Abstract
Marine picocyanobacteria are abundant photosynthetic organisms of global importance. They coexist in the ocean with cyanophages—viruses that infect cyanobacteria. Cyanophages carry many auxiliary metabolic genes acquired from their hosts that are thought to redirect host metabolism for the phage’s benefit1,2,3,4,5. One such gene is nblA, which is present in multiple cyanophage families2,6,7,8. Under nutrient deprivation cyanobacterial NblA is responsible for inducing proteolytic degradation of the phycobilisome9,10,11, the large cyanobacterial photosynthetic light-harvesting complex. This increases the pool of amino acids available for essential tasks11, serving as a survival mechanism12. Ectopic expression of different cyanophage nblA genes results in host pigment protein degradation6,8,13. However, the benefit of the virus-encoded NblA for cyanophages and the broader impact on the host are unclear. Here, using a recently developed genetic manipulation system for marine cyanophages14, we reveal that viral NblA significantly accelerates the cyanophage infection cycle, directs degradation of the host phycobilisome and other proteins, and reduces host photosynthetic light-harvesting efficiency. Metagenomic analysis revealed that cyanophages carrying nblA are widespread in the oceans and comprise 35% and 65% of oceanic T7-like cyanophages in surface and deep photic zones, respectively. Our results show a large benefit of NblA to the cyanophage, while it exerts a negative effect on the host photosynthetic apparatus and host photosynthesis. These findings suggest that cyanophage NblA has an adverse global impact on light harvesting by oceanic picocyanobacteria.
Similar content being viewed by others
Main
The light-harvesting antennae of most cyanobacteria are composed of phycobilisomes—gigantic membrane-associated soluble supercomplexes—located on the exterior side of the thylakoid membrane9,15. Cyanobacteria capture and funnel solar energy through their phycobilisomes towards the photosynthetic reaction centres16, where the primary energy-conversion reactions of photosynthesis occur. Typical phycobilisomes of marine Synechococcus are built of multiple subunits that absorb light at different wavelengths, including an allophycocyanin (APC) core and peripheral rods composed of disks of different phycobiliproteins, phycocyanin (PC) and phycoerythrins I and II (PEI, PEII), as well as linker proteins that connect rod subunits to each other and to the core and that attach the phycobilisome to the thylakoid membrane17,18. Each phycobiliprotein is composed of α and β subunits that form heterodimers10. These proteins bind to the pigment chromophores phycocyanobilin, phycoerythrobilin (PEB) and phycourobilin (PUB), which absorb light at different wavelengths19. Owing to their large size, phycobilisomes can comprise up to half of the soluble protein content of a cell15. Prochlorococcus is the most abundant photosynthetic organism on Earth20. The vast majority of Prochlorococcus lineages have a chlorophyll-based transmembranal light-harvesting complex rather than the typical phycobilisome21,22,23. However, Prochlorococcus does encode one or two subunits of phycoerythrin (PE)24,25, found at low levels in cells26. Notably, some low-light-adapted Prochlorococcus lineages from anoxic marine zones possess both chlorophyll-based and phycobilisome light-harvesting complexes20.
Viral NblA is beneficial for infection
Evidence collected over the past two decades suggests that cyanophages possess host-like auxiliary metabolic genes that are thought to influence key processes in host metabolism during infection and enhance phage replication1,2,3,4. These include genes that are thought to benefit the phage by maintaining essential cellular metabolic processes as well as those that impair host activities to redirect resources towards phage progeny production3,4. nblA encodes a proteolysis adaptor protein found in multiple cyanophage families2,6,7,8,13,27 and is particularly prevalent in T7-like cyanophages (family Autographiviridae). In nutrient-deprived cyanobacteria, the cyanobacterial NblA is responsible for inducing degradation of the phycobilisome complex, resulting in clear loss of pigmentation9,10,11. Here we use S-TIP37, a typical T7-like cyanophage isolated from the Red Sea that infects the open ocean Synechococcus sp. strain WH8109 (ref. 28), as our model system to study cyanophage-encoded nblA. We began by examining whether the cyanophage nblA gene is transcribed and translated during the infection process. We found high nblA transcript and protein levels from 2 h after infection onwards (Extended Data Fig. 1). This expression pattern was distinct to that of a host copy of the nblA gene, which displayed fairly consistent transcript and protein levels (Extended Data Fig. 1). The phage nblA expression pattern was similar to that of the phage DNA polymerase gene, and is therefore expressed with phage genes involved in DNA metabolism and replication and before expression of virion morphogenesis genes29,30.
Next, we assessed the influence of the nblA gene on cyanophage infection. For this we constructed nblA cyanophage deletion mutants (∆nblA) using the REEP approach14 (Methods). Infection of the host with the ∆nblA mutant phage revealed visibly less pigmentation loss compared with when the host cells were infected with the wild-type (WT) phage (Fig. 1a), indicating a clear phenotype for the ∆nblA mutant during infection.
a, Culture pigmentation phenotypes in uninfected Synechococcus sp. strain WH8109, and in WH8109 6 h after infection with the WT or the ∆nblA mutant S-TIP37 cyanophage. b, Cyanophage growth curves of the WT and ∆nblA mutant cyanophages. PFU, plaque-forming units. c, Changes in the abundances of different functional groups of phage proteins over time after infection with the WT (left) and ∆nblA (right) cyanophages. LFQ, label-free quantification. d, Cyanophage infection dynamics of the WT and ∆nblA mutant phages when infecting the WT host or the host ectopically expressing the cyanophage NblA. e, The apparent photochemical quantum yield of PSII (QYp) in cells infected with the WT or the ∆nblA cyanophage and in uninfected control cells after excitation of the phycobilisome at 495 nm (absorbance peak of PUB). Infection was performed at a multiplicity of infection (MOI) of 5. For b, linear regression formulas (Supplementary Data 2) are provided for the three near-linear segments of the cyanophage abundance (normalized to the maximum value) with time for WT (in blue) and ∆nblA (in red) cyanophages alongside the significance levels for the factor ‘infection type’ (type II Wald F-test); NS, not significant; ***false-discovery rate (FDR)-adjusted P < 0.001; exact FDR-adjusted P values for each interval: 0.60 (0–3 h), 2.0 × 10−18 (3–10 h), 8.7 × 10−5 (10–12 h). For c, genes were grouped into three clusters reflecting dominant gene functions. Linear models were fit for each cluster and a significant contribution of the factor ‘infection type’ (WT or mutant phage) was found only for the morphogenesis genes (analysis of variance (ANOVA) type II F-test, FDR-adjusted P = 7.1 × 10−8). Further statistical test details are provided in Supplementary Data 2. For b–e, data are mean ± s.d. of n = 3 independent biological experiments.
To investigate the importance of the cyanophage NblA to the infection process, we performed phage growth curve experiments. The ∆nblA mutant demonstrated a substantially longer latent period (the period of infection before the first release of phage progeny from cells) of 8–10 h compared with the 3 h latent period of the WT31 (Fig. 1b). In addition, synthesis of phage proteins increased with time during infection with the WT phage (Fig. 1c and Supplementary Data 1), whereas most phage protein production ceased 3 h after infection with the ∆nblA mutant, with significantly fewer virion morphogenesis proteins produced (Fig. 1c and Supplementary Data 2). These findings suggest a direct link between phage protein production and NblA expression and indicate that NblA is advantageous to the WT phage. As cyanobacterial NblAs are known to degrade phycobilisomes during nutrient stress9,10,11 and to release amino acids critical for cell survival11,12, we hypothesize that the phage NblA has a similar role during infection but the release of amino acids is used for building new virions.
The gene density in the region of nblA in the phage genome is high; therefore, to verify that the observed phenotype is a result of the lack of the nblA gene and is not due to potential side-effects caused by the deletion (Methods), we performed a rescue experiment. This was achieved by infecting Synechococcus sp. strain WH8109 cells ectopically expressing the cyanophage nblA gene under an inducible expression system32,33 with the ∆nblA mutant phage (Fig. 1d). Indeed, expression of the cyanophage NblA restored the timing of infection to that found for the WT cyanophage (Fig. 1d). The shorter latent period for the WT phage suggests that the nblA gene would improve phage fitness by allowing it to carry out more rounds of infection in a set time period. Our findings confirm that nblA has an important function in cyanophage infection and that it probably enhances phage fitness.
Cyanophage NblA would be expected to impact host photosynthesis if it directs phycobilisome degradation as phycobilisomes transfer energy to the photochemical reaction centres of the photosystems (PSI or PSII)16,34. We assessed the photosynthetic performance of PSII by measuring fluorescence at room temperature after exciting cultures with blue light (495 nm), which preferentially excites the most peripheral disks of the phycobilisome rods consisting of PEII. The apparent photochemical quantum yield of PSII (QYp) was then calculated. While uninfected cells maintained a steady PSII quantum yield, infection with both the WT and ∆nblA mutant phages resulted in a considerable decline (Fig. 1e). Notably, this decline reached low levels by 8 h after infection with the ∆nblA mutant, a delay of 4 h relative to infection with the WT cyanophage. This indicates that the cyanophage NblA negatively impacts the host’s PSII photochemical quantum yield by 50% over the period of infection. This is likely to have a substantial effect on the ability of the host cell to produce the energy and reducing equivalents necessary for many cellular processes. This diminished photosynthetic performance could be attributed to targeted phycobilisome degradation, accelerated cell lysis or a combination of both factors.
Viral NblA directs phycobilisome degradation
Cyanobacterial NblAs are proteolysis adapters that are known to be involved in phycobilisome degradation. To assess whether the cyanophage NblA has a similar effect on host phycobilisome integrity, we monitored whole-cell fluorescence emission spectra as a measure of the energy transfer pathway from phycobilisomes to PSII. Samples collected at different timepoints were excited at the wavelength for PUB, which binds mainly to the peripheral PEII disk of the phycobilisome, and the corresponding fluorescence emission spectra were measured. Infection with the WT cyanophage led to a rapid and substantial decrease in the chlorophyll a (Chl) peak with time (Fig. 2a and Extended Data Fig. 2a). This decline was first obvious at 3 h after infection (Fig. 2a), before the onset of cell lysis (Fig. 1b). By contrast, infection by the ∆nblA-knockout cyanophage led to a more gradual decrease in Chl fluorescence (Fig. 2a and Extended Data Fig. 2a). These findings indicate that energy transfer from PE to Chl was rapidly interrupted during infection with the WT phage, but considerably less so after infection with the ∆nblA-mutant phage.
a–c, Fluorescence emission spectra (a), absorbance spectra (b) and fractionated phycobilisomes on sucrose gradients (c) of uninfected Synechococcus sp. strain WH8109 cells (green), and WH8109 infected by the WT (blue) or ∆nblA mutant (red) S-TIP37 cyanophage at different timepoints after infection. In c, the lower-density (top) bands are disassembled phycobilisome subunits, and the higher-density (bottom) bands are intact phycobilisomes (Extended Data Fig. 3). d,e, Fluorescence (d) and absorbance spectra (e) of WH8109 cells 48 h after induced expression of the cyanophage NblA protein (blue) compared with the non-induced control (green). Fluorescence and absorbance spectra are representatives of five biological replicates. Mean and s.d. values are shown in Extended Data Fig. 2. Measurements were taken at room temperature, and peaks of Chl, PC, PE, PUB and PEB are indicated. Fluorescence measurements were normalized to the PE peak and absorbance measurements were normalized to the Chl peak. For a and d, the PE peak at 560 nm is made up of PEI and PEII, to which the PEB and PUB chromophores bind.
