Abstract
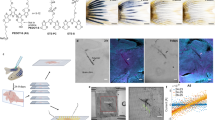
Developing bioelectronics capable of stably tracking brain-wide, single-cell, millisecond-resolved neural activity in the developing brain is critical for advancing neuroscience and understanding neurodevelopmental disorders. During development, the three-dimensional structure of the vertebrate brain arises from a two-dimensional neural plate1,2. These large morphological changes have previously posed a challenge for implantable bioelectronics to reliably track neural activity throughout brain development3,4,5,6,7,8,9. Here we introduce a tissue-level-soft, submicrometre-thick mesh microelectrode array that integrates into the embryonic neural plate by leveraging the tissue’s natural two-dimensional-to-three-dimensional reconfiguration. As organogenesis progresses, the mesh deforms, stretches and distributes throughout the brain, seamlessly integrating with neural tissue. Immunostaining, gene expression analysis and behavioural testing confirm no adverse effects on brain development or function. This embedded electrode array enables long-term, stable mapping of how single-neuron activity and population dynamics emerge and evolve during brain development. In axolotl models, it not only records neural electrical activity during regeneration but also modulates the process through electrical stimulation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The gene sequences used in the qPCR assays were obtained from Xenbase: https://www.xenbase.org/xenbase/. All the other data supporting the findings of this study are available within the paper, Extended Data, Supplementary Information and source data.
Code availability
All the code is available on GitHub at https://github.com/LiuLab-Bioelectronics-Harvard/Cyborg_Embryo.
References
Sanes, D. H. et al. Development of the Nervous System (Academic Press, 2011).
Smith, J. L. & Schoenwolf, G. C. Neurulation: coming to closure. Trends Neurosci. 20, 510–517 (1997).
Randlett, O. et al. Whole-brain activity mapping onto a zebrafish brain atlas. Nat. Methods 12, 1039–1046 (2015).
Keller, P. J. & Ahrens, M. B. Visualizing whole-brain activity and development at the single-cell level using light-sheet microscopy. Neuron 85, 462–483 (2015).
Alivisatos, A. P. et al. The brain activity map. Science 339, 1284–1285 (2013).
Stringer, C. et al. Spontaneous behaviors drive multidimensional, brainwide activity. Science 364, 255 (2019).
Fox, M. D. & Raichle, M. E. Spontaneous fluctuations in brain activity observed with functional magnetic resonance imaging. Nat. Rev. Neurosci. 8, 700–711 (2007).
Ahrens, M. B., Orger, M. B., Robson, D. N., Li, J. M. & Keller, P. J. Whole-brain functional imaging at cellular resolution using light-sheet microscopy. Nat. Methods 10, 413–420 (2013).
Hong, G. & Lieber, C. M. Novel electrode technologies for neural recordings. Nat. Rev. Neurosci. 20, 330–345 (2019).
Jun, J. J. et al. Fully integrated silicon probes for high-density recording of neural activity. Nature 551, 232–236 (2017).
Abbott, J. et al. A nanoelectrode array for obtaining intracellular recordings from thousands of connected neurons. Nat. Biomed. Eng. 4, 232–241 (2020).
Chiang, C. H. et al. Development of a neural interface for high-definition, long-term recording in rodents and nonhuman primates. Sci. Transl. Med. 12, eaay4682 (2020).
Musk, E. An integrated brain–machine interface platform with thousands of channels. J. Med. Internet Res. 21, e16194 (2019).
McDole, K. et al. In toto imaging and reconstruction of post-implantation mouse development at the single-cell level. Cell 175, 859–876 (2018).
Kasthuri, N. et al. Saturated reconstruction of a volume of neocortex. Cell 162, 648–661 (2015).
Pijuan-Sala, B. et al. A single-cell molecular map of mouse gastrulation and early organogenesis. Nature 566, 490–495 (2019).
Liu, J. et al. Syringe-injectable electronics. Nat. Nanotechnol. 10, 629–636 (2015).
Tian, B. et al. Macroporous nanowire nanoelectronic scaffolds for synthetic tissues. Nat. Mater. 11, 986–994 (2012).
