Abstract
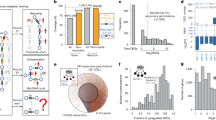
Metabolic flux, or the rate of metabolic reactions, is one of the most fundamental metrics describing the status of metabolism in living organisms. However, measuring fluxes across the entire metabolic network remains nearly impossible, especially in multicellular organisms. Computational methods based on flux balance analysis have been used with genome-scale metabolic network models to predict network-level flux wiring1,2,3,4,5,6. However, such approaches have limited power because of the lack of experimental constraints. Here, we introduce a strategy that infers whole-animal metabolic flux wiring from transcriptional phenotypes in the nematode Caenorhabditis elegans. Using a large-scale Worm Perturb-Seq (WPS) dataset for roughly 900 metabolic genes7, we show that the transcriptional response to metabolic gene perturbations can be integrated with the metabolic network model to infer a highly constrained, semi-quantitative flux distribution. We discover several features of adult C. elegans metabolism, including cyclic flux through the pentose phosphate pathway, lack of de novo purine synthesis flux and the primary use of amino acids and bacterial RNA as a tricarboxylic acid cycle carbon source, all of which we validate by stable isotope tracing. Our strategy for inferring metabolic wiring based on transcriptional phenotypes should be applicable to a variety of systems, including human cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Integrated peaks and mass distribution vector of isotope tracing data and DEGs of the RNA-seq experiments in this study can be downloaded from Supplementary Tables. Raw and processed data of RNA-seq (fastq files and count tables, respectively) are available through Gene Expression Omnibus accession GSE255866. All experimental source data are provided. Figures and results related to iMAT-WPS modelling can be reproduced using the iMAT-WPS GitHub repository (https://github.com/XuhangLi/iMAT-WPS). Mass spectrometry raw data are deposited in www.ebi.ac.uk/metabolights/MTBLS11741 and www.ebi.ac.uk/metabolights/MTBLS11742. Source data are provided with this paper.
Code availability
Source code for iMAT-WPS can be found at https://github.com/XuhangLi/iMAT-WPS.
References
Yilmaz, L. S. et al. Modeling tissue-relevant Caenorhabditis elegans metabolism at network, pathway, reaction, and metabolite levels. Mol. Syst. Biol. 16, e9649 (2020).
Colijn, C. et al. Interpreting expression data with metabolic flux models: predicting Mycobacterium tuberculosis mycolic acid production. PLoS Comput. Biol. 5, e1000489 (2009).
Sanchez, B. J. et al. Improving the phenotype predictions of a yeast genome-scale metabolic model by incorporating enzymatic constraints. Mol. Syst. Biol. 13, 935 (2017).
Lee, D. et al. Improving metabolic flux predictions using absolute gene expression data. BMC Syst. Biol. 6, 73 (2012).
O’Brien, E. J., Lerman, J. A., Chang, R. L., Hyduke, D. R. & Palsson, B. O. Genome-scale models of metabolism and gene expression extend and refine growth phenotype prediction. Mol. Syst. Biol. 9, 693 (2013).
Salvy, P. & Hatzimanikatis, V. The ETFL formulation allows multi-omics integration in thermodynamics-compliant metabolism and expression models. Nat. Commun. 11, 30 (2020).
Li, X. et al. Systems-level design principles of metabolic rewiring in an animal. Nature https://doi.org/10.1038/s41586-025-08636-5 (2025).
Gopalakrishnan, S. & Maranas, C. D. 13C metabolic flux analysis at a genome-scale. Metab. Eng. 32, 12–22 (2015).
Gopalakrishnan, S. & Maranas, C. D. Achieving metabolic flux analysis for S. cerevisiae at a genome-scale: challenges, requirements, and considerations. Metabolites 5, 521–535 (2015).
Hackett, S. R. et al. Systems-level analysis of mechanisms regulating yeast metabolic flux. Science 354, aaf2786 (2016).
