Abstract
The poly(ADP-ribose) polymerase inhibitor (PARPi) class of drugs represents a remarkable advance in the treatment of patients with homologous recombination-deficient tumours, but resistance remains a challenge1,2,3,4,5. Although most research has focused on the downstream consequences of PARPi exposure to tackle resistance, the immediate effect of PARP inhibition on the chromatin environment and its contribution to PARPi toxicity remains elusive. Here we show that PARP inhibition induces histone release from the chromatin. This presents a vulnerability of PARPi-resistant cancer cells, which require histone homeostasis mechanisms to sustain elevated DNA replication rates and survival. Through functional genetic screens, we identified NASP as a key factor in maintaining the stability of evicted histones via its TPR motifs. Loss of NASP renders tumour cells hypersensitive to PARPi treatment in vitro and in vivo, impairs replication fork progression and elevates levels of replication-associated DNA damage. Moreover, NASP acts together with the INO80 complex and the chaperoning activity of PARP1 to ensure efficient histone turnover and prevent the accumulation of lethal DNA damage. Collectively, our work reports on histone eviction as an immediate cellular response to PARPi treatment and provides a promising avenue for targeting histone supply pathways to overcome PARPi resistance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data required to evaluate the conclusions of this work are presented in the article or included in the supplementary material. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD061071. Raw data of the synthetic lethality screens are available at the European Nucleotide Archive under PRJEB86492, WT HAP1 cells have been previously reported24 and repli-ATAC data are accessible under PRJEB90052. Hi-C sequencing data have been deposited in the NCBI Gene Eexpression Omnibus database under GSE293482. Data for the FACS-based haploid genetic screen for NASP levels are also available on an interactive visualization platform (https://phenosaurus.nki.nl). Source data are provided with this paper.
References
Bryant, H. E. et al. Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature 434, 913–917 (2005).
Farmer, H. et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434, 917–921 (2005).
Lord, C. J. & Ashworth, A. PARP inhibitors: synthetic lethality in the clinic. Science 355, 1152–1158 (2017).
Dias, M. P., Moser, S. C., Ganesan, S. & Jonkers, J. Understanding and overcoming resistance to PARP inhibitors in cancer therapy. Nat. Rev. Clin. Oncol. 18, 773–791 (2021).
DiSilvestro, P. et al. Overall survival with maintenance olaparib at a 7-year follow-up in patients with newly diagnosed advanced ovarian cancer and a BRCA mutation: the SOLO1/GOG 3004 trial. J. Clin. Oncol. 41, 609–617 (2023).
Dobzhansky, T. Genetics of natural populations; recombination and variability in populations of Drosophila pseudoobscura. Genetics 31, 269–290 (1946).
Bridges, C. B. The origin of variations in sexual and sex-limited characters. Am. Nat. 56, 51–63 (1922).
Murai, J. et al. Trapping of PARP1 and PARP2 by clinical PARP inhibitors. Cancer Res. 72, 5588–5599 (2012).
Huang, D. & Kraus, W. L. The expanding universe of PARP1-mediated molecular and therapeutic mechanisms. Mol. Cell 82, 2315–2334 (2022).
Ray Chaudhuri, A. & Nussenzweig, A. The multifaceted roles of PARP1 in DNA repair and chromatin remodelling. Nat. Rev. Mol. Cell Biol. 18, 610–621 (2017).
Poirier, G. G., de Murcia, G., Jongstra-Bilen, J., Niedergang, C. & Mandel, P. Poly(ADP-ribosyl)ation of polynucleosomes causes relaxation of chromatin structure. Proc. Natl Acad. Sci. USA 79, 3423–3427 (1982).
Kruhlak, M. J. et al. Changes in chromatin structure and mobility in living cells at sites of DNA double-strand breaks. J. Cell Biol. 172, 823–834 (2006).
Ledermann, J. et al. Olaparib maintenance therapy in patients with platinum-sensitive relapsed serous ovarian cancer: a preplanned retrospective analysis of outcomes by BRCA status in a randomised phase 2 trial. Lancet Oncol. 15, 852–861 (2014).
Pujade-Lauraine, E. et al. Olaparib tablets as maintenance therapy in patients with platinum-sensitive, relapsed ovarian cancer and a BRCA1/2 mutation (SOLO2/ENGOT-Ov21): a double-blind, randomised, placebo-controlled, phase 3 trial. Lancet Oncol. 18, 1274–1284 (2017).