Xie, C. et al. Three-dimensional macroporous nanoelectronic networks as minimally invasive brain probes. Nat. Mater. 14, 1286–1292 (2015).
Dai, X., Zhou, W., Gao, T., Liu, J. & Lieber, C. M. Three-dimensional mapping and regulation of action potential propagation in nanoelectronics-innervated tissues. Nat. Nanotechnol. 11, 776–782 (2016).
Fu, T. M. et al. Stable long-term chronic brain mapping at the single-neuron level. Nat. Methods 13, 875–882 (2016).
Li, Q. et al. Cyborg organoids: implantation of nanoelectronics via organogenesis for tissue-wide electrophysiology. Nano Lett. 19, 5781–5789 (2019).
Le Floch, P. et al. Stretchable mesh nanoelectronics for 3D single-cell chronic electrophysiology from developing brain organoids. Adv. Mater. 34, 2106829 (2022).
Xu, T. et al. Characterization of the mechanical behavior of SU-8 at microscale by viscoelastic analysis. J. Micromech. Microeng. 26, 105001 (2016).
Gupta, P., Bera, M. & Maji, P. K. Nanotailoring of sepiolite clay with poly [styrene‐b‐(ethylene‐co‐butylene)‐b‐styrene]: structure–property correlation. Polym. Adv. Technol. 28, 1428–1437 (2017).
Chanthasopeephan, T., Desai, J. P. & Lau, A. C. W. Study of soft tissue cutting forces and cutting speeds. Stud. Health Technol. Inform. 98, 56–62 (2004).
Spruiell Eldridge, S. L. et al. A focal impact model of traumatic brain injury in Xenopus tadpoles reveals behavioral alterations, neuroinflammation, and an astroglial response. Int. J. Mol. Sci. 23, 7578 (2022).
Yoshino, J. & Tochinai, S. Successful reconstitution of the non‐regenerating adult telencephalon by cell transplantation in Xenopus laevis. Dev. Growth Differ. 46, 523–534 (2004).
Khodagholy, D. et al. NeuroGrid: recording action potentials from the surface of the brain. Nat. Neurosci. 18, 310–315 (2015).
Yang, X. et al. Bioinspired neuron-like electronics. Nat. Mater. 18, 510–517 (2019).
Liu, Y. et al. Soft and elastic hydrogel-based microelectronics for localized low-voltage neuromodulation. Nat. Biomed. Eng. 3, 58–68 (2019).
Minev, I. R. et al. Electronic dura mater for long-term multimodal neural interfaces. Science 347, 156–163 (2015).
Kozai, T. D. Y. et al. Ultrasmall implantable composite microelectrodes with bioactive surfaces for chronic neural interfaces. Nat. Mater. 11, 1065–1073 (2012).
Manita, S. & Ross, W. N. Synaptic activation and membrane potential changes modulate the frequency of spontaneous elementary Ca2+ release events in the dendrites of pyramidal neurons. J. Neurosci. 29, 7833–7845 (2009).
Ciarleglio, C. M. et al. Multivariate analysis of electrophysiological diversity of Xenopus visual neurons during development and plasticity. Elife 4, e11351 (2015).
Maden, M. Salamanders as key models for development and regeneration research. Methods Mol. Biol. 2562, 1 (2023).
Steinmetz, N. A. et al. Neuropixels 2.0: a miniaturized high-density probe for stable, long-term brain recordings. Science 372, eabf4588 (2021).
Schoonover, C. E., Ohashi, S. N., Axel, R. & Fink, A. J. P. Representational drift in primary olfactory cortex. Nature 594, 541–546 (2021).
Denker, M., Yegenoglu, A. & Grün, S. Collaborative HPC-enabled workflows on the HBP Collaboratory using the Elephant framework. Neuroinformatics 19, FZJ-2018-04114 (2018).
Cowley, B. et al. DataHigh: graphical user interface for visualizing and interacting with high-dimensional neural activity. J. Neural Eng. 10, 066012–066019 (2013).
Wei, X. et al. Single-cell stereo-seq reveals induced progenitor cells involved in axolotl brain regeneration. Science 377, eabp9444 (2022).