Whole-cell absorbance spectra and proteomic analysis provide a means to assess the levels of host phycobilisome subunits during infection. At 12 h after infection with the WT cyanophage, we observed a flattening of the peaks corresponding to both the PUB and PEB chromophores, which are attached to the PEI and PEII proteins. A less-prominent decrease in these peaks was observed during infection with the ∆nblA mutant (Fig. 2b), suggestive of NblA degradation of the PE proteins by the WT phage. Proteomic analysis during infection with the WT phage revealed significantly lower levels of the α subunits of PEII, PEI and PC rods (CpeA, MpeA and RpcA, respectively), as well as two rod linker proteins (MpeC and MpeE) relative to infection with the ∆nblA mutant at 4 h after infection (Supplementary Data 1). A 10–30% reduction was observed for some of the β subunits, but this was not statistically significant. By contrast, three linker proteins closer to the core35 that connect PC to APC and both PC and APC to the thylakoid membrane (ApcC, CpcG1, CpcG2) and one rod linker protein (CpeC) had higher protein levels during infection with the WT relative to the ∆nblA mutant (Supplementary Data 1). This may be indicative of attempts by the host to refurbish the phycobilisome core structure in response to the reduction in active phycobilisome rods. Combined, these findings provide further evidence for the role of cyanophage NblA in phycobilisome disassembly and degradation.
Next, we wanted to determine the fate of the antenna complex during infection. To achieve this, we isolated phycobilisomes from host cells 6 h after infection and compared them with those in uninfected cells. Fractionation of phycobilisomes on a sucrose gradient followed by fluorescence spectra analysis (Fig. 2c) enabled us to differentiate between assembled (bottom, high-density, pink bands) and disassembled complexes (top, low-density, orange bands) (Extended Data Fig. 3). While intact and disassembled phycobilisomes were apparent in uninfected host cells, only disassembled phycobilisomes were present in cells infected by the WT cyanophage (Fig. 2c). Cells infected by the ∆nblA mutant cyanophage were composed of both intact and degraded phycobilisomes (Fig. 2c), indicative of only partial phycobilisome disassembly.
The findings above are consistent with a role for NblA in disassembly and degradation of the phycobilisome during infection with the WT cyanophage. However, these findings could also be due, at least in part, to the beginning of cell lysis from 4 h onwards during infection with the WT phage. To more directly assess the impact of phage NblA on host phycobilisomes without the confounding myriad effects of phage infection, we ectopically expressed the viral NblA in host cells without infection. First, we measured whole-cell fluorescence spectra, which showed a decline in the energy transfer from PE to Chl, seen by a reduction in the Chl fluorescence peak after excitation at PUB relative to cells without induction of NblA expression (Fig. 2d). However, this decrease was smaller than that found during infection with the WT phage (Fig. 2a). Second, ectopic expression of the phage NblA resulted in some reduction in the PEB and PUB absorbance peaks (Fig. 2e). Third, we assessed whether the cyanophage NblA could restore phycobilisome disassembly during infection with the ∆nblA cyanophage mutant. Indeed, NblA expression led to a decrease in the Chl fluorescence peak when excited at PUB, declining to levels similar to those during infection with the WT cyanophage by 4 h after infection (Extended Data Fig. 4 (right)). Note that the partial decline in Chl fluorescence at 4 h after infection in the control without induction (Extended Data Fig. 4 (left)) is probably caused by leaky expression in this inducible system33. Taken together, our findings suggest a clear effect of the cyanophage NblA on energy transfer from PE to Chl, consistent with a role in phycobilisome disassembly and degradation during infection.
We next assessed the protein targets of the cyanophage NblA protein by following the production of new protein cleavages in host cells ectopically expressing the viral NblA. For this, we analysed proteins with newly produced N termini using mass spectrometry (MS)36 at 24 and 48 h after induction (at which time maximum protein induction occurs)33. Many new cleavages were observed (Supplementary Data 3). Among these, 201 cleavage events in 107 proteins were substantially more abundant (at least twofold) after viral NblA expression, including 81 cleavages in 59 proteins at 24 h and 103 cleavages in 59 proteins at 48 h after induction (Fig. 3 and Supplementary Data 3). These include several cleavages of phycobilisome subunits (Fig. 3a and Extended Data Fig. 5). At 24 h after induction, four phycobiliproteins were cleaved (Fig. 3b,c); the peripheral PEII disk α subunit (MpeA), the phycocyanin β subunit RpcB, the rod linker MpeE and the ApcE linker responsible for energy transfer from the APC core to the chlorophylls of the photosynthetic reaction centres in the thylakoid membranes9. At 48 h after induction, many of the new proteolytic cleavages were of phycobilisome subunits (Fig. 3a,b). These included the same three PEII, PEI and PC rod disk α subunits that displayed low levels during infection with the WT phage (MpeA, CpeA and RpcA) (see above) as well as the three PE rod disk β subunits (MpeB, CpeB and RpcB), one core protein (ApcB) and three linker proteins connecting between disks in the rods (CpeC, MpeD and MpeE) (Fig. 3b,c and Supplementary Data 3). These proteolytic cleavages spanned almost all phycobilisome rod proteins, and ApcB of the phycobilisome core (Fig. 3c). These results provide further evidence that the cyanophage NblA protein leads to the degradation and disassembly of the cyanobacterial phycobilisome antenna complex. They also suggest that the cyanophage NblA directs disassembly in a stepwise manner, initially to disconnect the host phycobilisome from the thylakoid membrane and to degrade peripheral PEII disks, followed by cleavage of the more internal PEI and PC disks. This is similar to the process known for cyanobacterial NblA during nutrient stress, in which phycobilisome degradation begins at the peripheral rod disks and moves inwards towards the phycobilisome core11. Notably, a previous study37 reported that infection by a cyanophage lacking a detectable nblA gene leads to the release of PE peptides, suggesting that cyanophages without nblA may have an alternative mechanism for degrading outer PE rods.
Proteomic analysis of newly cleaved proteins after cyanophage NblA expression in Synechococcus sp. strain WH8109 cells for 24 h (left) and 48 h (right), as compared to non-induced cells. n = 2 biologically independent experiments. a, The abundance ratios of neo-N-terminal peptides from phycobilisome proteins plotted against their MS signal intensities (which indicates the overall peptide abundance). The filled triangles indicate neo-N-terminal peptides with absolute log2-transformed ratios of ≥1 in both replicates. Upward-pointing and downward-pointing triangles represent peptides that are more and less abundant, respectively, after NblA expression relative to the control. b, The numbers of neo-N-terminal peptides and their corresponding proteins in different functional categories that showed consistent abundance changes after NblA expression (log2-ratio of ≥1 in both replicates; Extended Data Fig. 5). Proteins were categorized according to a modified classification of COG functional groups (Methods), with photosynthesis-related proteins other than phycobilisomes in ‘energy’, translation and DNA and RNA binding proteins in ‘information storage and processing’, cell division and porins in ‘cellular processes and signalling’ and transporters in ‘metabolism’. c, Phycobilisome cartoons showing proteolytic cleavages in phycobilisome subunits. The colours indicate phycobiliprotein cleavages in APC (turquoise), PC (blue), PEI (orange) and PEII (red), and linker proteins with cleavages between the discs, the core and the thylakoid membrane (various colours). Fully coloured discs indicate cleavages in both α and β subunits, semi-coloured discs or quarter-coloured cores represent cleavages only in α or β subunits d, Preferred amino acid cleavage motifs after ectopic expression of the cyanophage NblA. Sequence logos are for neo-N-terminal peptides with abundance ratios that were at least twofold higher in NblA-expressing cells. Residues passing two-sided t-tests are shown (uncorrected P ≤ 0.05), as implemented in IceLogo. Cleavage sites are marked with a dashed line.
Ectopic expression of the cyanophage NblA led to proteolytic processing of additional photosynthesis-related proteins beyond phycobilisome proteins (Fig. 3b, Extended Data Fig. 5 and Supplementary Data 3). These include those related to PSI (PsaC, PsaK), the PSII oxygen evolution complex (PsbO, PbsU) and carbon fixation (CcmK2, CsoS2). This suggests that the cyanophage NblA negatively impacts host photosynthesis through proteolytic degradation not only of phycobilisome components but also of photosystem proteins and proteins that are needed for carbon fixation. Moreover, a set of proteins with functions other than those related to photosynthesis was also cleaved as a result of expression of the cyanophage NblA (Fig. 3b and Extended Data Fig. 5). This includes proteins involved in translation, such as ribosomal proteins as well as RNA- or DNA-binding proteins at 24 h after induction (Fig. 3b and Supplementary Data 3). Translation initiation and elongation factors were also cleaved 48 h after induction (Fig. 3b and Supplementary Data 3). Other key cellular proteins cleaved include proteins involved in carbon metabolism, a subunit of the cellular RNA polymerase (RpoC2), the cell division protein FtsZ, as well as porin and transporter proteins (Fig. 3b and Supplementary Data 3). It is unclear whether host NblA proteins also direct the degradation of these proteins. Cleavage of these proteins would therefore disrupt multiple cellular processes during infection, in addition to photosynthesis.
In freshwater cyanobacteria, NblA proteins function by binding to phycobilisome subunits as well as to the ClpC chaperone responsible for recruiting the protease that induces proteolytic degradation of the phycobilisome11,38. As proteolytic enzymes often have preferred cleavage sites, we investigated cleavage sites after ectopic expression of the cyanophage NblA in the Synechococcus host (Fig. 3d). At 24 h after induction, we detected cleavages primarily between medium-sized, polar or partially polar residues (Met, Gln, Thr) and small, polar or nucleophilic residues (Ser, Thr, His), while, at 48 h after induction, an additional cleavage site was detected between Lys and Arg residues (Fig. 3d), providing an explanation for the additional cleaved proteins at 48 h after induction. This could be a result of NblA recruiting different proteases or the different substrates being cleaved by the same proteolytic system (presumably the Clp protease complex) with broad cleavage specificity.
Viral nblA is common in the oceans
T7-like cyanophages are very abundant in the oceans31,39. However, the prevalence of the nblA gene in this group is unclear. Thus, to understand the environmental importance of our findings, we first assessed the distribution of nblA genes in assembled genomes of both isolated and environmental T7-like cyanophages. Our search revealed that 46.4% of the complete non-redundant T7-like cyanophage genomes encode nblA genes (Extended Data Fig. 6). nblA-encoding cyanophages have been isolated on picocyanobacterial hosts from the three main lineages: marine Synechococcus subcluster 5.1 (Parasynechococcus40 in the broad sense), Synechococcus subcluster 5.2 (Cyanobium and related genera) and Prochlorococcus (the Prochlorococcus collective) (Fig. 4), even though the vast majority of the latter do not have phycobilisomes as their light-harvesting antennae21,23. We also found that the gene is present in a rare prophage residing in a Synechococcus subcluster 5.1 genome (AG-670-B23)41. Thus, cyanophage nblA is quite common in the genomes of T7-like cyanophages, infecting both Synechococcus and Prochlorococcus.
The maximum-likelihood phylogenetic tree is based on concatenation of trimmed amino acid alignments of nine core genes (Methods), is outgroup-rooted and was pruned to include the chosen genomes (the complete phylogeny of the cyanophages and the associated metadata are provided in Extended Data Fig. 5 and Supplementary Data 4 and 5). The genomes were subdivided into previously delineated clades A, B and C, as well as the newly defined minor clades: R (P-RSP2-like phages), S (S-SRP02-like phages) and T (no isolated representatives). For each representative, the genomic region containing the exonuclease gene and downstream of it is shown. Frequently appearing orthologous genes are indicated with colour. The colour of the phage names indicates the host lineage that they were isolated on, where known. Environmental genomes are indicated by asterisks. The solid circles mark branches with ultrafast bootstrap support values of ≥95.