Mateos-Gomez, P. A. et al. Mammalian polymerase θ promotes alternative NHEJ and suppresses recombination. Nature 518, 254–257 (2015).
Ceccaldi, R. et al. Homologous-recombination-deficient tumours are dependent on Polθ-mediated repair. Nature 518, 258–262 (2015).
Zatreanu, D. et al. Polθ inhibitors elicit BRCA-gene synthetic lethality and target PARP inhibitor resistance. Nat. Commun. 12, 3636 (2021).
Paes Dias, M. et al. Loss of nuclear DNA ligase III reverts PARP inhibitor resistance in BRCA1/53BP1 double-deficient cells by exposing ssDNA gaps. Mol. Cell 81, 4692–4708.e9 (2021).
Zhou, J. et al. A first-in-class polymerase θ inhibitor selectively targets homologous-recombination-deficient tumors. Nat. Cancer 2, 598–610 (2021).
Zong, D. et al. BRCA1 haploinsufficiency is masked by RNF168-mediated chromatin ubiquitylation. Mol. Cell 73, 1267–1281.e7 (2019).
Bouwman, P. et al. 53BP1 loss rescues BRCA1 deficiency and is associated with triple-negative and BRCA-mutated breast cancers. Nat. Struct. Mol. Biol. 17, 688–695 (2010).
Bunting, S. F. et al. 53BP1 inhibits homologous recombination in Brca1-deficient cells by blocking resection of DNA breaks. Cell 141, 243–254 (2010).
Boon, N. J. et al. DNA damage induces p53-independent apoptosis through ribosome stalling. Science 384, 785–792 (2024).
Blomen, V. A. et al. Gene essentiality and synthetic lethality in haploid human cells. Science 350, 1092–1096 (2015).
Adelman, C. A. et al. HELQ promotes RAD51 paralogue-dependent repair to avert germ cell loss and tumorigenesis. Nature 502, 381–384 (2013).
Setton, J. et al. Germline RAD51B variants confer susceptibility to breast and ovarian cancers deficient in homologous recombination. NPJ Breast Cancer 7, 135 (2021).
Mazouzi, A. et al. FIRRM/C1orf112 mediates resolution of homologous recombination intermediates in response to DNA interstrand crosslinks. Sci. Adv. 9, eadf4409 (2023).
Gibbs-Seymour, I., Fontana, P., Rack, J. G. M. & Ahel, I. HPF1/C4orf27 is a PARP-1-interacting protein that regulates PARP-1 ADP-ribosylation activity. Mol. Cell 62, 432–442 (2016).
Hammond, C. M. et al. DNAJC9 integrates heat shock molecular chaperones into the histone chaperone network. Mol. Cell 81, 2533–2548.e9 (2021).
Cook, A. J., Gurard-Levin, Z. A., Vassias, I. & Almouzni, G. A specific function for the histone chaperone NASP to fine-tune a reservoir of soluble H3–H4 in the histone supply chain. Mol. Cell 44, 918–927 (2011).
Osakabe, A. et al. Nucleosome formation activity of human somatic nuclear autoantigenic sperm protein (sNASP). J. Biol. Chem. 285, 11913–11921 (2010).
Hormazabal, J. et al. Chaperone mediated autophagy contributes to the newly synthesized histones H3 and H4 quality control. Nucleic Acids Res. 50, 1875–1887 (2022).
Richardson, R. T. et al. Characterization of the histone H1-binding protein, NASP, as a cell cycle-regulated somatic protein. J. Biol. Chem. 275, 30378–30386 (2000).
Liu, C. P. et al. Distinct histone H3–H4 binding modes of sNASP reveal the basis for cooperation and competition of histone chaperones. Genes Dev. 35, 1610–1624 (2021).
Bao, H. et al. NASP maintains histone H3–H4 homeostasis through two distinct H3 binding modes. Nucleic Acids Res. 50, 5349–5368 (2022).
Jaspers, J. E. et al. Loss of 53BP1 causes PARP inhibitor resistance in Brca1-mutated mouse mammary tumors. Cancer Discov. 3, 68–81 (2013).
