ribosomal peptide synthetase reveals novel inter-domain interactions. J. Biol. Chem. 296, 100432 (2021).
Bozhuyuk, K. A. J. et al. Modification and de novo design of non-ribosomal peptide synthetases using specific assembly points within condensation domains. Nat. Chem. 11, 653–661 (2019).
Nguyen, K. T. et al. Combinatorial biosynthesis of novel antibiotics related to daptomycin. Proc. Natl Acad. Sci. USA 103, 17462–17467 (2006).
Winn, M., Fyans, J. K., Zhuo, Y. & Micklefield, J. Recent advances in engineering nonribosomal peptide assembly lines. Nat. Prod. Rep. 33, 317–347 (2016).
Bozhuyuk, K. A. J. et al. De novo design and engineering of non-ribosomal peptide synthetases. Nat. Chem. 10, 275–281 (2018).
Bozhuyuk, K. A. J. et al. Evolution-inspired engineering of nonribosomal peptide synthetases. Science 383, eadg4320 (2024).
Mansour, B. & Gauld, J. W. Computational insights into amide bond formation catalyzed by the condensation domain of nonribosomal peptide synthetases. ACS Omega 9, 28556–28563 (2024).
Acknowledgements
This research was funded by grants to T.M.S. from Canadian Institutes for Health Research (grant PJT-178084) and to K.N.H. from the US National Science Foundation (CHE-2153972). P.M. acknowledges the National Natural Science Foundation of China (no. 22103060). Infrastructure at the McGill University Centre de Recherche en Biologie Strucutrale, used in this research, is supported by Fonds de Recherche du Québec (Health Sector) Research Centres grant 288558. We thank Q. Zhou for efforts with quantum mechanics/molecular mechanics; M. Eivaskhani, J. Reimer, A. Mantri, N. Frota and the other members of the Schmeing laboratory, K. Auclair, A. Berghuis, T. Sprules, N. Rogerson, G. Challis and A. Guarné for advice and discussions; A. Mittermaier for advice on statistics; and CLS beamline staff for facilitating data collection. The X-ray diffraction data collection described in this paper was performed using beamline CMCF-ID at the Canadian Light Source, a national research facility of the University of Saskatchewan, which is supported by the Canada Foundation for Innovation, the Natural Sciences and Engineering Research Council, the National Research Council, the Canadian Institutes of Health Research, the Government of Saskatchewan and the University of Saskatchewan. We thank C. Chalut and C. Guilhot for gift of E. coli BL21 (DE3) entD- cells. AEP1-encoding plasmid was provided by W. Bin.
Author information
Authors and Affiliations
Contributions
A.P. and T.M.S. designed, and A.P. performed, all chemistry, biochemistry and crystallography. K.M. performed CD experiments. K.N.H. and P.M. designed, and P.M. and Z.L. performed, computational simulations. T.M.S. supervised A.P.; K.N.H. supervised P.M.; and P.M. supervised Z.L. T.M.S., A.P. and K.N.H. wrote the manuscript with input from all of the authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Adam Balutowski, Andrew Gulick, Shiou-Chuan Tsai and Timothy Wencewicz for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Nonribosomal peptides and the NRPS synthetic cycle.
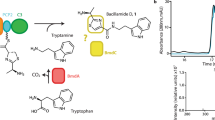
a. Representative therapeutics fully made by NRPSs, or for which NRPSs play a central synthetic role. The portion of gramicidin A that LgrA assembles is indicated by the grey box. b. The synthetic cycle of LgrA. Structures presented here represent the pre-condensation state (top left) and the post-condensation state (top right). Note that the small molecular substrates (fTHF, ATP, amino acids) are shown binding directly before they are required in the synthetic cycle, but the binding (as well as the adenylation reaction) can occur as soon as the relevant domain is free.