A close examination showed two types of nblA genes with differential distribution among the phylogenetic clades of T7-like cyanophages (Fig. 4 and Extended Data Figs. 6 and 7). nblA genes of one type are dominant and are present in clade B cyanophages (to which S-TIP37 belongs) and, to a lesser extent, in clade A. These nblA genes are consistently located downstream of the exonuclease gene and frequently have up to a 110-nucleotide overlap with it. nblA genes of another type are restricted to some P-RSP2-like cyanophages (designated here as clade R) and are located downstream of an HNH endonuclease gene. Neither nblA gene type was found in the genomes of cyanophages from the newly discovered clade C (Fig. 4).
To assess the relationships between NblA proteins from the T7-like cyanophages and those of their hosts, we performed a targeted search for NblA homologues among marine picocyanobacteria. Notably, we found nblA genes not only in marine Synechococcus, but also in many Prochlorococcus low-light-adapted ecotypes (Extended Data Figs. 7 and 8). While marine Synechococcus typically possesses several nblA genes, with, for example, Synechococcus sp. strain WH8109 having five intact genes and a pseudogene, Prochlorococcus strains have a single nblA gene. These findings indicate that nblA genes are also found in cyanobacteria that do not possess phycobilisomes as well as in the cyanophages that infect them.
NblAs from the marine picocyanobacteria are substantially divergent from the well-characterized freshwater cyanobacterial NblAs (as exemplified by the chromosomally encoded NblA from Nostoc sp. strain PCC 7120) with respect to sequence, yet their predicted structures preserve the classical dimeric NblA architecture (Extended Data Fig. 8). However, NblA from the T7-like cyanophages demonstrate higher overall sequence similarity to the well-characterized cyanobacterial NblAs rather than to those of their hosts (Extended Data Fig. 8). The T7-like cyanophage NblAs cluster together in a phylogenetic network, suggesting that they represent a monophyletic group (Extended Data Fig. 7). Together these findings suggest that the acquisition of nblA by T7-like cyanophages occurred in the distant evolutionary past. This is in contrast to nblA genes from freshwater cyanophages infecting Microcystis and Planktothrix that have high similarity to nblA genes from the corresponding cyanobacterial groups6,8,27.
We next assessed the relative abundance of T7-like cyanophages carrying the nblA gene in the oceans. We used the consistent genomic association of the dominant nblA type with the exonuclease gene, a core gene found in all T7-like cyanophages sequenced so far42,43 to examine the read data from the Global Ocean Viromes (GOV) metagenomic dataset. This enabled us to determine both the presence or absence of the nblA gene and the relative abundance of the different clades from the exonuclease gene. Our quantification strategy yielded general distribution patterns of T7-like cyanophages similar to previously reported studies. Cyanophages belonging to clades B and C dominated the global oceans31,42, while clade A and R phages were relatively rare (Fig. 5). The vast majority of cyanophages from clade B possess the nblA gene (Fig. 5), with around 72% of those at the surface and 89% of those deeper in the photic zone at the deep chlorophyll maximum (DCM) carrying the gene. The proportion of nblA-encoding cyanophages among the less abundant clade A group was much lower, averaging about 24% at the surface and 10% at the DCM. In accordance with the distribution of the nblA genes in complete genomes, none of the clade C and R cyanophages had nblA genes downstream of the exonuclease gene. As the nblA gene is located in a different genomic position in clade R, our quantification method could not be applied to them. Overall around 35% and 65% of the T7-like cyanophages (from clades A, B and C) encoded nblA in the surface and at the DCM, respectively. These findings indicate that nblA is widespread among the abundant clade B cyanophages both at the surface and deeper in the photic zone.
a,b, The relative abundance of T7-like cyanophages in surface waters (a) and at the DCM (b). Quantifications are based on fragments per kilobase per million total reads (FPKM) abundances for exonuclease genes with and without downstream nblA genes and assigned to one of the five clades of T7-like cyanophages. Raw data and exonuclease-containing contigs for read recruitment were retrieved from the Global Ocean Viromes 2.0 dataset (GOV 2.0). The insets show the relative abundances of the different clades of cyanophages across the data (based on the FPKM values added together) and serve as the legend for the colours on the maps. Map data are from Natural Earth.
Among the T7-like cyanophage isolates that encode nblA, half infect marine Synechococcus strains while the rest infect phycobilisome-less Prochlorococcus strains (Fig. 4). Both groups of cyanobacteria are prevalent in surface waters, whereas Prochlorococcus is considerably more abundant than marine Synechococcus at the DCM21,44. Thus, we anticipate that many of the clade B phages at the DCM infect Prochlorococcus. As the vast majority of Prochlorococcus lineages have a different type of light-harvesting antenna to the phycobilisomes of most cyanobacteria21,22,23, we expect that NblA of the phages infecting Prochlorococcus direct the degradation of other proteins. This possibility is supported by the finding that multiple low-light-adapted Prochlorococcus ecotypes encode an nblA gene (Extended Data Fig. 7), even though they do not have phycobilisomes. Furthermore, our proteomics data indicate that NblA-directed degradation is not restricted to phycobilisome proteins, even for a clade B phage that infects Synechococcus (Fig. 3a,b and Extended Data Fig. 5). Thus, other photosynthesis-related proteins and proteins involved in other cellular processes are probably targeted by many nblA-encoding cyanophages deep in the photic zone of the oceans.
Cyanobacteria acclimatize to low light to maximize their ability to harvest light. This is achieved through increasing the number and size of their antenna complexes45,46,47 as well as the number of their photosystems48. As such, more cyanobacterial resources are bound up in the photosynthetic apparatus at depth. Thus, we propose that the ability of cyanophages to degrade the photosynthetic apparatus deep in the photic zone would provide a considerable advantage through the release of amino acids for use in cyanophage progeny production and may explain why relatively more cyanophages at depth carry the nblA gene.
The general distribution of the dominant nblA type in T7-like cyanophages parallels that for the psbA gene (encoding the PSII reaction centre protein D1). Both genes are present in the majority of clade B cyanophages, sporadically in clade A and are absent from clade C. However, they do not always co-occur28 (Fig. 4 and Extended Data Fig. 6). nblA is more common in T7-like cyanophages than psbA, with 36% of the complete genomes (including environmentally assembled genomes) across all clades encoding psbA compared with 46% for nblA (see above). As psbA is considered to be one of the most common auxiliary metabolic genes in cyanophage genomes2,3,28,42,43, our results suggest that the cyanophage-encoded nblA is very common in oceanic environments as well.
An interesting aspect of T7-like cyanophages that carry both psbA and nblA is that psbA is thought to enable continued photosynthetic energy production during infection3,49,50 while nblA degrades the complex that harvests the light funnelled to the photosystems (this study). This raises the question of how these potentially conflicting functions for two different auxiliary metabolic genes in the same cyanophage can be reconciled. We propose that the two genes are most beneficial to the phage under different conditions. The psbA gene has been hypothesized to be more important under conditions of high light51,52 in which more photodamage to the D1 protein is expected. By contrast, nblA is probably more important to the phage under low-light conditions in which more of the cell’s resources are invested in building the photosynthetic apparatus (see above). Moreover, under high-light conditions, degradation of the phycobilisome by cyanophage NblA could serve to reduce photodamage to D17, such that photosynthesis can continue at some level despite less light being harvested by the phycobilisome. Thus, a balance between the activities of these two proteins is probably needed under high light.
The vast abundance of clade B T7-like cyanophages in the oceans encoding nblA raises the possibility that they have a global impact on cyanobacterial photosynthesis. This group of cyanophages is considerably more abundant deep in the photic zone than at the surface31,53, where a very large percentage of them encode nblA (Fig. 5). While it is difficult to quantify the global impact of a single gene, we have performed some rough calculations to estimate the potential effect of viral nblA on picocyanobacterial light harvesting. We base our calculation on the findings that the cyanophage gene caused a 50% reduction in PSII photosynthetic performance (Fig. 1e), that 1–15% of marine cyanobacterial cells are infected by T7-like cyanophages at a given time39,54,55 and that 35–65% of marine T7-like cyanophages encode nblA in different layers of the photic zone (Fig. 5 and Methods). We estimate that 0.2–1.1% and 1.6–4.9% of cyanobacterial photosynthesis in the surface and DCM layers, respectively, could be impacted by nblA. These estimations suggest that, collectively, T7-like cyanophages carrying the nblA gene have a global impact, reducing picocyanobacterial photosynthetic light harvesting by 0.2–5% in the upper oceans.
Concluding remarks
Auxiliary metabolic genes in viruses56,57 are a widespread phenomenon, especially in marine cyanophages1,2,3,4. Previous studies have investigated the putative function of several such genes using in vitro biochemical approaches58,59 or ectopic expression6,13,59,60. Here we combined a recently developed cyanophage genetic engineering system14, together with inducible ectopic expression33 and state-of-the-art N-terminal proteomics36 to directly examine the role of nblA, a gene encoding a small proteolysis adaptor, in cyanophage infection and its impact on the host photosynthetic apparatus. While NblA in cyanobacteria serves as a stress-response mechanism under nutrient deprivation and mediate the controlled degradation of the phycobilisome photosynthetic antenna9,10,11, we propose that its cyanophage counterpart is important not for cell survival, but for virion synthesis. Our results further reveal that the viral NblA directs degradation not only of host phycobilisomes, but also of a suite of other proteins, including important core photosynthesis and house-keeping proteins. Thus, this small auxiliary metabolic gene is likely to have a large impact on both the host’s light-harvesting efficiency, energy production and other essential cellular processes.
The implications of the nblA gene for the cyanophage are considerable, with an approximately threefold shorter infection cycle, suggestive of considerably improved fitness. These findings, together with genomic and metagenomic results of high abundance and widespread distribution of cyanophages carrying the nblA gene in the oceans (this study) and in freshwater ecosystems7,8,13 suggest that these cyanophages have a negative global effect of up to 5% on the amount of light harvested by oceanic cyanobacteria.
Methods
Cyanobacterial growth
Synechococcus sp. strain WH8109 was grown in artificial seawater (ASW) medium61, with modifications as described previously62. Cultures were grown at 21 °C under a light intensity of 20 µmol photons per m2 per s, under a 14 h–10 h light–dark cycle. Pour-plates were obtained using ASW medium with low-melting-point agarose at a final concentration of 0.28% with an additional 1 mM of sodium sulfite. A heterotrophic helper strain, Alteromonas sp. EZ55, was added to the pour-plate mixture for isolating Synechococcus colonies63.
qPCR with reverse transcription
For RNA extraction, a 1 ml culture sample was collected by centrifugation at 4 °C, 15,000g for 15 min, and the pellets were flash-frozen in liquid nitrogen. Cell pellets were thawed and incubated with lysozyme L6876-5G (Sigma-Aldrich) at a final concentration of 30 mM and 200 U of RNase inhibitor (Murine BioLabs) for 60 min at 37 °C. An equal volume of lysis buffer was added and cell debris was centrifuged at 4 °C, 16,000g for 1 min. Nucleic acids in the supernatant were precipitated with an equal volume of 95% ethanol and centrifuged as described above. RNA wash buffer (500 µl) from Monarch Total RNA Miniprep Kit (NEB, T2010) was added, and the sample was centrifuged again for 30 s. The supernatant was discarded and the step was repeated. DNase I reaction buffer (5 µl) and 4 U of DNase I from the TURBO DNA-free kit were added to the 45-µl sample and incubated for 10 min at 4 °C. The samples were incubated with 0.5 µl of 0.5 M of EDTA, pH 8.0, for 10 min at 75 °C.