Lust, K. et al. Single-cell analyses of axolotl telencephalon organization, neurogenesis, and regeneration. Science 377, eabp9262 (2022).
Takahashi, M. & Osumi, N. The method of rodent whole embryo culture using the rotator-type bottle culture system. J. Vis. Exp. 42, 2170 (2010).
Huang, Q. et al. Intravital imaging of mouse embryos. Science 368, 181–186 (2020).
Jouary, A., Haudrechy, M., Candelier, R. & Sumbre, G. A 2D virtual reality system for visual goal-driven navigation in zebrafish larvae. Sci. Rep. 6, 34015 (2016).
Le Floch, P. et al. 3D spatiotemporally scalable in vivo neural probes based on fluorinated elastomers. Nat. Nanotechnol. 19, 319–329 (2024).
Fowkes, F. M. Attractive forces at interfaces. Ind. Eng. Chem. 56, 40–52 (1964).
Karimi, K. et al. Xenbase: a genomic, epigenomic and transcriptomic model organism database. Nucleic Acids Res. 46, D861–D868 (2018).
Aguilera-Castrejon, A. et al. Ex utero mouse embryogenesis from pre-gastrulation to late organogenesis. Nature 593, 119–124 (2021).
Huang, Y., Wilkie, R. & Wilson, V. Methods for precisely localized transfer of cells or DNA into early postimplantation mouse embryos. J. Vis. Exp. 106, e53295 (2015).
Lohmiller, J. J., Swing, S. P. & Hanson, M. M. in The Laboratory Rat 3rd edn (eds Suckow, M. A. et al.) 157–179 (Elsevier, 2020).
Vczian, A. S. & Zuber, M. E. A simple behavioral assay for testing visual function in Xenopus laevis. J. Vis. Exp. 88, 51726 (2014).
Dong, W. et al. Visual avoidance in Xenopus tadpoles is correlated with the maturation of visual responses in the optic tectum. J. Neurophysiol. 101, 803–815 (2009).
Pratt, K. G. & Khakhalin, A. S. Modeling human neurodevelopmental disorders in the Xenopus tadpole: from mechanisms to therapeutic targets. Dis. Models Mech. 6, 1057–1065 (2013).
Klymkowsky, M. W. Whole-mount immunocytochemistry in Xenopus. Cold Spring Harb. Protoc. 2018, pdb-prot097295 (2018).
Hamilton, P. W. & Henry, J. J. Prolonged in vivo imaging of Xenopus laevis. Dev. Dyn. 243, 1011–1019 (2014).
Kamei, M. & Weinstein, B. M. Long-term time-lapse fluorescence imaging of developing zebrafish. Zebrafish 2, 113–123 (2005).
Kamei, M., Isogai, S., Pan, W. & Weinstein, B. M. Imaging blood vessels in the zebrafish. Methods Cell. Biol. 100, 27–54 (2010).
Chaure, F. J. et al. A novel and fully automatic spike-sorting implementation with variable number of features. J. Neurophysiol. 120, 1859–1871 (2018).
Chung, J. E. et al. A fully automated approach to spike sorting. Neuron 95, 1381–1394 (2017).
Buccino, A. P. et al. SpikeInterface, a unified framework for spike sorting. eLife 9, e61834 (2020).
Bartolo, R., Saunders, R. C., Mitz, A. R. & Averbeck, B. B. Information-limiting correlations in large neural populations. J. Neurosci. 40, 1668–1678 (2020).
Insel, N. & Barnes, C. A. Differential activation of fast-spiking and regular-firing neuron populations during movement and reward in the dorsal medial frontal cortex. Cereb. Cortex 25, 2631–2647 (2015).
Ogden, R. W. Non-linear Elastic Deformations (Courier, 1997).
Amar, M. B. & Goriely, A. Growth and instability in elastic tissues. J. Mech. Phys. Solids 53, 2284–2319 (2005).
Garaschuk, O., Hanse, E. & Konnerth, A. Developmental profile and synaptic origin of early network oscillations in the CA1 region of rat neonatal hippocampus. J. Physiol. 507, 219–236 (1998).