Lewis, N. E. et al. Omic data from evolved E. coli are consistent with computed optimal growth from genome-scale models. Mol Syst Biol 6, 390 (2010).
Wagner, A. et al. Metabolic modeling of single Th17 cells reveals regulators of autoimmunity. Cell https://doi.org/10.1016/j.cell.2021.05.045 (2021).
Huang, Y. et al. Characterizing cancer metabolism from bulk and single-cell RNA-seq data using METAFlux. Nat. Commun. 14, 4883 (2023).
Alghamdi, N. et al. A graph neural network model to estimate cell-wise metabolic flux using single-cell RNA-seq data. Genome Res. 31, 1867–1884 (2021).
Damiani, C. et al. Integration of single-cell RNA-seq data into population models to characterize cancer metabolism. PLoS Comput. Biol. 15, e1006733 (2019).
Zhang, H. et al. Worm Perturb-Seq: massively parallel whole-animal RNAi and RNA-seq. Preprint at bioRxiv https://doi.org/10.1101/2025.02.02.636107 (2025).
Yilmaz, L. S. & Walhout, A. J. A Caenorhabditis elegans genome-scale metabolic network model. Cell Syst. 2, 297–311 (2016).
Watson, E. et al. Interspecies systems biology uncovers metabolites affecting C. elegans gene expression and life history traits. Cell 156, 759–770 (2014).
Watson, E. et al. Metabolic network rewiring of propionate flux compensates vitamin B12 deficiency in C. elegans. eLife 5, e17670 (2016).
Bulcha, J. T. et al. A persistence detector for metabolic network rewiring in an animal. Cell Rep. 26, 460–468 (2019).
Ponomarova, O. et al. A D-2-hydroxyglutarate dehydrogenase mutant reveals a critical role for ketone body metabolism in Caenorhabditis elegans development. PLoS Biol. 21, e3002057 (2023).
Bartman, C. R., TeSlaa, T. & Rabinowitz, J. D. Quantitative flux analysis in mammals. Nat. Metab. 3, 896–908 (2021).
Beydoun, S. et al. An alternative food source for metabolism and longevity studies in Caenorhabditis elegans. Commun. Biol. 4, 258 (2021).
Orth, J. D., Thiele, I. & Palsson, B. O. What is flux balance analysis? Nat. Biotechnol. 28, 245–248 (2010).
Mahadevan, R. & Schilling, C. H. The effects of alternate optimal solutions in constraint-based genome-scale metabolic models. Metab. Eng. 5, 264–276 (2003).
Diot, C. et al. Bacterial diet modulates tamoxifen-induced death via host fatty acid metabolism. Nat. Commun. 13, 5595 (2022).
Perez, C. L. & Van Gilst, M. R. A 13C isotope labeling strategy reveals the influence of insulin signaling on lipogenesis in C. elegans. Cell Metab. 8, 266–274 (2008).
Braeckman, B. P., Houthoofd, K. & Vanfleteren, J. R. Intermediary metabolism. WormBook: The Online Review of C. elegans Biology https://doi.org/10.1895/wormbook.1.146.1 (2009).
Watts, J. L. & Ristow, M. Lipid and carbohydrate metabolism in Caenorhabditis elegans. Genetics 207, 413–446 (2017).
TeSlaa, T., Ralser, M., Fan, J. & Rabinowitz, J. D. The pentose phosphate pathway in health and disease. Nat. Metab. 5, 1275–1289 (2023).
Zhang, Z. et al. Serine catabolism generates liver NADPH and supports hepatic lipogenesis. Nat. Metab. 3, 1608–1620 (2021).
Britt, E. C. et al. Switching to the cyclic pentose phosphate pathway powers the oxidative burst in activated neutrophils. Nat. Metab. 4, 389–403 (2022).
Kuehne, A. et al. Acute activation of oxidative pentose phosphate pathway as first-line response to oxidative stress in human skin cells. Mol. Cell 59, 359–371 (2015).