Total RNA was reverse transcribed into cDNA using the LunaScript RT SuperMix Kit (New England Biolabs, E3010). The reaction mixture was prepared in a total volume of 20 µl, 4 µl of LunaScript RT SuperMix containing random hexamers and 16 µl of nuclease-free water, with the RNA sample added to reach the final volume. As a control, a similar mixture was prepared, but without addition of the reverse transcriptase enzyme. The reaction was incubated at 25 °C for 2 min, 55 °C for 10 min and 95 °C for 1 min.
Quantitative PCR (qPCR) reactions were prepared using LightCycler 480 SYBR Green I Master mix from Roche, combined with 0.2 µM of each primer and the nblA/rnpB DNA template. The reactions were run on the LightCycler 480 Real-Time PCR System. Cycle threshold fluorescence values for each reaction were determined using LightCycler 480 software. To quantify DNA copy numbers, a standard curve was generated by running tenfold serial dilutions of the template and correlating cycle threshold values to known DNA concentrations (Supplementary Data 6).
The genes of which the expression levels were analysed using qPCR were as follows: host nblA2 (Syncc8109_1607, see explanation below; primers: 5′-GCGATCAAGCGGTCAATCAAC-3′ and 5′-CTCTCTGCCGCACGTAGAGG-3′), host rnpB (Syncc8109_0157; primers: 5′-CATCGGCGGTGTGTTTCT-3′ and 5′-CAGGCTTGCTGGGT-3′), S-TIP37 nblA (primers: 5′-TTCCCGAGGCAGACAAGAG-3′ and 5′-TAATGGGATGGTGACTCGGC-3′), S-TIP37 DNA polymerase (STIP37_17B; primers: 5′-TGAGCTACTACGCAACAGGC-3′ and 5′-AGCGCGATCATTCAGGGAAG-3′). The Synechococcus sp. strain WH8109 nblA gene chosen for qPCR is the one that we reported previously6, although a more refined remote homology search with hhsearch64 using a custom NblA profile reveals that the genome carries four additional nblA-like genes and an nblA pseudogene. To clarify which homologue has the highest structural similarity to previously characterized proteins and is therefore most likely to have a function similar to that of freshwater cyanobacterial NblAs, we obtained the structures for the corresponding monomers with ColabFold65 and searched them against the Protein Data Bank (PDB) using Foldseek66. The best match was indeed obtained for NblA2 (Syncc8109_1607), with the highest similarity to the NblA protein from Nostoc sp. strain PCC 7120 (ref. 10; PDB: 1OJH chain E; probability, 0.94; TM-score, 0.6621).
Cyanophage-infection experiments
Before infection, Synechococcus sp. strain WH8109 was grown in liquid medium to mid-log growth of around 1 × 108 cells per ml. Infection experiments were initiated by adding the S-TIP37 cyanophage strains at an MOI of 5. Infection dynamics were determined by collecting samples at hourly time intervals during the initial 6 h, followed by sampling every 2 h thereafter. The samples were filtered through a 0.22 µm syringe filter and the filtrate containing free cyanophages was plated to determine the number of infective cyanophages using the plaque assay67 in semi-solid pour-plates (see above). Statistical analysis of the infection course was performed using segmented linear regression by selecting near-linear ranges in the infection curves and obtaining linear fits for the relative cyanophage abundance (obtained by dividing the abundance in the extracellular medium by its maximum across all experiments) with the factors time after inoculation, cyanophage type (WT or mutant) and their interaction, and replicate as random effects using lme4 (v.1.1-31)68. The significance of the model terms was tested using the Anova function from the car package (v.3.1-1)69. The distributions of the residuals were checked using QQ plots.
Spectral measurements
Absorbance spectra were measured at room temperature using a double-beam Shimadzu spectrophotometer equipped with deuterium and halogen lamps. Spectral data were obtained at a constant bandpass with a resolution of 1 nm. Absorbance spectra were normalized to chlorophyll (680 nm). Fluorescence emission spectra were collected using the Jobin-Yvon Horiba spectrofluorometer. The samples were placed into 1 ml micro quartz cuvettes and excited at 495 nm, and fluorescence emission was measured from 500 nm to 750 nm and normalized to the fluorescence intensity at the emission maximum of PEI (562–564 nm).
The apparent photochemical quantum yield of PSII (QYp), or efficiency of light use by PSII, was determined by measuring fluorescence emission after dark adaptation for 15 min, with and without the addition of (3-(3,4-dichlorophenyl)-1,1-dimethylurea (DCMU) to a final concentration of 10 µM. DCMU blocks electron transfer between the primary quinone electron acceptor (QA) and the secondary quinone electron acceptor (QB) on the acceptor side of PSII, causing the reaction centres to remain in a closed state70 The samples were excited at 495 nm and fluorescence emission was measured from 500 nm to 750 nm. The apparent PSII quantum yield was calculated as described previously71, from the onset of infection up to 10 h after infection, at which point all values fell below zero. QYP was calculated as QYP = (QYF+DCMU − QYF–DCMU)/QYF+DCMU. Absorbance-corrected fluorescence quantum yield (QYF) was calculated as QYF = F/f495, where F is the integrated area under the fluorescence emission spectrum and f495 is the fraction of incident light actually absorbed at 495 nm (\({f}_{495}=1-1{0}^{{A}_{495}};{A}_{495}\) is the measured absorbance at 495 nm).
Quantification of the absorption of the different pigments was performed by subtracting the background with the package baseline (v.1.3-4)72 using the modpolyfit method, identifying the peaks of PUB and PEB and chlorophyll and scaling the intensities to the intensity of the chlorophyll peak. The differences between the uninfected cells and cells infected with the WT and mutant cyanophages 12 h after infection were analysed with linear models and Tukey’s post hoc tests. Fluorescence emission spectra were scaled to the maximum intensity (intensity of the PE peak) to obtain the relative emission intensity of the chlorophyll peak. As the fluorescence measurements were performed repeatedly for the same batches, a mixed model was built to analyse the relative fluorescence of chlorophyll as a function of infection (uninfected cells and two types of cyanophages), time after infection and their interaction, taking into account the random effect of batch with lmer4. The distributions of the residuals were checked with QQ plots. The Anova function from the car package was used to obtain an analysis of deviance tables and P values for the model terms; Tukey’s test was performed as a post hoc test in both cases.
Phycobilisome extraction
Synechococcus sp. strain WH8109 cells were collected by centrifugation at 6,000g for 10 min at 4 °C, 6 h after infection, and kept at −80 °C. The pellets were thawed at room temperature, resuspended with 7.5 M potassium phosphate, pH 7.5, and homogenized. Cells were disrupted with a microfluidizer (Microfluidics, HC-2000) at 80 p.s.i. The supernatant was collected after centrifugation at 20,000g for 30 min at 4 °C and loaded onto a 0.25–1.25 M linear sucrose gradient in 7.5 M potassium phosphate buffer, pH 7.5. The samples were centrifuged in an ultracentrifuge at 170,000g for 18 h at 4 °C. The fractions were collected with a syringe and analysed by fluorometry at 77 K and spectrophotometry as described above.
Construction of S-TIP37 nblA-deletion mutant
The open reading frame (ORF) encoding nblA is 237 bp long (NCBI: MH540083.1, located at nucleotides 12467–12706) and was previously identified by us in the genome of S-TIP37 based on a match to our custom NblA protein profile6. Profile–profile searches with hhsearch using the HHpred server73 for this protein yield high-scoring matches to cyanobacterial NblA sequences and Pfam NblA profile (PF04485). Analogously, folding the protein with ColabFold65 produces a typical dimeric structure of NblA with the two monomers composed of two helices each and unstructured terminal regions (Extended Data Fig. 7).
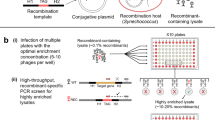
The nblA mutant was produced as described previously14. As phages gain random mutations readily, we constructed two independent S-TIP37 ∆nblA mutant cyanophages to ensure that the observed phenotype was directly related to the lack of the nblA gene. In brief, the nblA ORF was used to prepare a construct containing a recombination template for viral homologous recombination. We replaced 137 bp of the nblA gene from position 12,572 to 12,707 with a tag of 23 bp. As the nblA gene has an overlap of 94 nucleotides with the upstream exonuclease ORF exo, the region chosen for the deletion covered the part of nblA not overlapping this gene. However, this region covered the part of nblA overlapping a downstream ORF coding for a protein homologous to gp1.7, a nucleotide kinase, of the T7 Escherichia coli phage, resulting in deletion of 41 nucleotides at the 5′ end of the gene (this short gene is underannotated in T7-like cyanophage genomes and was not originally annotated in S-TIP37). The construct was cloned into the replicative pRL-proCAT14 plasmid and conjugated into Synechococcus sp. strain WH8109 (refs. 14,74). The strain expressing the recombination template was infected with WT S-TIP37. The lysate was filtered through a 0.22-µm syringe filter to remove cell debris. The presence of recombinant phages was verified by PCR with one primer for the inserted TAG sequence and one in the phage genome: 5′-TGGTGATCAGACCGATGGG-3′ (forward) and 5′-GAGCTCATAGCAAAGAAGACGTC-3′ (reverse).
Enrichment and PCR screening for recombinant cyanophages was performed in 96-well plates containing the Synechococcus host14. Wells containing recombinant cyanophages were filtered and plated on semi-solid medium. Single phages were plaque-purified twice and the whole genome of each purified mutant phage clone was sequenced alongside the corresponding WT. Illumina MiSeq trimmed read pairs were received and the genomes were assembled de novo with Spades (v.3.14.1)75. Both ∆nblA mutants had additional mutations in their genomes compared with their corresponding WT genomes (Supplementary Data 7 for the genome sequences). Non-synonymous mutations in the first mutant relative to the first WT had a T > C substitution at position 11142 (position according to reference genome MH540083.1) leading to a V208A mutation in the gene for the DNA polymerase nucleotidyl transferase subunit (NCBI: AXF42115.1) and an A > C substitution at position 37714 in gene 42, encoding a putative 2OG-Fe(ii) oxygenase (NCBI: AXF42102.1), leading to a K34T change. The second ∆nblA mutant had two non-synonymous differences compared with its WT in gene 40, encoding a putative tail fibre protein (NCBI: AXF42100.1): an A > C substitution at position 36856 in the WT genome leading to a D454A change (relative to the reference genome) and an A > G substitution at 36888 in the ∆nblA genome, leading to a N465D change. As both ∆nblA mutants had additional mutations compared with the WT, infection and spectral experiments were performed with both mutants and gave the same results. Results with the second mutant are shown. Rescue and proteomics experiments were performed with the second mutant. The mutant does not express the putative nucleotide kinase. The relevance of this gene to the phenotype was addressed in the rescue experiments through ectopic expression of nblA but not the putative kinase (see below section).
Ectopic expression of the cyanophage nblA
A theophylline translational induction system was used for expression of the cyanophage NblA protein in Synechococcus sp. strain WH8109 (refs. 32,33). A riboswitch sequence that theophylline binds to and the full-length S-TIP37 nblA gene were cloned into the pRL-proCAT replicative plasmid downstream of the rnpB promoter as described previously33 and were conjugated into Synechococcus sp. strain WH8109(refs. 14,74). The construct consists of the regions from nucleotides 12467 to 12703 relative to the reference S-TIP37 genome (GenBank: MH540083.1) and six additional amino acids that were added at the NblA C-terminus (Gly, Ser, Tyr, Ser, Val, Thr) during the cloning process. As nblA overlaps two neighbouring genes, the insert contains the last 95 nucleotides of the 3′ region of the putative exonuclease gene and the first 37 nucleotides of the 5′ region of the putative nucleotide kinase gene. The latter corresponds to 12 out of 94 amino acids of the protein, and proteomic analysis verified that this protein was not present in the induction experiments. As such, rescue experiments reinstated expression of NblA but not the putative nucleotide kinase.