Staaf, J. et al. Whole-genome sequencing of triple-negative breast cancers in a population-based clinical study. Nat. Med. 25, 1526–1533 (2019).
Harvey-Jones, E. et al. Longitudinal profiling identifies co-occurring BRCA1/2 reversions, TP53BP1, RIF1 and PAXIP1 mutations in PARP inhibitor-resistant advanced breast cancer. Ann. Oncol. 35, 364–380 (2024).
Nabet, B. et al. The dTAG system for immediate and target-specific protein degradation. Nat. Chem. Biol. 14, 431–441 (2018).
Stewart-Morgan, K. R. & Groth, A. Profiling chromatin accessibility on replicated DNA with repli-ATAC-seq. Methods Mol. Biol. 2611, 71–84 (2023).
Lim, P. X., Zaman, M., Feng, W. & Jasin, M. BRCA2 promotes genomic integrity and therapy resistance primarily through its role in homology-directed repair. Mol. Cell 84, 447–462.e10 (2024).
Xia, Y. et al. RNF8 mediates histone H3 ubiquitylation and promotes glycolysis and tumorigenesis. J. Exp. Med. 214, 1843–1855 (2017).
Singh, R. K., Kabbaj, M. H., Paik, J. & Gunjan, A. Histone levels are regulated by phosphorylation and ubiquitylation-dependent proteolysis. Nat. Cell Biol. 11, 925–933 (2009).
Shroff, M., Knebel, A., Toth, R. & Rouse, J. A complex comprising C15ORF41 and codanin-1: the products of two genes mutated in congenital dyserythropoietic anaemia type I (CDA-I). Biochem. J. 477, 1893–1905 (2020).
Hogan, A. K. et al. UBR7 acts as a histone chaperone for post-nucleosomal histone H3. EMBO J. 40, e108307 (2021).
Hauer, M. H. et al. Histone degradation in response to DNA damage enhances chromatin dynamics and recombination rates. Nat. Struct. Mol. Biol. 24, 99–107 (2017).
Tsukuda, T., Fleming, A. B., Nickoloff, J. A. & Osley, M. A. Chromatin remodelling at a DNA double-strand break site in Saccharomyces cerevisiae. Nature 438, 379–383 (2005).
Verma, P. et al. ALC1 links chromatin accessibility to PARP inhibitor response in homologous recombination-deficient cells. Nat. Cell Biol. 23, 160–171 (2021).
Hewitt, G. et al. Defective ALC1 nucleosome remodeling confers PARPi sensitization and synthetic lethality with HRD. Mol. Cell 81, 767–783.e11 (2021).
Gogola, E. et al. Selective loss of PARG restores PARylation and counteracts PARP inhibitor-mediated synthetic lethality. Cancer Cell 33, 1078–1093.e12 (2018).
Muthurajan, U. M. et al. Automodification switches PARP-1 function from chromatin architectural protein to histone chaperone. Proc. Natl Acad. Sci. USA 111, 12752–12757 (2014).
Challa, K. et al. Damage-induced chromatome dynamics link ubiquitin ligase and proteasome recruitment to histone loss and efficient DNA repair. Mol. Cell 81, 811–829.e6 (2021).
Campos, E. I. et al. The program for processing newly synthesized histones H3.1 and H4. Nat. Struct. Mol. Biol. 17, 1343–1351 (2010).
Wang, H., Walsh, S. T. & Parthun, M. R. Expanded binding specificity of the human histone chaperone NASP. Nucleic Acids Res. 36, 5763–5772 (2008).
Plessier, A. et al. Proteomic profiling of UV damage repair patches uncovers histone chaperones with central functions in chromatin repair. Preprint at bioRxiv https://doi.org/10.1101/2024.08.23.609352 (2024).
Lee, S. B. et al. Tousled-like kinases stabilize replication forks and show synthetic lethality with checkpoint and PARP inhibitors. Sci. Adv. 4, eaat4985 (2018).
Ali-Fehmi, R. et al. Analysis of the expression of human tumor antigens in ovarian cancer tissues. Cancer Biomark. 6, 33–48 (2010).
Poli, J., Gasser, S. M. & Papamichos-Chronakis, M. The INO80 remodeller in transcription, replication and repair. Phil. Trans. R. Soc. B https://doi.org/10.1098/rstb.2016.0290 (2017).