Blacklow, S. C., Raines, R. T., Lim, W. A., Zamore, P. D. & Knowles, J. R. Triosephosphate isomerase catalysis is diffusion controlled. Appendix: analysis of triose phosphate equilibria in aqueous solution by 31P NMR. Biochemistry 27, 1158–1167 (1988).
Jang, C., Chen, L. & Rabinowitz, J. D. Metabolomics and isotope tracing. Cell 173, 822–837 (2018).
Zecic, A., Dhondt, I. & Braeckman, B. P. The nutritional requirements of Caenorhabditis elegans. Genes Nutr. 14, 15 (2019).
Lee, Y. U. et al. Host-microbe interactions rewire metabolism in a C. elegans model of leucine breakdown deficiency. Nat. Metab. 6, 1584–1600 (2024).
Hui, S. et al. Glucose feeds the TCA cycle via circulating lactate. Nature 551, 115–118 (2017).
Hui, S. et al. Quantitative fluxomics of circulating metabolites. Cell Metab. 32, 676–688 e674 (2020).
Christodoulou, D. et al. Reserve flux capacity in the pentose phosphate pathway enables Escherichia coli’s rapid response to oxidative stress. Cell Syst. 6, 569–578 e567 (2018).
Kubagawa, H. M. et al. Oocyte signals derived from polyunsaturated fatty acids control sperm recruitment in vivo. Nat. Cell Biol. 8, 1143–1148 (2006).
Chen, L. et al. NADPH production by the oxidative pentose-phosphate pathway supports folate metabolism. Nat. Metab. 1, 404–415 (2019).
Nwosu, Z. C. et al. Uridine-derived ribose fuels glucose-restricted pancreatic cancer. Nature 618, 151–158 (2023).
Skinner, O. S. et al. Salvage of ribose from uridine or RNA supports glycolysis in nutrient-limited conditions. Nat. Metab. 5, 765–776 (2023).
Yuan, B., Inouye, K. E., Hotamisligil, G. S. & Hui, S. An organism-level quantitative flux model of mammalian energy metabolism. Preprint at bioRxiv https://doi.org/10.1101/2024.02.11.579776 (2024).
Gu, C., Kim, G. B., Kim, W. J., Kim, H. U. & Lee, S. Y. Current status and applications of genome-scale metabolic models. Genome Biol. 20, 121 (2019).
Przybyla, L. & Gilbert, L. A. A new era in functional genomics screens. Nat. Rev. Genet. 23, 89–103 (2022).
Replogle, J. M. et al. Mapping information-rich genotype-phenotype landscapes with genome-scale Perturb-seq. Cell 185, 2559–2575 (2022).
Zheng, X. et al. Massively parallel in vivo Perturb-seq reveals cell-type-specific transcriptional networks in cortical development. Cell 187, 3236–3248 (2024).
Santinha, A. J. et al. Transcriptional linkage analysis with in vivo AAV-Perturb-seq. Nature 622, 367–375 (2023).
King, Z. A. et al. BiGG models: a platform for integrating, standardizing and sharing genome-scale models. Nucleic Acids Res. 44, D515–D522 (2016).
Roux, A. E. et al. Individual cell types in C. elegans age differently and activate distinct cell-protective responses. Cell Rep. 42, 112902 (2023).
Acknowledgements
We thank members of the Walhout laboratory for discussion and critical reading of the manuscript. This work was supported by grants from the National Institutes of Health (grant nos. R35GM122502 and DK068429 to A.J.M.W. and R35GM145261 to E.A.S.). This work was supported by the National Key R&D Program of China (grant no. 2022YFA0806400 to J.Z.) and the National Natural Science Foundation of China (grant no. 32271215 to J.Z.).