Expression experiments were carried out with the exponentially growing WH8109 conjugant. Theophylline (0.3 mM), dissolved in ASW, was added to induce translation of the NblA protein. Expression of proteins under this theophylline-inducible system is leaky, with differences between induced and non-induced expression detected from 7.5 h after addition of theophylline that increased until 48 h after induction33. For rescue experiments, theophylline was added 8 h before infection with cyanophages. For assessing the effect of NblA on Synechococcus sp. strain WH8109 protein cleavage, samples were collected 24 and 48 h after induction. Induced samples were compared to non-induced samples. Each expression experiment was accompanied by a control construct carrying the pRL-proCAT plasmid containing the riboswitch but lacking a downstream gene.
Proteomic analyses
Protein extraction
Cyanobacteria were collected and washed three times using 50 mM HEPES, pH 7.5. After the final centrifugation, the cyanobacterial pellet was resuspended with 8 M guanidine hydrochloride (GuHCl), 100 mM HEPES, pH 7.5, and was heated at 95 °C for 10 min. The samples were then sonicated using the VialTweeter (Hielscher) system at maximum amplitude, 70% cycle time for 5 min, then heated for another 5 min at 95 °C to ensure maximal extraction and denaturation. Cellular debris was pelleted at 18,000g for 10 min, and the clear supernatant containing proteins was transferred to a new tube. Protein concentrations of each sample were measured with the BCA assay, before splitting each sample; 10 µg was taken for total proteome analysis and 50 µg was taken for N-terminome analysis.
Total proteomics sample preparation
Proteins were reduced with 5 mM dl-dithiothreitol (DTT) at 65 °C for 30 min then cooled to room temperature and alkylated with 12.5 mM chloroacetamide (CAA) for 30 min in the dark. The guanidinium concentration was diluted to 1 M using 100 mM HEPES, pH 8, and trypsin (Promega, V115A) was added at 1:100 (w/w) ratio overnight at 37 °C. Next, trypsin was quenched with 1% formic acid and the samples were desalted using C18 stage tips (Empore 66883-U) before subjecting them to MS analysis.
N-terminal proteomics sample preparation
N-terminal enrichment was performed using an optimized hydrophobic tagging-assisted N-termini enrichment (HYTANE) procedure as described previously36. Proteins were reduced and alkylated with DTT and CAA as described above. Different isotopes (heavy and light) of formaldehyde were then used to label primary amines in each pair of comparative samples. The labelling was done with 40 mM formaldehyde and 20 mM of sodium cyanoborohydride at 37 °C overnight. Leftover formaldehyde was quenched using 100 mM glycine for 1 h at 37 °C before mixing the heavy and light labelled samples together. The samples were diluted to reduce GuHCl concentration to 1 M using 100 mM HEPES, pH 8, before adding trypsin at 1:100 (w/w) and incubating overnight at 37 °C. The trypsin was then quenched with 1% (final) formic acid, and the samples were desalted using OASIS-HLB columns (Waters). The elution was done using 60% acetonitrile, 0.1% formic acid and the samples were dried using a speed-vac. The peptides were resuspended using 100 µl of 100 mM HEPES pH 7 and had undecanal solution (20 mg ml−1 prepared in ethanol) added to them at a w/w ratio of 1:50 (protein:undecanal), followed by the addition of sodium cyanoborohydride at a final concentration of 20 mM. Undecanal tagging was done for 2 h at 50 °C, renewing the sodium cyanoborohydride after 1 h. The samples were then centrifuged at 16,000g for 5 min and the supernatant was transferred to a new tube before drying in a Speed-Vac. The dried samples were resuspended in 500 µl of 2% acetonitrile, 0.1% formic acid and subjected again to the OASIS-HLB column. Elution was done using 60% acetonitrile, 0.1% formic acid, and the samples were dried in a Speed-Vac before being resuspended again and analysed using MS.
LC–MS analysis
Desalted samples were subjected to liquid chromatography (LC)–MS analysis using an Orbitrap Exploris 480 coupled with an EvoSep One HPLC. The samples were introduced onto the EvoTip, which was then washed twice with 20 μl of 0.1% formic acid. The washed peptides were kept wet by applying 150 μl of 0.1% formic acid atop the EvoTip until MS analysis. The samples loaded onto the EvoTips underwent chromatography separation on a 15 cm × 150 μm analytical column, filled with 1.9 μm C18 beads (EV1106). Peptides were separated over an 88-min gradient according to the manufacturer’s standard method. Full MS scans acquired in positive-ion mode, scanning from 300 to 1,800 m/z, were recorded at a resolution of 120,000 by a data-dependent mode, selecting the top 20 ions with high-energy collisional dissociation (HCD) fragmentation ion at a resolution of 17,500 and with dynamic exclusion enabled.
MS data analysis
For total proteomics, data analysis was conducted using FragPipe v.22.0 (https://fragpipe.nesvilab.org/) using DDA+ mode76 with the default label-free quantification–match between runs (LFQ-MBR) settings. The search enzyme was set to trypsin, allowing up to two missed cleavages. Variable modifications included methionine oxidation and protein N-terminal acetylation, while carbamidomethylation of cysteine was set as a fixed modification. For N-terminomics data, raw files were first converted to MzML format and then analysed using the Trans-Proteomic Pipeline (v.6.3)77 with the Comet search engine v.2023_01 rev2. The search enzyme was set to Semi-ArgC, allowing for up to two missed cleavages. Variable modifications included methionine oxidation and the mass difference between heavy and light dimethylation at lysine residues or the N termini of peptides. Fixed modifications included carbamidomethylation of cysteine and light dimethylation at lysine residues or peptide N termini.
Relative peptide quantifications were performed using XPRESS, with parameters set to mass tolerance of 20 ppm, a minimum of three chromatogram points for quantitation and the number of isotopic peaks to sum set to zero. Post-analysis table creation, cleavage motif extraction and ratio normalization were performed using an in-house script78. The search database included all Synechococcus sp. strain WH8109 proteins in UniProt (taxon ID: 166314, both reviewed and unreviewed) in addition to the vector added genes and standard contaminant proteins. For viral infection samples, this database was expanded to also include all Synechococcus T7-like phage S-TIP37 (taxon ID: 1332145) protein sequences from UniProt and the NblA sequence. In the analysis of the N-terminomics data, we considered three classes of blocked N-terminal peptides: ORF N termini, neo-N termini generated by internal proteolysis (SemiN1), and proteolysis-adjacent peptides truncated before the first arginine (SemiN2). Only NblA-induced cleavage events that were identified in both biological replicates and showed greater than twofold abundance increases in NblA-expressing cells compared with the control in each replicate were considered for downstream analysis. To bin host proteins into functional groups, we used the COG functional category assignments for Synechococcus sp. strain WH8109 proteins in the eggNOG database (v.5.0)79. The proteins were assigned to one of the broad functions based on a modified classification of COG functional groups as follows: phycobilisome components—phycobilisome; other photosynthesis-related genes and other proteins assigned to COG functional category C (energy production and conversion)—energy; COG functional categories J, A, K, L and B—information storage and processing; COG functional categories D, Y, V, T, M, N, Z, W, U and O—cellular processes and signalling; COG functional categories G, E, F, H, I, P and Q—metabolism; COG functional categories R, S and proteins assigned to multiple categories—other.
Phage proteins were classified into functional categories based on the functions of the majority of genes in three genomic and expression clusters29,43. Statistical analysis of the dynamics of abundance of the different functional groups of phage proteins in the total proteomics data as a function of time after infection onset was performed as follows. Genes were grouped into three clusters, with the cluster names reflecting functions of the majority of the corresponding genes: takeover (genes STIP37_1-12 in GenBank MH540083.1), DNA replication and metabolism (genes STIP37_13-23 (including nblA) and STIP37_55) and morphogenesis (genes STIP37_24-54). For each gene cluster, cumulative MS label-free quantification intensities were used as a response variable in a linear model with time after infection, infection type (by WT or ∆nblA mutant), and their interaction as predictor terms. The distributions of the residuals were checked with QQ plots. Detailed results of the statistical analysis are provided in Supplementary Data 2.
Bioinformatic analyses
Analysis of T7-like cyanophage genomes
Genomes of cultured T7-like cyanophages and outgroup phages (coliphage T7 and pelagiphages HTVC011P and HTVC019P) were collected from GenBank. Complete genomes and genomic fragments of environmental T7-like cyanophages and cyanoprophages were obtained from the JGI IMG/VR (v.4.1)80 and Global Ocean Virome (v.2.0) assemblies81. To collect genomic fragments containing exonuclease genes with sufficiently long downstream regions, a conserved region in the C-terminal part of the exonuclease (corresponding to residues 86–221 in the exonuclease protein from S-TIP37; RefSeq accession YP_009807515.1) was chosen and homologous regions from a reference set of cyanophages were used as queries for tblastn from NCBI blast (v.2.15.0)82. Fragments were retained when they had matches at least 400 bp long, identity of ≥40%, bit score of ≥400 and at least 500 bp of downstream sequence available. To collect genomes for phylogenetic analysis, out of the contigs with the exonuclease gene, we extracted sequences ≥30 kb long and dereplicated them with dRep (v.3.4.5)83 at the ANImf identity level of 0.95. Genes were predicted with Prodigal (v.2.6.3)84 and maximum-likelihood phylogeny was reconstructed with Phylophlan (v.3.0.2)85, Diamond (v.2.1.8)86, MAFFT (v.7.475)87, trimAl (v.1.4.1)88 and IQ-TREE (v.2.1.2)89) based on concatenated alignments of protein sequences of nine core genes: primase-helicase, exonuclease, portal protein (head-to-tail adaptor), head assembly protein, major capsid protein, tail tubular proteins A and B, and small and large terminase subunits. The genes were extracted by searching with Diamond blastp using representative protein sequences as queries from two distantly related cyanophages, P60 and S-SRP02: P60_gp14 (primase-helicase is missing from S-SRP02), P60_gp18 and SSRP02_p034, P60_gp26 and SSRP02_p038, P60_gp27 and SSRP02_p039, P60_gp28 and SSRP02_p040, P60_gp29 and SSRP02_p041, P60_gp30 and SSRP02_p042, P60_gp40 and SSRP02_p011, and P60_gp49 and SSRP02_p012, respectively. IQ-TREE was run with ‘-m LG --alrt 1000 -pers 0.2 -nstop 500’ with 1,000 ultrafast bootstrap replicates90, and the resulting tree was outgroup-rooted with the ingroup defined as the branch encompassing the known cyanophages. We also assessed the incidence and type of DNA polymerase in the genomes by searching the ORFs with blastp using the sequences of the exonuclease (YP_009807513.1) and polymerase (YP_009807514.1) subunits of the split polymerase of S-TIP37 with an E-value threshold of 10−10. Different T7-like cyanophage lineages were found to diverge in the presence and type of the DNA polymerase gene (Extended Data Fig. 6), and this gene was therefore not used as a phylogenetic marker. The cyanophage genomes were classified on the basis of the phylogeny into six clades: the previously recognized clades A (with a single-ORF DNA polymerase), B (a split DNA polymerase) and C (no DNA polymerase) and the newly defined clades R (for P-RSP2-like phages, single-ORF DNA polymerase), S (S-SRP02-like phages, no DNA polymerase) and T (represented by GOV contig Station52_DCM_ALL_NODE_78, no DNA polymerase) (Extended Data Fig. 6). Metadata of the genomes chosen to represent each of the clades are provided in Supplementary Data 4. Two outlier genomes were found to cause spurious clustering and were excluded from the phylogenetic analysis: IMGVR_UViG 3300032116_000204 and IMGVR_UViG 3300029337_000223. The incidence of nblA and psbA genes in the genomes was assessed by searching ORFs (between stop codons) with hmmsearch from HMMER (v.3.4)91 using a custom NblA profile (see below) and the PsbA profile TIGR01151.1 from NCBI Protein Family Models. A version of the tree focusing on representative genomes as shown in the main text was obtained by trimming the full tree. The trees were visualized with ggtree (v.3.2.0)92. The full phylogenetic tree without the outgroups is available in Supplementary Data 5.