Acknowledgements
J.L. acknowledges the support from the Startup fund from the School of Engineering and Applied Sciences, Harvard University; NIH/NIMH 1RF1MH123948; Aramont Fund for Emerging Science Research; and the William F. Milton Fund. J.L. and N.L. acknowledge the support from NSF/EFRI 2422348. H. Sheng acknowledges the support from Aramont Fund for Emerging Science Research. J.L.W. and J.L. acknowledge the support from NIH/NICHD 1R01HD115272. X.W. acknowledges support from Packard Fellowship for Science and Engineering, Merkin Institute Fellowship, Stanley Center gift and NIH New Innovator Award. We thank D. J. Needleman for discussions.
Author information
Authors and Affiliations
Contributions
J.L. and H. Sheng conceived of the idea. R.L., H. Sheng, P.L.F., W.W. and A.J.L. fabricated and characterized electronics. H. Sheng performed implantation and behaviour tests. H. Sheng and Q.L. applied immunofluorescence. H. Sheng, Z.L. and J.B. did the qPCR. H. Sheng and R.L. performed electrical recordings. L.J., Z.W., H. Sheng, S.L. and N.L. did the mechanical simulation. R.A.J.-C. housed frogs. H. Sheng, H.Z., Z.L., Y.H., X.T., D.S., H.Z. and H. Shen analysed data. J.L.W. advised on designing tail amputation experiments. H. Sheng and E.H. prepared illustrations. H. Sheng, J.L., T.S.B. and X.W. wrote the draft of the paper. All authors revised the paper. J.L. supervised the study.
Corresponding author
Ethics declarations
Competing interests
H. Sheng, R.L., A.J.L. and J.L. are on a patent application filed by Harvard University related to this work. X.W. is a scientific co-founder and equity holder of Stellaromics and Convergence Bio. P.L.F. and J.L. are co-founders of Axoft Inc. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Tal Dvir and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Test of implantation methods.
a, (Top) schematics showing an atomic force microscopy (AFM) setup for tissue elastic modulus measurement. (Bottom) elastic modulus of stage 15, 24, 32, 40, 48 Xenopus embryos, brain organoids, and cardiac organoids. Box plots indicate minimum, lower quartile, median, upper quartile, and maximum. Each dot represents a contact measurement. b, Photographs showing broken SU-8 mesh post-implantation (left) and the embryo before (middle) and after (right) mesh implantation, depicting damage to the embryo. c, BF microscopic images showing an embryo crushed by SU-8 meshes. d, BF microscopic images showing an embryo successfully implanted with a SEBS mesh. The dashed line circle highlights the portion of the mesh which remains exterior to the tadpole brain. e, Stress-strain curve of PFPE-DMA film with 8 kDa molecular weight, the blue dashed line indicates a linear relationship. f, Stretchability of 8 kDa PFPE-DMA and SU-8 films. g, Surface free energy of 8 kDa PFPE-DMA film and SU-8 films. h, Contact angle of phospholipid (cell membrane analog) on 8 kDa PFPE-DMA film, and on SU-8 film. Bar plots indicate mean ± s.e.m., each dot represents a sample, n = 5. i, Photographic images showing an embryo successfully implanted with a PFPE-DMA mesh. The dashed line circle highlights the portion of the mesh which remains exterior to the tadpole brain. j, Schematics showing how anchors fix the stretchable mesh electronics to the substrate, keeping the neural plate properly positioned during neurulation for device internalization, and keeping the stretchable mesh electrode array attached to the neural plate. The device’s initial dimensions and stretchability enable the stage 15 embryo to be slid under the device for implantation. k, The design of the stretchable mesh electronics showing the architecture of the stretchable mesh electrode array, stretchable serpentine interconnects, anchors, stretchable ribbons, and blockers. The blocker prevents the mesh electrodes from implanting into the caudal region of the neural plate. l, Schematics showing elongation of the neural tube during the embryo development of Xenopus laevis. The caudal region of the neural tube elongates to 3 times its initial length and forms the spinal cord while the cranial region elongates only 1.3 times its initial length and forms the brain.