Author information
Authors and Affiliations
Contributions
H.Z., X.L. and A.J.M.W. conceived the project and wrote the manuscript. H.Z. conducted most of the experiments. X.L. developed iMAT-WPS. H.Z. and X.L. jointly analysed the experimental data. L.S.Y. supervised the iMAT-WPS analysis and performed independent validations of the isotope balance calculations. H.Z. and L.T.T. conducted LC–MS measurements under the supervision of J.B.S. H.Z., R.L.N. and E.A.S. purified ribose-phosphate by using high-performance liquid chromatography. G.E.G. performed RNA-seq of animals fed with paraformaldehyde-killed bacteria. H.W. and B.Y. performed LC–MS analysis to study metabolomics for perturbations of genes in the Met/SAM cycle under the supervision of J.Z. A.J.M.W. supervised the entire study. The cofirst authorship order was determined by a coin flip. Both H.Z. and X.L. contributed equally and reserve the right to list their name first on their resumes.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Gregory Ducker, Ralf Sommer and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Steady-state isotope tracing in C. elegans and the labeling data for glycine and adenine tracing experiments.
a, Cartoon of two C. elegans propionate breakdown pathways. b, Bar graph of DEG numbers for perturbations of propionate breakdown genes. Data are from the metabolic-gene WPS study7. c, Bar graph of DEG numbers for perturbations of two representative genes in the canonical propionate breakdown and shunt pathways, with or without supplementation of vitamin B12, in a new set of WPS experiments. d, Diagram illustrating the steady-state isotope tracing procedure in C. elegans. Isotope-labeled metabolites were supplemented in the culture media to achieve ~10% labeling in vivo at steady state. This figure was created with BioRender.com. For a detailed protocol, please refer to the Supplementary Methods. e, Differential expression analysis comparing transcriptome profiles of gravid adult C. elegans fed with live versus PFA-killed HT115. Red dots represent genes identified as differentially expressed (DEGs) using the DESeq2 package in R, with adjustments for multiple comparisons via the Benjamini-Hochberg (BH) method (fold change (FC) > 2, adjusted P value (Padjust) < 0.01). f, g, Unnormalized isotope labeling data from [2-13C,15N]glycine and [8-13C]adenine tracing experiments for each biological replicate.
Extended Data Fig. 2 Dissecting the methionine degradation and regeneration paths within the Met/SAM cycle through WPS similarity analysis and isotope tracing experiments.
a, Zoomed-in view of two WPS similarity clusters, ‘Met/SAM cycle’ and ‘Lipid synthesis & V-ATPase’, in a 2-dimensional visualization of perturbations in the systematic metabolic-gene WPS dataset7. The visualization was generated in the referred study. Please note that although the cluster containing mthf-1, metr-1 and metr-1 genes was named ‘Met/SAM cycle’ in the referred study, these genes also belong to the folate cycle. Most Met/SAM cycle genes are actually in the ‘Lipid synthesis & V-ATPase’ cluster, leading to the flux wiring hypothesis stated in the main text. b, Fold change of substrate and product abundance upon RNAi of metabolic genes relative to vector control as measured by LC-MS (x-axis). Each bar represents the mean ± s.d. of experimental data, with individual points indicating the values from each biological replicate (n = 3). The P values were derived by comparing raw peak abundance data between RNAi and vector control samples using two-tailed Student’s t-test with ‘ns’ indicating not significant. c, Bar graph of DEG numbers of indicated genes, with and without supplementation of vitamin B12. We expect that, with vitamin B12 supplementation, flux through the cyclic path would increase, and flux through the linear path would decrease. Indeed, in the presence of vitamin B12, we observed a decrease in the number of DEGs in response to cbs-1 knockdown and an increase in the number of DEGs upon metr-1 RNAi. d, WPS similarity between perturbations of indicated genes and cbs-1 (left) or metr-1 (right). There was a decrease in WPS similarity between cbs-1 and other Met/SAM cycle genes (e.g., sams-1 and ahcy-1) and an increase in WPS similarity between metr-1 with mthf-1, indicating a decrease in linear pathway flux and an increase in cyclic pathway flux. e, GC-MS data of different methionine standards for confirming the identity of the m/z 176 peak, which represents the last four carbons of methionine. f, Unnormalized time-course data from [U-13C]methionine tracing experiments for each biological replicate with or without supplementation of 64 nM vitamin B12. g, The isotopologue distribution of methionine fragment (m/z 176) in [U-13C]methionine tracing experiments revealed a labeling pattern predominantly consisting of M + 4 or M + 3. Each bar represents the mean ± s.d. of experimental data and each dot represents a single time point in isotopic steady state (from 2 h to 9 h) of a biological replicate (n = 3). Data at steady state was normalized to the worth of 10% total labeling of methionine. h, i, Time-course data from [3-13C]serine and [2-13C,15N]glycine tracing experiments from three biological replicates with or without supplementation of 64 nM vitamin B12. Each curve shows the mean ± s.d. of labeling fractions from three biological replicates. Unnormalized data were used for making these curves. The P values were calculated using two-way ANOVA test.