Nishi, R. et al. Systematic characterization of deubiquitylating enzymes for roles in maintaining genome integrity. Nat. Cell Biol. 16, 1016–1026 (2014).
Zimmermann, M. et al. CRISPR screens identify genomic ribonucleotides as a source of PARP-trapping lesions. Nature 559, 285–289 (2018).
Reveron-Gomez, N. et al. Accurate recycling of parental histones reproduces the histone modification landscape during DNA replication. Mol. Cell 72, 239–249.e5 (2018).
Escobar, T. M. et al. Active and repressed chromatin domains exhibit distinct nucleosome segregation during DNA replication. Cell 179, 953–963.e11 (2019).
Li, N. et al. Parental histone transfer caught at the replication fork. Nature 627, 890–897 (2024).
Petryk, N. et al. MCM2 promotes symmetric inheritance of modified histones during DNA replication. Science 361, 1389–1392 (2018).
Carette, J. E. et al. Generation of iPSCs from cultured human malignant cells. Blood 115, 4039–4042 (2010).
Barazas, M. et al. Radiosensitivity is an acquired vulnerability of PARPi-resistant BRCA1-deficient tumors. Cancer Res. 79, 452–460 (2019).
Drean, A. et al. Modeling therapy resistance in BRCA1/2-mutant cancers. Mol. Cancer Ther. 16, 2022–2034 (2017).
Demichev, V., Messner, C. B., Vernardis, S. I., Lilley, K. S. & Ralser, M. DIA-NN: neural networks and interference correction enable deep proteome coverage in high throughput. Nat. Methods 17, 41–44 (2020).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Hughes, C. S. et al. Single-pot, solid-phase-enhanced sample preparation for proteomics experiments. Nat. Protoc. 14, 68–85 (2019).
MacLean, B. et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26, 966–968 (2010).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 259 (2015).
Flyamer, I. M. et al. Single-nucleus Hi-C reveals unique chromatin reorganization at oocyte-to-zygote transition. Nature 544, 110–114 (2017).
Brockmann, M. et al. Genetic wiring maps of single-cell protein states reveal an off-switch for GPCR signalling. Nature 546, 307–311 (2017).
Bhin, J. et al. Multi-omics analysis reveals distinct non-reversion mechanisms of PARPi resistance in BRCA1- versus BRCA2-deficient mammary tumors. Cell Rep. 42, 112538 (2023).
Iorio, F. et al. A landscape of pharmacogenomic interactions in cancer. Cell 166, 740–754 (2016).
Dreyer, J. et al. Acute multi-level response to defective de novo chromatin assembly in S-phase. Mol. Cell 84, 4711–4728.e10 (2024).
Haarhuis, J. H. I. et al. A Mediator–cohesin axis controls heterochromatin domain formation. Nat. Commun. 13, 754 (2022).
Luna-Vargas, M. P. et al. Enabling high-throughput ligation-independent cloning and protein expression for the family of ubiquitin specific proteases. J. Struct. Biol. 175, 113–119 (2011).
Acknowledgements
We thank members of the Jonkers laboratory for helpful discussions; E. de Wit and M. Martinovic for their assistance with the initial ATAC sequencing experiments; R. van der Weide for help with the analysis of the repli-ATAC sequencing; P. Kloosterman for bioinformatic assistance; C. Lutz for assistance with generation of lentiviruses; the NKI Flow cytometry facility, Genomics Core facility, the preclinical intervention unit of the Mouse Clinic for Cancer and Aging (MCCA) and the animal facility for technical support; and R. Chapman, M. Tarsounas and R. Beijersbergen for sharing reagents. Schematics were created using BioRender (https://biorender.com). Research at the Netherlands Cancer Institute is supported by institutional grants of the Dutch Cancer Society and the Dutch Ministry of Health, Welfare and Sport. The Jonkers and the Brummelkamp laboratories have received funding from the Oncode Institute, which is partially financed by the Dutch Cancer Society. Research in the Jonkers laboratory is funded by the Dutch Cancer Society (grants 14516 and 14949), Artios Pharma and the Swiss National Science Foundation (grant 320030M_219453). S.C.M. received a Boehringer Ingelheim Fonds PhD Fellowship. B.D.R. has received funding from the Dutch Cancer Society (12687) and the Dutch Research Council (NWO) (VI.C.202.098). O.B.B. and L.H. are supported by the Dutch NWO X-omics Initiative. A.M. is supported by an EMBO Long Term Fellowship (ALTF 158-2018). The Tutt and Lord laboratory is supported by Breast Cancer Now, Cancer Research UK and the Breast Cancer Research Foundation.