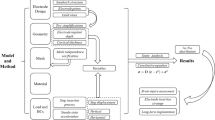
Extended Data Fig. 2 Mechanical simulation of stretchable mesh for brain implantation via embryo development.
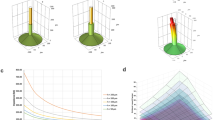
a, Snapshots of mechanical simulation of mesh-neural plate interaction (Fig. 1j, k), labeled with sequenced numbers. b, Snapshots of mechanical simulation procedure showing the stress distribution in the neural plates with and without stretchable mesh implanted. An embryo simulation without mesh implantation was used as a reference to calculate the additional stresses introduced by PFPE-DMA and SU-8 meshes. The red boxes highlight regions where the maximum stress was shown in (Fig. 1k).
Extended Data Fig. 3 Fabrication of PFPE-DMA-encapsulated stretchable mesh electronics.
a, b, Schematics showing the overlook (a) and section view (b) of the nitrogen chamber designed for use with the mask aligner in PFPE-DMA photopatterning. c, Schematic showing how the nitrogen chamber is used with mask aligner. d, Microscopic BF images showing representative high-resolution PFPE-DMA photolithography patterns made with the nitrogen chamber. e, f, Microscopic BF images showing the improved adhesion between Au interconnects and PFPE-DMA after inert gas plasma treatment. Dashed boxes highlight the sputtered regions on the PFPE-DMA layers. Without inert gas plasma treatment, Au interconnects peel off from the PFPE-DMA film after sputtering (e). With inert gas plasma treatment before sputtering, Au interconnects strongly bond to the PFPE-DMA film (f). g, Microscopic BF images showing the stretchable mesh electrode array region of PFPE-DMA device in fabrication steps corresponding to (Fig. 2c). Step 1 shows a homogeneous Ni layer. Step 2 is not included because the electrode array region does not have an SU-8 spacer. Step 3 shows Pt electrodes on the Ni layer. Electrodes are highlighted by red dashed circles. Steps 4-6 show sequential patterning of bottom PFPE-DMA, Au interconnects, and top PFPE-DMA layers. h, Schematics showing the post-fabrication steps of PFPE-DMA-encapsulated stretchable mesh electronics following (Fig. 2c). After fabrication, the device is soldered with a flexible flat cable (step 7) and bonded with a culture chamber (step 8). Then, the Ni layer is etched to release the device. Pt-black is electro-polymerized on electrodes to reduce electrode impedance. The device is washed with 0.1 × MMR and finally soaked in culture media (step 9).
Extended Data Fig. 4 Characterization of PFPE-DMA-encapsulated stretchable mesh electronics.
a, b, SEM images showing top views (a) and perspective views (b) of the stretchable mesh electrode array portion of the PFPE-DMA device. Each layer is pseudo-colored and labeled. c, BF image of a PFPE-DMA-encapsulated Au ribbon. d, (Left) atomic force microscopy (AFM) topography image of black dashed box-highlighted region in (c). (Right) height profiles of horizontal cross-sections highlighted in the left figure. e, BF images showing wrinkles of (left) straight and (right) serpentine PFPE-DMA-encapsulated interconnects. f, BF image of an Au interconnect without top PFPE-DMA passivation. g, (Left) AFM topography image of black dashed box-highlighted region in (f). (Right) height profiles of horizontal cross-sections highlighted in the left figure. h, SEM image showing perspective views of the PFPE-DMA stretchable mesh electronics after stretching and bending. i, SEM image of the dashed box-highlighted region in (h). j, SEM image showing cross-sections of PFPE-DMA-encapsulated Au interconnects, along the dashed line in (i). h-j, Each layer is pseudo-colored and labeled. k, Electrode impedance at 1 kHz in 37 °C PBS of PFPE-DMA mesh electronics before and after stretching and bending. Bar plots indicate mean ± s.e.m., each dot represents a single trial, two-tailed unpaired t-test, n = 4, ns, not significant. l, Resistance as a function of strain during the transverse stretch test of PFPE-DMA-encapsulated electronics. Red dots and line plots indicate mean ± s.d., and each gray dot and line plot represents one sample, n = 4. m, Electrochemical impedance spectroscopy of electrodes in stretchable mesh electronics with and without Pt-black coating. n, Electrode impedance at 1 kHz in 37 °C PBS of PFPE-DMA mesh electronics fabricated in different batches. Bar plots indicate mean ± s.e.m., each dot represents a sample, n = 4. o, Live cell ratio of wild-type rat cortical neurons after 10 days in vitro culture with PFPE-DMA mesh electronics, control, and with degradation products of PFPE-DMA. Bar plots indicate mean ± s.e.m., each dot represents a single trial, two-tailed unpaired t-test, n = 6, ns, not significant.