Extended Data Fig. 3 Directionally constrained reactions in the PFM and OFM.
(a-c), Randomizations (n = 10,000 in all) showing that: the percentage of nonresponsive genes conflicting with another responsive gene in the same reaction was significantly lower than random (a), total flux through reactions associated with no responsiveness in iMAT-WPS integration was significantly lower than random (b) and the objective value of WPS similarity integration (x-axis) based on real WPS data was significantly better optimized (minimized) than random (c). Red dashed lines indicate the percentage in the real WPS data and histograms in randomized data. The empirical P values are indicated. d, Bar graphs showing the number of directionally constrained reactions in the PFM. Grey indicates overlapped (identical) reactions in all five integrations; orange indicates overlapped reactions in four integrations except for the no-integration case (i.e., with only the biomass production constraint); and white indicates reactions that are different in individual integrations. e, f, Venn diagrams showing the number and overlap of directionally constrained reactions when different data were integrated, in the PFM (e) and OFM (f). g, Proportions of directionally constrained reactions in iCEL1314 model. We visualized the proportions for all reactions in the model (‘all’) and three mutually exclusive categories: ‘exchange’ for exchange reactions; ‘transport’ for transporter reactions; and ‘internal’ for all other reactions. h, Proportions of directionally constrained reactions in OFM ordered by pathways. Pathways are defined with the ‘subSystems’ annotation in the iCEL1314 model. Pathways enriched for directionally constrained and unconstrained reactions are colored in red and blue, respectively (BH-adjusted, one-sided hypergeometric test Padj < 0.01). The green dashed line indicates the cumulative number of directionally constrained reactions from left to right. i-j, Examples of flux solution space (upper and lower bounds) obtained by Flux Variability Analysis (FVA) for two different iCEL1314 reactions. The flux value in OFD, solution space of OFM and PFM are visualized together as indicated. The solution spaces obtained by different integration approaches, as indicated at the left of each figure, are compared side-by-side. Exp: integrating gene expression levels; resp: integrating WPS responsiveness; simi: integrating WPS similarity.
Extended Data Fig. 4 The sensitivity of iMAT-WPS predictions.
a, Recall of directionally constrained reactions with random subsampling of WPS data. WPS data were subsampled at different depths (i.e., number of perturbations included) for 30 times, followed by repeating iMAT-WPS using each subsample. Each dot represents the modeling result from one subsample. Y-axis shows the Jaccard index between directionally constrained reactions in full-data modeling and in a subsample analysis, giving the recall of original predictions. The box bounds the IQR divided by the median, and whiskers extend to a maximum of 1.5 × IQR beyond the box. b, An example reaction (RM04432, formula indicated) showing the distribution of OFD flux in subsampling analysis. OFD flux values are normalized to per unit bacterial uptake to facilitate cross comparisons. Each dot represents the flux of RM04432 reaction from one subsample. Any subsample flux that was within ± 30% of the original prediction was deemed ‘recalled’. The fraction of such ‘recalled’ predictions among the 30 repeats at each sampling depth is shown in the y-axis of the bottom plot. The box bounds the IQR divided by the median, and whiskers extend to a maximum of 1.5 × IQR beyond the box. c, Histogram showing the recall fraction (y-axis of the bottom plot in (b)) for all reactions at 80% subsampling depth.