Author information
Authors and Affiliations
Contributions
This study was conceptualized by S.C.M., A.M. and J.J. S.C.M. and A.M. performed the haploid genetic screens. S.C.M., A.K., J.R., E.P., M.L.K. and I.v.d.H. performed and analysed all cell line experiments. G.R., V.B. and F.M. generated and analysed the repli-ATAC sequencing data. O.B.B. and L.H. performed and analysed the proteomics experiments. S.d.S. performed and analysed the mass spectrometry-based analysis of histone PTMs under supervision of M.V.-A. A.F. and S.C. purified the recombinant proteins and performed surface plasmon resonance assays. J.H.I.H. and R.O. performed and analysed the Hi-C experiment under the supervision of B.D.R. J.S. and M.v.d.V. performed the in vivo intervention study. L.R.-M., S.J.P., A.N.J.T. and C.J.L. analysed NASP expression in the tumour transcriptomic data. D.J.V. and L.F.A.W. analysed NASP expression in the human cell line panels. S.C.M., A.M. and J.J. wrote the manuscript with input from all authors. The study was supervised by F.M., T.R.B., A.M. and J.J.
Corresponding authors
Ethics declarations
Competing interests
M.V.-A. is a shareholder of MoleQlar Analytics. T.R.B. is a co-founder and shareholder of Scenic Biotech B.V. J.J. has received research funding from Artios Pharma. C.J.L. receives and/or has received research funding from AstraZeneca, Merck KGaA, Artios, Neophore and FoRx; received consultancy, SAB membership or honoraria payments from FoRx, Syncona, Sun Pharma, Gerson Lehrman Group, Merck KGaA, Vertex, AstraZeneca, Tango Therapeutics, 3rd Rock, Ono Pharma, Artios, Abingworth, Tesselate, Dark Blue Therapeutics, Pontifax, Astex, Neophore, GlaxoSmithKline, Dawn Bioventures, Blacksmith Medicines, ForEx and Ariceum; and has stock in Tango, Ovibio, Hysplex, Tesselate and Ariceum. A.N.J.T. is or has been a consultant for AstraZeneca, Merck KGaA, Artios, Pfizer, Vertex, Gilead and MD Anderson Cancer Centre; and has received grant and/or research support from AstraZeneca, Myriad and Merck KgaA. C.J.L. and A.N.J.T. are named inventors on patents describing the use of DNA repair inhibitors and stand to gain from their development and use as part of the ICR ‘Rewards to Inventors’ scheme and also report benefits from this scheme associated with patents for PARP inhibitors paid to the research accounts for C.J.L. and A.N.J.T. at the Institute of Cancer Research. All other authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 HAP1 ΔBRCA1Δ53BP1 are PARP inhibitor resistant and PARPi treatment does not alter overall chromatin conformation.
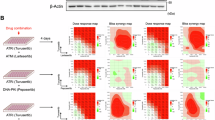
(a) Immunoblot depicting HAP1 cells of indicated genotypes, Actin is used as a loading control. (b) Clonogenic assay in HAP1 cells of indicated genotypes were exposed for seven days to indicated concentrations of olaparib, n = 3 (c) Immunofluorescence staining for RAD51 foci formation in HAP1 cells exposed to 1μg/mL doxycycline, 0.5 mM indole-3-acetic acid (IAA) and 100 ng/mL neocarcinostatin (NCS) for two hours respectively. Representative plots of experimental triplicates are shown. P value was calculated by a Mann-Whitney U test, ****P < 0.0001. (d) Hi-C contact matrices for chromosome 10 in HAP1 ΔBRCA1Δ53BP1 cells left untreated or exposed to 1 μM olaparib for 24 h. (e) Relative contact probability depicting the likelihood of a contact at increasing length scales is shown for indicated treatments and genotypes.
Extended Data Fig. 2 PARP inhibitor treatment results in a histone release irrespective of genetic background.