Picocyanobacterial genomes
Metadata and genome sequences for picocyanobacteria were downloaded from Cyanorak (v.2)93 and additional Prochlorococcus genomes20,25 were downloaded from Integrated Microbial Genomes & Microbiomes94. Genes were predicted with prodigal and GeneMarkS-2 (v.1.14_1.25)95, with the predictions merged using gffcompare (v.0.12.6)96.
Analysis of NblAs
The NblA HMM profile used to search for nblA genes in T7-like cyanophage genomes was obtained from an alignment of the previously released NblA sequences6 and manually curated NblA sequences from representative T7-like cyanophages. The searches for the NblA hits were performed without heuristic filters and the results were filtered using an empirically determined full score threshold of 20. The same NblA profile was also initially used to search for nblA genes in the protein sequences from picocyanobacteria. After finding that many of their NblAs yield sub-significant hits, we built a separate NblA profile specifically targeting picocyanobacterial NblAs based on the sequences collected in the first round of the search. Note that, due to the high divergence of the NblAs from picocyanobacteria, many of them avoided detection in our previous screen of cyanobacterial genomes using a general NblA protein profile6. The two NblA protein profiles are available in Supplementary Data 10.
A NeighborNet network was constructed to visualize similarity relationships between NblA proteins of different origins. NblAs from T7-like cyanophages and picocyanobacteria were combined with cyanobacterial sequences assigned in UniProt r.2025_297 to Pfam profile PF04485, as well as selected NblA sequences from non-T7-like cyanophages. To reduce redundancy, the sequences were clustered at 90% identity level with CD-HIT (v.4.8.1)98. The cluster representatives were aligned with MAFFT in automated mode, and trimAl was used to trim the alignment (strict mode, minimum column block size of 15). The resulting trimmed alignment was used as input to SplitsTree (v.4.17.0)99, which generated a NeighborNet network based on uncorrected distances. The network was visualized with tanggle (v.1.8.0)100.
Sequence logos were generated for different groups of NblAs using DiffLogo (v.2.26.0)101. The original MAFFT alignment was trimmed in a similar way to that above, but only regions outside of the core NblA domain were trimmed. The fasta file was converted to A3M format (matching columns with <50% gaps) and secondary structure predictions were added with tools from HH-suite (v.3.3.0)64. For the sequence logos, the remaining gaps were coded as the separate state X.
Representative structures of NblA dimers were predicted with ColabFold v.1.5.5 (based on AlphaFold2)65.
Analysis of the global distribution of the cyanophage nblAs
For the analysis of the distribution of nblA downstream of the exonuclease gene, we took all of the initially collected environmental contigs with T7-cyanophage type exonucleases and extracted regions containing the 3′ end of the exonuclease gene with up to 100 bp upstream and up to 1,000 bp downstream. These fragments were searched against exonuclease sequences from the genomes used for phylogeny using blastx and clades were assigned to fragments with best hits to cyanophages at identity level of ≥60%. The fragments coming from the GOV2.0 assemblies were used for recruitment of GOV raw data with bwa (v.0.7.17)102. Quantification was performed with featureCounts from subread (v.2.0.6)103. As the exonuclease and nblA genes frequently overlap, we selected strictly defined windows for read quantification instead of ORF boundaries: the exonuclease was represented by the above-mentioned conservative region at the 3′ end of the gene. For nblA, we chose the location corresponding to the core NblA region based on the hmmsearch matches. Genes assigned to the same cyanophage clade were grouped into metafeatures for quantification and the resulting per-station mapped read pair counts were converted to FPKM values by dividing them by the median feature length (in kb) and the total number of the reads per sample (in millions). The sequences used for recruitment and quantification results in tabular format are provided in Supplementary Data 9.
The bioinformatic workflow was implemented in snakemake104, Python and R.
Calculations of the ocean-wide impact of viral nblA genes on light harvesting by picocyanobacteria
To estimate the ocean-wide impact of viral nblA genes on light harvesting by picocyanobacteria, we used the following data: a 100% reduction in photosynthetic performance for 50% of the time attributable to the cyanophage nblA (Fig. 1e); 1–15% of marine cyanobacterial cells infected by T7-like cyanophages at a given time39,54,55; and 35% (surface waters) to 65% (DCM zone) of marine T7-like cyanophages encoding nblA (Fig. 5). The impact of viral NblA proteins on marine cyanobacterial photosynthesis in surface water was calculated as follows: 50% reduction in photosynthetic light-harvesting performance, with 1–6% of cells infected and 35% of T7-like cyanophages carrying nblA genes, giving an estimated impact of 0.175–1.05% on the photosynthetic performance of the picocyanobacteria. The impact on marine cyanobacterial photosynthesis at the DCM was estimated as follows: 50% reduction in photosynthetic light-harvesting performance, with 5–15% of cells infected and 65% of T7-like cyanophages carrying nblA genes, giving an estimated impact of 1.625–4.875% on the photosynthetic performance of picocyanobacteria.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data supporting the findings of this study are available in the Article and its Supplementary Information. The MS proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository under the following dataset identifiers: PXD057452 (whole proteome data) and PXD057454 (N-terminomics data).
Code availability
The code used for the bioinformatic analyses is available from the GitHub repository (https://github.com/BejaLab/T7_nblA).
References
Breitbart, M., Thompson, L. R., Suttle, C. A. & Sullivan, M. B. Exploring the vast diversity of marine viruses. Oceanography 20, 135–139 (2007).
Gao, E.-B., Huang, Y. & Ning, D. Metabolic genes within cyanophage genomes: implications for diversity and evolution. Genes 7, 80 (2016).
Puxty, R. J., Millard, A. D., Evans, D. J. & Scanlan, D. J. Shedding new light on viral photosynthesis. Photosynth. Res. 126, 71–97 (2015).
Zimmerman, A. E. et al. Metabolic and biogeochemical consequences of viral infection in aquatic ecosystems. Nat. Rev. Microbiol. 18, 21–34 (2020).
Tian, F. et al. Prokaryotic-virus-encoded auxiliary metabolic genes throughout the global oceans. Microbiome 12, 159 (2024).
Nadel, O. et al. An uncultured marine cyanophage encodes an active phycobilisome proteolysis adaptor protein NblA. Environ. Microbiol. Rep. 11, 848–854 (2019).
Yoshida, T. et al. Ma-LMM01 infecting toxic Microcystis aeruginosa illuminates diverse cyanophage genome strategies. J. Bacteriol. 190, 1762–1772 (2008).
Gao, E.-B., Gui, J.-F. & Zhang, Q.-Y. A novel cyanophage with a cyanobacterial nonbleaching protein A gene in the genome. J. Virol. 86, 236–245 (2012).
Collier, J. L. & Grossman, A. R. A small polypeptide triggers complete degradation of light-harvesting phycobiliproteins in nutrient-deprived cyanobacteria. EMBO J. 13, 1039–1047 (1994).
Bienert, R., Baier, K., Volkmer, R., Lockau, W. & Heinemann, U. Crystal structure of NblA from Anabaena sp. PCC 7120, a small protein playing a key role in phycobilisome degradation. J. Biol. Chem. 281, 5216–5223 (2006).
Sendersky, E. et al. The proteolysis adaptor, NblA, is essential for degradation of the core pigment of the cyanobacterial light-harvesting complex. Plant J. Cell Mol. Biol. 83, 845–852 (2015).
Schwarz, R. & Grossman, A. R. A response regulator of cyanobacteria integrates diverse environmental signals and is critical for survival under extreme conditions. Proc. Natl Acad. Sci. USA. 95, 11008–11013 (1998).
Ou, T., Gao, X.-C., Li, S.-H. & Zhang, Q.-Y. Genome analysis and gene nblA identification of Microcystis aeruginosa myovirus (MaMV-DC) reveal the evidence for horizontal gene transfer events between cyanomyovirus and host. J. Gen. Virol. 96, 3681–3697 (2015).
Shitrit, D. et al. Genetic engineering of marine cyanophages reveals integration but not lysogeny in T7-like cyanophages. ISME J. 16, 488–499 (2022).
Grossman, A. R., Schaefer, M. R., Chiang, G. G. & Collier, J. L. Environmental effects on the light-harvesting complex of cyanobacteria. J. Bacteriol. 175, 575–582 (1993).
Ueno, Y., Aikawa, S., Kondo, A. & Akimoto, S. Energy transfer in cyanobacteria and red algae: confirmation of spillover in intact megacomplexes of phycobilisome and both photosystems. J. Phys. Chem. Lett. 7, 3567–3571 (2016).
Ong, L. J. & Glazer, A. N. Phycoerythrins of marine unicellular cyanobacteria. I. Bilin types and locations and energy transfer pathways in Synechococcus spp. phycoerythrins. J. Biol. Chem. 266, 9515–9527 (1991).
Six, C. et al. Diversity and evolution of phycobilisomes in marine Synechococcus spp.: a comparative genomics study. Genome Biol. 8, R259 (2007).
Glazer, A. N. Light guides: directional energy transfer in a photosynthetic antenna. J. Biol. Chem. 264, 1–4 (1989).
Ulloa, O. et al. The cyanobacterium Prochlorococcus has divergent light-harvesting antennae and may have evolved in a low-oxygen ocean. Proc. Natl Acad. Sci. USA 118, e2025638118 (2021).
Partensky, F., Hess, W. R. & Vaulot, D. Prochlorococcus, a marine photosynthetic prokaryote of global significance. Microbiol. Mol. Biol. Rev. 63, 106–127 (1999).
Ting, C. S., Rocap, G., King, J. & Chisholm, S. W. Cyanobacterial photosynthesis in the oceans: the origins and significance of divergent light-harvesting strategies. Trends Microbiol. 10, 134–142 (2002).
Wiethaus, J., Busch, A. W. U., Dammeyer, T. & Frankenberg-Dinkel, N. Phycobiliproteins in Prochlorococcus marinus: biosynthesis of pigments and their assembly into proteins. Eur. J. Cell Biol. 89, 1005–1010 (2010).
Hess, W. R. et al. The photosynthetic apparatus of Prochlorococcus: insights through comparative genomics. Photosynth. Res. 70, 53–71 (2001).
Becker, J. W. et al. Novel isolates expand the physiological diversity of Prochlorococcus and illuminate its macroevolution. mBio 15, e03497-23 (2024).
Steglich, C., Mullineaux, C. W., Teuchner, K., Hess, W. R. & Lokstein, H. Photophysical properties of Prochlorococcus marinus SS120 divinyl chlorophylls and phycoerythrin in vitro and in vivo. FEBS Lett. 553, 79–84 (2003).
Meza-Padilla, I., McConkey, B. J. & Nissimov, J. I. Structural models predict a significantly higher binding affinity between the NblA protein of cyanophage Ma-LMM01 and the phycocyanin of Microcystis aeruginosa NIES-298 compared to the host homolog. Virus Evol. 10, veae082 (2024).
Dekel-Bird, N. P. et al. Diversity and evolutionary relationships of T7-like podoviruses infecting marine cyanobacteria. Environ. Microbiol. 15, 1476–1491 (2013).