Extended Data Fig. 5 Tissue expression patterns for reactions with predicted flux and the effects of WPS data integration on PPP flux prediction.
a. Tissue expression heatmaps for reactions with active OFD flux. The tissue expression levels (transcript per million, TPM) were obtained from a single-cell RNA-seq dataset for adult C. elegans52. These gene-level expression values were then converted to reaction-level TPM based on the corresponding GPR using a method we previously developed1 (Supplementary Methods). The left heatmap shows relative reaction expression across tissues ranging from 0 (not expressed) to 1 (highest expression). The right heatmap shows reaction-level log2(TPM) values. Rows represent all reactions with active flux in the OFD (normalized flux per unit bacterial uptake > 1e-5) and that had available expression data. The rows and columns are clustered by cosine similarity using the relative reaction expression matrix. Pathways associated with these reactions (rows) are annotated based on the ‘subSystems’ annotation in the iCEL1314 model and are indicated on the left (colored bins represent the associated pathways). The dashed box highlights energy-related reactions that are specifically enriched in the pharynx based on relative expression but are highly expressed in all tissues based on absolute TPM. b, Examining the effects of WPS responsiveness data for K07E3.4 or idh-1 on the flux prediction of reactions catalyzed by gspd-1 and gpi-1. The mRNA levels, WPS responsiveness and WPS similarity data were integrated with iCEL1314 through iMAT-WPS, while including or excluding the responsiveness data of the two indicated genes. c, Mean WPS similarity of cyclic PPP genes (gspd-1, tald-1, T25B9.9, tald-1, tkt-1, and gpi-1) compared to all other perturbations that yield significant transcriptional responses when knocked down in WPS experiments. This calculation used the cosine similarity matrix derived from the systematic metabolic-gene WPS study7. d,e, Similar to Extended Data Fig. 4b but for the two reactions shown in (b). The box bounds the IQR divided by the median, and whiskers extend to a maximum of 1.5 × IQR beyond the box.
Extended Data Fig. 6 [U-13C]glucose tracing and atom map of PPP and glycolysis/gluconeogenesis pathways.
a, Unnormalized time course data of g6p from [U-13C]glucose tracing experiments across each biological replicate revealed that the total labeling fraction of g6p did not exceed 20%. b, Time course data from [U-13C]glucose tracing experiments showing the labeling dynamics of various metabolites. Data were normalized to the worth of 10% of g6p total labeling. Each curve shows the mean ± s.d. from five biological replicates. c, Atom map depicting fully labeled g6p entering the TCA cycle and returning to g6p via the glycolysis/gluconeogenesis pathway, resulting in M + 1, M + 2, and M + 3 dominant and symmetric labeling of g6p. If glycolytic flux is dominant, g6p labeling pattern will be M + 6 dominant. Dashed line indicates multiple reactions. d, Atom map detailing the first cycle of PPP in [U-13C]glucose tracing experiments, illustrating how cyclic PPP can produce non-fully labeled g6p with an asymmetric labeling pattern. e, Fractional contribution (FC) of g6p to metabolites in the TCA cycle under RNAi treatments. Each bar represents the mean ± s.d. of experimental data and each dot indicates a biological replicate collected at 9 h post tracing. Sample sizes: n = 10 (vector), n = 4 (enol-1), n = 6 (dlat-1), n = 6 (T25B9.9) and n = 8 (tald-1). The P values were derived by comparing data between each RNAi and vector control using two-tailed Student’s t-test with **** indicating P < 0.0001 and ns as not significant. Exact P values are provided in the source data. f, g, Unnormalized time-course data of g6p from [1,2,3-13C]glucose or [4,5,6-13C]glucose tracing experiments across each biological replicate revealed that the total labeling fraction of g6p was around 3-4%.