Extended Data Fig. 5 Staining methods and extended immunofluorescence images.
a, b, Schematics showing the protocols for tissue clearing and whole-mount staining (a) and cryosection staining (b) to characterize brain tissue implanted with stretchable mesh electronics. c, Whole-mount-stained 3D reconstructed confocal fluorescence image of implanted mesh electronics showing that the mesh is embedded in the neural tissue. d, 3D reconstructed confocal fluorescence images of a whole-mount-stained cyborg tadpole whose device was implanted in the middle of neurulation. e, Confocal fluorescence images showing coronal sections of the fore-, mid-, and hindbrain of cyborg tadpoles fixed at 2-, 4- and 8-days post fertilization. In all images, DAPI labels cell nuclei, acetylated-tubulin labels basal bodies, R6G labels the device, and SRY-box transcription factor 2 (Sox2) labels neural stem cells. In fluorescence images of whole-mount staining samples, HuC/D labels neurons. In fluorescence images of cryosection staining samples, Myt1 labels neurons.
Extended Data Fig. 6 Experimental setup, trajectory analysis, and examples of behavior tests.
a, Schematics showing the setup for behavioral testing. In each test, a tadpole is placed in a clear tank, sitting on an upward-facing screen. The screen is programmed to display the appropriate stimulation pattern for the color preference, visual avoidance, and edge preference tests. In the color preference test, the screen alternated between displaying half white and half black for 40 s each. In the visual avoidance test, a black dot is directly controllable via a computer mouse. The operator moved the dot toward the tadpole. If the tadpole responded, the operator would proceed to initiate the next encounter. If the tadpole did not respond, the operator would initiate a new encounter after five seconds. In the edge preference test, the entire screen is white. The setup is placed inside a dark box to minimize light contamination. The interior of the box is coated black to minimize reflections from the screen. Tadpoles are recorded using a video camera pointed down on the tank through a hole in the top of the box. b-i, Time-lapse snapshots of a visual avoidance video showing the trajectory process of a behaving tadpole. The colored lines connect the position of the tadpole in adjacent frames to form a trajectory. Crosses are labeled in frames where the tadpole met the black dot. j-l, Representative traces of behavior test data (top) and corresponding analyzed data (bottom). j, (Top) representative trajectories of tadpole movement in a color preference test. The green dotted lines indicate the boundary between the black and white areas. The right and left areas are white and black from 0–40 s, and switch colors from 40–80 s. Dashed and solid lines represent the trajectories of tadpole movement from 0–40 s and 40–80 s, respectively. (Bottom) distance of the tadpoles to the black and white boundary. Dashed and solid lines represent the trajectories of tadpole movement from 0–40 s and 40–80 s, respectively. Gray color highlights the black side. k, (Top) representative trajectories of tadpole movement in a visual avoidance test. Green crosses indicate the locations where the tadpole encountered the black dots. (Bottom) distance between the tadpole and the black dot during the test. l, (Top) representative trajectories of tadpoles in an edge preference test. The green ring indicates the outer quarter radius of the container defined as the edge in the experiment. (Bottom) distance between the tadpole and the container center during the test. The green color indicates the edge region. The statistical results of behavior tests, including examples in (j-l), were presented in Fig. 3o–q.