Extended Data Fig. 2 Activity and structure of FAT–CAT.
a. Tripeptide synthesis assays of F1A1T1C2A2T2–CBmdB-M3 and F1A1T1–C2A2T2–CBmdB-M3 shows that the scar residues left by AEP1 ligation does not influence biosynthesis activity. CBmdB-M3 catalyses peptide bond formation of fVal-Gly-ppant-T2 with free tryptamine, releasing formyl-valinyl-glycyl-tryptamine, quantified by LC-MS. Note that although AEP1 ligation leaves an Asn-Gly-Leu scar, by splitting the dimoldular LgrA before residue Leu771 in the T1-C2 linker, only 2 extra residues are added to the AEP1 ligated FAT–CAT. The sequence of the ligated FAT–CAT becomes: …Val770-Asn770A-Gly770B-Leu771… The central value for the reactions represents the mean, while the standard deviation of the mean is represented by the error bars. Individual points of the triplicates (n = 3) are shown. Replicates presented in the same panel were performed with one preparation of protein each, but repeated preparations of the same proteins repeatedly show the same activity. Statistical significance between the wild-type and mutant variants of F1A1T1–C2A2T2–CBmdB-M3 was determined by a two-sided Student’s t-test. b. Comparison between pre-condensation F1A1T1-fVal–C2A2T2-Gly and post-condensation F1A1T1-SH–C2A2T2-fVal-Gly crystal structures. c. The C domains of F1A1T1-fVal–C2A2T2-Gly (green), F1A1T1-SH–C2A214 (light brown), and F1A1T1-fVal–C214 (olive) show the same lobe:lobe orientation. Relative motion of C domain lobes has been proposed as important for catalysis and predicated by normal mode analyses, and differential chemical footprinting of C domains observed in presence and absence of substrates8,10,23,41. However, different relative orientations of lobes has only been observed across different C domains. d. The latch loop (residues 1124–1138) and the floor loop (residues 1044–1057) of C2 is ordered and interacting with the C lobe, arguing that these elements do not change conformation to act directly in catalysis8.
Extended Data Fig. 3 Domain:domain interactions in dimodular LgrA.
Top: Comparison of a the low resolution, substrate-free F1A1T1C2A2T2 structure (6MFZ14), b F1A1T1-fVal–C2A2T2-Gly and c F1A1T1-fValC2Acore2 (6MFY14), shown individually (a-c) and pairwise (d,e). The structures are superimposed on F1Acore1. Bottom: Domain:domain interactions in F1A1T1-fVal–C2A2T2-Gly between (f) F1 and C2, (g) Asub1 and C2, and (h) C2 and Asub2. i. Tripeptide biosynthesis assay of C2 mutants E866A-E867A-E868A, K1002A-R1001A and E1107A-N974A. Measurements were carried out over n = 3 individual reactions in one experiment. Replicates presented in the same panel were performed with one preparation of protein each, but repeated preparations of the same proteins repeatedly show analogous activity levels. Individual points of the triplicates are shown. Each bar represents the average of the three measurements and the error bar is indicative of the standard deviation in the three replicates. Statistical significance between the wild-type and linker mutant variants was assessed by a two-sided Student’s t-test.
Extended Data Fig. 4 C domain interactions with aminoacyl-T domains.
a,b. Interactions of the LgrA C2 domain with donor fVal-T1 domain. At the donor entrance of the C2 tunnel, the ppant phosphate attached to T1 Ser729 forms a hydrogen bond with Thr1013, a well-conserved in LCL domain residue62. The ppant dimethyl moiety makes hydrophobic interactions with Tyr1015. This position shows strong conservation for aromatic residues in LCL domains62. The ppant also hydrogen bonds with the backbone amine of Asp1043 from the floor loop. c,d. At the donor entrance of the C2 tunnel, the ppant phosphate attached to T2 Ser1760 forms hydrogen bonds with the highly conserved Arg110662. Arg792 also forms hydrogen bonds with the backbone of the acceptor ppant. Notably, in structures lacking the acceptor T2 domain14, the side chains of Arg1106 and Arg792 extend into the tunnel, but are displaced in these full condensation complexes. Similarly, the arginine equivalent to Arg729 in fuscachelin A synthetase FscG-C3 was seen to move when an amino acyl ppant analogue binds4. Likewise, the arginine equivalent to Arg1106 do not obstruct the tunnel when acceptor ppants bind at the active site in FscG-C34 and holo-AB340325, but rather interact with the ppants. The subtle movements of these Arg residues suggest to us that this is a simple sidechain rearrangement upon acceptor binding. e. Polder map of the (acyl-)ppant moieties in F1A1T1-SH–C2A2T2-fVal-Gly. The map is displayed at 5 σ, and is not “carved” around acyl-ppant moieties.