Lindell, D. et al. Genome-wide expression dynamics of a marine virus and host reveal features of co-evolution. Nature 449, 83–86 (2007).
Huang, S., Sun, Y., Zhang, S. & Long, L. Temporal transcriptomes of a marine cyanopodovirus and its Synechococcus host during infection. MicrobiologyOpen 10, e1150 (2021).
Maidanik, I. et al. Cyanophages from a less virulent clade dominate over their sister clade in global oceans. ISME J. 16, 2169–2180 (2022).
Nakahira, Y., Ogawa, A., Asano, H., Oyama, T. & Tozawa, Y. Theophylline-dependent riboswitch as a novel genetic tool for strict regulation of protein expression in cyanobacterium Synechococcus elongatus PCC 7942. Plant Cell Physiol. 54, 1724–1735 (2013).
Tahan, R., Shitrit, D. & Lindell, D. A functional cyanophage thioredoxin increases competitive phage fitness. Preprint at bioRxiv https://doi.org/10.1101/2025.10.05.680603 (2025).
Biswas, A., Akhtar, P., Lambrev, P. H. & van Stokkum, I. H. M. Energy transfer from phycobilisomes to photosystem I at room temperature. Front. Plant Sci. 14, 1300532 (2024).
Kondo, K., Mullineaux, C. W. & Ikeuchi, M. Distinct roles of CpcG1-phycobilisome and CpcG2-phycobilisome in state transitions in a cyanobacterium Synechocystis sp. PCC 6803. Photosynth. Res. 99, 217–225 (2009).
Hanna, R. et al. In-depth characterization of apoptosis N-terminome reveals a link between caspase-3 cleavage and posttranslational N-terminal acetylation. Mol. Cell. Proteom. 22, 100584 (2023).
Ma, X., Coleman, M. L. & Waldbauer, J. R. Distinct molecular signatures in dissolved organic matter produced by viral lysis of marine cyanobacteria. Environ. Microbiol. 20, 3001–3011 (2018).
Karradt, A., Sobanski, J., Mattow, J., Lockau, W. & Baier, K. NblA, a key protein of phycobilisome degradation, interacts with ClpC, a HSP100 chaperone partner of a cyanobacterial Clp protease. J. Biol. Chem. 283, 32394–32403 (2008).
Carlson, M. C. G. et al. Viruses affect picocyanobacterial abundance and biogeography in the North Pacific Ocean. Nat. Microbiol. 7, 570–580 (2022).
Komárek, J., Johansen, J. R., Šmarda, J. & Strunecký, O. Phylogeny and taxonomy of Synechococcus-like cyanobacteria. Fottea 20, 171–191 (2020).
Berube, P. M. et al. Single cell genomes of Prochlorococcus, Synechococcus, and sympatric microbes from diverse marine environments. Sci. Data 5, 180154 (2018).
Cai, L. et al. Abundant and cosmopolitan lineage of cyanopodoviruses lacking a DNA polymerase gene. ISME J. 17, 252–262 (2023).
Labrie, S. J. et al. Genomes of marine cyanopodoviruses reveal multiple origins of diversity. Environ. Microbiol. 15, 1356–1376 (2013).
Flombaum, P. et al. Present and future global distributions of the marine Cyanobacteria Prochlorococcus and Synechococcus. Proc. Natl Acad. Sci. USA. 110, 9824–9829 (2013).
Sanfilippo, J. E., Garczarek, L., Partensky, F. & Kehoe, D. M. Chromatic acclimation in cyanobacteria: a diverse and widespread process for optimizing photosynthesis. Annu. Rev. Microbiol. 73, 407–433 (2019).
Kolodny, Y. et al. Phycobilisome light-harvesting efficiency in natural populations of the marine cyanobacteria Synechococcus increases with depth. Commun. Biol. 5, 727 (2022).
Garczarek, L., Hess, W. R., Holtzendorff, J., van der Staay, G. W. M. & Partensky, F. Multiplication of antenna genes as a major adaptation to low light in a marine prokaryote. Proc. Natl Acad. Sci. USA 97, 4098–4101 (2000).
Herranen, M., Tyystjärvi, T. & Aro, E.-M. Regulation of photosystem I reaction center genes in Synechocystis sp. strain PCC 6803 during light acclimation. Plant Cell Physiol. 46, 1484–1493 (2005).
Mann, N. H., Cook, A., Millard, A., Bailey, S. & Clokie, M. Marine ecosystems: bacterial photosynthesis genes in a virus. Nature 424, 741 (2003).
Lindell, D., Jaffe, J. D., Johnson, Z. I., Church, G. M. & Chisholm, S. W. Photosynthesis genes in marine viruses yield proteins during host infection. Nature 438, 86–89 (2005).
Bragg, J. G. & Chisholm, S. W. Modeling the fitness consequences of a cyanophage-encoded photosynthesis gene. PLoS ONE 3, e3550 (2008).
Hellweger, F. L. Carrying photosynthesis genes increases ecological fitness of cyanophage in silico. Environ. Microbiol. 11, 1386–1394 (2009).
Goldin, S., Hulata, Y., Baran, N. & Lindell, D. Quantification of T4-like and T7-like cyanophages using the polony method show they are significant members of the virioplankton in the North Pacific Subtropical Gyre. Front. Microbiol. 11, 1210 (2020).
Gilbert, N. E. et al. Seasonal enhancement of the viral shunt catalyzes a subsurface oxygen maximum in the Sargasso Sea. Preprint at bioRxiv https://doi.org/10.1101/2025.01.23.634377 (2025).
Shopen Gochev, C. et al. Cold surface waters of the sub-Antarctic Pacific Ocean support high cyanophage abundances and infection levels. Environ. Microbiol. 27, e70031 (2025).
Bidle, K. D. & Vardi, A. A chemical arms race at sea mediates algal host–virus interactions. Curr. Opin. Microbiol. 14, 449–457 (2011).
Anantharaman, K. et al. Sulfur oxidation genes in diverse deep-sea viruses. Science 344, 757–760 (2014).
Thompson, L. R. et al. Phage auxiliary metabolic genes and the redirection of cyanobacterial host carbon metabolism. Proc. Natl Acad. Sci. USA 108, E757–E764 (2011).
Dammeyer, T., Bagby, S. C., Sullivan, M. B., Chisholm, S. W. & Frankenberg-Dinkel, N. Efficient phage-mediated pigment biosynthesis in oceanic cyanobacteria. Curr. Biol. 18, 442–448 (2008).
Rihtman, B. et al. Cyanophage MazG is a pyrophosphohydrolase but unable to hydrolyse magic spot nucleotides. Environ. Microbiol. Rep. 11, 448–455 (2019).
Wyman, M., Gregory, R. P. F. & Carr, N. G. Novel role for phycoerythrin in a marine cyanobacterium, Synechococcus strain DC2. Science 230, 818–820 (1985).
Lindell, D., Padan, E. & Post, A. F. Regulation of ntcA expression and nitrite uptake in the marine Synechococcus sp. strain WH 7803. J. Bacteriol. 180, 1878–1886 (1998).
Morris, J. J., Kirkegaard, R., Szul, M. J., Johnson, Z. I. & Zinser, E. R. Facilitation of robust growth of Prochlorococcus colonies and dilute liquid cultures by “helper” heterotrophic bacteria. Appl. Environ. Microbiol. 74, 4530–4534 (2008).
Steinegger, M. et al. HH-suite3 for fast remote homology detection and deep protein annotation. BMC Bioinform. 20, 473 (2019).
Mirdita, M. et al. ColabFold: making protein folding accessible to all. Nat. Methods 19, 679–682 (2022).
van Kempen, M. et al. Fast and accurate protein structure search with Foldseek. Nat. Biotechnol. 42, 243–246 (2024).
Kirzner, S., Barak, E. & Lindell, D. Variability in progeny production and virulence of cyanophages determined at the single-cell level. Environ. Microbiol. Rep. 8, 605–613 (2016).
Bates, D., Mächler, M., Bolker, B. & Walker, S. Fitting linear mixed-effects models using lme4. J. Stat. Softw. 67, 1–48 (2015).
Fox, J. & Weisberg, S. An R Companion to Applied Regression (Sage, 2019).
Larom, S., Salama, F., Schuster, G. & Adir, N. Engineering of an alternative electron transfer path in photosystem II. Proc. Natl Acad. Sci. USA 107, 9650–9655 (2010).
Schoffman, H. & Keren, N. Function of the IsiA pigment–protein complex in vivo. Photosynth. Res. 141, 343–353 (2019).
Liland, K. H., Almøy, T. & Mevik, B.-H. Optimal choice of baseline correction for multivariate calibration of spectra. Appl. Spectrosc. 64, 1007–1016 (2010).
Söding, J., Biegert, A. & Lupas, A. N. The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res. 33, W244–W248 (2005).
Brahamsha, B. A genetic manipulation system for oceanic cyanobacteria of the genus Synechococcus. Appl. Environ. Microbiol. 62, 1747–1751 (1996).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Yu, F., Deng, Y. & Nesvizhskii, A. I. MSFragger-DDA+ enhances peptide identification sensitivity with full isolation window search. Nat. Commun. 16, 3329 (2025).
Deutsch, E. W. et al. Trans-Proteomic Pipeline: robust mass spectrometry-based proteomics data analysis suite. J. Proteome Res. 22, 615–624 (2023).
Hanna, R., Rozenberg, A., Lavy, T. & Kleifeld, O. Increasing the coverage of the N-terminome with LysN amino terminal enrichment (LATE). Methods Enzymol. 686, 1–28 (2023).
Huerta-Cepas, J. et al. eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res. 47, D309–D314 (2019).
Camargo, A. P. et al. IMG/VR v4: an expanded database of uncultivated virus genomes within a framework of extensive functional, taxonomic, and ecological metadata. Nucleic Acids Res. 51, D733–D743 (2023).
Gregory, A. C. et al. Marine DNA viral macro- and microdiversity from pole to pole. Cell 177, 1109–1123 (2019).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinform. 10, 421 (2009).
Olm, M. R., Brown, C. T., Brooks, B. & Banfield, J. F. dRep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication. ISME J. 11, 2864–2868 (2017).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinform. 11, 119 (2010).
Asnicar, F. et al. Precise phylogenetic analysis of microbial isolates and genomes from metagenomes using PhyloPhlAn 3.0. Nat. Commun. 11, 2500 (2020).
Buchfink, B., Reuter, K. & Drost, H.-G. Sensitive protein alignments at tree-of-life scale using DIAMOND. Nat. Methods 18, 366–368 (2021).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Hoang, D. T., Chernomor, O., von Haeseler, A., Minh, B. Q. & Vinh, L. S. UFBoot2: improving the ultrafast bootstrap approximation. Mol. Biol. Evol. 35, 518–522 (2018).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Xu, S. et al. Ggtree: a serialized data object for visualization of a phylogenetic tree and annotation data. iMeta 1, e56 (2022).
Garczarek, L. et al. Cyanorak v2.1: a scalable information system dedicated to the visualization and expert curation of marine and brackish picocyanobacteria genomes. Nucleic Acids Res. 49, D667–D676 (2021).
Chen, I.-M. A. et al. IMG/M v.5.0: an integrated data management and comparative analysis system for microbial genomes and microbiomes. Nucleic Acids Res. 47, D666–D677 (2019).
Lomsadze, A., Gemayel, K., Tang, S. & Borodovsky, M. Modeling leaderless transcription and atypical genes results in more accurate gene prediction in prokaryotes. Genome Res. 28, 1079–1089 (2018).
Pertea, G. & Pertea, M. GFF Utilities: GffRead and GffCompare. F1000Research 9, ISCB Comm J-304 (2020).