Extended Data Fig. 7 Experimental setup, raw data, and reference comparison of continuous electrophysiology in Xenopus embryonic brain development.
a, Schematics showing the oxygen anesthetic system used to minimize tadpole movement during culture for recording. The system mixes the anesthetic media with fresh oxygen as previously reported55 to minimize the effects of anesthesia on tadpole development. b, Schematics showing the recording setup for electrophysiological experiments. During recording, the culture chamber is placed in a Faraday cage on a grounded optic table. The input/output of the implanted mesh electronics is connected to a recording system using a flexible flat cable (FFC) connector. A Pt probe is placed in the culture media as ground. c, Raw data of continuous recordings shown in (Fig. 4). d-i, Reference comparison of continuous electrophysiology. Distribution plots showing comparisons of oscillation signal width (d) and interval (e); calcium-wave like signal width (f) and interval (g); spike width (h) and interval (i). Reference data is as follows: reference 166, reference 234, and reference 335. Reference 3 did not include the corresponding dataset for spike intervals, so it is not included in (i). The results in (d-i) are determined from signals collected from three cyborg tadpoles.
Extended Data Fig. 8 Analysis of continuous electrophysiology in Xenopus embryonic brain development.
a-g, Pharmacological tests. a, Dynamics of calcium wave-like signals during the pharmacological test. Representative voltage traces from the cyborg tadpole under serial pharmacological test conditions of no drug, cyanquixaline (CNQX)/[2 R]-amino-5-phosphonopentanoate (APV), wash of CNQX/APV, and CNQX/APV/ tetrodotoxin (TTX). b, The change of wave number per minute during the pharmacological test. Bar plots indicate mean ± s.d., each dot represents a recording trial, two-tailed unpaired t-test, n = 3, No drug vs APV/CNQX, ****, p < 0.0001; APV/CNQX vs. Wash, ***, p = 0.0004; Wash vs. APV/CNQX/TTX, ***, p = 0.0004. c-g, Dynamics of spike-like signals during the pharmacological test. c, Statistical summary of the firing rate changes under different conditions. The tadpole was treated with APV/CNQX, washed, then CNQX/APV/ TTX in series. Bar plots indicate mean ± s.d., each dot represents a recording trial, two-tailed unpaired t-test, n = 3, No drug vs. APV/CNQX, *, p = 0.0343; APV/CNQX vs. Wash, **, p = 0.0064; Wash vs. APV/CNQX/TTX, **, p = 0.0024. d, Representative voltage traces from the cyborg tadpole under serial pharmacological test conditions of no drug, bicuculline (BIC)/picrotoxin (PTX), washed, followed by BIC/PTX/TTX. e, f, Zoomed-in views of the signal highlighted by the green (e) and magenta (f) boxes in (d). g, Statistical summary of the firing rate changes under different conditions. The tadpole was treated with BIC/PTX, washed, then BIC/PTX/TTX in series. Bar plots indicate mean ± s.d., each dot represents a recording trial, two-tailed unpaired t-test, n = 3, No drug vs. BIC/PTX, *, p = 0.0146; BIC/PTX vs. Wash, *, p = 0.0202; Wash vs. BIC/PTX/TTX, *, p = 0.0356. h-o, Correlation of single-unit action potential signals with the corresponding electrode position. h, Representative voltage traces from a cyborg tadpole showing single-unit action potentials. i, j, Zoomed-in views of the signal highlighted by green (i) and magenta (j) boxes in (h). k, Mean spike superimposed on all spikes from the same unit, sorted by spike sorting of data in (h). l-o, Confocal fluorescence images of the cyborg tadpole brain slice showing DAPI (l), HuC/D (m), BF (n) and overlaid (o) channels. The white dashed circles highlight the position of the electrode that recorded the voltage trace in (h). p, Correlation coefficient between channels of stages 24, 26, stage 40 local field potential and stage 40 spike signals. Positive correlation corresponds to a coefficient of 1, negative to -1, and no correlation to 0. White dots represent the lower quartile, median, upper quartile from bottom to top. Each translucent dot represents a sample, two-tailed unpaired t-test, ****, p < 0.0001. q, Standard deviation of time delay between channels of stages 24, 26, and 40 local field potential signals. Lower standard deviations indicate greater synchronization between channels. Upper and lower whiskers of the box plots represent maximum and minimum. Top, middle, and bottom lines of the box plots represent upper quartile, median and lower quartile. Each white dot represents a sample, two-tailed unpaired t-test, n ≥ 5, Stage 24 vs. Stage 26, *, p = 0.0102; Stage 26 vs. Stage 40, *, p = 0.0162; Stage 24 vs. Stage 40, ***, p = 0.0003. r-t, Propagating wave signals in stage 20 embryonic brain. r, Schematic of the cyborg tadpole at developmental stage 20. s, Representative voltage traces from four channels in the cyborg tadpole at stage 20. t, Zoomed-in views of the signals highlighted by dashed lines in (s).