Extended Data Fig. 7 Assessing the symmetry of g6p labeling pattern using GC-MS.
a, GC-MS fragments of g6p were identified using various g6p standards. The m/z 271 peak was identified to be a fragment encompassing all six carbons of g6p (M + 0). The m/z 204 was determined to be a fragment containing the second and third carbons ([C2-C3]) of g6p (M + 0). Meanwhile, m/z 370 was pinpointed as a fragment covering the fourth to the sixth carbons ([C4-C6]) of g6p (M + 0). These fragment identities were confirmed based on the mass shifts of various isotope-labeled standards as shown in different peak colors. b, Isotopologue distributions of various metabolites from identical [U-13C]glucose tracing samples (collected at 6, 9 and 12 h post tracing) were compared using LC-MS and GC-MS analyses. Each bar represents the mean ± s.d. of experimental data and each dot represents a single time point in a biological replicate (n = 3). Shapes indicate biological replicates. The results show that the data obtained from GC-MS are consistent with data from LC-MS, evidenced by the similar levels in average labeling and their variances across replicates. c, The left cartoon illustrates that in the absence of cyclic PPP flux, the fraction of [C2-C3] (M1) is expected to be lower than or equal to that of [C4-C6] (M1 + M2). Detailed reasoning can be found in Supplementary Methods. The bar plot shows that gene knockdown in the PPP, but not in the glycolysis/gluconeogenesis pathway, markedly affects the asymmetric labeling pattern of g6p. Each bar represents the mean ± s.d. of experimental data and each dot indicates a biological replicate collected at 9 h post tracing. Sample sizes: n = 10 (vector), n = 4 (enol-1), n = 6 (dlat-1), n = 6 (T25B9.9) and n = 8 (tald-1). The P values were derived by comparing data between each red and blue bar (P = 5.50 × 10−12 (vector), P = 4.74 × 10−4 (enol-1), P = 4.15 × 10−10 (dlat-1), P = 2.38 × 10−3 (tald-1), and P = 9.29 × 10−1 (T25B9.9)).
Extended Data Fig. 8 RNA fuels central carbon metabolism and the effects of glucose supplementation on fluxes in central carbon metabolism.
a, Unnormalized time-course data of ribose-1-phosphate (r1p) labeling from [ribose-13C5]adenosine tracing experiments across each biological replicate revealed that the total labeling fraction of r1p was around 20%. Based on the data, 6 to 9 h was identified as steady state time window. b, The labeling dynamics of r1p and g6p from [ribose-13C5]adenosine tracing experiments. Data were normalized to the worth of 10% of total labeling of r1p. c, A labeling ratio of approximately 100% for r1p over AMP suggests an efficient exchange flux between AMP and r1p. Each curve in (b) and (c) shows the mean ± s.d. from three biological replicates. d, The fractional contribution of g6p to r1p. The samples were purified by HPLC to enrich r1p (Supplementary Methods). e, The fractional contribution of r1p to 3-phosphoglycerate (3pg), analyzed in the context of vector control RNAi and RNAi targeting genes within the PPP. The P values were calculated using two-tailed Student’s t-test. f, The fractional contribution of r1p to metabolites in TCA cycle (aspartate as a proxy for oaa). g, The labeling pattern of g6p under supplementation of various concentration of [U-13C]glucose. Data were normalized to the worth of 10% of total labeling of g6p. h, The bar plot shows the influence of glucose supplementation on the asymmetric labeling pattern of g6p. Notably, the fraction of g6p [C2-C3] (M1) was still two-fold higher than that of g6p [C4-C6] (M1 + M2) with 250 mM [U-13C]glucose supplementation. i, The fractional contribution of g6p to metabolites in central carbon metabolism. The contribution of g6p to serine increases slightly, suggesting an upregulation in de novo serine synthesis flux. In contrast, the contribution to L-alanine (a proxy for pyruvate) and metabolites within the TCA cycle decreases significantly. In (d-i), each bar represents the mean ± s.d. of experimental data and each dot represents a steady-state time point (9 h) from one replicate. Sample sizes: n = 5 for (d) and n = 4 for (e-i).