(a) Schematic depiction of the subcellular fractionations performed throughout the manuscript. The schematic was created using BioRender (https://biorender.com). (b-d) Relative abundance of indicated marker proteins across different subcellular fractions in comparison to whole-cell extracts (WCE). (e) Subcellular fractionation was performed in HAP1 ΔBRCA1Δ53BP1 cells and levels of histone H2A-H2B and H3K4me3 were measured. Related to Fig. 1a, PCNA serves as a loading control. (f, g) Quantification of alterations in H2A-H2B and H3-H4 isoforms in indicated cellular fractions upon olaparib treatment compared to unchallenged conditions. Each dot represents the mean of a biological triplicate, related to Fig. 1b. (h) Levels of indicated histone PTMs in subcellular fractions upon olaparib treatment. Post-nucleosomal marks are labeled in blue and pre-nucleosomal marks in red. Empty H3-H4 tetramer was loaded to control for antibody specificity. (i-n) Subcellular fractionations in HAP1 WT (i), HAP1 BRCA1-AID cells exposed to 1μg/mL doxycycline, 0.5 mM IAA (j), SUM149PT(k), SUM149PT-BRCA1 reverted cells (l), RPE-hTERT (m) and human mammary epithelial cells (HMEC-hTERT, n). All cell lines were exposed to the indicated PARPi for 24 h before collection. Data show a representative of biological triplicates, except (k, l) which were performed in duplicates.
Extended Data Fig. 3 Haploid genetic screens identify known and novel modulators of PARP inhibitor response.
(a) Scatter plots displaying the ratios (sense versus total insertions) for two biological replicates of ΔBRCA1Δ53BP1 cell clones passaged in the presence or absence of 125 nM olaparib for 10 days. Each dot represents one gene and hits scoring differentially compared to the wild type are marked in dark blue. (b) Gene ontology terms of the genes identified in untreated and olaparib treated conditions in ΔBRCA1Δ53BP1 cells. (c) Heatmap depicting DNA repair genes identified in the genetic screens. INO80 complex members are marked in bold. (d) Immunoblot depicting HAP1 cells transduced with sgRNAs targeting DNAJC9 or a non-targeting control. (e, f) Clonogenic assay (e) and quantification (f) of HAP1 cells with indicated genotypes exposed to different concentrations of olaparib for seven days. Scans and mean ± SEM of a representative experiment from three biological duplicates is depicted.
Extended Data Fig. 4 NASP binding to H3 is crucial to maintain histone stability upon PARPi treatment.
(a) NASP levels quantified by mass spectrometry across different subcellular fractions. Data are represented as the relative abundance compared to whole cell extract (WCE), mean of three biological replicates. Significance is calculated by a two-tailed t-test, **P < 0.01. (b, c) Levels of H2A-H2B (b) and H3-H4 (c) in indicated subcellular fractions upon olaparib exposure in HAP1 ΔBRCA1Δ53BP1 cells of indicated genotype quantified by mass spectrometry compared to untreated conditions, related to Fig. 2b. Data for WT cells are also already displayed in Extended Data Fig. 2f,g. (d) KB1P-G3 Δ53BP1 cells were transduced with indicated GFP-tagged histone variants and analysed by immunoblotting. (e) WT or ΔNASP cells were treated with 1 μM olaparib for 24 h and with 10 μM MG132 or 200 nM bafilomycin for the final 6 h and subjected to subcellular fractionation. Whole cell extract (WCE) is depicted, related to Fig. 2f. (f) Schematic depiction of the NASP isoforms, somatic (sNASP) and testicular (tNASP). (g, h) Co-immunoprecipitation in HAP1 ΔBRCA1Δ53BP1 cells expressing a doxycycline-inducible construct of GFP-sNASP, treated as indicated with 2μg/mL doxycycline and 1 μM olaparib for 24 h. After collection, cells were fractioned and co-IP was performed in Fraction B. Representative images of three biological replicates are displayed, asterisk indicates an unspecific band.
Extended Data Fig. 5 Loss of NASP sensitizes cells to PARP inhibition irrespective of genetic background.