Extended Data Fig. 5 T domain conformations and interactions.
a. Superimposition of all observations of LgrA T1, in thiolation13,14, formylation13 and condensation donation state14. The C terminus of helix 4 shows minor variability and there is relatively modest differences in ppant moiety positions. b. Superimposition of all observations of LgrA T2, all in the condensation state. c. Superimposition of all observations of LgrA T1:C2 interactions14. (i) Comparing superimposition by (i) T1 and by (ii) C2 shows that small variations in C2 binding by T1 (mainly minor angle shifts of T1 around the C2 donor site interaction surface (highlighted in ii)), are compensated for by small changes in ppant conformations (highlighted in i), to allow fVal delivery to the active site. c. Superimposition of one example each of structurally characterized, cognate interaction co-complexes of T domains with each type of bone fide NRPS domain42: LgrA A113, LgrA F113, LgrA C2, TqaA fungal terminal cyclization domain12, GrsA E domain76, EntF thioesterase domain77, CAR reductase domain78 and Mru_0351 archaeal reductase domain79. e. F1A1T1-fVal–C2A2T2-Gly with all residues within 4 Å of the partner domains shown as spheres. The donor binding site is exclusively made from residues from the C-lobe of C2. The acceptor binding site is mainly made from residues from the N-lobe of C2. One residue from each Asub is close to its neighbouring T domain (yellow). f,g. The donor binding site being part of the C-lobe of C2 and the acceptor binding site being mainly part of the N-lobe of C2 means that useful-but-very-simplified domain bubble diagrams (f) can be misleading: They could be taken to imply that upstream and downstream T domains bind at the portions of the C domains that are proximal to them in primary sequence. However, the interactions are mainly with the portions of the C domain which are more distal in primary sequence (g). This means any cut point in module-swap bioengineering between Tn−1 and Tn would disturb all native Tn-1: Cn interactions and most native Cn: Tn interactions. h,i. For structures in d, the T domain residues within 4 Å of their partner domain residues, mapped onto (h) the primary sequence and (i) the structure of LgrA T1 as Cα spheres. T domains use an extensively overlapping surface for all their known interactions with partner domains.
Extended Data Fig. 6 C domain mutants and pH profiles.
a. CD of wild-type and mutant C2A2T2 showing active site mutations not to affect global folding. b. DSF in quadruplicate (n = 4) of wild-type and mutant C2A2T2 shows decreased Tm for H907A and D912A compared to wild-type and other mutants. Tm values (inflexion point of DSF) are plotted in °C: Wild-type: 43.25 ± 0.14; H907A: 37.54 ± 0.69; H908A: 43.34 ± 0.20; D912A: 38.15 ± 0.91; G913A: 45.32 ± 0.09; L914P: 45.85 ± 0.40. Statistical significance between wild-type and variants were calculated using a two-sided Student’s t-test (wild-type versus H907A (p = 0.0003; ***), H908A (p = 0.5067; n.s), D912A (p = 0.0013; **), G913A (p = 0.000002; ****), L914P (p = 0.0002;***)). Central values represent the mean, and error bars represent the standard deviation of the mean. c. His907 points away from His908 and hydrogen bonds with its own backbone amine, the Ile909 backbone amine and the Gln797 sidechain. d. Asp912 hydrogen bonds with the backbone amines of Leu914, Ser915 and Phe1057, and the side chains of Ser915 and Arg1045. e,f. Initial tripeptide synthesis rates at pHs 5.7 to 8.7 for F1A1T1–C2A2T2–CBmdB-M3 and the L914P mutant, where condensation by C2 is rate limiting. g,h. Rates plotted as peak area (mAU280nm⋅min) per hour. Each data point represents the average of the initial rate of n = 3 replicates collected over three independent experiments. To collect all the data two preparations of protein were necessary, for each variant (wild type and L914P). The central value plotted represents the mean initial rate and the error bars the standard deviation in the initial rates. The error is taken to be the standard deviation between rates determined by individual time course experiments, represented by error bars. Trend lines are shown as a visual aid only. Statistical significance between initial rates at all pH points were determined using two-sided Student’s t-tests.
Extended Data Fig. 7 Quantum mechanical calculations for different conditions and mutants.