Extended Data Fig. 9 Soft and stretchable high-density mesh electrode array for tracking neural activities.
a, Schematics showing the electron-beam (e-beam) lithographic fabrication of PFPE-DMA-encapsulated stretchable mesh electronics with a 32-channel high-density electrode array. First, a Ni layer is deposited on a blank silicon oxide wafer as a sacrificial layer. A SU-8 layer is patterned as a spacer, Pt electrodes are photolithographically patterned (step 1). Then, the bottom PFPE-DMA (step 2), Au interconnects (step 3), and top PFPE-DMA layer (step 4) are lithographically patterned. Au layer is patterned by e-beam lithography. PFPE-DMA layers are patterned by photolithography. Zoomed-in images show the details of the electrode arrays (highlighted in blue dashed boxes) and individual electrodes (highlighted in red dashed boxes). b, Photographic images showing the free-floating 32-channel high-density electrode array during stretching, bending, and twisting. c, BF microscopic image showing a representative 128-channel electrode array. d, Zoomed-in view of the red dashed box-highlighted region in (c) showing the high-density electrodes and interconnects. e, Zoomed-in view of the blue dashed box-highlighted region in (d) showing the stretchable design. f, Zoomed-in view of the green dashed box-highlighted region in (e) showing the individual electrodes and interconnects. g, Single-unit waveform centroids (n = 20 neurons) from the continuous 5-day recording of the cyborg axolotl embryo in (Fig. 5). Centroid computed using spatial average across electrode positions weighted by the square of the mean waveform amplitude at each electrode. Grey patterns indicate the positions and sizes of the mesh electrodes. h, Single-unit waveform centroid displacement throughout the 5-day recording in (Fig. 5). Centroids from the same day are labeled with the same color. Centroids for the same units across different days are connected by lines. Grey patterns indicate the positions and sizes of the mesh electrodes. i, Average displacement of single-unit centroids across different days of the 5-day recording in (Fig. 5). Grey contours indicate quintile boundaries of the distribution of centroid position displacement.
Extended Data Fig. 10 Single-unit action potential in axolotl embryonic brain development.
a, Representative average single-unit waveforms recorded from the same neurons overlaid with extremum electrodes over the 5-day recording period in (Fig. 5). b, UMAP visualization of units sorted from the continuous recording of the tail-amputated cyborg axolotl embryo, as shown in (Fig. 6). c, Firing rates of units recorded immediately before and after tail amputation of cyborg axolotl embryo, as shown in (Fig. 6). Bar plots indicate mean ± s.d., with individual dots representing firing rate of individual units, two-tailed unpaired t-test, n = 30, ****, p < 0.0001. d, Normalized presence of units sorted from continuous recording of the tail-amputated axolotl embryo, as shown in (Fig. 6).
Supplementary information
Supplementary Information (download PDF )
Supplementary Discussion, Figs. 1–8 and Tables 1–3.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sheng, H., Liu, R., Li, Q. et al. Brain implantation of soft bioelectronics via embryonic development. Nature 642, 954–964 (2025). https://doi.org/10.1038/s41586-025-09106-8
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09106-8
This article is cited by
-
Conforming strategies for bioelectronics on arbitrary surfaces
npj Biosensing (2025)
-
Polymer Materials for Stretchable Electronics Encapsulation
Chemical Research in Chinese Universities (2025)