(a) Clonogenic assay in ΔBRCA1Δ53BP1 HAP1 cells exposed to specified concentrations of olaparib. Quantification is depicted in Fig. 3b. (b to d) HAP1 ΔBRCA1Δ53BP1 cells were exposed to indicated PARPi for seven days and viability was determined. (e) Immunoblot depicting HAP1 single cell clones carrying an isoform specific knockout for tNASP and a complete NASP knockout cell line. (f) Colony formation assay in WT HAP1 cells, lacking tNASP and full NASP knockout cells exposed to indicated concentrations of olaparib. Cellular viability was assessed seven days after seeding. (g) Immunblot depicting NASP levels in WT HAP1 and ΔNASP single cell clones. (h, i) Clonogenic assay (h) and quantification (i) in HAP1 wild type cells exposed to different concentrations of olaparib for seven days. (j, l, n) SUM149PT ΔSHLD2, SUM149PT-BRCA1rev and PEO4 cells transduced with non-targeting (NT) or sgRNAs targeting NASP were analysed for NASP levels. (k, m, o) Cells displayed in (j, l, n) were plated in a clonogenic assay and exposed to indicated concentrations of olaparib for ten days. (p, q) Related to Fig. 3j. Plot depicting tumor outgrowth in the mammary fat pad of individual animals (p) and animals pooled into experimental groups (q). Animals were sacrificed once tumor size reached >1500mm3. (a-o) Representative experiments of three biological replicates are depicted as mean ± SEM.
Extended Data Fig. 6 NASP loss results in sensitivity to DNA-protein adducts and NASP expression is high irrespective of PARPi response.
(a-h) Quantification of colony formation assays in HAP1 ΔBRCA1Δ53BP1 or KB1P-G3 Δ53BP1 cells treated with indicated DNA damaging agents for seven days. HU=hydroxyurea, IR=ionizing radiation, MMS= methylmethanesulfonate. Data are displayed as mean ± SEM and representatives of three independent replicates are shown. (i) Normalised NASP mRNA expression in 211 HRDetect classified breast cancers from the SCAN-B study37. Data are depicted as expression data from individual cancers and line depicts the median, *p < 0.001, Mann-Whitney test. (j) NASP mRNA expression in patient matched BRCA1/2-mutant breast tumors isolated before (pre) or after (post) olaparib treatment from 11 patients described as part of the BTBC study38. Normalised expression levels (log2CPM) for NASP are shown. Statistical significance was assessed using a paired t-test. (k, l) NASP gene expression levels in Brca1/2;p53-deficient mouse mammary tumors grouped according to their PARP inhibitor response76. (m, n) NASP gene expression levels in human cancer cell lines correlated with their response to olaparib (m) and talazoparib (n).
Extended Data Fig. 7 Loss of NASP results in increased chromatin accessibility at replication forks.
(a) Western blot of HAP1 cells carrying a degradation tag (dTAG) at the C-terminus of NASP treated with 1 μM dTAG-13 for 18 h or left untreated. (b) Immunofluorescence staining for NASP and HA-signal in NASP-dTAG cells left untreated or treated with 1 μM dTAG-13 for 18 h. (c) Ratio between spiked-in Drosophila melanogaster and human EdU reads of dechromatinized DNA samples and repli-ATAC samples. Dots represent biological replicates and data are used to control for effects of EdU incorporation in processed repli-ATAC-seq samples. (d) Accessibility changes upon dTAG-13 treatment across different chromatin regions in HAP1 NASP-dTAG cells. Representative of two biological replicates is shown. (e, f) Quantification (e) and representative plots (f) of cell cycle profiles of HAP1 cells stained with DAPI. Cells were left untreated or treated with 1 μM olaparib for 24 h and collected or recovered for an additional 24 h. Each dot represents one biological replicate, data are displayed as mean ± SEM. P value was calculated by a two-tailed t test, *P < 0.05. (g) Immunoblot depicting HAP1 cells of indicated genotypes transduced with a non-targeting (NT) guide RNA or a sgRNA targeting 53BP1. (h) Cells depicted in (g) were subjected to a colony formation assay and viability was assessed after seven days. (i) RAD51 foci formation was quantified in HAP1 ΔBRCA1Δ53BP1 cells treated with 1 μM olaparib for 24 h and recovered for indicated times. Graph displays the mean number of foci per cell and the 10 and 90th percentile. n > 1500 cells per condition.
Extended Data Fig. 8 Loss of known NASP interactors does not confer PARP inhibitor sensitivity.