The theozyme Gibbs free energies and electronic energies of each species are shown for diagram in different conditions and mutants; Hirshfeld Charges of each fragment along the reaction pathway are also shown. 3D structures are shown below. For clarity, nonpolar H atoms are omitted except on Ns and on −CH2 group. Curved lines represent groups omitted in the computations. The backbone atoms of the protein chain were constrained in the theozyme calculations. a. The calculated information for no solvent (gas phase) and in CPCM water solvent (implicit solvent), only showing TS, which has free energy lower than that of Int (Fig. 4a). The other species in these conditions are show in Fig. 4a and not redrawn here. b. His908 imidazole replaced by −CH3; c. −NH− of Gly913 is replaced by −CH2 − .
Extended Data Fig. 8 Molecular dynamics simulations.
a,b. Distances of H908 δ or ε nitrogen (N) to the hydrogens (H1, H2) on the α-amine nucleophile, during simulations. Large graphs plot distances versus time, small graphs on right show populations of distances over the full simulation, and average distances the simulations are listed left of the caption. In a, His908 δ nitrogen is protonated, in b, the ε nitrogen is protonated. The C···N distance that eventually leads to an amide bond is constrained to 2.2 Å. The most populated structure in the first 100 ns (green) and in the full 500 ns (cyan) are shown in a and the most populated structure in the 500 ns simulation is shown in b. In this MD, the substrates and separate from the catalytic residues. This separation is not unusual for enzyme with spacious active sites. A recent study featuring MD of substrates manually placed into tightly packed areas of a C domain showed low root mean square deviation for 30 ns85, but the manual substrate placement is at odds with known binding sites and T domain substrate delivery, which may compromise analysis. c,d. The most populated 3D structures (left) and bond distances between the reactants and key residues (right) during 200 ns simulations, starting from Int1 of the theozyme calculations. Simulation c has the H(Acceptor)···N(H908) distance unconstrained during the simulation, while in d, the distance is constrained the 2.2 Å distance seen in the theozyme Int. In all these simulations, His907 and Asp912 stay pointed away from the reactive atoms and His908. Note the brief increase in d3 in panel d replica 1 is from a transient movement of Gln787 away from the active site, not His907 towards His908.
Extended Data Fig. 9 Comparisons with other C domain structures.
This figure shows comparisons with some of the most informative ligand-bound structures of other C domains. a-f. Comparison of F1A1T1-fVal–C2A2T2-Gly with acceptor-bound fuscachelin A synthetase C4 complexes. A thioether analogue of Gly-ppant (ethylamine-ppant)4 (a), and Leu-NHppant30 (c) bind FscG-C3 similarly to each other and to how LgrA-C2 binds Gly-NHppant. However, in FscG-C3, the nucleophile interacts with Glu2702, reaching from the donor side of the active site. This interaction could not be maintained in the presence of a donor substrate as its position overlaps with that of fVal-NHppant in the overlays (e,f) and the end of the donor tunnel in general. Interestingly, this Glu2702 in FscG-C3 one of less common cases (along with AmbB11) where the final position of the HHxxxDG motif is not a Gly but a side-chain bearing amino acid. Gly913 is 83% conserved in sequences of as LCL or DCL domains and 79% conserved in all C domains. Small differences in the approach of the donor ppants compared to that seen in LgrA would be needed to avoid clashing with the sidechain. g. A comparison with the a small molecule acceptor fused to the first C domain of calcium dependant antibiotic synthetase9. Note that this amide-containing acceptor analogue shown in g, and the thioether analogue in a are both shown to be competent for reaction4,9, whereas the amide containing acceptor analogue in c is reported to be non-reaction competent30. The positions of amide and thioester analogues studied in reference4 are so similar that both provide equivalent structural insight.
Supplementary information
Supplementary Information (download DOCX )
Supplementary Discussion, Supplementary Methods, Supplementary Table 1 and Supplementary Figs. 1–12.
Supplementary Data (download DOCX )
Uncropped gel presented in Fig. 1.
Supplementary Data (download ZIP )
Source data for Supplementary Figs. 2, 4, 6, 8, 10 and 11.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pistofidis, A., Ma, P., Li, Z. et al. Structures and mechanism of condensation in non-ribosomal peptide synthesis. Nature 638, 270–278 (2025). https://doi.org/10.1038/s41586-024-08417-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-024-08417-6
This article is cited by
-
Evolutionary repurposing of a metabolic thiolase complex enables antibiotic biosynthesis
Nature Communications (2026)
-
Pseudokinases can catalyse peptide cyclization through thioether crosslinking
Nature Chemistry (2026)