The UniProt Consortium. UniProt: the Universal Protein Knowledgebase in 2025. Nucleic Acids Res. 53, D609–D617 (2025).
Li, W. & Godzik, A. Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–1659 (2006).
Huson, D. H. & Bryant, D. Application of phylogenetic networks in evolutionary studies. Mol. Biol. Evol. 23, 254–267 (2006).
Schliep, K. et al. Tanggle: visualization of phylogenetic networks (Bioconductor, 2023); https://doi.org/10.18129/B9.bioc.tanggle.
Nettling, M. et al. DiffLogo: a comparative visualization of sequence motifs. BMC Bioinform. 16, 387 (2015).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at arxiv.org/abs/1303.3997 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Mölder, F. et al. Sustainable data analysis with Snakemake. F1000Research 10, 33 (2021).
Sabehi, G. & Lindell, D. The P-SSP7 cyanophage has a linear genome with direct terminal repeats. PLoS ONE 7, e36710 (2012).
Acknowledgements
We thank F. Salama for biophysical guidance; M. Suissa-Szlejf for help in setting up the phycobilisome isolation gradients; R. Edrei for help with spectroscopy measurements; G. Sabehi for discussions on infection experiments; S. Goldin for help with qPCR calculations; S. Larom for technical support; N. Keren and C. Kranzler for discussions on photosynthesis experiments; N. Keren for help in performing spectroscopy; and N. Frankenberg-Dinkel for comments on the manuscript. This work was funded by the European Research Council (ERC Advanced Grant 321647 to O.B. and ERC Consolidator Grant 646868 to D.L.), the Nancy and Stephen Grand Technion Energy Program (GTEP), Israel Science Foundation grants to O.B. (143/18) and O.K. (1515/23), and the Simons Foundation Life Science Award 735081 to D.L.; O.B. holds the Louis and Lyra Richmond Chair in Life Sciences and D.L. holds the Dresner Chair in Life Sciences and Medicine.
Author information
Authors and Affiliations
Contributions
O.N. conceived the project, performed viral work, phycobilisome isolation and experimentation and, together with O.B., O.K. and D.L., designed the experiments. I.P. performed phage growth curve experiments. A.R. performed bioinformatic and statistical analyses. R.H. and O.K. performed proteomic analyses. D.S., R.T. and D.L. developed the methods for constructing the mutant phage and the cyanobacterial NblA expression strain. O.B. coordinated the project. O.N., A.R., O.B., O.K. and D.L. wrote the paper, which was revised and approved by all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Frédéric Partensky and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Expression of the cyanophage nblA gene during S-TIP37 infection of Synechococcus sp. strain WH8109.
a, Transcript levels of S-TIP37 cyanophage nblA and DNA polymerase genes and one of the host nblA genes during infection with WT cyanophage determined by qRT-PCR. Transcript levels are normalized to the rnpB gene. Among the host nblA genes, nblA2 was chosen as the gene coding for the NblA protein with the highest structural similarity to previously characterized NblAs (see Methods). b, Protein mass spectrometry intensity levels of the S-TIP37 cyanophage NblA, DNA polymerase, and host NblA6 proteins during infection with WT (left) or ∆nblA mutant (right) cyanophages, determined by LC-MS/MS analysis. NblA6 was the only host NblA protein detected. Data are presented as mean ± standard deviation of three independent biological replicates, n = 3.
Extended Data Fig. 2 Spectroscopic characteristics of cells infected with wild-type and ∆nblA mutant S-TIP37 cyanophages.
a, Relative absorbance of PUB and PEB at 12 h post-infection. Relative absorbance was determined as follows (see the panel to the left): the absorbance spectra were background-subtracted (red line) and the intensities were scaled to the height of the Chl peak. UI, uninfected, KO, infection with the ∆nblA mutant, and WT, infection with the wild-type phage. b, Relative chlorophyll fluorescence at different time points during infection. The samples were excited at 495 nm (excitation maximum of PUB), emission spectra were obtained and the height of the chlorophyll peak relative to the PE peak was calculated (see the panel to the left). The PE peak at 560 nm is made up of PEI and PEII, to which the PEB and PUB chromophores bind. In both panels, values in the pairwise comparisons indicate p-values from the Tukey’s (two-sided) post-hoc tests and asterisks denote significant pairs of differences: *** — p-value < 0.001 (after FDR adjustment in panel b). See Suppl. Data File 2 for further statistical details and exact p-values. Boxplots reflect minimum-maximum range (vertical line), interquartile range (rectangle) and median (horizontal line). The results are based on five biological replicates of each condition: uninfected cells and cells infected either with the ∆nblA (KO) or wild-type (WT) phage, sampled repeatedly at different time points for measurement. For further results of the statistical analyses refer to Suppl. Data File 2. Representative spectra are shown in Fig. 2a,b.
Extended Data Fig. 3 Fluorescence spectral analysis of fractionated phycobilisomes.
Fluorescence emission spectra at 77 K of fractionated phycobilisomes from uninfected Synechococcus sp. strain WH8109. The lower density, top band (see Fig. 2c), contains disassembled phycobilisome complexes as seen by the single fluorescence peak corresponding to the excited PE protein. In comparison, the higher density, bottom band (see Fig. 2c), contains assembled phycobilisome complexes characterized by the presence of three phycobiliprotein peaks which indicates intact energy transfer from PE to PC and APC.
Extended Data Fig. 4 Fluorescence emission spectra upon ectopic expression of cyanophage NblA in Synechococcus sp. strain WH8109 during infection.
Fluorescence emission spectra of uninfected cells (green), cyanobacteria infected by the WT cyanophage (blue) and by the ∆nblA mutant cyanophage (red) at different time points post-infection. The host strain carries a plasmid for inducible expression of the cyanophage nblA gene. Ectopic expression was induced in the host 8 h prior to addition of the phages (NblA induction, the panels to the right). Control experiments (the panels to the left) were performed with the same host strain but without the addition of the inducer. Measurements were taken at room temperature and normalized using the PE peak.
Extended Data Fig. 5 Cleavage and proteome changes upon ectopic expression of cyanophage NblA in Synechococcus sp. strain WH8109.
Proteomic analysis of cleavage events (a and b) and total protein abundance changes (c and d) following ectopic expression of the cyanophage NblA in Synechococcus sp. strain WH8109 cells for 24 h (a and c) and 48 h (b and d), compared to non-induced controls. In both analyses, n = 2 biologically independent experiments. The top panels show the abundance ratios of neo-N-terminal peptides identified by N-terminomics, calculated as the ratio of signal intensities between induced and non-induced samples, reflecting changes in peptide abundance. The bottom panels show the abundance ratios for proteins in the total proteomic analysis based on the intensities of tryptic peptides. i.e. Each data point represents an individual peptide or protein. The values represent averages of two biological replicates. Proteins are categorized according to a modified classification of COG functional groups (see Methods). Filled triangles indicate neo-N-terminal peptides and proteins with absolute log2-ratios ≥1 in both replicates. Upward-pointing and downward-pointing triangles represent peptides more and less abundant, respectively, upon NblA ectopic expression relative to the control. Red asterisks indicate significant deviations of the category-wise mean ratios from zero (two-sided one-sample t-test) after FDR adjustment across all comparisons: *** — adjusted p-value ≤ 0.001, ** — adjusted p-value ≤ 0.01, see Suppl. Data File 2 for details of the statistics and the exact p-values.
Extended Data Fig. 6 Phylogeny and distribution of nblA genes among T7-like cyanophages.
Maximum likelihood phylogenetic tree was built based on concatenation of protein alignments of nine core genes coding for: primase-helicase, exonuclease, portal protein (head-tail adaptor), head assembly, major capsid protein, tail tubular proteins A and B, and small and large terminase subunits. Phage genomes used in the phylogeny were recruited from among cultured T7-like cyanophages and from environmental sources: Global Ocean Virome v. 2 assemblies and JGI IMG/VR v. 4.1 (see Suppl. Data File 4). The environmental genomes were recruited by searching for the exonuclease gene, selecting genomes at least 30 kb long, dereplicating them at 0.95 ANImf identity level and requiring the presence of at least eight core genes. The tree was outgroup-rooted by inclusion of coliphage T7, pelagiphage HTVC011P, pelagiphage HTVC019P and environmental phages falling outside of the cyanophage cluster. The pie chart summarizes the numbers of non-redundant phage genomes with and without nblA in each clade (notice that no clade C, S or T genomes were found to code for nblA). Phages were categorized into previously recognized clades A (P60-like), B (S-TIP37-like) and C (P-SCSP1a/2-like) and newly-delineated clades R (P-RSP2-like), S (S-SRP02-like) and T (Ga0063588_100037_PP-like, no cultured representatives). Presence and type of DNA polymerase genes is indicated with the clades dominated by phages falling in one of the following categories: no DNA polymerase gene (clades C, S, T), DNA polymerase coded in a single ORF (clades A and R) and a split DNA polymerase gene105 with the exonuclease domain coded in a separate ORF (clade B). Clade representatives are coloured with labels according to the host’s phylogenetic affiliation when known. Environmental genomes are indicated by asterisks. Note that each of the three picocyanobacterial lineages represents an assemblage of several genus-level taxa. The incidence of nblA (circles) and psbA (squares) genes is indicated with symbols at the tips of the corresponding branches and the nblA genes of two categories are differentiated: those located downstream of the exonuclease gene (as in S-TIP37) and the nblA gene downstream of an HNH endonuclease gene (in a subclade of clade R). Solid circles mark branches with ultrafast bootstrap support of ≥95. See Suppl. Data File 7 for the source data.
Extended Data Fig. 7 Phylogenetic network of NblA proteins from the T7-like cyanophages, marine picocyanobacteria and other cyanophages and cyanobacteria.
The NblA protein sequences were aligned, trimmed and a NeighborNet network was obtained based on uncorrected distances. The proteins are categorized by the corresponding cyanophage or cyanobacterial group. NblA was found in the low-light adapted Prochlorococcus ecotypes LLII/LLIII, LLIV, LLVII, LVIII, AMZIA, AMZIB, and AMZII but not in LLI and AMZIII nor any of the high-light-adapted ecotypes. NblAs from selected cyanophages and model cyanobacteria with the NblAs from S-TIP37, Synechococcus sp. strain WH8109 and Prochlorococcus sp. strain SS120, are highlighted in bold. The network and trimmed sequences of cluster representatives in NEXUS format are available in Suppl. Data File 8 and metadata and untrimmed sequences and aligned sequences of the core NblA region are available in Suppl. Data File 11.
Extended Data Fig. 8 Structures and protein profiles of NblA proteins from T7-like cyanophages and cyanobacteria.
Left: Dimeric structures of selected representatives of the NblA proteins. All of the structures, except that of the NblA from Nostoc sp. strain PCC 7120 [PDB:1OJH10] were obtained with ColabFold (AlphaFold2)65. Right: Sequence logos generated based on a trimmed alignment of clustered NblA sequences. Numbers of cluster representatives per group are provided in parentheses. Secondary structure predictions based on the whole alignment of all NblA types are shown at the bottom (H: helix, C: coil) with the height proportional to the prediction score. Positions known to mediate phycobiliprotein binding38 are highlighted with red-framed boxes: protein profile positions 47 (residues L51 in Nostoc sp. strain PCC 7120 and L64 in cyanophage S-TIP37) and 49 (residues K53 in PCC 7120 and T66 in S-TIP37).
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Nadel, O., Hanna, R., Rozenberg, A. et al. Viral NblA proteins negatively affect oceanic cyanobacterial photosynthesis. Nature 648, 434–442 (2025). https://doi.org/10.1038/s41586-025-09656-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09656-x