(a) Western blots of HAP1 cells with indicated genotypes. (b-d) Colony formation assays in HAP1 cells, cells were exposed to olaparib for seven days and viability was measured. (e) Western blot of HAP1 cells transduced with non-targeting (NT) or a sgRNA targeting HAT1. (f) Cells depicted in (e) were plated in a colony formation assay and exposed to indicated concentrations of olaparib for seven days.
Extended Data Fig. 9 Genetic loss of PARP1 reduces proliferation of ΔNASP cells.
(a) Western blot of nuclear extracts, related to Fig. 5c. (b) KB1P-G3 Δ53BP1 cells were treated with 0.01% MMS and 100 nM talazoparib (talaz.), recovered as depicted and subjected to subcellular fractionation and the chromatin fraction was analysed for PARP1 recruitment. (c) Whole cell extracts of HAP1 ΔBRCA1Δ53BP1 cells treated with 1 μM olaparib for 24 h or left untreated. One hour before collection, cells were treated with 0.01% MMS and 1 μM PARG inhibitor as indicated. Levels of Poly-ADP-ribosylation were analysed in whole cell extracts by immunoblotting and H3 served as a loading control. (d) Colony formation assay in HAP1 cells exposed to olaparib alone or supplemented with 1 μM PARGi PDDX-004. Mean ± SEM is depicted and a representative of two independent biological replicates is shown. (e) HAP1 single cell knockout clones of displayed genotype were assessed for NASP and PARP1 levels. (f) Clonogenic survival assay of the cells depicted in (e). Data are depicted as mean ± SEM and a representative of three biological replicates is depicted. (g) Clonogenic outgrowth of HAP1 cells of indicated genotypes in unchallenged conditions after seven days. P value was calculated by a two-tailed t test, *P < 0.05, **P < 0.01 and ***P < 0.001. (h) HAP1 cells of indicated genotype were monitored for their proliferation over seven days using Incucyte Live Cell imaging. Confluency per well is depicted as a readout. (i, j) Levels of γH2AX (i) and cleaved caspase-3 (j) were assessed in HAP1 cells using flow cytometry. Data are depicted as mean ± SD, P value was calculated by a two-tailed t test, *P < 0.05, **P < 0.01 and ***P < 0.001. (k) Subcellular fractionation of HAP1 cells of indicated genotypes upon 24 h of camptothecin treatment. Nuclei fraction is depicted and soluble fraction B is shown in Fig. 5h. All experiments were performed in triplicates, except (h) was performed in duplicates.
Extended Data Fig. 10 PARP1 and H3 interact in the soluble fraction.
(a) Cells were treated for 24 h with 2μg/mL doxycycline and/or 1 μM olaparib, fraction B was isolated from HAP1 ΔBRCA1Δ53BP1 cells overexpressing GFP-H3.1 and co-immunoprecipitation was performed against GFP. Representative blots of three biological replicates are shown. (b) Gels of the purified recombinant proteins stained with Coomassie. (c) Western blot of recombinant PARP1 protein and Coomassie staining of the corresponding gel. (d) In untreated conditions, NASP ensures the storage of pre- and postnucleosomal H3-H4 tetramers to promote ongoing DNA replication, while PARP1 possibly promotes reestablishment of the chromatin through its chaperoning function. PARP inhibitor (PARPi) treatment leads to a release of histones from chromatin by the INO80 complex. H3-H4 tetramers are subsequently stabilized and shielded from proteasomal degradation through the chaperone activity of NASP. Additionally, the reestablishment of chromatin boundaries is impaired due to inactivated PARP1. In the absence of NASP, chromatin-released and newly synthesized histones remain unprotected and are degraded. As a result, exposure of NASP knockout cells to PARPi leads to a lack of histone supply, caused by proteasomal degradation of chromatin released histones, culminating in failure of DNA replication, genome instability, and cell death. The schematic in panel d was created using BioRender (https://biorender.com).
Supplementary information
Supplementary Information (download PDF )
Supplementary Fig. 1 and Supplementary Tables 5–7.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Moser, S.C., Khalizieva, A., Roehsner, J. et al. NASP modulates histone turnover to drive PARP inhibitor resistance. Nature 645, 1071–1080 (2025). https://doi.org/10.1038/s41586-025-09414-z
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09414-z