Extended Data Fig. 9 Isotope tracing with various amino acid or propionate tracers.
a, Unnormalized labeling fractions of time-course data using different fully labeled tracers. For the amino acid tracing, the labeling dynamics of the corresponding tracer metabolite was displayed, whereas for propionate tracing, 3-Hydroxypropionate (3HP) was used as a proxy for the tracer. The data revealed that the total labeling ratio did not exceed 20% for any of the tracers. b, The dynamics of normalized labeling of citrate from various tracing experiments. c, The dynamics of normalized labeling of 3HP from tracing experiments with amino acids potentially involved in propionate production. d, The total labeling fraction of tyrosine in [U-13C]phenylalanine tracing experiment. (b-d), The labeling fractions were normalized to a worth of 10% total labeling in the corresponding tracer metabolite. Each curve in (b-d) shows the mean ± s.d. from three biological replicates. e, Left panel shows the pathway cartoon for hydroxymethylglutaryl-CoA (hmgcoa) breakdown and dolichol phosphate (dolp) and ubiquinone-9 (q9) synthesis. Bar plot on the right shows the predicted flux through the hmgcoa breakdown reaction, RM01360, using the default (0.01) and a 10-fold lowered (0.001) epsilon value for q9 and dolp synthesis reactions in iMAT-WPS. This reaction flux is used as a proxy of the leucine breakdown flux that eventually fuels the TCA cycle.
Extended Data Fig. 10 [U-13C]pyruvate tracing data.
a, Unnormalized time course data for pyruvate, L-alanine and lactate from [U-13C]pyruvate tracing experiments across each biological replicate. The data indicated that the total labeling fraction of pyruvate was approximately 5% and L-alanine exhibited the smallest variance, and hence, the highest data quality. b, Time-course data from [U-13C]pyruvate tracing experiments revealed the labeling dynamics of various metabolites. Data were normalized to the worth of 10% of total labeling of L-alanine. Each curve shows the mean ± s.d. from three biological replicates. c, Bar plot displaying the isotopologue distribution of steady-state data for pyruvate, g6p, citrate, malate, and aspartate. The network cartoon depicts the flux status of pyruvate metabolism reactions deduced from the labeling pattern (Supplementary Note 6) with grey dashed line indicating negligible flux and black solid line indicating significant flux. d, The fractional contribution of metabolites in TCA cycle (aspartate as a proxy for oaa) to g6p. For (c, d), each bar represents the mean ± s.d. of experimental data and each dot is a single time point at 6 or 9 h in one biological replicate (n = 3 for both (c) and (d)).
Supplementary information
Supplementary Information (download PDF )
This file contains Supplementary Notes 1–6, Methods and References.
Supplementary Tables (download ZIP )
Supplementary Tables 1–12.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, H., Li, X., Tseyang, L.T. et al. A systems-level, semi-quantitative landscape of metabolic flux in C. elegans. Nature 640, 194–202 (2025). https://doi.org/10.1038/s41586-025-08635-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-08635-6
This article is cited by
-
Worm Perturb-Seq: massively parallel whole-animal RNAi and RNA-seq
Nature Communications (2025)
-
Systems-level design principles of metabolic rewiring in an animal
Nature (2025)
-
Harnessing microalgae for bioproducts: innovations in synthetic biology
World Journal of Microbiology and Biotechnology (2025)